Abstract
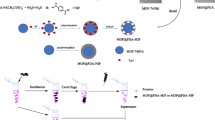
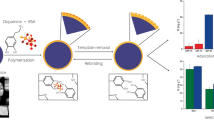
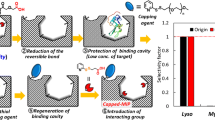
Here, we introduce a new type of molecular imprinted polymer (MIP) with immobilized assistant recognition polymer chains (ARPCs) to create effective recognition sites and with bacterial cloned protein as template for adsorbing the low content target protein from cell extract. In this work, cloned pig cyclophilin 18 (pCyP18), a peptidyl-prolyl cis/trans-isomerase, was used as template. The template protein was selectively assembled with ARPCs from their library, which consists of numerous limited length polymer chains with randomly distributed recognition sites of positively charged amino groups and immobilizing sites. These assemblies were adsorbed by porous microsphers and immobilized on them. After removing the template, binding sites complementary to the target protein in size, shape and the position of recognition groups were exposed, and their confirmation was preserved by the cross-linked structure. The synthesized MIP was used to adsorb the cellular pCyP18, and its proportional content was enriched more than hundred times. The extended experiment on imprinting bovine serum albumin (BSA) with ARPCs shows that this method is also suitable for large protein.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Anderson L, Sellergren B, Mosbach K. Imprinting of amino acid derivatives in macroporous polymers. Tetrahedron Lett, 1984, 25: 5211–5214
Vlatakis G, Andersson L I, Mueller R et al. Drug assay using antibody mimics made by molecular imprinting. Nature, 1993, 361: 645–647
Wulff G. Molecular imprinting in cross-linked materials with the aid of molecular templates-a way towards artificial antibodies. Angew Chem Int Ed Engl, 1995, 34: 1812–1832
Wulff G. Enzyme-like catalysis by molecularly imprinted polymers. Chem Rev, 2002, 102: 1–27
Shi F, Liu Z, Wu G L et al. Surface imprinting in layer-by-layer nanostructured films. Advanced Functional Materials, 2007, 17(11): 1821–1827
Bossi A, Piletsky S A, Piletska E V et al. Surface-grafted molecularly imprinted polymers for protein recognition. Anal Chem, 2001, 73: 5281–5286
Venton D L, Gudipati E. Influence of protein on polysiloxane polymer formation: Evidence for induction of complementary protein-polymer interaction. Biochim Biophys Acta, 1995, 1250: 126–136
Hjèrten J, Liao J L, Nakazato K, et al. Gels mimicking antibodies in there selective recognition of proteins. Chromatography, 1997, 44: 227–234
Kempe M, Glad M, Mosbach K. An approach towards surface imprinting using the enzyme ribonuclease A. J Molrecogn, 1995, 8: 35–39
Burow, M, Minoura, N. Molecular imprinting: Synthesis of polymer particles with antibody-like binding characteristics for glucose oxidase. Biochem Biophys Res Commun, 1996, 227: 419–422
Hirayama K, Burow M, Morikawa Y, et al. Synthesis of polymercoated silica particles with specific recognition sites for glucose oxidase by the molecular imprinting technique. Chem Lett, 1998, 8: 731–732
Guo T Y, Xia Y Q, Hao G J, et al. Adsorptive separation of hemoglobin by molecularly imprinted chitosan beads. Biomaterials, 2004, 25: 5905–5912
Shi H, Tsai W B, Ferrari S, et al. Templateimprinted nanostructured surfaces for protein recognition. Nature, 1999, 398: 593–597
Rachkov A, Minoura N. Recognition of oxytocin and oxytocin-related peptides in aqueous media using a molecularly imprinted polymer synthesized by the epitope approach. J Chromatography A, 2000, 889: 111–118
Nishino H, Huang C S, Shea K J. Selective protein capture by epitope imprinting. Angew Chem Int Ed, 2006, 45: 2392–2396
Zhao Z, Wang C H, Guo M J, et al. Molecular imprinted polymer with cloned bacterial protein template enriches authentic target in cell extract. FEBS Lett, 2006, 580: 2750–2754
Haupt K. Creating a good impression. Nat Biotechnol, 2002, 20: 884–885
Guo M J, Zhao Z, Fan Y G, et al. Protein-imprinted polymer with immobilized assistant recognition polymer chains. Biomaterials, 2006, 27: 4381–4387
Handschumacher R E, Harding M W, Rice J, et al. Cyclophilin: A specific cytosolic binding protein for cyclosporin A. Science, 1984, 226: 544–547
Ke H, Zydowsky L D, Liu J, et al. Crystal structure of recombinant human T-cell cyclophilin A at 2.5 A resolution. Proc Natl Acad Sci USA, 1991, 88: 9483–9487
Nowick J S, Lam K S, Khasanova T V, et al. An unnatural amino acid that induces β-sheet folding and interaction in peptides. J Am Chem Soc, 2004, 124: 4972–4973
Zhen Y, Wan S, Liu Y, et al. A series of polymer-clay nanocomposites were prepared by the simultaneous. Macromol Chem Phys, 2005, 206: 607–612
Wang Y, Han R F, Wu D, et al. The binding of FKBP23 to BiP modulates BiP’s ATPase activity with its PPIase activity. BBRC, 2007, 354: 315–320
Bloom H, Beier H, Gross H. Improved silver staining of plant proteins, RNA and DNA in polyacrylamide gels. Electrophoresis, 1987, 8: 93–99
Borgeson C E, Bowman B J. Isolation and characterization of the Neurospora crassa endoplasmic reticulum. J Bacteriol, 1983, 156: 362–368
Chen C, Ma H, Wang Y, et al. Binding of FKBP23 to BiP in ER shown by gel filtration chromatograhy. Z Naturforsch, 2007, 62c: 133–137
Zhang X B, Wang Y, Li H, et al. The mouse FKBP23 binds to BiP in ER and the binding of C-terminal domain is interrelated with Ca2+ concentration. FEBS Lett, 2004, 559: 57–60
Bradford M M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein dye binding. Anal Biochem, 1976, 72: 248–254
Niu J, Shi F, Liu Z, et al. Reversible Disulfide cross-linking in layerby-layer films: Preassembly enhanced loading and pH/reductant dually controllable release. Langmuir, 2007, 23(11): 6377–6384
Author information
Authors and Affiliations
Corresponding author
Additional information
Supported by the National Natural Science Foundation of China (Grant No. 20674040) and National Basic Research Program of China (Grant No. 2006CB705703)
About this article
Cite this article
Long, Y., Sun, Y., Wang, Y. et al. Molecular imprinted polymer with positively charged assistant recognition polymer chains for adsorption/enrichment of low content target protein. Chin. Sci. Bull. 53, 2617–2623 (2008). https://doi.org/10.1007/s11434-008-0374-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11434-008-0374-6