Abstract
The CRISPR/Cas9 system is a revolutionary gene editing technology that combines simplicity of use and efficiency of mutagenesis. As this technology progresses toward human therapies, valid concerns including off-target mutations and immunogenicity must be addressed. One approach to address these issues is to minimize the presence of the CRISPR/Cas9 components by maintaining a tighter temporal control of Cas9 endonuclease and reducing the time period of activity. This has been achieved to some degree by delivering the CRISPR/Cas9 system via pre-formed Cas9 + gRNA ribonucleoprotein (RNP) complexes. In this review, we first discuss the molecular modifications that can be made using CRISPR/Cas9 and provide an overview of current methods for delivering Cas9 RNP complexes both in vitro and in vivo. We conclude with examples of how Cas9 RNP delivery may be used to target neuroinflammatory processes, namely in regard to viral infections of the central nervous system and neurodegenerative diseases. We propose that Cas9 RNP delivery is a viable approach when considering the CRISPR/Cas9 system for both experimentation and the treatment of disease.

Graphical Abstract
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
CRISPR/Cas9 gene editing technology has become an invaluable scientific tool and has great potential the treatment of disease. This technology allows for editing of genomic regions by targeting a DNA nuclease (Cas9) to a specific DNA sequence resulting in a double-stranded break in the DNA (Doudna and Charpentier 2014). The sequence specific targeting is achieved using guide RNA (gRNA) that is complementary to the target sequence (Jinek et al. 2012). The host’s cellular repair mechanisms then act on the “broken” DNA and the repaired DNA is modified from the original sequence (or it is cut again by the nuclease). For example, non-homologous end joining (NHEJ) is one mechanism of repairing DNA breaks and it results in insertions and deletions (indels) in the region which can change the function of the target gene (Cong et al. 2013). Another repair mechanism is homology dependent repair (HDR), which incorporates new genetic material by way of a donor template (e.g. an exogenously provided DNA construct or homologous chromosome) containing homologous arms to the site affected by the double stranded break (Chang et al. 2013; Kim and Kim 2014). Thus CRISPR/Cas9 technology provides a means to specifically manipulate the genome of almost any organism.
Originally characterized as a bacterial immune system, investigators have adapted the two key components- Cas9 endonuclease and the original guide RNA (gRNA) for use in other model organisms (Jinek et al. 2012). This has been achieved primarily by delivery of genetically-encoded components, via plasmid-based transfection or lentiviral transduction (Shalem et al. 2014; Wang et al. 2014); however, newer forms of Cas9 are compatible with other vectors such as adeno-associated virus vectors (Ran et al. 2015; Swiech et al. 2015). The packaging and delivery of CRISPR/Cas9 by viral vectors produces robust, and long term expression of Cas9 and gRNA. Unfortunately, a growing body of evidence suggest DNA based methods of delivery may lead to negative effects within the targeted cell population, including the unintentional integration of Cas9/gRNA genetic material (Cradick et al. 2013; Fu et al. 2013), the heightened occurrence of off target mutation due to its long-term or constitutive expression (Hsu et al. 2013), and the development of an immune response (Chew et al. 2016). Thus, alternative delivery methods of CRISPR/Cas9 are needed to aid in the translation of this technology into human therapies.
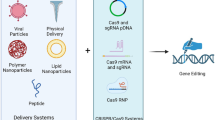
An established approach of CRISPR/Cas9 delivery uses a pre-formed ribonucleoprotein (RNP) Cas9 + gRNA complex. Seminal work by Kim et al. demonstrate that this protein-based delivery of CRISPR/Cas9 offers many benefits including transient activity of Cas9, rapid evidence of mutagenesis, decreased incidence of off-target mutation, and increased viability of cell populations when compared to plasmid based methods (Kim et al. 2014). Many of these attributes address the concerns of CRISPR/Cas9-based gene editing. This manuscript will review the current uses and methods of Cas9 delivery as a ribonucleoprotein complex both in vitro and in vivo which include i.) direct injection of Cas9 RNP, ii.) Cas9 RNP delivery by electroporation, iii.) Cas9 RNP modification by penetrating peptides, iv.) Cas9 packaging by cationic lipoplexes/nanoparticles, and v.) Cas9 RNP packaging by extracellular vesicles. The use of Cas9 RNPs to modulate neuroinflammatory processes will also be discussed.
Mechanisms and Applications of CRISPR-Based Genome Editing
Non-Homologous End Joining (NHEJ)
The Cas9 protein from S. pyogenes, as well as its orthologs from other bacteria, uses sequence homology provided by a guide RNA molecule cofactor to bind to and produce a double-strand break at a defined location within a double-stranded DNA molecule (Cong et al. 2013; Jinek et al. 2013; Mali et al. 2013). As described above, when this technology is used to produce effects on a eukaryotic genome, the break is typically resolved by NHEJ. The “error-prone” NHEJ process typically results in a deletion or insertion of nucleotides at the break site (Fig. 1). This type of repair typically leads to an insertion or deletion of nucleotides (indel) around the break site. An indel occurring within the coding region of a gene frequently produces a change in the protein sequence or causes a frameshift. However, it should be noted that the stochastic nature of this process creates heterogeneity, both within a single cell containing two or more target loci to create a complex mutant heterozygote and within a population of similarly treated cells. Despite the stochastic nature of the genotypes within an otherwise isogenic population, the use of CRISPR for NHEJ-mediated gene disruption has become commonplace using various modalities including plasmid and viral vectors, as well pre-formed ribonucleoprotein complexes of Cas9 protein and gRNA (i.e.RNPs) (Cong et al. 2013; Jinek et al. 2013; Ran et al. 2013; Wang et al. 2013; Zallar et al. 2018)
Examples of genome modification using CRISPR/Cas9. a A depiction of a Cas9-induced DSB being resolved by NHEJ processes to produce an indel. b The same DSB as in A, being resolved by homology-directed repair using a repair template consisting of a novel sequence flanked by homologous arms. c Cas9-BE variant modifying a single base within the gRNA targeting region without making a double-stranded break
The use of viral vectors to deliver CRISPR for gene disruption in the rodent brain has been demonstrated for protein coding genes and miRNAs (Murlidharan et al. 2016; Swiech et al. 2015; Yamaguchi et al. 2018). RNPs have used with oocytes and germlines (Nakagawa et al. 2016; Paix et al. 2017) and zygotes (Teixeira et al. 2018) to create genome-edited/transgenic lines. Other studies have used RNPs (Campbell et al. 2019; Jacobi et al. 2017; Liu et al. 2015; Mangeot et al. 2019; Montagna et al. 2018). In fact, Mangeot et al. (2019) have demonstrated that RNP-loaded viral-like particles can be used for genome editing in vivo by successfully mutating the Hpd gene in the mouse liver. Methods of RNP delivery are discussed in more detail in sections below.
In addition to exploiting the NHEJ process to create targeted indel mutations, progress has been made towards achieving more predictable outcomes by using homology directed repair and base editing.
