Abstract
Lung cancer is the most common cancer worldwide. Up to 85% of lung cancer cases are diagnosed as non-small cell lung cancer (NSCLC). The effectiveness of NSCLC treatment is expected to be improved through the implementation of robust and specific biomarkers. MicroRNAs (miRNAs) are small, non-coding molecules that play a key role in the regulation of basic cellular processes, including differentiation, proliferation and apoptosis, by controlling gene expression at the post-transcriptional level. Deregulation of miRNA activity results in the loss of homeostasis and the development of a number of pathologies, including lung cancer. During lung carcinogenesis, miRNAs exhibit dual regulatory function: they act as oncogenes to promote cancer development or as tumour suppressors. Unique miRNA sequences have been detected in malignant tissues and corresponding healthy tissues. Furthermore, stable forms of tumour-related miRNAs are detectable in the peripheral blood of patients with NSCLC. The potential benefits of using extracellular miRNAs present in body fluids as part of the diagnostic evaluation of cancer include low invasiveness (compared with tumour cell/tissue sampling), and the repeatability and ease of obtaining the specimens. Apart from the diagnostic applications of altered miRNA expression profiles, the dual regulatory role of miRNA in cancer might drive the further development of personalised therapies in NSCLC. The clinical usefulness of miRNA expression analysis to predict the efficacy of various treatment strategies including surgery, radio- and chemotherapy, and targeted therapies has been evaluated in NSCLC. Also, the capacity of a single miRNA to regulate the expression of multiple genes simultaneously presents an opportunity to use these small molecules in personalised therapy as individualised therapeutic tools.

Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Lung cancer is the most common cancer worldwide and causes over 1.6 million deaths per year [1]. Up to 85% of lung cancer cases are diagnosed as non-small cell lung cancer (NSCLC). High mortality of NSCLC results from the fact that a majority of patients are diagnosed with advanced disease, when the possibility of offering potentially curative surgical treatment is limited. Five-year survival rates are greatly improved when the disease is found while still localized; unfortunately, only 16% of lung cancers are diagnosed at this early stage. For advanced stages with metastatic tumours the 5-year survival rate is only 4% [2]. Key problems are the lack of effective tools and methods for early detection of NSCLC and its resistance to the majority of the currently used therapies.
The past two decades have seen considerable progress in research on the underlying molecular mechanisms of lung carcinogenesis and the recognition of NSCLC as a disease with complex genetics. This gave hope for realistic chances to successfully implement the concept of personalised medicine in the management of lung cancer. The personalisation of diagnostic and therapeutic approaches implies the correct sub-classification of a tumour type based on unique histological and molecular features determining the choice of treatment. The final objective of a personalised approach is to improve a disappointing overall survival in NSCLC.
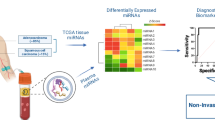
One prerequisite for the development of personalised medicine is the identification of robust biomarkers to guide clinical decision-making [3, 4]. In this context, the potential of small, non-coding microRNA (miRNA) molecules has rapidly become apparent. The clinical applicability of miRNAs in the diagnosis and treatment of NSCLC is currently under evaluation (Fig. 1). Although no miRNA biomarker has been validated and approved for cancer diagnostics to date, there are numerous in vitro and in vivo studies that demonstrate great clinical potential of these molecules.
Potential clinical applications of miRNAs as biomarkers for diagnosis and treatment of NSCLC. The expression of biomarker miRNA, which may consist of a single or multiple miRNA species, might be effectively evaluated in either tumour tissue obtained by biopsy or blood specimens (plasma, serum) collected in a minimally invasive manner as a so-called liquid biopsy before, during and after the treatment. Validated miRNA signatures of lung cancer subtypes could serve as an auxiliary tool in the diagnostic classification of the disease. Circulating miRNA biomarkers detectable in plasma/serum might greatly aid in the differential diagnosis of malignant and benign lung nodules through preselecting the patients for further more expensive or invasive procedures. The serial blood collection during the treatment also offers a unique opportunity for therapy effectiveness monitoring in real time by tracing the dynamic changes in expression levels of selected miRNA biomarkers
The need for a personalised approach in the management of screen-detected nodules has been recently emphasized by the National Lung Screening Trial (NLST), the first study to show a statistically significant 20% reduction in lung cancer mortality among high-risk individuals screened with low-dose computed tomography (LDCT) scans, when compared to chest X-ray [5]. However, a high rate of false positive results associated with LDCT screening opened a debate over the cost-effectiveness of LDCT screening programs and the potential harms related to overdiagnosis and radiation-induced cancers. In view of personalised medicine, the potential role for specific, miRNA-based biomarkers to complement the radiological modalities and increase the total sensitivity and specificity of the lung cancer screening process is currently being investigated [6, 7].
The last decade identified a number of genetic alterations in NSCLC as useful predictive biomarkers and assigned a permanent position to molecular biology, together with histology and radiology, in the selection of optimal treatment strategies for lung cancer patients. In the era of ‘theranostics’, therapeutics and diagnostics have been meaningfully combined to achieve personalised pharmacotherapy. For NSCLC, much of the work in recent years has focussed on mutations of the epidermal growth factor receptor (EGFR) and on the abnormal fusions of the anaplastic lymphoma kinase (ALK) or the c-ros oncogene 1 (ROS1) genes. While conventional chemotherapy remains a gold standard in the management of advanced NSCLC patients without druggable genetic abnormalities, patients whose tumours harbouring specific alterations in EGFR or ALK/ROS1 genes are adequate for the targeted therapy, offering a prolonged progression-free survival as compared to chemotherapy [8]. However, the relatively rapid acquisition of resistance to such treatments that is observed in virtually all patients significantly limits their utility and remains a substantial challenge to the clinical management of advanced lung cancers and the further development of targeted therapies. Since molecular mechanisms of resistance have been identified, new strategies to overcome or prevent the development of resistance have emerged, including the regulation of specific signalling pathways by epigenetic mechanisms [9].
Finally, advances in the understanding of immune evasion strategies used by tumours enabled the development of new immunotherapies and culminated in positive results with checkpoint inhibitors in randomized clinical trials. Several programmed death-1 (PD-1) and programmed death ligand-1(PD-L1) inhibitors have been approved by the Food and Drug Administration (FDA) to treat metastatic NSCLC. Still, many questions remain to be addressed on immunotherapy, regarding the optimal schedule of treatment, identifying proper predictive biomarkers and co-targeting of the other key modulators of tumour immune response [10]. Recently, various miRNAs have been found to target key cancer-related immune pathways, which seem involved in the secretion of immunosuppressive or immunostimulating factors by cancer or immune cells [11].
The pronounced role that miRNAs have across human diseases, including all cancer types, led to the development of new therapeutic strategies through identification and validation of miRNAs that are causally involved in the disease process and the effective regulation of target-miRNA function by a drug. Recently, the clinical trial on “Miravirsen” (SPC3649), a synthetic oligonucleotide complementary to miR-122 which can sequester and inhibit the activity of this miRNA, has been extended to long-term phase 2 study for patients with chronic hepatitis C virus genotype 1 infection [12]. This shows some promise for the successful implementation of currently developed miRNA-based therapeutics for malignant diseases, which currently are still mostly evaluated in early preclinical phases. The aim of this review is to highlight the most promising studies reported to date that investigate the clinical applicability of miRNAs, either as a biomarker or therapeutics, in lung cancer treatment.
2 miRNA Biogenesis and Function
miRNAs are a class of non-coding, endogenous, single-stranded small RNA molecules composed of 19–22 nucleotides. miRNAs function as regulators of gene expression in both plant and animal cells [13, 14]. It is believed that in mammals including humans, miRNAs may regulate more than 50% of all protein-encoding genes [15]. Ever since their discovery, the number of newly identified human miRNAs has been increasing constantly, and more than 2500 known sequences have been identified thus far [16]. This corresponds to approximately 1–4% of all expressed genes in humans, and thus, miRNAs are currently considered as one of the largest classes of gene regulators [17–19]. miRNA molecules can regulate genes at the post-transcriptional level by specifically recognising and affecting messenger RNA (mRNA), depending on the degree of homology with the targeted sequence [20].
The standard miRNA biogenesis pathway consists of two cleavage events, one nuclear and one cytoplasmic [21]. After the cleavage, primary miRNAs are processed into active mature miRNAs through a series of biochemical steps, and miRNA expression can be regulated at each step of biogenesis pathway (Fig. 2).
Biogenesis of miRNA. miRNAs are transcribed by RNA polymerase II into primary miRNAs (pri-miRNAs), which are several-fold larger than mature miRNAs (usually 100–1000 nucleotides). In the nucleus, pri-miRNAs are processed into precursor miRNAs (pre-miRNAs) consisting of approximately 60–120 nucleotides by a ribonuclease (RNase) III displaying endonuclease activity (the Drosha RNase) and the protein DGCR8 (Pasha). Subsequently, pre-miRNAs are folded into the characteristic hairpin structure and transported by RAN-GTP-dependent exportin 5 to the cytoplasm, where they are further processed by the Dicer RNase III, to target miRNA sequences of 19–22 nucleotides. Mature miRNA initially has the form of an asymmetric duplex of two strands of miRNA:miRNA*. Usually, the strand that contains the less thermostable 5′-terminus is packaged into a protein complex (RISC) whose main component is an AGO protein. The miRISC complex can then act in the cell or be secreted into the extracellular space inside extracellular vesicles (EVs) or as complexes with RNA-binding AGO proteins or high-density lipoproteins. There are two ways for EVs to be secreted from donor cells: (1) microvesicles are directly shed from the cell membrane; and (2) the intraluminal endosomal vesicles are released from the multivesicular body (MVB) into the extracellular space to become exosomes [22–35].
Genes encoding miRNAs are often clustered and are not only present in exons but also in introns and untranslated regions (UTRs) [36, 37]. This configuration of transcription units allows for simultaneous formation of both miRNA and mRNA transcripts. The organisation of the genes encoding miRNAs allows for the activity of polymerases II and III which are both involved in transcribing genes encoding small RNAs [38, 39]. However, regions encoding pre-miRNA sequences have been shown to contain approximately 2000 single nucleotide polymorphisms (SNPs) which may affect miRNA–mRNA interactions [40]. Genetic alterations in miRNA sequences are likely to affect their regulatory activity and, consequently, a number of cellular processes including carcinogenesis [41]. The rs11614913 (C → T) SNP in pre-miR-192a2 has been linked to a higher risk of NSCLC [42–44].
Interestingly, Czubak et al. [45] showed several miRNA genes (miR-30d, miR-21, miR-17 and miR-155), as well as two miRNA biogenesis genes, DICER1 and DROSHA, to be frequently amplified in tumour tissue specimens from 254 NSCLC patients. Moreover, the copy number variation of DICER1 and DROSHA correlated well with their expression and the survival of NSCLC patients. Upregulation of the expression of DROSHA and DICER1 decreases or increases the survival, respectively. This study demonstrated that gene copy number variation may be an important mechanism of upregulation/downregulation of miRNAs in lung cancer and suggest an oncogenic role for DROSHA.
Abnormal regulation of miRNA expression has been shown to interfere with important cellular processes, such as differentiation, proliferation and apoptosis, resulting in the loss of homoeostasis and the development of a number of diseases including tumours [46]. Some miRNAs can act as oncosuppressors, while others act as oncogenes that stimulate the growth of tumours [47]. miRNAs that are overexpressed in malignant cells (oncomiRs), such as miR-21, act as oncogenes that promote the development of tumours by negatively regulating tumour suppressor genes and/or genes that control cellular processes such as differentiation and apoptosis. miRNAs that are downregulated in cancer, such as let-7, function as oncosuppressors and can suppress tumour development by regulating oncogenes and/or genes involved in the cell cycle. The various miRNA expression patterns are unique for specific tissue types. These molecules are either over- or underexpressed depending on the tumour type [36, 48].
2.1 Extracellular miRNA
Studies conducted by several research groups have confirmed the presence of miRNAs in various body fluids in humans, including serum [49–51], plasma [49, 50], saliva [52], urine [53], milk [54], cerebrospinal fluid [55] and seminal fluid [56]. In cancer, there is a distinct relationship between the type of biological material and the original location of the neoplasm (e.g. urine – bladder cancer, cerebrospinal fluid – brain tumour etc.) which may be potentially significant to the development of a new class of non-invasive diagnostic tests based on extracellular nucleic acids. Accordingly, tumour-related miRNAs have been found in bronchoalveolar lavage fluid [57, 58], pleural effusion fluid [59, 60] and blood plasma/serum [15, 51, 61] from lung cancer patients.
The mechanisms of how miRNAs can be released from cells include both cell death (necrosis and apoptosis) and active secretion [62]. In the latter, miRNAs are secreted inside exosomes and other extracellular vesicles (EVs) or as complexes with RNA-binding AGO proteins or high-density lipoproteins (Fig. 2). Studies investigating miRNAs in the blood have confirmed their high stability in both plasma and serum [51]. Biochemical analyses have shown that extracellular miRNAs present in the blood are resistant to RNases and are exceptionally stable at extreme pH values and temperatures [49, 50]. This resistance is suggested to be the result of miRNA selective packaging (e.g. within exosomes or EV) and association with RNA binding proteins such as AGO, nucleophosmin (NPM1) or ribosomal proteins which provide a robust protective effect on miRNAs from RNases activity [63]. Nevertheless, it is not entirely clear how miRNAs manage their stability.
In a study by Weber et al. [64], the total amount of RNA isolated from various body fluids was found to range widely from 113 to 48,240 μg/L. Plasma, cerebrospinal fluid, pleural effusion fluid and urine contained less RNA than seminal fluid, saliva and bronchoalveolar lavage fluid did. The numbers of various miRNA sequences detected by real-time reverse transcription polymerase chain reaction (RT-qPCR) ranged from 204 in urine to up to 458 in saliva. The absolute total amount of extracellular RNA in plasma was estimated to be in the low nanomolar range. The concentration of extracellular miRNA in the plasma of healthy donors was estimated to be approximately 100 fM [64, 65]. The concentrations of individual miRNAs are therefore thought to be a fraction of this value. Significantly increased or decreased release of certain miRNAs into the circulation is a characteristic of individual tumour types including lung cancer [66].
