Abstract
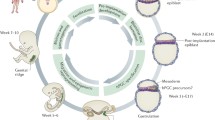
Primordial germ cells (PGCs) are precursors of both male and female gametes as fundamental materials for organism development. The transcriptome, methylome, and chromatin accessibility profiles of PGCs in both mice and humans have been recently reported. However, little is known about the characteristics of PGCs at the protein levels, which directly exert cellular functions. Here, we construct landscapes of both proteome and 3D spatial distribution of mouse PGCs at E11.5, E13.5 and E16.5 days, the three critical developmental windows for PGCs' sex differentiation, female meiosis initiation and male mitotic arrest. In each developmental stage of PGCs, nearly 2,000–3,000 proteins are identified, among which specific functional pathways such as oxidative phosphorylation, DNA damage repair, and meiotic cell cycle are involved for different events during PGCs development. Interestingly, by 3D modeling we find that PGCs spatially cluster into around 1,300 nests in genital ridge at E11.5 and the nest number is not increased by the exponential proliferation of PGCs. Comparative analysis of our proteomic data with published transcriptomic data does not show a close correlation, meaning that the practically executive factors are beyond the transcriptome. Thus, our work offers a valuable resource for the systematic investigations of PGC development at protein level and spatial map.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Agarwal, A., Mulgund, A., Hamada, A., and Chyatte, M.R. (2015). A unique view on male infertility around the globe. Reprod Biol Endocrinol 13, 37.
Bhattacharyya, T., Walker, M., Powers, N.R., Brunton, C., Fine, A.D., Petkov, P.M., and Handel, M.A. (2019). Prdm9 and meiotic cohesin proteins cooperatively promote DNA double-strand break formation in mammalian spermatocytes. Curr Biol 29, 1002–1018.e7.
Brennan, J., and Capel, B. (2004). One tissue, two fates: molecular genetic events that underlie testis versus ovary development. Nat Rev Genet 5, 509–521.
Ewen, K.A., and Koopman, P. (2010). Mouse germ cell development: from specification to sex determination. Mol Cell Endocrinol 323, 76–93.
Gkountela, S., Li, Z., Vincent, J.J., Zhang, K.X., Chen, A., Pellegrini, M., and Clark, A.T. (2013). The ontogeny of cKIT+ human primordial germ cells proves to be a resource for human germ line reprogramming, imprint erasure and in vitro differentiation. Nat Cell Biol 15, 113–122.
Godin, I., Wylie, C., and Heasman, J. (1990). Genital ridges exert long-range effects on mouse primordial germ cell numbers and direction of migration in culture. Development 108, 357–363.
Guo, F., Yan, L., Guo, H., Li, L., Hu, B., Zhao, Y., Yong, J., Hu, Y., Wang, X., Wei, Y., et al. (2015). The transcriptome and DNA methylome landscapes of human primordial germ cells. Cell 161, 1437–1452.
Guo, H., Hu, B., Yan, L., Yong, J., Wu, Y., Gao, Y., Guo, F., Hou, Y., Fan, X., Dong, J., et al. (2017). DNA methylation and chromatin accessibility profiling of mouse and human fetal germ cells. Cell Res 27, 165–183.
Hayashi, K., de Sousa Lopes, S.M.C., and Surani, M.A. (2007). Germ cell specification in mice. Science 316, 394–396.
Hayashi, Y., Otsuka, K., Ebina, M., Igarashi, K., Takehara, A., Matsumoto, M., Kanai, A., Igarashi, K., Soga, T., and Matsui, Y. (2017). Distinct requirements for energy metabolism in mouse primordial germ cells and their reprogramming to embryonic germ cells. Proc Natl Acad Sci USA 114, 8289–8294.
Hikabe, O., Hamazaki, N., Nagamatsu, G., Obata, Y., Hirao, Y., Hamada, N., Shimamoto, S., Imamura, T., Nakashima, K., Saitou, M., et al. (2016). Reconstitution in vitro of the entire cycle of the mouse female germ line. Nature 539, 299–303.