Homology-Directed Repair (HDR)
The earliest demonstration of a pre-determined outcome from CRISPR-based DSBs was made by providing a “repair template” along with the Cas9 and gRNA components (Wang et al. 2013). While the structural details of the repair template can vary based on the model organism’s target cell type and delivery mechanism, in general, there is a common structural layout of the template. For example, there is a region of novel or mutant sequence flanked by segments that are identical or highly homologous to that of the termini of the induced DSB (aka homologous arms). These regions of homology are used by recombinational machinery of the cell to bridge the gap between the DSB termini, filling the interval spanning the break with the novel sequence (Fig. 1b). Double-stranded DNA as well as single-stranded DNA with arms and novel sequences of varying lengths have been used to generate base changes and codon substitutions as well as insertions of epitope tags or entire transgene cassettes at precise locations.
One caveat with this method is that it requires the delivery of an additional component, i.e. the repair template DNA, to the target cell. Once all elements are delivered to a cell, there is no guarantee that a cell will use homology-directed repair processes unless some form of selection is built into the design. DNA that is repaired by NHEJ, for example, will create indels within the population or possibly within the same cell. For example, the resulting DNA at a target site within a single cell may be heterozygous with a novel sequence on one chromosome and a deletion on the other; both differing from the original target sequence.
Despite this caveat to HDR following CRISPR/Cas9 exposure, modifications in vivo have been successfully performed. For example, clinically relevant alleles causing alpha-1 antitrypsin deficiency (AATD) in the liver have been repaired in mice (Shen et al. 2018) adding epitope tags to endogenous genes (Aird et al. 2018; Nishiyama et al. 2017), and knocking in large DNA fragments (Bak and Porteus 2017) have all been successful using various combinations of transgenic animals and viral vectors to deliver the necessary components (Mizuno et al. 2018). The use of virus-like particles for the delivery of Cas9 RNPs and a repair template has also been reported (Mangeot et al. 2019), where a single-stranded DNA donor molecule was adhered or complexed to the exterior of the particle using polybrene prior to delivery. Future studies that further optimize the co-delivery of repair template will be essential for exploiting HDR for in vivo genome editing and repair.
Base Editing
CRISPR-Base edit (CRISPR-BE) seeks to circumvent the variable outcomes associated with the processing of Cas9-induced double-strand breaks by avoiding the chromosomal cleavage all together. The binding activity of Cas9 to specific DNA sequences in an RNA-directed manner is separable from the DNA domains. These two domains can be independently rendered non-functional by specific point mutations (D10A and H840A, respectively) to create “nickases” (Cas9n) which only break one strand of the DNA target site or mutated simultaneously to encode a cleavage-dead protein, dCas9 (Jinek et al. 2012). These variants can then be appended on the both the amino- and carboxyl- ends with orthogonal functional domains, and when combined with a guide RNA, will localize these new effectors to a specific genomic locus.
When paired in this manner with the cytidine deaminase (APOBEC1) and uracil-N-glycosylase inhibitor (UGI), Cas9-BE can make specific modifications that result in the conversion of C-G basepairs into T-A basepairs (Komor et al. 2017). The same group also engineered an adenosine deaminase to enable editing of A-T basepairs into G-C basepairs (Gaudelli et al. 2017). Others have extended the versatility this technology by datamining the available genome data intent on cataloging and evaluating all potential gRNAs that could produce an early stop codon in their target gene when paired with Cas9-BE reagents (Billon et al. 2017; Kuscu et al. 2017).
CRISPR-BE has been applied for in vivo editing by direct injection of expression plasmids and ribonucleotide protein complexes into mouse embryos (Kim et al. 2017) and xenopus embryos (Park et al. 2017). The adaptation of base editing tools for delivery in virus-like particles has not yet been reported, however the modifications needed to produce Cas9-BE ribonucleoprotein complexes from existing regents seems straightforward.
Methods of Cas9 RNP Delivery
The utilization of Cas9 RNP varies with both RNP assembly and the method of delivery. Fig. 2 shows a summary schematic of both processes that will be described. Table 1 summarizes the relative efficiency of each method when compared to previous models of CRISPR/Cas9 delivery.
Direct Injection of Cas9 RNP
Early work with Cas9 RNP’s involved direct injection into Caenorhabditis elegans (Cho et al. 2013; Paix et al. 2015). In this method, the ribonucleoprotein complex was created using purified Cas9 (commercially acquired or prepared through purification methods (Kim et al. 2014)) and in vitro transcribed gRNA, which will associate by mixing of Cas9 and gRNA in the appropriate buffers. The purified Cas9, as with Cas9 expressed from a DNA source in eukaryotic cells, has a nuclear localization signal which facilitates its transfer to the nucleus. After assembly the RNP complex can be used directly for injection.
Studies utilizing this method have been highly successful. In C. elegans, investigators have targeted the genes dpy-3 and dpy-10, mutations of which led to an easily observable phenotype. In experiments targeting dpy-3, investigators injected Cas9 ribonucleoprotein into cytoplasm of the syncytial germline of P0 generation hermaphrodites. Progeny (F1 and F2) were analyzed for mutations and were confirmed to have site specific mutation within the dpy-3 region as shown by the T7 Endonuclease I assay (which is used to determine indels) and sequencing, in addition to exhibiting the “dumpy” phenotype (Cho et al. 2013).
For targeted repair of dpy-10, investigators used a single-strand oligodeoxyribonucleotide and HDR to introduce a missense mutation to create a “roller” phenotype. They observed that the incidence of roller phenotypes using Cas9 RNP delivery is a significant improvement (50%) versus their previously observed experiments that utilized plasmid based delivery (11%) (Paix et al. 2015). Thus, direct injection of Cas9 RNP is shown to be suitable for both gene disruption and homology directed repair.
Overall, the method of Cas9 RNP delivery by direct injection continues to be utilized and is translatable to other scientific models including human cell lines/primary cells (Lin et al. 2014), mice (Hirano et al. 2016), crustaceans (Kumagai et al. 2017), and others.
Cas9 RNP Delivery by Electroporation
Electroporation of Cas9 RNP has primarily been used for its delivery to a wide variety of cell types in vitro. The assembly of the Cas9 + gRNA complex is made by co-incubation of purified Cas9 and in vitro transcribed gRNA. The Cas9 RNP is prepared in a solution of electroporation compatible buffer. A cell pellet is resuspended in electroporation buffer and combined with the Cas9 RNP in an electroporation cuvette. Electroporation is performed, after which cells can be returned to culture and assayed for genomic modifications, protein expression and functional changes.
As noted, the electroporation procedure is commonplace in producing CRISPR/Cas9 mediated mutagenesis in vitro. In one example, investigators utilized this method to produce transgenic mice through the electroporation of mouse zygotes with the Cas9 RNP complex (Chen et al. 2016; Qin et al. 2015; Teixeira et al. 2018). Collectively, the studies describe increased mutations, recombinations and viable offspring using this method when compared to Cas9 RNP microinjections. Interestingly, Chen et al. identified that a 10 ms pulse length and 16 μM Cas9 RNP concentration allowed for 100% bi-allelic edited embryos, but at a cost to embryo viability (30% viability). Ultimately, the investigators found that a pulse duration of 3 ms with a Cas9 RNP concentration of 8 μM combined the benefits of 67% bi-allelic editing efficiency with 60% of embryos developing to viable offspring. Additionally, in experiments comparing mRNA injection to RNP delivery, an increase both editing efficiency (60 to 80%) and viability (10% to 50%) were observed.