EV-derived miRNAs function in cell-cell communication and play roles in various biological processes including immune system regulation, inflammation and tumour development [67]. Exosomes can be secreted by most types of cancers including lung cancer. One notable feature of cancer cells is that they produce exosomes in greater amounts than normal cells do, and this feature can be a useful diagnostic biomarker [68]. Rabinowits et al. [67] evaluated circulating exosomal miRNA levels of patients with lung adenocarcinoma and compared them with those of patients without lung cancer, showing that the miRNA signatures of exosomes paralleled those of the miRNA expression profiles of the originating tumour cells. Exosomes from tumours (tumour-derived exosomes) have protumorigenic functions and can promote cancer stimulatory activities such as proliferation, extracellular matrix remodelling, migration, invasion, angiogenesis and contribute in the metastatic cascade [69]. It is postulated that EV-derived miRNAs are therefore potentially better disease biomarkers than other forms of circulating miRNAs.
miRNA levels and profiles in bodily fluids may reflect not only the body’s physiological status but, more importantly, various pathological conditions [70]. Changes in the expression profiles of a few or multiple extracellular miRNAs may be a useful diagnostic markers for the early detection and identification of the tumour type and a prognostic and/or predictive marker for establishing prognosis and treatment [62]. The potential benefits of using extracellular miRNAs present in bodily fluids, sampled via so-called liquid biopsies, as a part of the diagnostic evaluation of cancer include low invasiveness compared with tumour cell/tissue sampling, repeatability and ease of obtaining specimens. In addition, analysis of plasma/serum as a reservoir of miRNAs released by tumour cells from different parts of the primary tumour or from various locations in the body (distant metastases) evades the issues encountered with cellular and molecular tumour heterogeneity [71]. A single tumour tissue sample obtained by biopsy is not fully representative of the molecular changes occurring in developing lung cancers, nor does it reflect the diversity of tumour clones found in distant metastases. In this case, analysis of extracellular miRNAs should not only enable disease monitoring but also allow for effective treatment monitoring, which seems to be a key factor in improving prognosis [72]. Extracellular miRNA sequences might also be an early marker of recurrence after radical treatment.
2.2 Methods of miRNA Expression Analysis
A reliable expression analysis of miRNAs, particularly extracellular miRNAs, in bodily fluids still remains an analytical challenge because of the unique characteristics of miRNA molecules (small size, high homology among miRNAs within the same family and low concentrations in body fluids), the lack of standardized methodologies, and the different detection methods and sensitivities of the various commercial reagent kits and systems [73–75]. Currently, the methods most commonly used for the analysis of miRNA expression include RT-qPCR, microarrays and next-generation sequencing (NGS). Table 1 summarises the key advantages and potential limitations of each of these methods. In terms of equipment, reagents and labour costs, RT-qPCR seems to be the most suitable technique for clinical diagnostics. This method is much cheaper than NGS and does not require extensive technical facilities and specialised bioinformatics staff to analyse the results. While NGS is commonly regarded as the future of molecular biology and genomics, it is undoubtedly also the future of clinical diagnostics, as it allows simultaneous analysis of a large number of DNA/RNA sequences in multiple samples. However, the multiple advantages PCR offers make it the current gold standard for molecular tumour diagnostics.
The low repeatability of testing in terms of selecting the optimal panel of miRNAs as NSCLC biomarkers results from a lack of standard methodologies and frequent errors during the planning stage of experiments (e.g. incorrect selection or poor representativeness of the patient groups, incorrect processing and storage of the specimens and insufficient statistical power of the study). There is also no consensus on the normalisation method used for the resulting data, which is why all steps of an analytical procedure must be validated and standardised before any miRNA-based diagnostic method can be implemented into clinical practice [76].
3 miRNA as Potential Biomarkers in NSCLC
miRNA expression profiles are highly specific to individual types of cells, tissues and organs [48]. Unique miRNA sequences are detected in corresponding healthy and malignant tissues [77]. More than 50% of all miRNAs in humans are located in sensitive regions of chromosomes that undergo amplification, deletion and translocation during carcinogenesis [78, 79]. Attempts to determine the order of the epigenetic and genetic changes that occur during lung carcinogenesis have shown that the earliest alterations, which take place during the hyperplastic and metaplastic stages, include loss of heterozygosity on chromosomes 3p and 9p and microsatellite instability and deregulation of telomerase expression; all of these changes are progressive. In dysplastic lesions, chromosomal aberrations (aneuploidy), chromosomal deletions, methylation, mutations in suppressor genes (e.g. TP53, FHIT, RB1, MYC) and telomerase activation are also observed. At the pre-invasive carcinoma stage (carcinoma in situ), mutations in important oncogenes (e.g. KRAS and HER2/neu) are detected. Invasive lung carcinoma is characterised by the presence of many various genetic and epigenetic alterations in tumour cells [80]. This leads to deregulated expression of certain miRNAs [81], resulting in significant changes in their concentrations and compositions and, consequently, their activities (described as miRNA under- or overexpression), and these miRNAs are potential biomarkers of clinical relevance [82, 83]. Previous studies have confirmed the usefulness of miRNAs as biomarkers of NSCLC [48, 84–86].
Two meta-analyses by He et al. [87] (510 patients and 465 healthy volunteers) and by Wang et al. [88] (2121 patients and 1582 volunteers) provided an insight into the overall diagnostic performance of miRNA and explored the influential factors that may affect the diagnostic accuracy of miRNA in NSCLC. In both analyses, panels composed of several miRNAs offered a much higher diagnostic value than single miRNAs and showed a much higher application potential as prospective biomarkers of NSCLC. In the meta-analysis conducted by He et al. [87], a single miRNA biomarker had a sensitivity of 78.3% in detection of early-stage NSCLC, whereas a miRNA panel had a sensitivity of 83%. Similar results were reported by Wang et al. [88], who obtained a 77% sensitivity and 71% specificity for a single miRNA, and 83% sensitivity and 82% specificity for multiple miRNAs. It is essential to comment, however, that the approximately 80% sensitivity reported in those studies is of limited diagnostic use without further stratification of the risk groups.
Importantly, several trials have shown promise for using multiple miRNA signatures as biomarkers for screening or evaluation of suspected cancer in high-risk groups. In the MILD screening trial, investigators evaluated the diagnostic performance of plasma microRNAs as complementary biomarkers for LDCT screening in a cohort of current or former smokers, older than 50 years [6]. They retrospectively evaluated plasma miRNA signatures in samples from 939 participants, including 69 patients with lung cancer and 870 disease-free individuals. The diagnostic performance of miRNAs for lung cancer detection was 87% for sensitivity and 81% for specificity; a negative predictive value (NPV) of 99% and the combination of both miRNA and LDCT resulted in a fivefold reduction of the LDCT false positive rate. Similarly, another group of investigators used a 13 miRNA signature on 1115 participants (heavy smokers, older than 50 years) in the COSMOS lung cancer screening trial and reported sensitivity and specificity of 79.2% and 75.9%, respectively, with an NPV of 99% [88]. A negative result was found in 810 out of the 1067 (76%) individuals without lung cancer and in 10 of the 48 (21%) individuals with lung cancer, and the authors suggested that the miRNA test could be used as a first-line screening tool in high-risk individuals.
However, the usefulness of miRNAs extends beyond the early detection of NSCLC [89, 90]. miRNA panels offer the possibility to differentiate NSCLC from benign lesions and to determine the histological tumour type from a tissue sample [87, 91], with sensitivities and specificities within the range of 60–100%. In their analysis, Boeri et al. [89] suggested that specific miRNAs detectable in plasma samples collected from healthy smokers with an average exposure of 40 pack-years allowed the identification of those with NSCLC 1–2 years before the first symptoms of lung cancer became evident and allowed determination of their prognoses. Cazzoli et al. [92] screened 742 miRNAs isolated from circulating exosomes and identified four miRNAs (miR-378a, miR-379, miR-139-5p and miR-200b-5p) as screening markers to segregate lung adenocarcinoma and granuloma patients, from healthy former smokers (AUC = 0.908; P < 0.001). Then they identified six miRNAs (miR-151a-5p, miR-30a-3p, miR-200b-5p, miR-629, miR-100 and miR-154-3p) that segregated lung adenocarcinoma patients and lung granuloma patients (AUC = 0.760; P < 0.001). Early detection of lung cancer using specific miRNAs, which in most cases is not possible with conventional imaging methods, offers prospects that more sensitive diagnostic algorithms will be developed. In contrast to studies investigating the usefulness of miRNAs as diagnostic and prognostic markers, relatively little attention has been devoted to verification of the predictive significance of miRNAs in the treatment of NSCLC.
3.1 miRNA as a Noninvasive Predictor for Recurrence and Survival After Surgical Treatment of Lung Cancer
Surgery remains the only potentially curative modality for early-stage NSCLC patients. However, 30% to 55% of patients with NSCLC develop recurrence and die of their disease despite curative resection. While recurrence most commonly occurs at distant sites [93], there has also been a report concerning the underestimation of the frequency of local recurrence [94]. There is an urgent need for the identification of new predictive factors to improve long-term survival rates. Intensive follow-up is also important to reduce lung cancer mortality by the early detection of recurrences after surgery. The clinical value of miRNA expression in resected NSCLC to identify patients at high risk of relapse after surgery is currently being evaluated.
Leidinger et al. [95] followed the plasma levels of miRNAs in five lung cancer patients starting prior to surgery and ending 18 months after surgery, with blood taken at 3-month intervals (eight different time points). In terms of the quantitative changes in miRNA abundance in plasma samples, the fewest miRNAs were detected 6 months after cancer resection, while the most miRNAs at about 2 weeks after cancer resection that probably reflected the extensive tissue trauma and healing processes. A correlation analysis of all miRNAs revealed 17 miRNAs that showed a general decrease over time while 16 miRNAs showed an increase. Importantly, this data clearly contradicts the idea of a general decrease of circulating miRNAs after tumour surgery (a decrease was found for specific miRNAs only) indicating that fluctuating patterns of miRNA levels during the follow-up are patient-specific. miR-486-5p was detected in all five samples before surgery and showed a rather strong increase in expression over time. In a further study, the same group observed a significantly increased number of miRNAs detectable in the plasma of eight NSCLC patients developing metastases with respect to 18 subjects without progressive disease (on average, 316 versus 286 miRNAs, respectively; P = 0.0096) [96].
It is worth noting that downregulated plasma miR-486 levels after surgery have recently been associated with prolonged recurrence-free survival of NSCLC patients [97]. Consistently, in the study of Hu et al. [98], the serum levels of the oncogenic miR-486 and miR-30d were significantly increased while those of the oncosuppressor miRNAs (miR-499 and miR-1) were significantly decreased in the group of NSCLC patients with a shorter overall survival (23.97 months), as compared with patients with longer survival (47.17 months). Also, Le et al. [99] reported that high expressions of miR-21 and miR-30d in preoperative sera were independently correlated with shorter overall survival in a cohort of 82 lung cancer patients (log-rank test for miR-21 and miR-30d: P = 0.0498 and P = 0.0019, respectively). The findings mentioned above imply that changes of selected miRNA levels in plasma or serum in response to surgery can be a promising blood-based marker for predicting local recurrence of tumours and survival in NSCLC patients after surgery. Still, this data needs to be verified in an independent study on a larger cohort of lung cancer patients, and using uniform methodology.
3.2 miRNAs as Predictive Biomarkers in NSCLC Radiotherapy
Radiotherapy, through ionising radiation, aims to destroy tumours by causing the production of free radicals in the neoplastic cell (particularly, but not only, on the DNA) [100]. The natural biological heterogeneity of NSCLC is one of the major factors contributing to the diversity of responses to radiotherapy observed in patients with this particular tumour type. Resistance of lung cancer to ionising radiation may result from chronic tumour hypoxia, disturbed DNA damage response (DDR) pathways, deregulation of pathways responsible for cell survival and cell death via constitutive activation of growth factor receptors (e.g. EGFR), mutations in oncogenes (e.g. KRAS) or tumour suppressors (e.g. PTEN), and the presence of radiation-resistant cancer stem cells with a high regenerative and proliferative potential in hypoxic areas of the tumour [101]. If it becomes possible to determine the degree of tumour sensitivity/resistance to radiotherapy, clinicians could use this treatment modality with greater accuracy by optimising the selection of radiation dose and irradiation site, and by limiting radiation toxicity. Some studies have focused on the regulation of radiosensitivity by miRNAs and its clinical implications for radiotherapy, most of them performing cell culture experiments only (Table 2). In summary, many in vitro studies have shown that sensitivity or resistance to ionising radiations can be regulated by an increase or reduction of specific miRNA levels and the result depends on the cell type, specific miRNA up- or downregulated, and pathway triggered or silenced by downstream effects. Targeting specific miRNAs could enhance radiosensitivity or limit radioresistance, thus producing more relevant clinical effects in response of the cancer patient to radiotherapy.
Only few studies used patient specimen data, defining radiosensitivity and radioresistance miRNA signatures. Wang et al. [110] found 12 differently expressed miRNAs in radiotherapy-sensitive and radiotherapy-resistant NSCLC samples (n = 30). When comparing with resistant patients, five miRNAs (miR-126, miR-let-7a, miR-495, miR-451 and miR-128b) were significantly upregulated and seven miRNAs (miR-130a, miR-106b, miR-19b, miR-22, miR-15b, miR-17-5p and miR-21) were downregulated in the radiotherapy-sensitive group. Overexpression of miR-126 increased the radiosensitivity of lung cancer cells through the PI3K-Akt pathway.