Hill, P.W.S., Leitch, H.G., Requena, C.E., Sun, Z., Amouroux, R., Roman-Trufero, M., Borkowska, M., Terragni, J., Vaisvila, R., Linnett, S., et al. (2018). Epigenetic reprogramming enables the transition from primordial germ cell to gonocyte. Nature 555, 392–396.
Jameson, S.A., Natarajan, A., Cool, J., DeFalco, T., Maatouk, D.M., Mork, L., Munger, S.C., and Capel, B. (2012). Temporal transcriptional profiling of somatic and germ cells reveals biased lineage priming of sexual fate in the fetal mouse gonad. PLoS Genet 8, e1002575.
Jiao, X., Ke, H., Qin, Y., and Chen, Z.J. (2018). Molecular genetics of premature ovarian insufficiency. Trends Endocrinol Metab 29, 795–807.
Katari, S., Aarabi, M., Kintigh, A., Mann, S., Yatsenko, S.A., Sanfilippo, J. S., Zeleznik, A.J., and Rajkovic, A. (2018). Chromosomal instability in women with primary ovarian insufficiency. Human Reprod 33, 531–538.
Krämer, A., Green, J., Pollard Jr, J., and Tugendreich, S. (2014). Causal analysis approaches in Ingenuity Pathway Analysis. Bioinformatics 30, 523–530.
Kumar, L., and E Futschik, M. (2007). Mfuzz: a software package for soft clustering of microarray data. Bioinformation 2, 5–7.
Lei, L., and Spradling, A.C. (2013). Mouse primordial germ cells produce cysts that partially fragment prior to meiosis. Development 140, 2075–2081.
Li, W., Germain, R.N., and Gerner, M.Y. (2019). High-dimensional cell-level analysis of tissues with Ce3D multiplex volume imaging. Nat Protoc 14, 1708–1733.
Liao, Y., Wang, J., Jaehnig, E.J., Shi, Z., and Zhang, B. (2019). WebGestalt 2019: gene set analysis toolkit with revamped UIs and APIs. Nucleic Acids Res 47, W199–W205.
Lin, Y., Gill, M.E., Koubova, J., and Page, D.C. (2008). Germ cell-intrinsic and -extrinsic factors govern meiotic initiation in mouse embryos. Science 322, 1685–1687.
Martínez-Arroyo, A.M., Medrano, J.V., Remohí, J., and Simón, C. (2014). Germ line development: lessons learned from pluripotent stem cells. Curr Opin Genet Dev 28, 64–70.
Matsuoka, S., Ballif, B.A., Smogorzewska, A., McDonald, E.R., Hurov, K. E., Luo, J., Bakalarski, C.E., Zhao, Z., Solimini, N., Lerenthal, Y., et al. (2007). ATM and ATR substrate analysis reveals extensive protein networks responsive to DNA damage. Science 316, 1160–1166.
Pacheco, S., Maldonado-Linares, A., Marcet-Ortega, M., Rojas, C., Martínez-Marchal, A., Fuentes-Lazaro, J., Lange, J., Jasin, M., Keeney, S., Fernández-Capetillo, O., et al. (2018). ATR is required to complete meiotic recombination in mice. Nat Commun 9, 2622.
Paigen, K., and Petkov, P. (2010). Mammalian recombination hot spots: properties, control and evolution. Nat Rev Genet 11, 221–233.
Pepling, M.E., and Spradling, A.C. (2001). Mouse ovarian germ cell cysts undergo programmed breakdown to form primordial follicles. Dev Biol 234, 339–351.
Sabour, D., Araúzo-Bravo, M.J., Hübner, K., Ko, K., Greber, B., Gentile, L., Stehling, M., and Schöler, H.R. (2011). Identification of genes specific to mouse primordial germ cells through dynamic global gene expression. Hum Mol Genet 20, 115–125.
Saitou, M., and Yamaji, M. (2010). Germ cell specification in mice: signaling, transcription regulation, and epigenetic consequences. REPRODUCTION 139, 931–942.
Saitou, M., and Yamaji, M. (2012). Primordial germ cells in mice. Cold Spring Harb Perspect Biol 4, a008375.