Other investigators have used Cas9 RNP electroporation on human immune cells including T cells and B cells. The genetic modification of immune cells is an attractive approach to create unique targeted therapies to specific disease states. In primary T cells, Cas9 RNP electroporation has been used to decrease the expression of the HIV co-receptor CXCR4 to combat HIV infection, and the surface receptor PD-1 to enhance immune cell clearance of cancers (Schumann et al. 2015). B cell editing offered similar results, including high editing efficiency (upwards of 80%) and the ability to modify both undifferentiated and differentiated B cell populations (Wu et al. 2018).
Cas9 Modification and Delivery by Penetrating Peptides
Another approach to Cas9 RNP delivery involves the modification of the Cas9 protein itself using cell-penetrating peptides. These peptides allow for the delivery of cargo (DNA, RNA, protein, small molecules) across the cellular membrane through passive “energy-independent” or active endocytic pathways. The energy-independent pathways involve disruption of the cellular membrane, enabling the direct translocation of the cargo into the cytosol. This type of membrane disruption is often associated with antimicrobial peptides, which are typically positively charged amphipathic molecules capable of direct interaction with phospholipid groups of cellular membranes (Hancock and Scott 2000; Zasloff 2002). The endocytic pathway consists of a subset of cellular pathways (e.g. caveolae-mediated endocytosis, clathrin-depending endocytosis, macropinocytosis) for moving substances from outside the cell to the interior.
All of these subsets follow a basic mechanism encompassing membrane binding, membrane folding and invagination, endocytosis of a host, membrane-derived vesicle containing the cargo, and endosomal escape, leading to the delivery of the cargo into the cytosol (Gestin et al. 2017).
In one study, investigators modified the N-terminus of Cas9 with increasing numbers of Simian vacuolating virus 40 nuclear localization sequences (SV40-NLS, 0x, 1x, 2x, 4x, 7x-NLS) to allow for penetration of the Cas9 RNP complex into target cells (Staahl et al. 2017). The investigators used a tdTomato reporter model, where the reporter was proceeded by a loxP flaked SV40 polyA sequence (i.e. a stop cassette). The stop cassette was then targeted by unique gRNA sequences to cause its excision, thereby allowing for the transcription of tdTomato. The investigators observed that increasing the number of nuclear localization sequences up to 4x-NLS significantly enhanced the expression of tdTomato in neural progenitor cells. However, the 7x-NLS modification did not provide an increase in tdTomato, suggesting limitations to the modifications that can be made to Cas9. Furthermore, injection of 4x-NLS Cas9 RNP into the brain parenchyma resulted in tdTomato expression co-localizing with neuronal populations, suggesting that this method of penetration could be used in vivo.
Another group of investigators applied a different approach using cell penetrating peptides by co-incubating the preformed Cas9 RNP with an engineered cell penetrating peptide (HIV-Tat variant PTD4) combined with an endosomal leakage domain (CM18) to make 6His-CM18-PTD4 (Del'Guidice et al. 2018). Co-incubation and administration of the Cas9 RNP complex successfully edited the HPRT and DNMT1 genes in Jurkat and primary natural killer cells.
Cas9 RNP in Combination with Nanoparticles
Nanoparticles are nano-scale (less than 100 nm) inorganic materials that possess unique characteristics e.g. conductance, melting temperature, malleability (Laura Soriano et al. 2018). One group has successfully combined a modified Cas9 RNP with a gold nanoparticle (Mout et al. 2017). The use of nanoparticles, including the gold nanoparticle (AuNP) has been shown to successfully deliver protein components for immune and cancer therapies (Almeida et al. 2014). Cas9 was first modified to have increasing numbers (0, 5, 10, 20) of glutamate residues (E-tag) on the N-terminus to allow for a localized negative charge on the Cas9 protein, facilitating the interaction with the AuNPs. Increasing the number of E-tags, subsequently enhanced nanoassembly and improved direct delivery of Cas9 into the cytoplasm of the target cells resulting in roughly 30% editing efficiency.
Another group performed Cas9-initiated, homology-directed repair using a nanoassembly containing a AuNP, Cas9 RNP, a donor DNA molecule, and an endosomal-disruptive polymer PAsp(DET), resulting in the formation of what they termed “CRISPR-Gold” (Lee et al. 2017). They show successful gene editing and homology directed repair in human embryonic stem cells, induced pluripotent stem cells, and a mouse model of Duchenne muscular dystrophy. In the mouse model, muscle function was improved in the CRISPR-Gold treated animals and analysis of inflammatory markers indicated that no aberrant inflammatory effects were observed as a consequence of CRISPR-Gold treatment. The lack of an inflammatory response supports further evaluation of Cas9 RNP nanoassembilies for in vivo applications.
Cas9 RNP Packaged into Microvesicles
Recently, the discovery and study of extracellular vesicles has revealed their importance in biological functions including the communication between organ systems, packaging and delivery of macromolecules (e.g. DNA, RNA, protein), and a role in disease pathogenesis (Robbins and Morelli 2014; Skog et al. 2008; van Niel et al. 2018). The microvesicle is a specific type of extracellular vesicle which is formed through budding of the cellular membrane, creating vesicles ranging from 100 to 1000 nm in diameter (Kalra et al. 2012). Packaging Cas9 RNP into a microvesicle usually involves a transfection of a producer cell line (such as HEK293FT cells) with at least 3 components; Cas9, the chosen guide RNA, and a microvesicle inducing protein. After this step, microvesicles containing Cas9 RNP that are released into the media can be purified, concentrated, and subsequently used to deliver Cas9 RNP to the recipient cell or organism.
Two recent papers describe the use of microvesicles formed by expressing vesicular stomatitis virus protein g (VSV-G) in HEK293 producers cells to cause microvesicle formation and packaging of Cas9 RNP’s. Using an EGFP reporter protein as a target, Montagana et al. (Montagna et al. 2018) confirm effective packaging and editing utilizing both wild type Cas9 and Cas9 nickase, suggesting that microvesicles can package at least two gRNA’s simultaneously. Additionally, the investigators edited EGFP expression in induced pluripotent stem cells and the heart of an EGFP expressing mouse, confirming the ability of microvesicle-based delivery in vitro and in vivo.
Additionally, VSV-G based microvesicles have been used to deliver Cas9 RNP to microglial cells harboring an HIV provirus (Campbell et al. 2019). Using a microglia model with a stably integrated provirus, our lab demonstrated that microvesicle delivery of a Cas9 RNP targeted to the HIV long terminal repeat (LTR) resulted in both mutation of the viral promoter and excision of the HIV provirus. This led to decreased proviral activity including decreased expression of the HIV viral protein Nef after simulation with pro-inflammatory factors such as lipopolysaccharide and tumor necrosis factor alpha. Furthermore, we observed a dose-dependency of Cas9 RNPs supporting need for pharmacological studies of Cas9 RNPs in microvesicle.
In conclusion, the five described methodologies show that Cas9 RNP delivery confers improved efficiency of genome modification (see Table 1), transient period of Cas9 activity, low cellular toxicity and minimal off-target mutagenic effects. All of which suggest that utilizing Cas9 RNPs may be a viable approach for CRISPR/Cas9 based therapies.