Bi et al. [111] profiled circulating miRNAs in serum samples from 100 unresectable NSCLC patients treated with definitive radiotherapy. The serum miR-885/miR-7 signature was identified as a significant predictor for overall survival in the training set (n = 50), which was validated by the validation set (n = 50, P = 0.02). This signature remained significant (P = 0.04) after adjustment for total tumour volume and Karnofsky performance status, the only two significant clinical factors in a univariate analysis. In the group treated with high-dose radiation (>70 Gy, n = 45), low-risk patients had a significantly longer overall survival than high-risk patients (70.7 vs. 18.8 months, P = 0.007); while in the low-dose radiation group (≤70 Gy, n = 55), no significant association was observed.
Chen et al. [112] identified 14 miRNAs associated with radioresistant genes (miR-153-3p, miR-1-3p, miR-613, miR-372-3p, miR-302e, miR-495-3p, miR-206, miR-520a-3p, miR-328-3p, miR-520b, miR-1297, miR-520d-3p, miR-193a-3p and miR-520e) and five miRNAs associated with radiosensitive genes (let-7c-5p, miR-98-5p, miR-203a-3p, miR-137 and miR-34c-5p) on the basis of miRNA profiles screened from NSCLC cell lines with different radiosensitivities. Next they correlated the expression of candidate miRNAs in the plasma of 54 NSCLC patients with their radiotherapy response to identify circulating radiosensitivity biomarkers in NSCLC. Four miRNAs (miR-98-5p, miR-302e, miR-495-3p and miR-613) demonstrated a higher expression in responders (complete response + partial response, 15 cases) than in non-responders (stable disease + progressive disease, 39 cases). Based on each cut-point, objective response rate was higher in the miR-high group than in the miR-low group. No miRNA showed correlation with survival.
3.3 miRNAs and Resistance to Lung Cancer Chemotherapies
The available studies suggest that miRNAs not only regulate the response to chemotherapy but are also regulated by chemotherapeutic agents. Thus, the use of specific agents directly affecting the activities of specific miRNAs, which may lead to the development of a new cancer treatment strategy, is as important as identifying miRNAs that are useful in monitoring the course of traditional chemotherapy. The most recent reports suggest that a patient’s response to a chemotherapeutic agent is accompanied by changes in the expression of specific miRNAs, which clearly implies that these molecules can be used as predictive biomarkers (Fig. 3) [113].
miRNAs involved in mechanisms of resistance to various chemotherapeutic strategies and TRAIL-based therapy in NSCLC. Some miRNAs act as oncogenes by downregulating the genes involved in pro-apoptotic pathways thus promoting the survival of tumour cells. Therapeutic silencing of those miRNAs could potentially sensitize tumour cells to the drugs. Other miRNas function as tumour suppressors that target the genes promoting tumour cell survival. Induced overexpression of these miRNAs might increase the therapeutic effect that depends on the apoptosis of the tumour cells
3.3.1 Platinum
Cui et al. [114] proposed miR-125b as a potential negative predictive biomarker of the response to cisplatin treatment. In a study investigating 260 patients with advanced NSCLC (stages IIIA–IV), high serum expression of this miRNA was correlated with a poorer treatment response. Furthermore, high serum expression of miR-125b in patients receiving cisplatin-based chemotherapy was associated with shorter survival. Many currently ongoing studies are based on the analysis of miRNAs in NSCLC cell cultures as in vitro models of the mechanisms of sensitivity and resistance of this tumour to chemotherapeutic agents. Gao et al. [115] demonstrated that transfection of A549 lung adenocarcinoma cells (resistant to platinum-based chemotherapy) with synthetic antisense oligonucleotides targeting miR-21 (anti-miR-21) resulted in a considerable increase in the expression of the PTEN tumour suppressor and a decrease in the expression of the anti-apoptotic BCL2. miR-21 expression was also associated with the shorter disease-free survival in platinum-based chemotherapy-resistant patients. In another study, overexpression of miR-513a-3p, induced in the A549 cell line by transfection with miRNA mimics (synthetic oligonucleotide sequences that mimic the body’s endogenous miRNA), promoted cisplatin-dependent apoptosis of cells previously resistant to this chemotherapeutic agent [116]. Interestingly, miR-513a-3p negatively regulated the production of glutathione S-transferase by cancer cells, which was linked to resistance to cisplatin [117, 118]. These results demonstrate that miRNAs may not only be used as markers of the response to cisplatin but also as potential treatment tools that stimulate the sensitivity of lung cancer cells to this chemotherapeutic agent.
3.3.2 Taxanes
Rui et al. [119] found a significant association between the level of miR-200b expression in SPC-A1 lung adenocarcinoma cells and the resistance of these cells to docetaxel. As a result of induced overexpression of miR-200b in tumour cells, decreased resistance to this chemotherapeutic agent was observed, which was explained by suppression of proliferation and augmentation of the apoptotic pathway [120]. In another in vitro study, miR-200b was suggested as a chemosensitivity restorer to docetaxel therapy in lung adenocarcinoma cells by targeting E2F3. This transcription factor is critical for the maintenance of regular cell cycle progression [120]. Also miR-100 has been shown to have an impact on increasing the chemosensitivity of SPC-A1 cancer cells to docetaxel by reducing the expression of PLK1 [121]. The inverse correlation between miR-100 and PLK1 expression was also detected in nude mice SPC-A1/DTX tumour xenografts and clinical lung adenocarcinoma tissues and was proved to be related to the in vivo response to docetaxel. In other study conducted by Cui et al. [122], the restoration of let-7c had an ability to reverse the chemoresistance of docetaxel-resistant lung adenocarcinoma cells owing to direct targeting of BCL-xL and inactivation of Akt phosphorylation both in vitro and in vivo.
A study investigating the expression of EZH2, a gene overexpressed in NSCLC, showed that miR-101 was capable of inhibiting the growth of tumour cells by inducing the apoptotic pathway associated with the therapeutic effects of paclitaxel [123]. To determine the effects of miR-101 in lung cancer cells, the cells were transfected with miRNA mimics. This led to a decrease in the proliferation and invasiveness of the tumour cells by sensitizing the NSCLC cell to paclitaxel, which was partly due to decreased expression of EZH2. Results of in vitro studies on the established lung cancer cell line A549 demonstrate that an important role in the mechanism of resistance to paclitaxel is played by miR-16 and miR-17, whose expression profiles were significantly correlated with the resistance of tumour cells to this chemotherapeutic agent [124, 125]. In another in vitro study, resistance of the same cell line to paclitaxel was significantly associated with miR-135a expression [126]. In both in vitro and in vivo models, researchers observed that inhibition of miR-135a expression led to re-sensitization of previously resistant NSCLC cells to paclitaxel and caused the cells to undergo apoptosis. Expression of miR-135a has also been linked to the activity of the APC gene, which is involved in cancer development. Shen et al. [127] showed that miR-137 also has a potential role in drug resistance of lung cancer cells. After the repression of miR-137 in A549 cells, cell growth, migration and cell survival were increased. Importantly, induced overexpression of miR-137 underlined a tumour suppressive role of this miRNA in chemosensitivity by the inhibition of cell growth and angiogenesis in vivo. In a xenograft mouse model, miR-186 also showed tumour growth inhibitory functions [128]. This miRNA directly targeted MAPT and the chemosensitizing function of miR-186 was partially caused by the induction of the p53-mediated apoptotic pathway.
3.4 miRNAs as Key Modulators of Tumour Immune Response in Lung Cancer Patients
Advances in our understanding of the mechanisms responsible for the regulation of tumour-directed immune response have led to the development of immune checkpoint inhibitors, currently an established therapeutic option for patients with advanced NSCLC. These agents target molecular pathways orchestrated by the programmed cell death protein-1 (PD-1)/programmed cell death ligand-1 (PD-L1) interaction. PD-L1 expression is directly involved in evasion of the immune response by cancer cells [10]. Its binding to the PD-1 receptor on T cells promotes a dual inhibition mechanism: firstly, by inducing apoptosis in antigen-specific T cells and secondly, by simultaneously reducing regulatory T cell (Treg) apoptosis. Thus, via this PD-1/PD-L1 interaction, the tumour is able to induce the anergy of T cells and avoid the recognition by the immune system [129]. Clinical trials have demonstrated significant clinical effects of PD-1 or PD-L1 blockade in advanced NSCLC patients [130]. Currently, there are three checkpoint inhibitors of the PD-1/PD-L1 pathway approved by the Food and Drug Administration (FDA) to treat advanced disease: pembrolizumab (approved for PD-L1-positive NSCLC), nivolumab and atezolizumab (regardless of PD-L1 expression status). Inhibitors targeting other immune checkpoints, such as anti-CTLA-4 antibodies, have not resulted in benefits from single-agent response.
Recently, a molecular link between the evasion of the immune response by lung cancer cells and the miRNA function has been identified. Chen et al. [131] demonstrated that miR-200 suppressed the epithelial to mesenchymal transition (EMT) process by targeting PD-L1 and thus delaying cancer progression in a mouse model. As shown before, miR-200 expression is downregulated in highly metastatic cancer cells [132, 133]. By inducing its expression, a reversed EMT phenotype was induced with abolished invasion and metastasis formation. miR-200 family members (arranged in two genomic cluster: miR-200a/200b/429 and miR-200c/141) directly target EMT-inducing transcription factors such as zinc finger E-box-binding homeobox 1 (ZEB1) [131]. ZEB1 regulates the miR-200 family expression by repressing the transcription of both miR-200 loci. In cancer cells, the miR-200/ZEB1 axis controls the expression of multiple genes involved in migration, invasion and metastatic growth at distant sites. Thus, miR-200 and ZEB1 form a double-negative feedback loop and function as a key regulatory axis of the EMT program. ZEB1 as the transcriptional repressor of miR-200 plays a critical role in the upregulation of PD-L1 expression on tumour cells followed by CD8+ T cell immunosuppression and metastasis.
It has also been observed that miR-200 expression negatively correlates with PD-L1 expression, particularly in tumours with a mesenchymal phenotype. This finding implicates the potential usefulness of miR-200 expression as a predictive biomarker for checkpoint inhibitor therapy in NSCLC. On the basis of evidence presented above, lung cancer patients presenting an adenocarcinoma subtype with upregulated PD-L1, mesenchymal expression pattern and low miR-200 expression would particularly benefit from the treatment with PD-1/PD-L1 inhibitors.
PD-L1 can also be regulated by p53 via the miR-34 family, which directly binds to the 3′UTR of the PD-L1 transcript [134]. NSCLC patients with high miR-34a/p53 and low PD-L1 levels are characterized by higher survival rates than those with low miR-34a/p53 and high PD-L1 levels. These findings have potential clinical application and further studies are needed to confirm the usage of miR-34a/p53 expression as a predictive biomarker for immunotherapy. Novel mechanisms underlying the regulation of tumour immune evasion by specific miRNAs are currently investigated.
3.5 miRNAs as Modulators of TRAIL-Based Therapy
Tumour necrosis factor (TNF)-related apoptosis inducing ligand (TRAIL) is a cytokine and a member of the TNF family that is being tested in clinical trials as a powerful anticancer agent. Although TRAIL had shown clinical efficacy in a subset of NSCLC patients, acquired resistance to this anticancer agent undermines its therapeutic value. The mechanism of this resistance is still not fully understood.
In 2008, Garofalo et al. [85] reported that NSCLC cells overexpressing miR-221/222 were TRAIL-resistant and showed an increase in migration and invasion capabilities. Later on, the same group [135] demonstrated that miR-34a and miR-34c, which are downregulated in NSCLC cell lines, could play a significant role in lung carcinogenesis by modulating the expression of PDGFR-α/β and thereby regulating TRAIL-induced cell death sensitivity. Another miRNA that can be involved in TRAIL resistance is miR-24 [136]. This miRNA directly downregulates the expression of XIAP and induces sensitivity to TRAIL-based therapy in TRAIL-resistant lung cancer cell line. Joshi et al. [137] showed that enforced expression of miR-148a sensitized cells to TRAIL and reduced lung carcinogenesis both in vitro and in vivo through the downregulation of matrix metalloproteinase 15 (MMP15) and Rho-associated kinase 1 (ROCK1). Although miRNAs may lead to re-sensitization of cancer cells to TRAIL therapy, there are also miRNAs which play the opposite role and promote TRAIL resistance. Expression of miR-21, miR-30c and miR-100 in NSCLC has been related to acquired TRAIL resistance [138]. Accordingly, continuous exposure to TRAIL caused acquired resistance to this agent and activated miR-21, miR-30c and miR-100 transcripts.
3.6 miRNAs as Biomarkers for Targeted Therapies in NSCLC
NSCLC is a challenging diagnostic target due to the considerable diversity of neoplastic clones within the tumour and the difficult access to good-quality and informative diagnostic material [139]. As the disease progresses, the profile of molecular markers changes as a result of the genetic alterations within the tumour [140]. miRNAs are associated with tumour progression, suggesting their potential applications for targeted treatment monitoring.
Targeted therapy with tyrosine kinase inhibitors (TKIs) is currently the most common form of personalised treatment of NSCLC. Somatic mutations in exons 18 to 21 of the EGFR gene are the most important molecular predictive marker whose clinical value in the diagnosis of NSCLC has been confirmed [141]. EGFR mutations occur in about 30–50% of Asian and approximately 10–20% of Caucasian NSCLC patients. In contrast to standard chemotherapy, TKIs selectively inhibit tumour cell growth by affecting the intracellular domain of EGFR [142]. Targeted therapy using TKIs, which either bind to EGFR reversibly (erlotinib, gefitinib) or irreversibly (afatinib, osimertinib), is the clinical standard in the personalised treatment of NSCLC for eligible patients. The greatest limitation of the efficacy of reversible EGFR TKIs is the resistance to these agents that is acquired by most patients during treatment, with a median time to progression of 9 months [143].