Seisenberger, S., Andrews, S., Krueger, F., Arand, J., Walter, J., Santos, F., Popp, C., Thienpont, B., Dean, W., and Reik, W. (2012). The dynamics ofgenome-wide DNA methylation reprogramming in mouse primordial germ cells. Mol Cell 48, 849–862.
Shen, C., Li, M., Zhang, P., Guo, Y., Zhang, H., Zheng, B., Teng, H., Zhou, T., Guo, X., and Huo, R. (2018). A comparative proteome profile of female mouse gonads suggests a tight link between the electron transport chain and meiosis initiation. Mol Cell Proteomics 17, 31–42.
Shiloh, Y. (2001). ATM and ATR: networking cellular responses to DNA damage. Curr Opin Genet Dev 11, 71–77.
Strome, S., and Updike, D. (2015). Specifying and protecting germ cell fate. Nat Rev Mol Cell Biol 16, 406–416.
Subramanian, A., Tamayo, P., Mootha, V.K., Mukherjee, S., Ebert, B.L., Gillette, M.A., Paulovich, A., Pomeroy, S.L., Golub, T.R., Lander, E.S., et al. (2005). Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102, 15545–15550.
Sung, P., Krejci, L., Van Komen, S., and Sehorn, M.G. (2003). Rad51 recombinase and recombination mediators. J Biol Chem 278, 42729–42732.
Sybirna, A., Wong, F.C.K., and Surani, M.A. (2019). Genetic basis for primordial germ cells specification in mouse and human: Conserved and divergent roles of PRDM and SOX transcription factors. Curr Top Dev Biol 135, 35–89.
Tam, P.P., and Snow, M.H. (1981). Proliferation and migration of primordial germ cells during compensatory growth in mouse embryos. J Embryol Exp Morphol 64, 133–147.
Tan, H., and Tee, W.W. (2019). Committing the primordial germ cell: An updated molecular perspective. WIREs Syst Biol Med 11, e1436.
Wen, L., Liu, Q., Xu, J., Liu, X., Shi, C., Yang, Z., Zhang, Y., Xu, H., Liu, J., Yang, H., et al. (2020). Recent advances in mammalian reproductive biology. Sci China Life Sci 63, 18–58.
Widger, A., Mahadevaiah, S.K., Lange, J., ElInati, E., Zohren, J., Hirota, T., Pacheco, S., Maldonado-Linares, A., Stanzione, M., Ojarikre, O., et al. (2018). ATR is a multifunctional regulator of male mouse meiosis. Nat Commun 9, 2621.
Yamaguchi, S., Hong, K., Liu, R., Inoue, A., Shen, L., Zhang, K., and Zhang, Y. (2013). Dynamics of 5-methylcytosine and 5-hydroxymethylcytosine during germ cell reprogramming. Cell Res 23, 329–339.
Yokobayashi, S., Liang, C.Y., Kohler, H., Nestorov, P., Liu, Z., Vidal, M., van Lohuizen, M., Roloff, T.C., and Peters, A.H.F.M. (2013). PRC1 coordinates timing of sexual differentiation of female primordial germ cells. Nature 495, 236–240.
Zhou, Z., Wang, L., Ge, F., Gong, P., Wang, H., Wang, F., Chen, L., and Liu, L. (2018). Pold3 is required for genomic stability and telomere integrity in embryonic stem cells and meiosis. Nucleic Acids Res 46, 3468–3486.
Acknowledgements
This work was supported by the National Key Research and Development Program of China (2017YFC1001501, 2016YFC1000604, 2018YFC1004002) and the Foundation for Innovative Research Groups of National Natural Science Foundation of China (81521002).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Compliance and ethics
The author(s) declare that they have no conflict of interest. The institutional and national guide for the care and use of laboratory animals was followed.
Electronic Supplementary Material
Rights and permissions
About this article
Cite this article
Wang, P., Miao, Y., Li, XH. et al. Proteome landscape and spatial map of mouse primordial germ cells. Sci. China Life Sci. 64, 966–981 (2021). https://doi.org/10.1007/s11427-020-1762-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-020-1762-2