Targeting Neuroinflammation with Cas9 RNPs
Virus Mediated Neuroinflammation
In the central nervous system, microglia are the primary cellular reservoir for HIV and have received much attention for their role in the neuroinflammation associated with an HIV-infected brain (Chen et al. 2017; Ghorpade et al. 2005; Heaton et al. 2011). As the resident macrophage of the brain, microglia can be productively infected with HIV and harbor latent, integrated proviral genomes. Over time, both replication competent virus and viral proteins can be released from infected cells, leading to damage of surrounding neuronal populations which manifest into cognitive, motor, and behavioral abnormalities (Zhou and Saksena 2013). Furthermore, once the virus has infected the brain environment, it is no longer within the reach of many anti-retroviral drugs which have sub optimal penetration into the CNS due to the blood brain barrier, leading to active viral production even in the case of undetectable viral load in the periphery (Letendre et al. 2008). Indeed, studies confirm that HIV strains isolated from the CNS can have a different clonal expansion than from other tissue sites (Schnell et al. 2010), emphasizing the need for new strategies to target microglia to effectively diminish viral outgrowth and subsequent pathogenesis.
Unfortunately, an efficient and reproducible method to genetically manipulate microglia remains elusive, which may be due in part to their ability to sense and react to foreign DNA (Jeffries and Marriott 2017; Reinert et al. 2016). Thus, Cas9 RNP’s may serve as a useful tool in this regard, bypassing the need for DNA delivery and shielding the gRNA within the ribonucleoprotein. Preliminary data from our lab has shown that microvesicle-based, CherryPicker Red containing “gesicles” can deliver Cas9 protein to primary microglia populations both in vitro and in vivo (Fig. 3). Application of gesicles to primary microglia cultures exhibit co-localization of the gesicle marker CherryPicker Red with Iba-1 positive cells (Fig. 3a). Additionally, gesicles were stereotaxically injected into the striatum of rats and co-localization of CherryPicker Red and Iba-1 fluorescence was observed on brain sections. While there were CherryPicker Red positive Iba-1 cells, the red fluorescence also co-localizes with other cellular populations, suggesting the association of gesicles to a wide array of cell types (Fig. 3b). These data suggest that microvesicles have potential as method to deliver Cas9 RNP’s targeting the HIV provirus in microglia populations.
Microvesicle delivery of CherryPicker Red to primary microglia in vitro and in vivo. Microvesicles labeled with CherryPicker red protein were used to monitor the association with primary rat microglia (Iba-1 positive cells). a Immunocytochemistry of Iba-1 (green), CherryPicker Red (Red) and merged image with DAPI (nuclear stain; blue) 24 h post-application of microvesicles directly to cultured microglia. b Immunofluorescent labeling of rat striatum for Iba-1 (green), CherryPicker Red (Red) and merged image with DAPI stain. Brains were processed 24 h after intrastriatal injection with CherryPicker Red labeled microvesicles. Scale bar = 50 μM
Another viral-based neuroinflammatory disease is progressive multifocal leukoencephalopathy, which occurs from reactivation of the human polyomavirus JC. This disorder is commonly associated with HIV infection and can result in the disruption of myelin formation, inflammation and ultimately death (Berger 2011). Investigators have utilized plasmid based CRISPR/Cas9 delivery to target the large T antigen (T-Ag) of this virus in order to disrupt proviral reactivation and transcription. Targeting the gene regions of T-Ag resulted in decreased T-Ag expression which in turn decreased viral activation via a JCVL promotor driven luciferase reporter (Wollebo et al. 2015). To date, no data is available that displays Cas9 RNP delivery to oligodendrocyte populations. The development of this technology would be beneficial for not only polyomavirus JC infection, but to other demyelinating diseases such as Multiple Sclerosis.
Neuroinflammatory and Neurodegenerative Diseases
CRISPR/Cas9 technology can be targeted other cells in the CNS including neurons, astrocytes, and oligodendrocytes to potentially be used as a treatment for neurodegenerative and neuroinflammatory diseases. Neuroinflammation can occur in many neurological diseases and injuries, for example brain trauma, ischemia, viral infection, and neurodegeneration. Thus, targeting the underlying genetic components to these disease states may ultimately decrease inflammatory processes. For example, in Huntington’s disease, a mutant form of the huntingtin gene contains excessive (greater than 36) CAG tri-nucleotide repeats. When transcribed, an elongated, mutant form of the protein is produced that ultimately leads to protein aggregation, inflammation, and neuronal toxicity. Plasmid-based nucleofection of CRISPR/Cas9 components (5 μg Cas9 Nickase; 2 × 5 μg gRNA in ~1 × 106 to 5 × 105 cells) has been used to cause partial excision of CAG repeats (Dabrowska et al. 2018) in cell lines and patient-derived fibroblasts. This approach led to a 70–80% decrease in levels of total huntingtin protein.
The pathogenesis of Alzheimer’s Disease continues to be uncovered, with aspects including amyloid beta cleavage, apolipoprotein ε4 allele inheritance, and tau/neurofibrillary tangles and chronic inflammation all contributing to neuronal death and progressive dementia (Dionisio-Santos et al. 2019). CRISPR/Cas9 have recently been used to target amyloid precursor protein and its subsequent cleavage. For example, CRISPR/Cas9-mediated alterations to the C-terminus of amyloid precursor protein (APP) have been introduced to promote splicing towards the neuroprotective α cleavage form (Sun et al. 2019). Using lentivirus based delivery approaches in both wild type iPSCs and cells harboring the familial Alzheimer’s mutation (London mutation) the investigators observed a significant reduction of full length APP, with an increase in secreted APPα (neuroprotective form) by western blot and a decrease in Aβ40/42 (pathogenic form) by ELISA. The same approach has been performed in familial forms of Alzheimer’s stemming from the Swedish mutation of amyloid precursor protein (György et al. 2018). In fibroblasts, electroporation of ~ 1 × 106 cells (with 20–50% transfection efficiency) led to decreased Aβ40/42 expression by 50%. In vivo, the investigators utilized Tg2576 mice which harbor the Swedish mutation. AAV injection (104–105 genome copies per cell) of Cas9/gRNAs into the hippocampus resulting in a ~1.3% indel formation. These studies illustrate the potential for CRISPR/Cas9 use in AD.
For neurodegenerative diseases associated with known genomic mutations that cause altered gene expression, such as Huntington’s disease or Alzheimer’s disease, Cas9 RNP delivery may offer therapeutic hope. Much of Huntington’s disease affects the basal ganglia, therefore stereotaxic injections of Cas9 RNP’s (by cell penetrating peptides, nanoparticles, etc.) targeting the CAG repeats may serve as a therapeutic approach enabling long lasting effects with temporary expression of Cas9. Alzheimer’s disease will likely require a more ubiquitous delivery of Cas9 in the brain. One method that may offer a minimally invasive means of delivering RNPs broadly or to a specific area of the brain is focused ultrasound (FUS) coupled with microbubbles for disrupting the blood-brain-barrier (BBB) (Song et al. 2018). FUS has been used to transiently open the BBB and delivery a variety of macromolecules including RNA, DNA and proteins (Munoz et al. 2018). It may be possible to conjugate RNPs in some form to microbubbles then use FUS to deliver them to specific brain regions (Slagle et al. 2018). Research and development are needed for Cas9 RNPs capable of specifically targeting resident cells of the brain when administered systemically.