The aberrant expression of specific miRNAs or miRNA families has serious consequences resulting in abnormal regulation of key components of signalling pathways in tumour cells (Fig. 4). The dual activity of miRNAs (oncogenic or suppressor) in carcinogenesis seems to be an important factor affecting the efficacy of targeted therapies, including EGFR TKIs in lung cancer [144]. Abnormal regulation of gene expression by miRNAs triggers alternative (collateral) signalling pathways or activates downstream signalling mediators, bypassing the pathway blocked by EGFR TKIs. As a result of the feedback between miRNA levels and the activity of the genes targeted by the given miRNA, including oncogenes and tumour suppressors, miRNA expression may change dynamically, indicating the current status of the cell’s genetic activity. Identification of miRNA profiles involved in the mechanisms of the cellular response to EGFR TKIs may open a new direction of research to develop a more effective personalised therapy for patients with NSCLC [145]. It could also significantly increase the efficacy of disease and treatment monitoring.
Non-invasive approaches, usually based on plasma or serum samples, showed great potential for monitoring EGFR-TKI treatment in recent years. Several circulating miRNA signatures are associated with response to EGFR-TKIs in NSCLC. Zhang et al. [146] analysed the expression of 20 miRNAs in the plasma of 105 non-smoking female patients with lung adenocarcinoma (stage I–IV). They found miR-122 to be differently expressed between wild and mutant EGFR carriers (P = 0.018). After adjusting for stage, the associations of miR-16, miR-20b, miR-195, miR-122 and miR-486-3p with EGFR mutation status were evident in advanced stage (P = 0.019, 0.047, 0.041, 0.033, or 0.017, respectively). Plasma levels of miR-195 and miR-122 expression were also associated with overall survival in the patients, especially in those with advanced stage (HR = 0.23, 95% CI 0.07–0.84; and HR = 0.22, 95% CI 0.06–0.77).
Shen et al. [147] reported that the expression levels of serum miR-21 and miR-10b were much higher in 128 radically resected NSCLC patients with EGFR mutation (n = 60) than those without mutation (n = 68). In both univariate and multivariate analyses, gefitinib treatment was associated with a significant improvement in overall survival in patients with reduced miR-21 expression. Thus, miR-21 expression emerged as an independent predictor of the response to EGFR-TKI.
In another study, upregulation of five selected miRNAs (miR-21, miR-122, miR-195, miR-125b, miR-25) correlated significantly with EGFR mutation in both tumour tissues and matched plasma of 150 NSCLC patients (all P < 0.001) [148]. The discriminatory power of predicting EGFR mutation using plasma miRNAs was demonstrated by the AUC (area under the curve) of 0.869 (P < 0.001, 95% CI 0.808–0.930).
Wang et al. [149] studied the role of miRNAs in primary resistance to EGFR-TKIs in advanced NSCLC patients with EGFR mutation. First, they found 153 miRNAs that were differentially expressed between the sensitive and resistant groups in a training cohort of 20 advanced NSCLC patients with EGFR exon 19 deletions treated with first-line EGFR-TKIs. Then, three miRNAs (miR-21, miR-27a and miR-218) were verified to have significantly higher expression (P = 0.011, 0.011, 0.026, respectively) in the resistant group compared to the sensitive group in a validation cohort (n = 34). This is the first study that identified a potential association between miR-21 and primary resistance to EGFR-TKIs in EGFR-mutant patients.
Interestingly, another study showed that miR-21 is involved in acquired resistance of EGFR-TKI in NSCLC [150]. The serum miR-21 expression in 25 advanced NSCLC patients treated with EGFR-TKI was significantly higher at the time of acquiring resistance than at baseline (P < 0.01). Moreover, the authors provided mechanistic evidence based on animal models that miR-21 can mediate EGFR-TKI resistance by downregulating PTEN and PDCD4, and activating the PI3K/Akt pathway. PTEN is a tumour suppressor protein which negatively regulates the PI3K/Akt pathway by converting phosphatidylinositol 3,4,5-triphosphate (PIP3) to phosphatidylinositol 4,5-bisphosphate (PIP2) [151]. Allelic loss or mutation of PTEN is common in many human malignancies and PTEN abnormality-associated Akt activation (PI3K/Akt pathway) has been known to play an important role in chemotherapy resistance.
About 20–44% of treated NSCLC patients acquire resistance to EGFR-TKIs through phenotypic changes of the tumour cells undergoing EMT, possibly as a result of altered epigenetic regulation [152]. It has been shown that EMT-related acquired resistance to EGFR-TKIs in NSCLC is driven by increased ZEB1, which is negatively regulated by miR-200c [153]. Interestingly, clinical results by Li et al. [154] showed that wild-type EGFR patients with high miR-200c expression (n = 26) could benefit more from EGFR-TKIs than those with low miR-200c expression (n = 40), while no such effect was observed in the mutant EGFR subgroup (n = 73). Patients with high miR-200c expression had significantly better clinical outcomes than those with low expression in terms of progression-free survival (5.0 months [95% CI 1.41–8.59] vs. 1.2 months [95% CI 0.89–1.51), P = 0.001), overall survival (9.6 months [95% CI 4.27–14.93] vs. 5.0 months [95% CI 3.90–6.10], P = 0.037), and a numerically higher objective response rate (11.5% vs. 2.5%, P = 0.292). This study implies that miR-200c overexpression might be a potential predictive biomarker for the outcome of EGFR-TKIs in advanced NSCLC patients with wild-type EGFR.
The available literature pertaining to the role of miRNAs in the sensitivity and resistance to molecularly targeted therapies other than EGFR TKIs is very sparse. Currently, it is recommended that newly diagnosed patients with advanced lung adenocarcinoma are tested for EGFR mutations and ALK fusions [155, 156]. Additionally many clinical laboratories are now routinely testing for alterations in genes such as ROS1, RET, MET, BRAF and HER2 which have shown initial promise in tailored cancer treatment.
Chromosomal rearrangements of ALK are present in 3–7% of NSCLC. The resulting ALK fusions, such as EML4-ALK, function as potent oncogenic drivers and lead to a state of oncogene addiction [156]. In the clinic, this phenomenon underlies the marked responsiveness of ALK-positive tumours to crizotinib, a multitargeted tyrosine kinase inhibitor (TKI) of ALK, ROS1 and c-MET [157]. Despite the impressive efficacy of crizotinib in the treatment of ALK-positive lung cancer, acquired resistance eventually develops in the majority of patients [158]. In a recent study, Gao et al. [159] reversed EMT in a crizotinib-resistant NSCLC cell line through overexpression of miR‑200c targeting ZEB1, thus providing further evidence on the role of miRNAs in mediating sensitivity of tumour cells to TKIs.
The MET/hepatocyte growth factor (HGF) pathway has been identified as a potential therapeutic target in multiple solid tumours, including NSCLC [156]. In NSCLC, the most common mechanism for aberrant MET signalling is overexpression of HGF and HGFR. Importantly, MET amplification has been identified as a mechanism for acquired resistance to EGFR tyrosine kinase inhibition in a subset (5–20%) of patients with activating EGFR mutations through ERBB3-dependent activation of the PI3K pathway. There are a number of MET tyrosine kinase inhibitors currently undergoing testing in early phase clinical trials [160]. Although crizotinib is FDA-approved for ALK-translocated NSCLC, it also has in vitro activity against MET.
The miR-34 family consists of three members: miR-34a, miR-34b and miR-34c; all of them negatively regulate the MET gene [161]. All members of the miR-34 family, targeting more than 77 target mRNAs, were shown to suppress tumour growth and metastasis by inhibiting the processes that stimulate cancer development, including cell cycle progression, EMT, metastasis, stemness and by promoting the processes that inhibit carcinogenesis, such as apoptosis [162]. miR-34b/c (and miR-199) contributes to control MET activity by binding to the 3′UTR of MET [163]. The transfection of cells with miR-34b/c precursors showed a significant reduction of MET protein even in cells displaying overexpression and MET gene amplification; furthermore the transfected cells were unable to migrate and to scatter (to break intercellular junctions) in response to HGF. Conversely, the inhibition of endogenous miR-34b/c by use of antagomiRs resulted in increased expression of MET.
The members of the miR-221/222 cluster are among the oncogenic miRNAs that target MET [161]. Garofalo et al. [164] studied miRNAs implicated in EGFR resistance and identified eight miRNAs that are regulated by both EGFR and MET (miRNAs: 221/222, 30b/c, 21, 29a/c, 100). Then they demonstrated that gefitinib treatment triggers programmed cell death through the downregulation of miR-30b/c and miR-221/222 in sensitive NSCLC cells. MET overexpression is able to induce resistance to gefitinib through the upregulation of miR-30b/c and miR-221/222 making the EGFR inhibition alone ineffective. They also showed that this resistance could be overcome using MET inhibitors or anti-miRNA 221/222 and anti-30c which strongly increase gefitinib sensitivity in xenograft mouse models in vivo.
4 miRNAs as Therapeutic Agents in NSCLC Treatment
The pathogenesis of lung cancer is characterised by extensive epigenetic and genetic changes that include mutations, rearrangements, changes in gene expression, and chromosomal amplifications and deletions. Therefore, an ideal targeted therapy should focus on many key coding and/or regulatory sequences involved in the process of carcinogenesis. A significant advantage of using miRNAs as potential targets and/or treatment agents is their ability to regulate the expression of a number of genes and non-coding sequences that are associated with a single pathway or are involved in the regulation of parallel signalling pathways that control tumour development [165]. Compared with treatment based on small interfering RNAs (siRNAs), each of which can only silence one gene, the use of miRNAs as treatment agents seems much more beneficial. For example, multiple genes in the EGFR signalling pathway, including MAPK, STAT3 and AKT2, are all potential targets of miR-124 [166].
Therapies based on miRNAs are being developed with two objectives. The first involves inhibition of specific miRNA molecules if their expression levels are pathologically elevated. The second involves supplementation of those miRNAs whose cellular expression is insufficient or lacking. miRNA activity may be regulated by inhibiting or supplementing a specific miRNA sequence or by modulating the expression of the miRNA coding sequence. In the latter method, the treatment agents are mainly siRNAs and genetically engineered expression vectors that encode small hairpin RNAs. The potential application of each of the proposed strategies depends on the tumour type, tumour stage and, most likely, many other previously unknown clinicopathological factors [167].
Oligonucleotides that act as miRNA antagonists (anti-miRs) are being designed to disrupt the biogenesis of miRNAs, resulting in increased expression of suppressor genes. miRNA mimics, which are also referred to as replacement therapy for miRNAs, are characterised by another mechanism of action. The decrease in the activity of suppressor miRNAs during cancer progression may be compensated for by miRNA mimics that have the same actions as those of the native molecules.
4.1 Preclinical Studies
It is estimated that only about 200 out of approximately 2500 mature human miRNAs registered in miRBase have sufficiently high expression to be feasible targets for mechanistic studies or therapeutic purposes [168]. Recent studies have focused on aberrant expression of the let-7 and miR-34 families in lung tumour tissues when compared to the normal lung tissue, which can indicate the involvement of those miRNAs in lung carcinogenesis [169].
Several studies have shown that downregulated expression of let-7, one of the earliest discovered miRNAs, was associated with poor prognosis in lung cancer patients [170–173]. This association was later proposed to be due to the let‑7‑mediated inhibition of RAS, which is a critical oncogene that is involved in lung cancer development [174]. Induced overexpression of let-7 in lung adenocarcinoma A549 cells with mutated KRAS showed cancer cell growth inhibition and reduction in cell cycle progression by modulation of CDC25A, CDK6 and cyclin D2 [175]. Let-7 suppresses tumour growth in xenograft models of human lung cancer [176, 177]. Mirna Therapeutics is currently developing let-7 as a potential miRNA replacement treatment for cancer [168].
miR-34 is another master tumour suppressor miRNA, the downregulation of which is largely investigated in lung cancer [178, 179]. miR-34 family members (a, b and c) are responsible for cell cycle control, apoptosis and cell senescence via the repression of several targets involved in carcinogenesis like BCL2, MYC and MET genes [180]. Corresponding to the studies conducted by Garofalo et al. [135], an important therapeutic application of miR-34 in lung cancer therapy was further supported by studies on the effects of miR-34 replacement on carcinogenesis in a mouse model [181].
Wiggins et al. [182] demonstrated that the transfection of tumour cells with synthetic miR-34a resulted in tumour growth suppression in a mouse model. The negative influence of this miRNA on the expression of genes determining tumour cell survival (c-MET and BCL2) was also confirmed. Importantly, accumulation of this miRNA in tumour tissues was observed along with low nephro-, cardio- and hepatotoxicity. Similarly, Trang et al. [183] reported a significant antineoplastic effect (tumour mass reduction by 60%) by two miRNA mimics, miR-34a and let-7, administered intravenously as a complex with a neutral lipid emulsion, which was preferentially taken up by lung tissue.
The therapeutic approach presented in another preclinical work on miR-34 showed that this miRNA directly represses the checkpoint molecule PD-L1 (programmed death ligand 1) in a mouse model of non-squamous lung cancer [134]. Therapeutic delivery of MRX34 (miR-34 mimic) promoted the TILs (tumour-infiltrating lymphocytes, CD8+) and reduced CD8 + PD1+ immune cells. This novel mechanism, in which a tumour can evade the immune system, is regulated by the p53/miR-34/PD-L1 axis and may be crucial for developing a new therapeutic approach for lung cancer immunotherapy.
miR-21 is an oncogenic miRNA and overexpressed in various types of human tumours including lung cancer [47]. Aberrant expression of miR-21 is considered to contribute to the malignant phenotype, affect proliferation, apoptosis and metastasis [184]. Mechanistically, overexpression of miR‑21 leads to the suppression of several key tumour suppressor genes, such as PTEN, TPM1 and PDCD4 [168]. Thus, miR‑21 was identified as one of the oncomiRs whose inhibition may have therapeutic benefits.
Seike et al. [185] reported that miRNA microarray data showed higher levels of miR-21 in EGFR-mutant cases, and in vitro analyses using NSCLC cell lines showed that activated EGFR signalling upregulated miR-21 expression. A statistically significant positive correlation was observed between miR-21 expression levels and p-EGFR levels in NSCLC cell lines. Furthermore, treatment with the EGFR-TKI inhibited miR-21 expression in two NSCLC cell lines with elevated p-EGFR, providing a mechanistic link between an activated EGFR signalling pathway and the aberrant upregulation of miR-21.