Pain and Neuroinflammation
An acute injury often results in the direct activation of peripheral neurons at the site of injury and indirect activation of central neurons that convey sensations of pain. If the pain does not resolve over time, a persistent condition of “chronic pain” can occur that often has debilitating consequences to the affected individual. In the United States in 2016, an estimated 20% of adults (50 million) had chronic pain and 8% of adults (20 million) had high-impact chronic pain (Dahlhamer et al. 2018). One of the primary factors associated with chronic pain is neuroinflammation (Ji et al. 2014). Current approaches to treating chronic pain often involve trying to reduce the associated neuronal activity both peripherally and centrally as well as try to reduce inflammatory processes. For example, non-steroidal anti-inflammatories (NSAIDs) and opioids are routinely used for pain management and work by reducing inflammation and suppressing the sensation of pain. Although these approaches have some efficacy, long term use of prescription opioids causes tolerance which decreases opioid potency and increases user dependence. As a consequence, excessive prescriptions of opioids propagate the current opioid addiction epidemic (Volkow and McLellan 2016). There is an immediate need for alternate strategies to treating chronic pain.
The CRISPR/Cas9 system presents new opportunities to modify the expression of genes involved in neuroinflammation and chronic pain. For example, lentiviral delivery of CRISPR/Cas9 to rat dorsal root ganglion decreased expression the voltage-gated sodium channel Nav1.6 and reduced spontaneous pain and mechanical allodynia (Qin et al. 2017). CRISPR/Cas9 has also been used to create neurofibromatosis type 1 model in rats by intrathecal delivery of plasmid encoding Cas9 and gRNA to neurofibromin (Moutal et al. 2017). CRISPR/Cas9 has also been successfully used to reduce expression of proinflammatory cytokines associated with neuroinflammation as well. We propose that using extracellular vesicles engineered to selectively deliver Cas9/gRNA RNP to knockout modulators of pain transmission and neuroinflammation such as inflammatory mediators and pain-related receptors and channels may provide an alternative treatment to chronic pain. However, the consequences of removing a particular gene or genes from a particular cell type must be thoroughly investigated to ensure the such modifications do not worsen or create new challenges for a patient. More importantly, the use of CRISPR/Cas9 RNPs to treat any human disease through genomic modification must continue to be approached with extreme caution and rigorous pre-clinical evaluation. As research in this area progresses, the use of RNPs for transient delivery Cas9 should be evaluated in models of pain and neuroinflammation.
Enhancements in Cas9 RNP Delivery
A benefit to Cas9 RNP utilization is the transient expression of Cas9, which has been reported to undergo degradation by 24 h using various delivery methods including nucleofection (Kim et al. 2014), cell penetrating peptides (Del'Guidice et al. 2018), and microvesicles (Campbell et al. 2019; Montagna et al. 2018). This is in contrast to DNA based delivery methods, whereby Cas9 is expressed for up to 72 h by plasmid transfection (Kim et al. 2014).
Currently, efforts are being made to further decrease the half-life of Cas9 protein through protein modifications. One group of investigators tagged Cas9 with ubiquitin-proteasomal degradation signals and found significantly decreased Cas9 expression after 24 h using plasmid delivery (Tu et al. 2017). Another group of investigators decreased the half-life of Cas9 in neuronal cells by appending a Geminin tag that is decreased in G1/G0 phase of cell division. They showed that Geminin-tagged Cas9 expression is reduced both in vitro and in vivo after intrastriatal injection (Yang et al. 2018). Future experiments to combine Cas9 RNP delivery mechanisms with these short half-life variants of Cas9 could be used to minimize the duration of Cas9 activity yet still produce a phenotypic effect that would be therapeutically meaningful.
Additionally, several other factors can be improved upon. First, target specificity needs to be increased to optimally deliver Cas9 RNP. One approach is to alter the membrane proteins of the microvesicle by engineering the producer cells to express ligands that promote cell specific interactions and uptake. As our data show, “gesicles” can effectively deliver active protein cargo such as Cre recombinase to a wide variety of cell types (Fig. 4). Additional engineering of this microvesicle to provide cell specific effects would further enhance its viability in future therapeutics. Furthermore, systemic injection of microvesicles containing Cas9 RNP can be performed to ascertain the bioavailability, spread and immune response to this delivery method. A Cas9 dependent fluorescent reporter mouse similar to Staahl et al. (2017) would be a useful model to evaluate Cas9 delivery in vivo.
Microvesicle delivery of Cre recombinase to cell lines of differing tissue origin. Human cell lines of different origins were transfected with a Cre-dependent, nuclear localized, red to green fluorescent protein construct. The presence of GFP denotes functional Cre recombinase delivery and nuclear translocation. Microvesicle “gesicles” containing CherryPicker Red and Cre recombinase were applied to cells and GFP expression was visualized after 24 h. Left panel: brightfield image of all cell lines; Middle panel: Cherry fluorescence denoting both gesicle delivery (small red dots) and red to green transfected cells; Right panel: GFP fluorescence
Concluding Remarks
In this review, we have provided a brief overview of how CRISPR/Cas9 is being used to alter target sequences in the genomes of eukaryotic organisms. We focused on the current methods of Cas9 delivery as a ribonucleoprotein complex to alter gene function with examples related to neuroinflammation. Although using CRISPR/Cas9 can be considered a form of gene therapy, the use of Cas9 RNPs provides a pharmacological approach to gene therapy where the transgenic components used to alter genetic makeup of a cell is present transiently (hours to days) compared to viral or plasmid based gene therapies that can sustain transgene expression for weeks, months and years. The method of Cas9 RNP delivery has huge appeal as a therapeutic including reduced chance of off target effects and decreased chance of developing a strong immune response to the Cas9. The latter concern was recently reviewed as CRISPR/Cas9 delivery by viral vectors continues to be considered for gene therapy applications (Crudele and Chamberlain 2018). Lastly, the engineering of non-viral particles to deliver Cas9 RNPs to specific cells has immense therapeutic potential and should be given consideration when embarking on CRISPR/Cas9-related research. As the field continues to move towards engineered particles for cell-specific targeting, standardized methods for comparing the efficiencies of Cas9 RNP delivery systems are needed at the single cell level if a therapeutically viable approach is to be realized.
References
Aird EJ, Lovendahl KN, St Martin A, Harris RS, Gordon WR (2018) Increasing Cas9-mediated homology-directed repair efficiency through covalent tethering of DNA repair template. Commun Biol 1:54
Almeida JP, Figueroa ER, Drezek RA (2014) Gold nanoparticle mediated cancer immunotherapy. Nanomedicine 10:503–514
Bak RO, Porteus MH (2017) CRISPR-mediated integration of large gene cassettes using AAV donor vectors. Cell Rep 20:750–756
Berger JR (2011) The clinical features of PML. Cleve Clin J Med 78(Suppl 2):S8–S12
Billon P, Bryant EE, Joseph SA, Nambiar TS, Hayward SB, Rothstein R, Ciccia A (2017) CRISPR-Mediated Base editing enables efficient disruption of eukaryotic genes through induction of STOP codons. Mol Cell 67:1068–1079.e4
Campbell LA, Coke LM, Richie CT, Fortuno LV, Park AY, Harvey BK (2019) Gesicle-mediated delivery of CRISPR/Cas9 ribonucleoprotein complex for inactivating the HIV provirus. Mol Ther 27:151–163
Chang N, Sun C, Gao L, Zhu D, Xu X, Zhu X, Xiong JW, Xi JJ (2013) Genome editing with RNA-guided Cas9 nuclease in zebrafish embryos. Cell Res 23:465–472
Chen S, Lee B, Lee AY, Modzelewski AJ, He L (2016) Highly efficient mouse genome editing by CRISPR ribonucleoprotein electroporation of zygotes. J Biol Chem 291:14457–14467
Chen NC, Partridge AT, Sell C, Torres C, Martín-García J (2017) Fate of microglia during HIV-1 infection: from activation to senescence? Glia 65:431–446
Chew WL, Tabebordbar M, Cheng JK, Mali P, Wu EY, Ng AH, Zhu K, Wagers AJ, Church GM (2016) A multifunctional AAV-CRISPR-Cas9 and its host response. Nat Methods 13:868–874
Cho SW, Lee J, Carroll D, Kim JS (2013) Heritable gene knockout in Caenorhabditis elegans by direct injection of Cas9-sgRNA ribonucleoproteins. Genetics 195:1177–1180
Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, Hsu PD, Wu X, Jiang W, Marraffini LA, Zhang F (2013) Multiplex genome engineering using CRISPR/Cas systems. Science 339:819–823
Cradick TJ, Fine EJ, Antico CJ, Bao G (2013) CRISPR/Cas9 systems targeting β-globin and CCR5 genes have substantial off-target activity. Nucleic Acids Res 41:9584–9592
Crudele JM, Chamberlain JS (2018) Cas9 immunity creates challenges for CRISPR gene editing therapies. Nat Commun 9:3497
Dabrowska M, Juzwa W, Krzyzosiak WJ, Olejniczak M (2018) Precise excision of the CAG tract from the huntingtin gene by Cas9 Nickases. Front Neurosci 12:75
Dahlhamer J, Lucas J, Zelaya C, Nahin R, Mackey S, DeBar L, Kerns R, Von Korff M, Porter L, Helmick C (2018) Prevalence of chronic pain and high-impact chronic pain among adults - United States, 2016. MMWR Morb Mortal Wkly Rep 67:1001–1006
Del'Guidice T, Lepetit-Stoffaes JP, Bordeleau LJ, Roberge J, Théberge V, Lauvaux C, Barbeau X, Trottier J, Dave V, Roy DC, Gaillet B, Garnier A, Guay D (2018) Membrane permeabilizing amphiphilic peptide delivers recombinant transcription factor and CRISPR-Cas9/Cpf1 ribonucleoproteins in hard-to-modify cells. PLoS One 13:e0195558
Dionisio-Santos DA, Olschowka JA, O'Banion MK (2019) Exploiting microglial and peripheral immune cell crosstalk to treat Alzheimer's disease. J Neuroinflammation 16:74
Doudna JA, Charpentier E (2014) Genome editing. The new frontier of genome engineering with CRISPR-Cas9. Science 346:1258096
Fu Y, Foden JA, Khayter C, Maeder ML, Reyon D, Joung JK, Sander JD (2013) High-frequency off-target mutagenesis induced by CRISPR-Cas nucleases in human cells. Nat Biotechnol 31:822–826
Gaudelli NM, Komor AC, Rees HA, Packer MS, Badran AH, Bryson DI, Liu DR (2017) Programmable base editing of a•T to G•C in genomic DNA without DNA cleavage. Nature 551:464–471
Gestin M, Dowaidar M, Langel Ü (2017) Uptake mechanism of cell-penetrating peptides. Adv Exp Med Biol 1030:255–264
Ghorpade A, Persidsky Y, Swindells S, Borgmann K, Persidsky R, Holter S, Cotter R, Gendelman HE (2005) Neuroinflammatory responses from microglia recovered from HIV-1-infected and seronegative subjects. J Neuroimmunol 163:145–156
György B, Lööv C, Zaborowski MP, Takeda S, Kleinstiver BP, Commins C, Kastanenka K, Mu D, Volak A, Giedraitis V, Lannfelt L, Maguire CA, Joung JK, Hyman BT, Breakefield XO, Ingelsson M (2018) CRISPR/Cas9 mediated disruption of the Swedish APP allele as a therapeutic approach for early-onset Alzheimer's disease. Mol Ther Nucleic Acids 11:429–440
Hancock RE, Scott MG (2000) The role of antimicrobial peptides in animal defenses. Proc Natl Acad Sci U S A 97:8856–8861
Heaton RK, Franklin DR, Ellis RJ, McCutchan JA, Letendre SL, Leblanc S, Corkran SH, Duarte NA, Clifford DB, Woods SP, Collier AC, Marra CM, Morgello S, Mindt MR, Taylor MJ, Marcotte TD, Atkinson JH, Wolfson T, Gelman BB, McArthur JC, Simpson DM, Abramson I, Gamst A, Fennema-Notestine C, Jernigan TL, Wong J, Grant I (2011) HIV-associated neurocognitive disorders before and during the era of combination antiretroviral therapy: differences in rates, nature, and predictors. J Neuro-Oncol 17:3–16
Hirano H, Gootenberg JS, Horii T, Abudayyeh OO, Kimura M, Hsu PD, Nakane T, Ishitani R, Hatada I, Zhang F, Nishimasu H, Nureki O (2016) Structure and engineering of Francisella novicida Cas9. Cell 164:950–961
Hsu PD, Scott DA, Weinstein JA, Ran FA, Konermann S, Agarwala V, Li Y, Fine EJ, Wu X, Shalem O, Cradick TJ, Marraffini LA, Bao G, Zhang F (2013) DNA targeting specificity of RNA-guided Cas9 nucleases. Nat Biotechnol 31:827–832
Jacobi AM, Rettig GR, Turk R, Collingwood MA, Zeiner SA, Quadros RM, Harms DW, Bonthuis PJ, Gregg C, Ohtsuka M, Gurumurthy CB, Behlke MA (2017) Simplified CRISPR tools for efficient genome editing and streamlined protocols for their delivery into mammalian cells and mouse zygotes. Methods 121-122:16–28
Jeffries AM, Marriott I (2017) Human microglia and astrocytes express cGAS-STING viral sensing components. Neurosci Lett 658:53–56
Ji RR, Xu ZZ, Gao YJ (2014) Emerging targets in neuroinflammation-driven chronic pain. Nat Rev Drug Discov 13:533–548
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E (2012) A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337:816–821