Recently, Microlin Bio, Inc. announced positive results from a preclinical lung cancer study using its lead anti-miRNA candidate in a murine lung cancer model [186]. Anti-miR-21 (AM-21) was delivered using the company’s novel QTsome® (QT) delivery platform, which is composed of a combination of two cationic lipids with tertiary and quaternary amine head groups, respectively. The company claimed that animals in the 1 mg/kg IV QT/AM-21 treated group displayed significant tumour regression or no tumour growth while untreated animals exhibited rapid tumour growth. Median survival was prolonged following treatment with QT/AM-21 from 21 days in the untreated group to 33 days in the 1 mg/kg QT/AM-21 treatment group. Moreover, there were no obvious signs of treatment-based toxicity. A summary of preclinical studies for miRNA-based treatment is given in Table 3.
4.2 Clinical Studies
On the basis of the promising results of preclinical studies, the miR-34 mimic was the first therapeutic miRNA agent to enter clinical trials in phase I testing in patients with primary liver cancer or metastatic cancer that has spread to the liver [179]. The leading therapeutic, MRX34, was a lipid-formulated miR-34 mimic under development by Mirna Therapeutics [191]. A study enrolled patients with unresectable primary liver cancer or advanced metastatic cancer (e.g. NSCLC, SCLC) with or without liver involvement or haematologic malignancies. MRX34 was given as intravenous liposomal injection. In September 2016, the clinical trial was terminated because of multiple immune-related severe adverse events (SAEs) observed in patients dosed with MRX34 over the course of the trial [192]. To date, there are no other registered clinical trials investigating the therapeutic use of miRNAs in lung cancer.
4.3 Problems to Overcome
Therapies based on miRNAs have encountered a number of obstacles. First, the multitargeted action of miRNAs, which might potentate the efficacy of miRNA-based drugs, also carries the risk of off-target effects resulting in frequently occurring and severe adverse events in other organs [193]. Additionally, while miRNA mimicry increases the levels of a miRNA that is lost during disease progression, systemic delivery of such miRNA mimics can also result in uptake by non-target tissues that normally do not express the miRNA of interest, resulting in potential off-target effects [194]. Thus, the development of an effective and safe system to deliver miRNAs into tumour cells or their environment is going to be important to prevent unwanted side effects of this therapeutic approach.
One of the methods of introducing exogenous miRNA molecules into tumour tissues involves transfection using a viral vector. Transfection of synthetic let-7 molecules into murine tumour cells caused the tumour mass to decrease [187]. Unfortunately, using modified viral vectors in therapy is still considered controversial by the medical community because of the risk of viral DNA integration into undesirable locations in the host genome, with the resulting transformation of healthy somatic and germ line cells in the body. Also, the level of expression of the exogenous gene is usually too low to mount a full treatment effect [195]. Another method of transferring specific miRNA into the lung tumour tissue is a cationic lipid-based miRNA delivery system, which had good efficiency both in vitro and in vivo, as was reported by Wu et al. [189, 196]. Most challenges to overcome in liposome-based therapies are related to toxicity due to charged liposomes, inability to escape the immune system, and low stability [197, 198]. Recently, as an alternative to the liposomal or nanoparticle-based methods of miRNA delivery, the EDV™ nanocells (EDVs) have been employed [199, 200]. This delivery system developed by EnGeneIC Ltd (Sydney, Australia) comprises nonviable minicells 400 ± 20 nm in diameter, produced by de-repressing polar sites of cell division in bacteria [201]. Once loaded, EDVs are coated with a bispecific antibody (BsAB), where one arm is available for binding to a receptor expressed on the surface of cancer cells. Following intravenous administration, EDVs tend to accumulate in the tumour vasculature then bind to overexpressed target receptors on tumour cells and are thought to become involved in the endocytosis process. EDVs not only deliver toxic payloads to tumours, they stimulate the adaptive immune system to augment the antitumour response.
The low in vitro stability of RNA is another problem. Unmodified RNA molecules are degraded by RNases and are excreted in the urine [202]. When RNA molecules were administered via the tail vein in mice, they were removed from the bloodstream within less than 30 min [183]. Effective in vivo silencing of specific miRNAs which expression increased pathologically requires the use of anti-miRs characterised by improved biological stability, highly specific miRNA binding and optimal pharmacokinetic properties, which are achieved by chemical modification of the RNA molecule [194]. In the case of miRNA mimics, in vivo stability is ensured by the double-stranded structure of these molecules and by protection of the leading strand, which is identical to the mimicked miRNA, by the addition of fluorine to cytosine and uracil nucleotides (2′-FC and 2’-FU, respectively) [203]. This modification does not interfere with the recognition of miRNA mimics by the native RISC complex of the recipient cell.
5 Summary
Nearly two decades have elapsed since the discovery of miRNAs, and many studies have since focused on the possibility of using these small regulatory molecules as biomarkers for various cancer types, including NSCLC. As a result of the involvement of miRNAs in carcinogenesis during all stages, these molecules could be used not only as specific biomarkers of cancer (diagnostic biomarkers) but also as dynamic markers of the tumour status before (prognostic biomarkers) and during treatment (predictive biomarkers). Extracellular miRNAs, which are secreted in a stable form by tumour cells into the blood and other body fluids during the early stages of lung cancer development, seem to be particularly useful in clinical practice. Their principal advantage is the ease by which their quantitative and qualitative changes can be monitored in real time and at every stage of the disease, without causing patients discomfort by invasive collection of specimens for testing.
Tremendous technological progress has been made over the past decade in the methodology of miRNA detection and identification, particularly in high-throughput techniques, such as NGS. This has marked a new era of translational research enabling simultaneous objective analysis of multiple miRNAs in the context of identifying epigenetic profiles in lung cancer. Bioinformatics tools have played an equally important role, enabling the analysis of raw data, the classification of miRNA families and the prediction of potential target genes.
In light of the existing expectations regarding the effective use of miRNAs in the diagnosis and treatment of lung cancer, there is an urgent need to continue basic research investigating the biogenesis and functions of miRNA, as well as translational research to verify the possibilities of their practical applications as biomarkers in the clinic. Although these small, non-coding RNA particles clearly possess many desirable properties to become “ideal” biomarkers for lung cancer diagnosis and treatment effectiveness monitoring, only few of all the known miRNAs will actually pass strict validation processes to reach clinical approval.
Also, the concept of using miRNAs in the personalised therapy of lung cancer, which involves modulating the expression of specific miRNAs that regulate the activity of genes involved in carcinogenesis, seems realistic providing that several problems of key importance are solved. First, effective delivery of miRNA modulators to the cell type or tissue of interest should be developed. Next, better understanding the side effects and potential off-target effects under physiologically normal and diseased conditions in vivo is essential in order to bring miRNA therapeutics closer to the clinic. Furthermore, extensive preclinical studies are required to determine the optimal level of inhibition for a given miRNA target. Finally, well-designed clinical trials need to be conducted to test which patients are more likely to benefit from miRNA-based therapy. As the number of studies evaluating the applicability of miRNA for lung cancer diagnosis, prognosis and treatment is increasing continuously, one can expect the true potential of miRNAs as either biomarkers and/or therapeutics to come to light shortly in coming years.
References
Torre LA, Siegel RL, Jemal A. Lung cancer statistics. Adv Exp Med Biol. 2016;893:1–19. doi:10.1007/978-3-319-24223-1_1.
Howlader N, Noone AM, Krapcho M, et al. SEER Cancer Statistics Review 1975–2013. In: National Cancer Institute. 2015. http://seer.cancer.gov/csr/1975_2013/. Accessed Based on November 2015 SEER data submission, posted to the SEER web site, April 2016 accessed: 2.01.2016.
Fabbri M. MicroRNAs and cancer: towards a personalized medicine. Curr Mol Med. 2013;13(5):751–6.
Saumet A, Mathelier A, Lecellier CH. The potential of microRNAs in personalized medicine against cancers. Biomed Res Int. 2014;2014:642916. doi:10.1155/2014/642916.
National Lung Screening Trial Research Team, Aberle DR, Adams AM, et al. Reduced lung-cancer mortality with low-dose computed tomographic screening. N Engl J Med. 2011;365(5):395–409. doi:10.1056/NEJMoa1102873.
Sozzi G, Boeri M, Rossi M, et al. Clinical utility of a plasma-based miRNA signature classifier within computed tomography lung cancer screening: a correlative MILD Trial Study. J Clin Oncol. 2014;32(8):768–73. doi:10.1200/JCO.2013.50.4357.
Montani F, Marzi MJ, Dezi F, et al. miR-Test: a blood test for lung cancer early detection. J Natl Cancer Inst. 2015;107(6):djv063. doi:10.1093/jnci/djv063.
Chan BA, Hughes BG. Targeted therapy for non-small cell lung cancer: current standards and the promise of the future. Translat Lung Cancer Res. 2015;4(1):36–54. doi:10.3978/j.issn.2218-6751.2014.05.01.
Sin TK, Wang F, Meng F, et al. Implications of MicroRNAs in the treatment of gefitinib-resistant non-small cell lung cancer. Int J Mol Sci. 2016;17(2):237. doi:10.3390/ijms17020237.
Schvartsman G, Ferrarotto R, Massarelli E. Checkpoint inhibitors in lung cancer: latest developments and clinical potential. Ther Adv Med Oncol. 2016;8(6):460–73. doi:10.1177/1758834016661164.
Paladini L, Fabris L, Bottai G, Raschioni C, Calin GA, Santarpia L. Targeting microRNAs as key modulators of tumor immune response. J Exp Clin Cancer Res. 2016;35(1):103. doi:10.1186/s13046-016-0375-2.
Gebert LFR, Rebhan MAE, Crivelli SEM, Denzler R, Stoffel M, Hall J. Miravirsen (SPC3649) can inhibit the biogenesis of miR-122. Nucleic Acids Research. 2014;42(1):609-621. doi:10.1093/nar/gkt852.
Baek D, Villen J, Shin C, Camargo FD, Gygi SP, Bartel DP. The impact of microRNAs on protein output. Nature. 2008;455(7209):64–71. doi:10.1038/nature07242.
Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116(2):281–97.
Turchinovich A, Weiz L, Burwinkel B. Extracellular miRNAs: the mystery of their origin and function. Trends Biochem Sci. 2012;37(11):460–5. doi:10.1016/j.tibs.2012.08.003.
Chou CH, Chang NW, Shrestha S, et al. miRTarBase 2016: updates to the experimentally validated miRNA-target interactions database. Nucleic Acids Res. 2016;44(D1):D239–47. doi:10.1093/nar/gkv1258.
Chang S, Johnston Jr RJ, Frokjaer-Jensen C, Lockery S, Hobert O. MicroRNAs act sequentially and asymmetrically to control chemosensory laterality in the nematode. Nature. 2004;430(7001):785–9. doi:10.1038/nature02752.
Lim LP, Glasner ME, Yekta S, Burge CB, Bartel DP. Vertebrate microRNA genes. Science. 2003;299(5612):1540. doi:10.1126/science.1080372.
Berezikov E, Guryev V, van de Belt J, Wienholds E, Plasterk RH, Cuppen E. Phylogenetic shadowing and computational identification of human microRNA genes. Cell. 2005;120(1):21–4. doi:10.1016/j.cell.2004.12.031.
Giovannetti E, Erozenci A, Smit J, Danesi R, Peters GJ. Molecular mechanisms underlying the role of microRNAs (miRNAs) in anticancer drug resistance and implications for clinical practice. Crit Rev Oncol Hematol. 2012;81(2):103–22. doi:10.1016/j.critrevonc.2011.03.010.
Macfarlane LA, Murphy PR. MicroRNA: biogenesis, function and role in cancer. Curr Genomics. 2010;11(7):537–61. doi:10.2174/138920210793175895.
Place RF, Li LC, Pookot D, Noonan EJ, Dahiya R. MicroRNA-373 induces expression of genes with complementary promoter sequences. Proc Natl Acad Sci U S A. 2008;105(5):1608–13. doi:10.1073/pnas.0707594105.
Ketting RF, Fischer SE, Bernstein E, Sijen T, Hannon GJ, Plasterk RH. Dicer functions in RNA interference and in synthesis of small RNA involved in developmental timing in C. elegans. Genes Dev. 2001;15(20):2654–9. doi:10.1101/gad.927801.
Sun T, Kalionis B, Lv G, Xia S, Gao W. Role of exosomal noncoding RNAs in lung carcinogenesis. Biomed Res Int. 2015;2015:125807. doi:10.1155/2015/125807.
Lee Y, Ahn C, Han J, et al. The nuclear RNase III Drosha initiates microRNA processing. Nature. 2003;425(6956):415–9. doi:10.1038/nature01957.
Basyuk E, Suavet F, Doglio A, Bordonne R, Bertrand E. Human let-7 stem-loop precursors harbor features of RNase III cleavage products. Nucleic Acids Res. 2003;31(22):6593–7.
Zeng Y, Cullen BR. Sequence requirements for micro RNA processing and function in human cells. RNA. 2003;9(1):112–23.
Cullen BR. Transcription and processing of human microRNA precursors. Mol Cell. 2004;16(6):861–5. doi:10.1016/j.molcel.2004.12.002.
Gregory RI, Yan KP, Amuthan G, et al. The microprocessor complex mediates the genesis of microRNAs. Nature. 2004;432(7014):235–40. doi:10.1038/nature03120.
Lee YS, Nakahara K, Pham JW, et al. Distinct roles for drosophila dicer-1 and dicer-2 in the siRNA/miRNA silencing pathways. Cell. 2004;117(1):69–81.
Cai XZ, Hagedorn CH, Cullen BR. Human microRNAs are processed from capped, polyadenylated transcripts that can also function as mRNAs. RNA. 2004;10(12):1957–66. doi:10.1261/rna.7135204.
Denli AM, Tops BB, Plasterk RH, Ketting RF, Hannon GJ. Processing of primary microRNAs by the microprocessor complex. Nature. 2004;432(7014):231–5. doi:10.1038/nature03049.
Yi R, Qin Y, Macara IG, Cullen BR. Exportin-5 mediates the nuclear export of pre-microRNAs and short hairpin RNAs. Genes Dev. 2003;17(24):3011–6. doi:10.1101/gad.1158803.
Hutvagner G, McLachlan J, Pasquinelli AE, Balint E, Tuschl T, Zamore PD. A cellular function for the RNA-interference enzyme Dicer in the maturation of the let-7 small temporal RNA. Science. 2001;293(5531):834–8. doi:10.1126/science.1062961.
Bohnsack MT, Czaplinski K, Gorlich D. Exportin 5 is a RanGTP-dependent dsRNA-binding protein that mediates nuclear export of pre-miRNAs. RNA. 2004;10(2):185–91. doi:10.1261/rna.5167604.
Lee Y, Jeon K, Lee JT, Kim S, Kim VN. MicroRNA maturation: stepwise processing and subcellular localization. EMBO J. 2002;21(17):4663–70.
Isik M, Korswagen HC, Berezikov E. Expression patterns of intronic microRNAs in Caenorhabditis elegans. Silence. 2010;1(1):5. doi:10.1186/1758-907X-1-5.
Lee Y, Kim M, Han J, et al. MicroRNA genes are transcribed by RNA polymerase II. EMBO J. 2004;23(20):4051–60. doi:10.1038/sj.emboj.7600385.
Shomron N, Levy C. MicroRNA-biogenesis and Pre-mRNA splicing crosstalk. J Biomed Biotechnol. 2009;2009:594678. doi:10.1155/2009/594678.
Liu C, Zhang F, Li T, et al. MirSNP, a database of polymorphisms altering miRNA target sites, identifies miRNA-related SNPs in GWAS SNPs and eQTLs. BMC Genomics. 2012;13:661. doi:10.1186/1471-2164-13-661.
de la Chapelle A, Jazdzewski K. MicroRNAs in thyroid cancer. J Clin Endocrinol Metab. 2011;96(11):3326–36. doi:10.1210/jc.2011-1004.
Hu Z, Chen J, Tian T, et al. Genetic variants of miRNA sequences and non-small cell lung cancer survival. J Clin Invest. 2008;118(7):2600–8. doi:10.1172/JCI34934.
Tian T, Shu Y, Chen J, et al. A functional genetic variant in microRNA-196a2 is associated with increased susceptibility of lung cancer in Chinese. Cancer Epidemiol Biomark Prev. 2009;18(4):1183–7. doi:10.1158/1055-9965.EPI-08-0814.
Yuan Z, Zeng X, Yang D, Wang W, Liu Z. Effects of common polymorphism rs11614913 in Hsa-miR-196a2 on lung cancer risk. PLoS One. 2013;8(4):e61047. doi:10.1371/journal.pone.0061047.
Czubak K, Lewandowska MA, Klonowska K, et al. High copy number variation of cancer-related microRNA genes and frequent amplification of DICER1 and DROSHA in lung cancer. Oncotarget. 2015;6(27):23399–416. doi:10.18632/oncotarget.4351.
Croce CM. Causes and consequences of microRNA dysregulation in cancer. Nat Rev Genet. 2009;10(10):704–14. doi:10.1038/nrg2634.
Volinia S, Calin GA, Liu CG, et al. A microRNA expression signature of human solid tumors defines cancer gene targets. Proc Natl Acad Sci U S A. 2006;103(7):2257–61. doi:10.1073/pnas.0510565103.
Lu J, Getz G, Miska EA, et al. MicroRNA expression profiles classify human cancers. Nature. 2005;435(7043):834–8. doi:10.1038/nature03702.
Chen X, Ba Y, Ma L, et al. Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res. 2008;18(10):997–1006. doi:10.1038/cr.2008.282.
Gilad S, Meiri E, Yogev Y, et al. Serum microRNAs are promising novel biomarkers. PLoS One. 2008;3(9):e3148. doi:10.1371/journal.pone.0003148.
Mitchell PS, Parkin RK, Kroh EM, et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci U S A. 2008;105(30):10513–8. doi:10.1073/pnas.0804549105.
Park NJ, Zhou H, Elashoff D, et al. Salivary microRNA: discovery, characterization, and clinical utility for oral cancer detection. Clin Cancer Res. 2009;15(17):5473–7. doi:10.1158/1078-0432.CCR-09-0736.
Hanke M, Hoefig K, Merz H, et al. A robust methodology to study urine microRNA as tumor marker: microRNA-126 and microRNA-182 are related to urinary bladder cancer. Urol Oncol Semin Ori. 2010;28(6):655–61. doi:10.1016/j.urolonc.2009.01.027.
Kosaka N, Izumi H, Sekine K, Ochiya T. microRNA as a new immune-regulatory agent in breast milk. Silence. 2010;1:7.
Cogswell JP, Ward J, Taylor IA, et al. Identification of miRNA changes in Alzheimer’s disease brain and CSF yields putative biomarkers and insights into disease pathways. J Alzheimers Dis. 2008;14(1):27–41.
Hanson EK, Lubenow H, Ballantyne J. Identification of forensically relevant body fluids using a panel of differentially expressed microRNAs. Anal Biochem. 2009;387(2):303–14. doi:10.1016/j.ab.2009.01.037.
Rodriguez M, Silva J, Lopez-Alfonso A, et al. Different exosome cargo from plasma/bronchoalveolar lavage in non-small-cell lung cancer. Genes Chromosom Cancer. 2014;53(9):713–24. doi:10.1002/gcc.22181.
Molina-Pinelo S, Pastor MD, Suarez R, et al. MicroRNA clusters: dysregulation in lung adenocarcinoma and COPD. Eur Respir J. 2014;43(6):1740–9. doi:10.1183/09031936.00091513.
Wang T, Lv M, Shen S, et al. Cell-free microRNA expression profiles in malignant effusion associated with patient survival in non-small cell lung cancer. PLoS One. 2012;7(8):e43268. doi:10.1371/journal.pone.0043268.
Xie L, Chen X, Wang L, et al. Cell-free miRNAs may indicate diagnosis and docetaxel sensitivity of tumor cells in malignant effusions. BMC Cancer. 2010;10:591. doi:10.1186/1471-2407-10-591.
Zen K, Zhang CY. Circulating microRNAs: a novel class of biomarkers to diagnose and monitor human cancers. Med Res Rev. 2012;32(2):326–48. doi:10.1002/med.20215.
Schwarzenbach H, Nishida N, Calin GA, Pantel K. Clinical relevance of circulating cell-free microRNAs in cancer. Nat Rev Clin Oncol. 2014;11(3):145–56. doi:10.1038/nrclinonc.2014.5.
Turchinovich A, Weiz L, Langheinz A, Burwinkel B. Characterization of extracellular circulating microRNA. Nucleic Acids Res. 2011;39(16):7223–33. doi:10.1093/nar/gkr254.
Weber JA, Baxter DH, Zhang S, et al. The microRNA spectrum in 12 body fluids. Clin Chem. 2010;56(11):1733–41. doi:10.1373/clinchem.2010.147405.
Akat KM, Moore-McGriff D, Morozov P, et al. Comparative RNA-sequencing analysis of myocardial and circulating small RNAs in human heart failure and their utility as biomarkers. Proc Natl Acad Sci U S A. 2014;111(30):11151–6. doi:10.1073/pnas.1401724111.
Hayes J, Peruzzi PP, Lawler S. MicroRNAs in cancer: biomarkers, functions and therapy. Trends Mol Med. 2014;20(8):460–9. doi:10.1016/j.molmed.2014.06.005.
Rabinowits G, Gercel-Taylor C, Day JM, Taylor DD, Kloecker GH. Exosomal microRNA: a diagnostic marker for lung cancer. Clin Lung Cancer. 2009;10(1):42–6. doi:10.3816/CLC.2009.n.006.
Hannafon BN, Ding W-Q. Intercellular communication by exosome-derived microRNAs in cancer. Int J Mol Sci. 2013;14(7):14240–69. doi:10.3390/ijms140714240.
Frydrychowicz M, Kolecka-Bednarczyk A, Madejczyk M, Yasar S, Dworacki G. Exosomes - structure, biogenesis and biological role in non-small-cell lung cancer. Scand J Immunol. 2015;81(1):2–10. doi:10.1111/sji.12247.
Ardekani AM, Naeini MM. The role of MicroRNAs in human diseases. Avicenna J Med Biotechnol. 2010;2(4):161–79.
Esposito A, Criscitiello C, Locatelli M, Milano M, Curigliano G. Liquid biopsies for solid tumors: understanding tumor heterogeneity and real time monitoring of early resistance to targeted therapies. Pharmacol Ther. 2016;157:120–4. doi:10.1016/j.pharmthera.2015.11.007.
Zandberga E, Kozirovskis V, Abols A, Andrejeva D, Purkalne G, Line A. Cell-free microRNAs as diagnostic, prognostic, and predictive biomarkers for lung cancer. Genes Chromosom Cancer. 2013;52(4):356–69. doi:10.1002/gcc.22032.
Kroh EM, Parkin RK, Mitchell PS, Tewari M. Analysis of circulating microRNA biomarkers in plasma and serum using quantitative reverse transcription-PCR (qRT-PCR). Methods. 2010;50(4):298–301. doi:10.1016/j.ymeth.2010.01.032.
Zampetaki A, Mayr M. Analytical challenges and technical limitations in assessing circulating MiRNAs. Thromb Haemost. 2012;108(10):592–8. doi:10.1160/TH12-02-0097.
Schwarzenbach H, da Silva AM, Calin G, Pantel K. Data normalization strategies for MicroRNA quantification. Clin Chem. 2015;61(11):1333–42. doi:10.1373/clinchem.2015.239459.
Tiberio P, Callari M, Angeloni V, Daidone MG, Appierto V. Challenges in using circulating miRNAs as cancer biomarkers. Biomed Res Int. 2015;2015:10.
Ferdin J, Kunej T, Calin GA. Non-coding RNAs: identification of cancer-associated microRNAs by gene profiling. Technol Cancer Res Treat. 2010;9(2):123–38.
Calin GA, Sevignani C, Dumitru CD, et al. Human microRNA genes are frequently located at fragile sites and genomic regions involved in cancers. Proc Natl Acad Sci U S A. 2004;101(9):2999–3004. doi:10.1073/pnas.0307323101.
Zhang BH, Pan XP, Cobb GP, Anderson TA. microRNAs as oncogenes and tumor suppressors. Dev Biol. 2007;302(1):1–12. doi:10.1016/j.ydbio.2006.08.028.
Hirsch FR, Franklin WA, Gazdar AF, Bunn Jr PA. Early detection of lung cancer: clinical perspectives of recent advances in biology and radiology. Clin Cancer Res. 2001;7(1):5–22.
Esquela-Kerscher A, Slack FJ. Oncomirs - microRNAs with a role in cancer. Nat Rev Cancer. 2006;6(4):259–69. doi:10.1038/nrc1840.
Yu SL, Chen HY, Chang GC, et al. MicroRNA signature predicts survival and relapse in lung cancer. Cancer Cell. 2008;13(1):48–57. doi:10.1016/j.ccr.2007.12.008.
Dvinge H, Git A, Graf S, et al. The shaping and functional consequences of the microRNA landscape in breast cancer. Nature. 2013;497(7449):378–82. doi:10.1038/nature12108.
Nasser MW, Datta J, Nuovo G, et al. Down-regulation of micro-RNA-1 (miR-1) in lung cancer. Suppression of tumorigenic property of lung cancer cells and their sensitization to doxorubicin-induced apoptosis by miR-1. J Biol Chem. 2008;283(48):33394–405. doi:10.1074/jbc.M804788200.
Garofalo M, Quintavalle C, Di Leva G, et al. MicroRNA signatures of TRAIL resistance in human non-small cell lung cancer. Oncogene. 2008;27(27):3845–55. doi:10.1038/onc.2008.6.
Hayashita Y, Osada H, Tatematsu Y, et al. A polycistronic microRNA cluster, miR-17-92, is overexpressed in human lung cancers and enhances cell proliferation. Cancer Res. 2005;65(21):9628–32. doi:10.1158/0008-5472.CAN-05-2352.
He W-J, Li W-H, Jiang B, Wang Y-F, Xia Y-X, Wang L. MicroRNAs level as an initial screening method for early-stage lung cancer: a bivariate diagnostic random-effects meta-analysis. Int J Clin Exp Med. 2015;8(8):12317–26.
Wang H, Wu S, Zhao L, Zhao J, Liu J, Wang Z. Clinical use of microRNAs as potential non-invasive biomarkers for detecting non-small cell lung cancer: a meta-analysis. Respirology. 2015;20(1):56–65. doi:10.1111/resp.12444.
Boeri M, Verri C, Conte D, et al. MicroRNA signatures in tissues and plasma predict development and prognosis of computed tomography detected lung cancer. Proc Natl Acad Sci U S A. 2011;108(9):3713–8. doi:10.1073/pnas.1100048108.
Hennessey PT, Sanford T, Choudhary A, et al. Serum microRNA biomarkers for detection of non-small cell lung cancer. PLoS One. 2012;7(2):e32307. doi:10.1371/journal.pone.0032307.
Solomides CC, Evans BJ, Navenot JM, Vadigepalli R, Peiper SC, Wang ZX. MicroRNA profiling in lung cancer reveals new molecular markers for diagnosis. Acta Cytol. 2012;56(6):645–54. doi:10.1159/000343473.
Cazzoli R, Buttitta F, Di Nicola M, Malatesta S, Marchetti A, Pass HI. MicroRNAs derived from circulating exosomes as non-invasive biomarkers for screening and diagnose lung cancer. J Thorac Oncol. 2013;8(9):1156–62. doi:10.1097/JTO.0b013e318299ac32.