Jinek M, East A, Cheng A, Lin S, Ma E, Doudna J (2013) RNA-programmed genome editing in human cells. Elife 2:e00471
Kalra H, Simpson RJ, Ji H, Aikawa E, Altevogt P, Askenase P, Bond VC, Borràs FE, Breakefield X, Budnik V, Buzas E, Camussi G, Clayton A, Cocucci E, Falcon-Perez JM, Gabrielsson S, Gho YS, Gupta D, Harsha HC, Hendrix A, Hill AF, Inal JM, Jenster G, Krämer-Albers EM, Lim SK, Llorente A, Lötvall J, Marcilla A, Mincheva-Nilsson L, Nazarenko I, Nieuwland R, Nolte-'t Hoen EN, Pandey A, Patel T, Piper MG, Pluchino S, Prasad TS, Rajendran L, Raposo G, Record M, Reid GE, Sánchez-Madrid F, Schiffelers RM, Siljander P, Stensballe A, Stoorvogel W, Taylor D, Thery C, Valadi H, van Balkom BW, Vázquez J, Vidal M, Wauben MH, Yáñez-Mó M, Zoeller M, Mathivanan S (2012) Vesiclepedia: a compendium for extracellular vesicles with continuous community annotation. PLoS Biol 10:e1001450
Kim H, Kim JS (2014) A guide to genome engineering with programmable nucleases. Nat Rev Genet 15:321–334
Kim S, Kim D, Cho SW, Kim J, Kim JS (2014) Highly efficient RNA-guided genome editing in human cells via delivery of purified Cas9 ribonucleoproteins. Genome Res 24:1012–1019
Kim K, Ryu SM, Kim ST, Baek G, Kim D, Lim K, Chung E, Kim S, Kim JS (2017) Highly efficient RNA-guided base editing in mouse embryos. Nat Biotechnol 35:435–437
Komor AC, Zhao KT, Packer MS, Gaudelli NM, Waterbury AL, Koblan LW, Kim YB, Badran AH, Liu DR (2017) Improved base excision repair inhibition and bacteriophage Mu gam protein yields C:G-to-T:A base editors with higher efficiency and product purity. Sci Adv 3:eaao4774
Kumagai H, Nakanishi T, Matsuura T, Kato Y, Watanabe H (2017) CRISPR/Cas-mediated knock-in via non-homologous end-joining in the crustacean Daphnia magna. PLoS One 12:e0186112
Kuscu C, Parlak M, Tufan T, Yang J, Szlachta K, Wei X, Mammadov R, Adli M (2017) CRISPR-STOP: gene silencing through base-editing-induced nonsense mutations. Nat Methods 14:710–712
Laura Soriano M, Zougagh M, Valcárcel M, Ríos Á (2018) Analytical nanoscience and nanotechnology: where we are and where we are heading. Talanta 177:104–121
Lee K, Conboy M, Park HM, Jiang F, Kim HJ, Dewitt MA, Mackley VA, Chang K, Rao A, Skinner C, Shobha T, Mehdipour M, Liu H, Huang WC, Lan F, Bray NL, Li S, Corn JE, Kataoka K, Doudna JA, Conboy I, Murthy N (2017) Nanoparticle delivery of Cas9 ribonucleoprotein and donor DNA. Nat Biomed Eng 1:889–901
Letendre S, Marquie-Beck J, Capparelli E, Best B, Clifford D, Collier AC, Gelman BB, JC MA, JA MC, Morgello S, Simpson D, Grant I, Ellis RJ, Group C (2008) Validation of the CNS penetration-effectiveness rank for quantifying antiretroviral penetration into the central nervous system. Arch Neurol 65:65–70
Lin S, Staahl BT, Alla RK, Doudna JA (2014) Enhanced homology-directed human genome engineering by controlled timing of CRISPR/Cas9 delivery. Elife 3:e04766
Liu J, Gaj T, Yang Y, Wang N, Shui S, Kim S, Kanchiswamy CN, Kim JS, Barbas CF (2015) Efficient delivery of nuclease proteins for genome editing in human stem cells and primary cells. Nat Protoc 10:1842–1859
Mali P, Yang L, Esvelt KM, Aach J, Guell M, DiCarlo JE, Norville JE, Church GM (2013) RNA-guided human genome engineering via Cas9. Science 339:823–826
Mangeot PE, Risson V, Fusil F, Marnef A, Laurent E, Blin J, Mournetas V, Massouridès E, Sohier TJM, Corbin A, Aubé F, Teixeira M, Pinset C, Schaeffer L, Legube G, Cosset FL, Verhoeyen E, Ohlmann T, Ricci EP (2019) Genome editing in primary cells and in vivo using viral-derived Nanoblades loaded with Cas9-sgRNA ribonucleoproteins. Nat Commun 10:45
Mizuno N, Mizutani E, Sato H, Kasai M, Ogawa A, Suchy F, Yamaguchi T, Nakauchi H (2018) Intra-embryo gene cassette Knockin by CRISPR/Cas9-mediated genome editing with adeno-associated viral vector. iScience 9:286–297
Montagna C, Petris G, Casini A, Maule G, Franceschini GM, Zanella I, Conti L, Arnoldi F, Burrone OR, Zentilin L, Zacchigna S, Giacca M, Cereseto A (2018) VSV-G-enveloped vesicles for traceless delivery of CRISPR-Cas9. Mol Ther Nucleic Acids 12:453–462
Mout R, Ray M, Yesilbag Tonga G, Lee YW, Tay T, Sasaki K, Rotello VM (2017) Direct cytosolic delivery of CRISPR/Cas9-ribonucleoprotein for efficient gene editing. ACS Nano 11:2452–2458
Moutal A, Yang X, Li W, Gilbraith KB, Luo S, Cai S, François-Moutal L, Chew LA, Yeon SK, Bellampalli SS, Qu C, Xie JY, Ibrahim MM, Khanna M, Park KD, Porreca F, Khanna R (2017) CRISPR/Cas9 editing of Nf1 gene identifies CRMP2 as a therapeutic target in neurofibromatosis type 1-related pain that is reversed by (S)-Lacosamide. Pain 158:2301–2319
Munoz F, Aurup C, Konofagou EE, Ferrera VP (2018) Modulation of brain function and behavior by focused ultrasound. Curr Behav Neurosci Rep 5:153–164
Murlidharan G, Sakamoto K, Rao L, Corriher T, Wang D, Gao G, Sullivan P, Asokan A (2016) CNS-restricted transduction and CRISPR/Cas9-mediated gene deletion with an engineered AAV vector. Mol Ther Nucleic Acids 5:e338
Nakagawa Y, Sakuma T, Nishimichi N, Yokosaki Y, Yanaka N, Takeo T, Nakagata N, Yamamoto T (2016) Ultra-superovulation for the CRISPR-Cas9-mediated production of gene-knockout, single-amino-acid-substituted, and floxed mice. Biol Open 5:1142–1148
Nishiyama J, Mikuni T, Yasuda R (2017) Virus-mediated genome editing via homology-directed repair in mitotic and Postmitotic cells in mammalian brain. Neuron 96:755–768.e5
Paix A, Folkmann A, Rasoloson D, Seydoux G (2015) High efficiency, homology-directed genome editing in Caenorhabditis elegans using CRISPR-Cas9 ribonucleoprotein complexes. Genetics 201:47–54
Paix A, Folkmann A, Seydoux G (2017) Precision genome editing using CRISPR-Cas9 and linear repair templates in C. elegans. Methods 121-122:86–93
Park DS, Yoon M, Kweon J, Jang AH, Kim Y, Choi SC (2017) Targeted Base editing via RNA-guided cytidine deaminases in Xenopus laevis embryos. Mol Cell 40:823–827
Qin W, Dion SL, Kutny PM, Zhang Y, Cheng AW, Jillette NL, Malhotra A, Geurts AM, Chen YG, Wang H (2015) Efficient CRISPR/Cas9-mediated genome editing in mice by zygote electroporation of nuclease. Genetics 200:423–430