Uramoto H, Tanaka F. Recurrence after surgery in patients with NSCLC. Translat Lung Cancer Res. 2014;3(4):242–9. doi:10.3978/j.issn.2218-6751.2013.12.05.
Boyd JA, Hubbs JL, Kim DW, Hollis D, Marks LB, Kelsey CR. Timing of local and distant failure in resected lung cancer: implications for reported rates of local failure. J Thorac Oncol. 2010;5(2):211–4. doi:10.1097/JTO.0b013e3181c20080.
Leidinger P, Keller A, Backes C, Huwer H, Meese E. MicroRNA expression changes after lung cancer resection: a follow-up study. RNA Biol. 2012;9(6):900–10. doi:10.4161/rna.20107.
Leidinger P, Galata V, Backes C, et al. Longitudinal study on circulating miRNAs in patients after lung cancer resection. Oncotarget. 2015;6(18):16674–85. doi:10.18632/oncotarget.4322.
Li W, Wang Y, Zhang Q, et al. MicroRNA-486 as a biomarker for early diagnosis and recurrence of non-small cell lung cancer. PLoS One. 2015;10(8):e0134220. doi:10.1371/journal.pone.0134220.
Hu Z, Chen X, Zhao Y, et al. Serum microRNA signatures identified in a genome-wide serum microRNA expression profiling predict survival of non-small-cell lung cancer. J Clin Oncol. 2010;28(10):1721–6. doi:10.1200/JCO.2009.24.9342.
Le HB, Zhu WY, Chen DD, et al. Evaluation of dynamic change of serum miR-21 and miR-24 in pre- and post-operative lung carcinoma patients. Med Oncol. 2012;29(5):3190–7. doi:10.1007/s12032-012-0303-z.
Cellini F, Morganti AG, Genovesi D, Silvestris N, Valentini V. Role of microRNA in response to ionizing radiations: evidences and potential impact on clinical practice for radiotherapy. Molecules. 2014;19(4):5379–401. doi:10.3390/molecules19045379.
Begg AC, Stewart FA, Vens C. Strategies to improve radiotherapy with targeted drugs. Nat Rev Cancer. 2011;11(4):239–53. doi:10.1038/nrc3007.
Weidhaas JB, Babar I, Nallur SM, et al. MicroRNAs as potential agents to alter resistance to cytotoxic anticancer therapy. Cancer Res. 2007;67(23):11111–6.
Lee KM, Choi EJ, Kim IA. microRNA-7 increases radiosensitivity of human cancer cells with activated EGFR-associated signaling. Radiother Oncol. 2011;101(1):171–6. doi:10.1016/j.radonc.2011.05.050.
Duan W, Xu Y, Dong Y, Cao L, Tong J, Zhou X. Ectopic expression of miR-34a enhances radiosensitivity of non-small cell lung cancer cells, partly by suppressing the LyGDI signaling pathway. J Radiat Res. 2013;54(4):611–9. doi:10.1093/jrr/rrs136.
Cortez MA, Valdecanas D, Niknam S, et al. In vivo delivery of miR-34a sensitizes lung tumors to radiation through RAD51 regulation. Mol Ther Nucleic Acids. 2015;4:e270. doi:10.1038/mtna.2015.47.
Salim H, Akbar NS, Zong D, et al. miRNA-214 modulates radiotherapy response of non-small cell lung cancer cells through regulation of p38MAPK, apoptosis and senescence. Br J Cancer. 2012;107(8):1361–73. doi:10.1038/bjc.2012.382.
Ma W, Ma CN, Zhou NN, Li XD, Zhang YJ. Up-regulation of miR-328-3p sensitizes non-small cell lung cancer to radiotherapy. Sci Rep. 2016;6:31651. doi:10.1038/srep31651.
Tian F, Han Y, Yan X, et al. Upregulation of microrna-451 increases the sensitivity of A549 cells to radiotherapy through enhancement of apoptosis. Thorac Cancer. 2016;7(2):226–31. doi:10.1111/1759-7714.12318.
Liu YJ, Lin YF, Chen YF, Luo EC, Sher YP, et al. (2013) MicroRNA-449a Enhances Radiosensitivity in CL1-0 Lung Adenocarcinoma Cells. PLOS ONE 8(4): e62383. doi: 10.1371/journal.pone.0062383.
Wang XC, Du LQ, Tian LL, et al. Expression and function of miRNA in postoperative radiotherapy sensitive and resistant patients of non-small cell lung cancer. Lung Cancer. 2011;72(1):92–9. doi:10.1016/j.lungcan.2010.07.014.
Bi N, Schipper M, Stanton P, Wang W, Kong F-M. Serum miRNA signature to identify a patient’s resistance to high-dose radiation therapy for unresectable non-small cell lung cancer. J Clin Oncol. 2013;31(Suppl:abstr 7580).
Chen X, Xu Y, Liao X, et al. Plasma miRNAs in predicting radiosensitivity in non-small cell lung cancer. Tumour Biol. 2016;37(9):11927–36. doi:10.1007/s13277-016-5052-8.
Ponomaryova AA, Morozkin ES, Rykova EY, et al. Dynamic changes in circulating miRNA levels in response to antitumor therapy of lung cancer. Exp Lung Res. 2016;42(2):95–102. doi:10.3109/01902148.2016.1155245.
Cui E-H, Li H-J, Hua F, et al. Serum microRNA 125b as a diagnostic or prognostic biomarker for advanced NSCLC patients receiving cisplatin-based chemotherapy. Acta Pharmacol Sin. 2013;34(2):309–13. doi:10.1038/aps.2012.125.
Gao W, Lu X, Liu L, Xu J, Feng D, Shu Y. MiRNA-21: a biomarker predictive for platinum-based adjuvant chemotherapy response in patients with non-small cell lung cancer. Cancer Biol Ther. 2012;13(5):330–40. doi:10.4161/cbt.19073.
Zhang X, Zhu J, Xing R, et al. miR-513a-3p sensitizes human lung adenocarcinoma cells to chemotherapy by targeting GSTP1. Lung Cancer. 2012;77(3):488–94. doi:10.1016/j.lungcan.2012.05.107.
Ishii T, Fujishiro M, Masuda M, Teramoto S, Matsuse T. A methylated oligonucleotide induced methylation of GSTP1 promoter and suppressed its expression in A549 lung adenocarcinoma cells. Cancer Lett. 2004;212(2):211–23. doi:10.1016/j.canlet.2004.03.001.
Wang J, Zhang J, Zhang L, et al. Expression of P-gp, MRP, LRP, GST-pi and TopoIIalpha and intrinsic resistance in human lung cancer cell lines. Oncol Rep. 2011;26(5):1081–9. doi:10.3892/or.2011.1405.
Rui W, Bing F, Hai-Zhu S, Wei D, Long-Bang C. Identification of microRNA profiles in docetaxel-resistant human non-small cell lung carcinoma cells (SPC-A1). J Cell Mol Med. 2010;14(1–2):206–14. doi:10.1111/j.1582-4934.2009.00964.x.
Feng B, Wang R, Song HZ, Chen LB. MicroRNA-200b reverses chemoresistance of docetaxel-resistant human lung adenocarcinoma cells by targeting E2F3. Cancer. 2012;118(13):3365–76. doi:10.1002/cncr.26560.
Feng B, Wang R, Chen LB. MiR-100 resensitizes docetaxel-resistant human lung adenocarcinoma cells (SPC-A1) to docetaxel by targeting Plk1. Cancer Lett. 2012;317(2):184–91. doi:10.1016/j.canlet.2011.11.024.
Cui SY, Huang JY, Chen YT, et al. Let-7c governs the acquisition of chemo- or radioresistance and epithelial-to-mesenchymal transition phenotypes in docetaxel-resistant lung adenocarcinoma. Mol Cancer Res. 2013;11(7):699–713. doi:10.1158/1541-7786.MCR-13-0019-T.
Zhang JG, Guo JF, Liu DL, Liu Q, Wang JJ. MicroRNA-101 exerts tumor-suppressive functions in non-small cell lung cancer through directly targeting enhancer of zeste homolog 2. J Thorac Oncol. 2011;6(4):671–8. doi:10.1097/JTO.0b013e318208eb35.
Chatterjee A, Chattopadhyay D, Chakrabarti G. MiR-16 targets Bcl-2 in paclitaxel-resistant lung cancer cells and overexpression of miR-16 along with miR-17 causes unprecedented sensitivity by simultaneously modulating autophagy and apoptosis. Cell Signal. 2015;27(2):189–203. doi:10.1016/j.cellsig.2014.11.023.
Chatterjee A, Chattopadhyay D, Chakrabarti G. miR-17-5p downregulation contributes to paclitaxel resistance of lung cancer cells through altering beclin1 expression. PLoS One. 2014;9(4):e95716. doi:10.1371/journal.pone.0095716.
Holleman A, Chung I, Olsen RR, et al. miR-135a contributes to paclitaxel resistance in tumor cells both in vitro and in vivo. Oncogene. 2011;30(43):4386–98. doi:10.1038/onc.2011.148.
Shen H, Wang L, Ge X, Jiang CF, et al. MicroRNA-137 inhibits tumor growth and sensitizes chemosensitivity to paclitaxel and cisplatin in lung cancer. Oncotarget. 2016;7(15):20728–42. doi:10.18632/oncotarget.8011.
Ye J, Zhang Z, Sun L, Fang Y, Xu X, Zhou G. miR-186 regulates chemo-sensitivity to paclitaxel via targeting MAPT in non-small cell lung cancer (NSCLC). Mol BioSyst. 2016;12(11):3417–24. doi:10.1039/c6mb00576d.
Ahmadzadeh M, Johnson LA, Heemskerk B, et al. Tumor antigen-specific CD8 T cells infiltrating the tumor express high levels of PD-1 and are functionally impaired. Blood. 2009;114(8):1537–44. doi:10.1182/blood-2008-12-195792.
Brahmer JR, Pardoll DM. Immune checkpoint inhibitors: making immunotherapy a reality for the treatment of lung cancer. Cancer Immunol Res. 2013;1(2):85–91. doi:10.1158/2326-6066.CIR-13-0078.
Chen L, Gibbons DL, Goswami S, et al. Metastasis is regulated via microRNA-200/ZEB1 axis control of tumor cell PD-L1 expression and intratumoral immunosuppression. Nat Commun. 2014;5:5241. doi:10.1038/ncomms6241.
Gibbons DL, Lin W, Creighton CJ, et al. Contextual extracellular cues promote tumor cell EMT and metastasis by regulating miR-200 family expression. Genes Dev. 2009;23(18):2140–51. doi:10.1101/gad.1820209.
Gibbons DL, Lin W, Creighton CJ, et al. Expression signatures of metastatic capacity in a genetic mouse model of lung adenocarcinoma. PLoS One. 2009;4(4):e5401. doi:10.1371/journal.pone.0005401.
Cortez MA, Ivan C, Valdecanas D, et al. PDL1 Regulation by p53 via miR-34. J Natl Cancer Inst. 2016;108(1):djv303.
Garofalo M, Jeon Y-J, Nuovo GJ, et al. MiR-34a/c-dependent PDGFR-α/β downregulation inhibits tumorigenesis and enhances TRAIL-induced apoptosis in lung cancer. PLoS One. 2013;8(6):e67581. doi:10.1371/journal.pone.0067581.
Xie Y, Tobin LA, Camps J, et al. MicroRNA-24 regulates XIAP to reduce the apoptosis threshold in cancer cells. Oncogene. 2013;32(19):2442–51. doi:10.1038/onc.2012.258.
Joshi P, Jeon YJ, Lagana A, et al. MicroRNA-148a reduces tumorigenesis and increases TRAIL-induced apoptosis in NSCLC. Proc Natl Acad Sci U S A. 2015;112(28):8650–5. doi:10.1073/pnas.1500886112.
Jeon YJ, Middleton J, Kim T, et al. A set of NF-kappaB-regulated microRNAs induces acquired TRAIL resistance in lung cancer. Proc Natl Acad Sci U S A. 2015;112(26):E3355–64. doi:10.1073/pnas.1504630112.
Chorostowska-Wynimko J, Szpechcinski A. The impact of genetic markers on the diagnosis of lung cancer: a current perspective. J Thorac Oncol. 2007;2(11):1044–51. doi:10.1097/JTO.0b013e318158eed4.
Chorostowska-Wynimko J, Skronski M, Szpechcinski A. Markery molekularne we wczesnej diagnostyce raka płuca - fakty i nadzieje. Onkol Inf. 2011;8(3):152–9.
Skronski M, Szpechcinski A, Chorostowska-Wynimko J. Współczesne metody wykrywania mutacji genu EGFR jako czynnika predykcyjnego dla terapii ukierunkowanej molekularnie chorych na niedrobnokomórkowego raka płuca - czy istnieje “złoty standard” diagnostyczny? Pneumonol Alergol Pol. 2014;82(3):311–22.
Chorostowska-Wynimko J, Skroński M, Szpechcinski A. Molekularne markery prognostyczne i predykcyjne w diagnostyce niedrobnokomórkowego raka płuca. Onkol Inf. 2011;8(3):160–7.
Sequist LV, Waltman BA, Dias-Santagata D, et al. Genotypic and histological evolution of lung cancers acquiring resistance to EGFR inhibitors. Sci Transl Med. 2011;3(75):75ra26. doi:10.1126/scitranslmed.3002003.
Ricciuti B, Mecca C, Cenci M, et al. miRNAs and resistance to EGFR-TKIs in EGFR-mutant non-small cell lung cancer: beyond ‘traditional mechanisms’ of resistance. Ecancermedicalscience. 2015;9:569. doi:10.3332/ecancer.2015.569.
Zhaohui G, Jie Y, Jingqiu L, et al. Novel insights into the role of MicroRNA in lung cancer resistance to treatment and targeted therapy. Curr Cancer Drug Targets. 2014;14(3):241–58. doi:10.2174/1568009614666140305104845.
Zhang H, Su Y, Xu F, Kong J, Yu H, Qian B. Circulating microRNAs in relation to EGFR status and survival of lung adenocarcinoma in female non-smokers. PLoS One. 2013;8(11):e81408. doi:10.1371/journal.pone.0081408.