Qin S, Jiang F, Zhou Y, Zhou G, Ye P, Ji Y (2017) Local knockdown of Nav1.6 relieves pain behaviors induced by BmK I. Acta Biochim Biophys Sin Shanghai 49:713–721
Ran FA, Hsu PD, Lin CY, Gootenberg JS, Konermann S, Trevino AE, Scott DA, Inoue A, Matoba S, Zhang Y, Zhang F (2013) Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 154:1380–1389
Ran FA, Cong L, Yan WX, Scott DA, Gootenberg JS, Kriz AJ, Zetsche B, Shalem O, Wu X, Makarova KS, Koonin EV, Sharp PA, Zhang F (2015) In vivo genome editing using Staphylococcus aureus Cas9. Nature 520:186–191
Reinert LS, Lopušná K, Winther H, Sun C, Thomsen MK, Nandakumar R, Mogensen TH, Meyer M, Vægter C, Nyengaard JR, Fitzgerald KA, Paludan SR (2016) Sensing of HSV-1 by the cGAS-STING pathway in microglia orchestrates antiviral defence in the CNS. Nat Commun 7:13348
Robbins PD, Morelli AE (2014) Regulation of immune responses by extracellular vesicles. Nat Rev Immunol 14:195–208
Schnell G, Price RW, Swanstrom R, Spudich S (2010) Compartmentalization and clonal amplification of HIV-1 variants in the cerebrospinal fluid during primary infection. J Virol 84:2395–2407
Schumann K, Lin S, Boyer E, Simeonov DR, Subramaniam M, Gate RE, Haliburton GE, Ye CJ, Bluestone JA, Doudna JA, Marson A (2015) Generation of knock-in primary human T cells using Cas9 ribonucleoproteins. Proc Natl Acad Sci U S A 112:10437–10442
Shalem O, Sanjana NE, Hartenian E, Shi X, Scott DA, Mikkelson T, Heckl D, Ebert BL, Root DE, Doench JG, Zhang F (2014) Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343:84–87
Shen S, Sanchez ME, Blomenkamp K, Corcoran EM, Marco E, Yudkoff CJ, Jiang H, Teckman JH, Bumcrot D, Albright CF (2018) Amelioration of Alpha-1 antitrypsin deficiency diseases with genome editing in transgenic mice. Hum Gene Ther 29:861–873
Skog J, Würdinger T, van Rijn S, Meijer DH, Gainche L, Sena-Esteves M, Curry WT, Carter BS, Krichevsky AM, Breakefield XO (2008) Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nat Cell Biol 10:1470–1476
Slagle CJ, Thamm DH, Randall EK, Borden MA (2018) Click conjugation of cloaked peptide ligands to microbubbles. Bioconjug Chem 29:1534–1543
Song KH, Harvey BK, Borden MA (2018) State-of-the-art of microbubble-assisted blood-brain barrier disruption. Theranostics 8:4393–4408
Staahl BT, Benekareddy M, Coulon-Bainier C, Banfal AA, Floor SN, Sabo JK, Urnes C, Munares GA, Ghosh A, Doudna JA (2017) Efficient genome editing in the mouse brain by local delivery of engineered Cas9 ribonucleoprotein complexes. Nat Biotechnol 35:431–434
Sun J, Carlson-Stevermer J, Das U, Shen M, Delenclos M, Snead AM, Koo SY, Wang L, Qiao D, Loi J, Petersen AJ, Stockton M, Bhattacharyya A, Jones MV, Zhao X, McLean PJ, Sproul AA, Saha K, Roy S (2019) CRISPR/Cas9 editing of APP C-terminus attenuates β-cleavage and promotes α-cleavage. Nat Commun 10:53
Swiech L, Heidenreich M, Banerjee A, Habib N, Li Y, Trombetta J, Sur M, Zhang F (2015) In vivo interrogation of gene function in the mammalian brain using CRISPR-Cas9. Nat Biotechnol 33:102–106
Teixeira M, Py BF, Bosc C, Laubreton D, Moutin MJ, Marvel J, Flamant F, Markossian S (2018) Electroporation of mice zygotes with dual guide RNA/Cas9 complexes for simple and efficient cloning-free genome editing. Sci Rep 8:474
Tu Z, Yang W, Yan S, Yin A, Gao J, Liu X, Zheng Y, Zheng J, Li Z, Yang S, Li S, Guo X, Li XJ (2017) Promoting Cas9 degradation reduces mosaic mutations in non-human primate embryos. Sci Rep 7:42081
van Niel G, D'Angelo G, Raposo G (2018) Shedding light on the cell biology of extracellular vesicles. Nat Rev Mol Cell Biol 19:213–228
Volkow ND, McLellan AT (2016) Opioid abuse in chronic pain--misconceptions and mitigation strategies. N Engl J Med 374:1253–1263
Wang H, Yang H, Shivalila CS, Dawlaty MM, Cheng AW, Zhang F, Jaenisch R (2013) One-step generation of mice carrying mutations in multiple genes by CRISPR/Cas-mediated genome engineering. Cell 153:910–918
Wang T, Wei JJ, Sabatini DM, Lander ES (2014) Genetic screens in human cells using the CRISPR-Cas9 system. Science 343:80–84
Wollebo HS, Bellizzi A, Kaminski R, Hu W, White MK, Khalili K (2015) CRISPR/Cas9 system as an agent for eliminating polyomavirus JC infection. PLoS One 10:e0136046
Wu CM, Roth TL, Baglaenko Y, Ferri DM, Brauer P, Zuniga-Pflucker JC, Rosbe KW, Wither JE, Marson A, Allen CDC (2018) Genetic engineering in primary human B cells with CRISPR-Cas9 ribonucleoproteins. J Immunol Methods 457:33–40
Yamaguchi H, Hopf FW, Li SB, de Lecea L (2018) In vivo cell type-specific CRISPR knockdown of dopamine beta hydroxylase reduces locus coeruleus evoked wakefulness. Nat Commun 9:5211
Yang S, Li S, Li XJ (2018) Shortening the half-life of Cas9 maintains its gene editing ability and reduces neuronal toxicity. Cell Rep 25:2653–2659.e3
Zallar LJ, Tunstall BJ, Richie CT, Zhang YJ, You ZB, Gardner EL, Heilig M, Pickel J, Koob GF, Vendruscolo LF, Harvey BK, Leggio L (2018) Development and initial characterization of a novel ghrelin receptor CRISPR/Cas9 knockout wistar rat model. Int J Obes
Zasloff M (2002) Antimicrobial peptides of multicellular organisms. Nature 415:389–395
Zhou L, Saksena NK (2013) HIV associated neurocognitive disorders. Infect Dis Rep 5:e8
Acknowledgements
This research was supported by the Intramural Research Program of the NIH, NIDA.
Funding
This study was funded by the National Institute on Drug Abuse Intramural Research Program
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare no conflict of interest.
Animal Studies
All animal studies were carried out following the Guide for the Care and Use of Laboratory Animals as adopted and promulgated by the U.S. National Institutes of Health and approved by the National Institute on Drug Abuse IRP internal review board.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Campbell, L.A., Richie, C.T., Maggirwar, N.S. et al. Cas9 Ribonucleoprotein Complex Delivery: Methods and Applications for Neuroinflammation. J Neuroimmune Pharmacol 14, 565–577 (2019). https://doi.org/10.1007/s11481-019-09856-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11481-019-09856-z