Shen Y, Tang D, Yao R, et al. microRNA expression profiles associated with survival, disease progression, and response to gefitinib in completely resected non-small-cell lung cancer with EGFR mutation. Med Oncol. 2013;30(4):750. doi:10.1007/s12032-013-0750-1.
Zhao Q, Cao J, Wu Y-C, et al. Circulating miRNAs is a potential marker for gefitinib sensitivity and correlation with EGFR mutational status in human lung cancers. Am J Cancer Res. 2015;5(5):1692–705.
Wang S, Su X, Bai H, et al. Identification of plasma microRNA profiles for primary resistance to EGFR-TKIs in advanced non-small cell lung cancer (NSCLC) patients with EGFR activating mutation. J Hematol Oncol. 2015;8:127. doi:10.1186/s13045-015-0210-9.
Li B, Ren S, Li X, et al. MiR-21 overexpression is associated with acquired resistance of EGFR-TKI in non-small cell lung cancer. Lung Cancer. 2014;83(2):146–53. doi:10.1016/j.lungcan.2013.11.003.
Kim SM, Kim JS, Kim JH, et al. Acquired resistance to cetuximab is mediated by increased PTEN instability and leads cross-resistance to gefitinib in HCC827 NSCLC cells. Cancer Lett. 2010;296(2):150–9. doi:10.1016/j.canlet.2010.04.006.
Cheng X, Chen H. Tumor heterogeneity and resistance to EGFR-targeted therapy in advanced nonsmall cell lung cancer: challenges and perspectives. Onco Targets Ther. 2014;7:1689–704. doi:10.2147/OTT.S66502.
Yoshida T, Song L, Bai Y, et al. ZEB1 mediates acquired resistance to the epidermal growth factor receptor-tyrosine kinase inhibitors in non-small cell lung cancer. PLoS One. 2016;11(1):e0147344. doi:10.1371/journal.pone.0147344.
Li J, Li X, Ren S, et al. miR-200c overexpression is associated with better efficacy of EGFR-TKIs in non-small cell lung cancer patients with EGFR wild-type. Oncotarget. 2014;5(17):7902–16. doi:10.18632/oncotarget.2302.
Lindeman NI, Cagle PT, Beasley MB, et al. Molecular testing guideline for selection of lung cancer patients for EGFR and ALK tyrosine kinase inhibitors: guideline from the College of American Pathologists, International Association for the Study of Lung Cancer, and Association for Molecular Pathology. J Thorac Oncol. 2013;8(7):823–59. doi:10.1097/JTO.0b013e318290868f.
Morgensztern D, Campo MJ, Dahlberg SE, et al. Molecularly targeted therapies in non-small-cell lung cancer annual update 2014. J Thorac Oncol. 2015;10(1 Suppl 1):S1–63. doi:10.1097/JTO.0000000000000405.
Kwak EL, Bang YJ, Camidge DR, et al. Anaplastic lymphoma kinase inhibition in non-small-cell lung cancer. N Engl J Med. 2010;363(18):1693–703. doi:10.1056/NEJMoa1006448.
Maione P, Sacco PC, Sgambato A, Casaluce F, Rossi A, Gridelli C. Overcoming resistance to targeted therapies in NSCLC: current approaches and clinical application. Ther Adv Med Oncol. 2015;7(5):263–73. doi:10.1177/1758834015595048.
Gao HX, Yan L, Li C, Zhao LM, Liu W. miR-200c regulates crizotinib-resistant ALK-positive lung cancer cells by reversing epithelial-mesenchymal transition via targeting ZEB1. Mol Med Rep. 2016;14(5):4135–43. doi:10.3892/mmr.2016.5770.
Gelsomino F, Facchinetti F, Haspinger ER, et al. Targeting the MET gene for the treatment of non-small-cell lung cancer. Crit Rev Oncol/Hematol. 2014;89(2):284–99. doi:10.1016/j.critrevonc.2013.11.006.
Brighenti M. MicroRNA and MET in lung cancer. Ann Translat Med. 2015;3(5):68. doi:10.3978/j.issn.2305-5839.2015.01.26.
Rokavec M, Li H, Jiang L, Hermeking H. The p53/miR-34 axis in development and disease. J Mol Cell Biol. 2014;6(3):214–30. doi:10.1093/jmcb/mju003.
Migliore C, Petrelli A, Ghiso E, et al. MicroRNAs impair MET-mediated invasive growth. Cancer Res. 2008;68(24):10128–36. doi:10.1158/0008-5472.CAN-08-2148.
Garofalo M, Romano G, Di Leva G, et al. EGFR and MET receptor tyrosine kinase-altered microRNA expression induces tumorigenesis and gefitinib resistance in lung cancers. Nat Med. 2011;18(1):74–82. doi:10.1038/nm.2577.
Ling H, Fabbri M, Calin GA. MicroRNAs and other non-coding RNAs as targets for anticancer drug development. Nat Rev Drug Discov. 2013;12(11):847–65. doi:10.1038/nrd4140.
Uhlmann S, Mannsperger H, Zhang JD, et al. Global microRNA level regulation of EGFR-driven cell-cycle protein network in breast cancer. Mol Syst Biol. 2012;8:570. doi:10.1038/msb.2011.100.
Bader AG, Brown D, Stoudemire J, Lammers P. Developing therapeutic microRNAs for cancer. Gene Ther. 2011;18(12):1121–6. doi:10.1038/gt.2011.79.
Li Z, Rana TM. Therapeutic targeting of microRNAs: current status and future challenges. Nat Rev Drug Discov. 2014;13(8):622–38. doi:10.1038/nrd4359.
Fortunato O, Boeri M, Verri C, Moro M, Sozzi G. Therapeutic use of MicroRNAs in lung cancer. Biomed Res Int. 2014;2014:8. doi:10.1155/2014/756975.
Takamizawa J, Konishi H, Yanagisawa K, et al. Reduced expression of the let-7 microRNAs in human lung cancers in association with shortened postoperative survival. Cancer Res. 2004;64(11):3753–6. doi:10.1158/0008-5472.CAN-04-0637.
Raponi M, Dossey L, Jatkoe T, et al. MicroRNA classifiers for predicting prognosis of squamous cell lung cancer. Cancer Res. 2009;69(14):5776–83. doi:10.1158/0008-5472.CAN-09-0587.
Landi MT, Zhao Y, Rotunno M, et al. MicroRNA expression differentiates histology and predicts survival of lung cancer. Clin Cancer Res. 2010;16(2):430–41. doi:10.1158/1078-0432.CCR-09-1736.
Xia Y, Zhu Y, Zhou X, Chen Y. Low expression of let-7 predicts poor prognosis in patients with multiple cancers: a meta-analysis. Tumour Biol. 2014;35(6):5143–8. doi:10.1007/s13277-014-1663-0.
Johnson SM, Grosshans H, Shingara J, et al. RAS is regulated by the let-7 MicroRNA family. Cell. 2005;120(5):635–47. doi:10.1016/j.cell.2005.01.014.
Johnson CD, Esquela-Kerscher A, Stefani G, et al. The let-7 microRNA represses cell proliferation pathways in human cells. Cancer Res. 2007;67(16):7713–22. doi:10.1158/0008-5472.CAN-07-1083.
Esquela-Kerscher A, Trang P, Wiggins JF, et al. The let-7 microRNA reduces tumor growth in mouse models of lung cancer. Cell Cycle. 2008;7(6):759–64. doi:10.4161/cc.7.6.5834.
Kumar MS, Erkeland SJ, Pester RE, et al. Suppression of non-small cell lung tumor development by the let-7 microRNA family. Proc Natl Acad Sci U S A. 2008;105(10):3903–8. doi:10.1073/pnas.0712321105.
Gallardo E, Navarro A, Viñolas N, et al. miR-34a as a prognostic marker of relapse in surgically resected non-small-cell lung cancer. Carcinogenesis. 2009;30(11):1903–9. doi:10.1093/carcin/bgp219.
Bader AG. miR-34 – a microRNA replacement therapy is headed to the clinic. Front Genet. 2012;3:120. doi:10.3389/fgene.2012.00120.
Bommer GT, Gerin I, Feng Y, et al. p53-mediated activation of miRNA34 candidate tumor-suppressor genes. Curr Biol. 2007;17(15):1298–307. doi:10.1016/j.cub.2007.06.068.
Kasinski AL, Slack FJ. miRNA-34 prevents cancer initiation and progression in a therapeutically resistant K-ras and p53-induced mouse model of lung adenocarcinoma. Cancer Res. 2012;72(21):5576–87. doi:10.1158/0008-5472.CAN-12-2001.
Wiggins JF, Ruffino L, Kelnar K, et al. Development of a lung cancer therapeutic based on the tumor suppressor microRNA-34. Cancer Res. 2010;70(14):5923–30. doi:10.1158/0008-5472.CAN-10-0655.
Trang P, Wiggins JF, Daige CL, et al. Systemic delivery of tumor suppressor microRNA mimics using a neutral lipid emulsion inhibits lung tumors in mice. Mol Ther. 2011;19(6):1116–22. doi:10.1038/mt.2011.48.
Selcuklu SD, Donoghue Mark TA, Spillane C. miR-21 as a key regulator of oncogenic processes. Biochem Soc Trans. 2009;37(4):918.
Seike M, Goto A, Okano T, et al. MiR-21 is an EGFR-regulated anti-apoptotic factor in lung cancer in never-smokers. Proc Natl Acad Sci U S A. 2009;106(29):12085–90. doi:10.1073/pnas.0905234106.
Lee R. Announces positive results from preclinical lung cancer study using lead anti-microRNA candidate. 2015. http://www.microlinbio.com/Microlin_PR_9-24-15.pdf. Accessed 12 Nov 2016.
Trang P, Medina PP, Wiggins JF, et al. Regression of murine lung tumors by the let-7 microRNA. Oncogene. 2010;29(11):1580–7. doi:10.1038/onc.2009.445.
Kasinski AL, Kelnar K, Stahlhut C, et al. A combinatorial microRNA therapeutics approach to suppressing non-small cell lung cancer. Oncogene. 2015;34(27):3547–55. doi:10.1038/onc.2014.282.
Wu Y, Crawford M, Mao Y, et al. Therapeutic delivery of MicroRNA-29b by cationic lipoplexes for lung cancer. Mol Ther Nucleic Acids. 2013;2:e84. doi:10.1038/mtna.2013.14.
Bacchi CE, Ciol H, Queiroga EM, Benine LC, Silva LH, Ojopi EB. Epidermal growth factor receptor and KRAS mutations in Brazilian lung cancer patients. Clinics. 2012;67(5):419–24.
Bouchie A. First microRNA mimic enters clinic. Nat Biotechnol. 2013;31(7):577. doi:10.1038/nbt0713-577.
Product Development Pipeline. http://www.mirnarx.com/pipeline/mirna-pipeline.html. Accessed 12 Nov 2016.
Pichler M, Calin GA. MicroRNAs in cancer: from developmental genes in worms to their clinical application in patients. Br J Cancer. 2015;113(4):569–73. doi:10.1038/bjc.2015.253.
van Rooij E, Kauppinen S. Development of microRNA therapeutics is coming of age. EMBO Mol Med. 2014;6(7):851–64. doi:10.15252/emmm.201100899.
Chira S, Jackson CS, Oprea I, et al. Progresses towards safe and efficient gene therapy vectors. Oncotarget. 2015;6(31):30675–703. doi:10.18632/oncotarget.5169.
Wu Y, Crawford M, Yu B, Mao Y, Nana-Sinkam SP, Lee LJ. MicroRNA delivery by cationic lipoplexes for lung cancer therapy. Mol Pharm. 2011;8(4):1381–9. doi:10.1021/mp2002076.
Bharali DJ, Khalil M, Gurbuz M, Simone TM, Mousa SA. Nanoparticles and cancer therapy: a concise review with emphasis on dendrimers. Int J Nanomedicine. 2009;4:1–7.
Immordino ML, Dosio F, Cattel L. Stealth liposomes: review of the basic science, rationale, and clinical applications, existing and potential. Int J Nanomedicine. 2006;1(3):297–315.
Reid G, Kao SC, Pavlakis N, et al. Clinical development of TargomiRs, a miRNA mimic-based treatment for patients with recurrent thoracic cancer. Epigenomics. 2016;8(8):1079–85. doi:10.2217/epi-2016-0035.
Reid G, Williams M, Kirschner MB, et al. Abstract 3976: targeted delivery of a synthetic microRNA-based mimic as an approach to cancer therapy. Cancer Res. 2015;75(15 Supplement):3976. doi:10.1158/1538-7445.am2015-3976.
MacDiarmid JA, Mugridge NB, Weiss JC, et al. Bacterially derived 400 nm particles for encapsulation and cancer cell targeting of chemotherapeutics. Cancer Cell. 2007;11(5):431–45. doi:10.1016/j.ccr.2007.03.012.
Soutschek J, Akinc A, Bramlage B, et al. Therapeutic silencing of an endogenous gene by systemic administration of modified siRNAs. Nature. 2004;432(7014):173–8.
Chiu YL, Rana TM. siRNA function in RNAi: a chemical modification analysis. RNA. 2003;9(9):1034–48.
Acknowledgements
The authors acknowledge English language assistance from textcheck. For a certificate, please see http://www.textcheck.com/certificate/qwqF5f.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This article has been prepared under the research project granted by the Polpharma Scientific Foundation.
Conflict of Interest
The authors declare no conflict of interest.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution-NonCommercial 4.0 International License (http://creativecommons.org/licenses/by-nc/4.0/), which permits any noncommercial use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Florczuk, M., Szpechcinski, A. & Chorostowska-Wynimko, J. miRNAs as Biomarkers and Therapeutic Targets in Non-Small Cell Lung Cancer: Current Perspectives. Targ Oncol 12, 179–200 (2017). https://doi.org/10.1007/s11523-017-0478-5
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11523-017-0478-5