Abstract
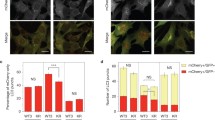
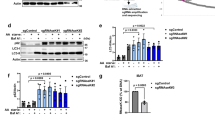
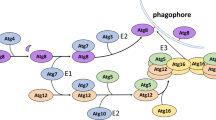
Autophagy is an evolutionarily conserved lysosome-based degradation process. Atg5 plays a very important role in autophagosome formation. Here we show that Atg5 is required for biogenesis of late endosomes and lysosomes in an autophagy-independent manner. In Atg5−/− cells, but not in other essential autophagy genes defecting cells, recycling and retrieval of late endosomal components from hybrid organelles are impaired, causing persistent hybrid organelles and defective formation of late endosomes and lysosomes. Defective retrieval of late endosomal components from hybrid organelles resulting from impaired recruitment of a component of V1-ATPase to acidic organelles blocks the pH-dependent retrieval of late endosomal components from hybrid organelles. Lowering the intracellular pH restores late endosome/lysosome biogenesis in Atg5−/− cells. Our data demonstrate an unexpected role of Atg5 and shed new light on late endosome and lysosome biogenesis.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Luzio JP, Mullock BM, Pryor PR, Lindsay MR, James DE, Piper RC. Relationship between endosomes and lysosomes. Biochem Soc T, 2001, 29: 476–480
Luzio JP, Rous BA, Bright NA, Pryor PR, Mullock BM, Piper RC. Lysosome-endosome fusion and lysosome biogenesis. J Cell Sci, 2000, 113: 1515–1524
Gruenberg J, Stenmark H. The biogenesis of multivesicular endosomes. Nat Rev Mol Cell Biol, 2004, 5: 317–323
Luzio JP, Pryor PR, Bright NA. Lysosomes: fusion and function. Nat Rev Mol Cell Biol, 2007, 8: 622–632
Luzio JP, Rous BA, Bright NA, Pryor PR, Mullock BM, Piper RC. Lysosome-endosome fusion and lysosome biogenesis. J Cell Sci, 2000, 113(Pt 9): 1515–1524
Saftig P, Klumperman J. Lysosome biogenesis and lysosomal membrane proteins: trafficking meets function. Nat Rev Mol Cell Biol, 2009, 10: 623–635
Bright NA, Gratian MJ, Luzio JP. Endocytic delivery to lysosomes mediated by concurrent fusion and kissing events in living cells. Curr Biol: CB, 2005, 15: 360–365
Luzio JP, Pryor PR, Gray SR, Gratian MJ, Piper RC, Bright NA. Membrane traffic to and from lysosomes. Biochem Soc Symp, 2005, 77-86
Klionsky DJ, Emr SD. Autophagy as a regulated pathway of cellular degradation. Science, 2000, 290: 1717–1721
Mizushima N. Autophagy: process and function. Genes Dev, 2007, 21: 2861–2873
Xie Z, Klionsky DJ. Autophagosome formation: core machinery and adaptations. Nat Cell Biol, 2007, 9: 1102–1109
Tian Y, Li Z, Hu W, Ren H, Tian E, Zhao Y, Lu Q, Huang X, Yang P, Li X, Wang X, Kovács AL, Yu L, Zhang H. C. elegans screen identifies autophagy genes specific to multicellular organisms. Cell, 2010, 141: 1042–1055
Levine B, Mizushima N, Virgin HW. Autophagy in immunity and inflammation. Nature, 2011, 469: 323–335
Deretic V. Links between autophagy, innate immunity, inflammation and Crohn’s disease. Dig Dis, 2009, 27: 246–251
Mizushima N, Noda T, Yoshimori T, Tanaka Y, Ishii T, George MD, Klionsky DJ, Ohsumi M, Ohsumi Y. A protein conjugation system essential for autophagy. Nature, 1998, 395: 395–398
Codogno P, Meijer AJ. Atg5: more than an autophagy factor. Nat Cell Biol, 2006, 8: 1045–1047
Hirst J, Futter CE, Hopkins CR. The kinetics of mannose 6-phosphate receptor trafficking in the endocytic pathway in HEp-2 cells: the receptor enters and rapidly leaves multivesicular endosomes without accumulating in a prelysosomal compartment. Mol Biol Cell, 1998, 9: 809–816
Wassmer T, Attar N, Harterink M, van Weering JR, Traer CJ, Oakley J, Goud B, Stephens DJ, Verkade P, Korswagen HC, Cullen PJ. The retromer coat complex coordinates endosomal sorting and dynein-mediated transport, with carrier recognition by the trans-Golgi network. Dev Cell, 2009, 17: 110–122
Cullen PJ, Korswagen HC. Sorting nexins provide diversity for retromer-dependent trafficking events. Nat Cell Biol, 2012, 14: 29–37
van Weering JR, Verkade P, Cullen PJ. SNX-BAR-mediated endosome tubulation is coordinated with endosome maturation. Traffic, 2012, 13: 94–107
Kane PM. Disassembly and reassembly of the yeast vacuolar H(+)-ATPase in vivo. J Biol Chem, 1995, 270: 17025–17032
Kane PM, Parra KJ. Assembly and regulation of the yeast vacuolar H(+)-ATPase. J Exp Biol, 2000, 203: 81–87
Parton RG, Dotti CG, Bacallao R, Kurtz I, Simons K, Prydz K. pH-induced microtubule-dependent redistribution of late endosomes in neuronal and epithelial cells. J Cell Biol, 1991, 113: 261–274
Burkhardt JK, Wiebel FA, Hester S, Argon Y. The giant organelles in beige and Chediak-Higashi fibroblasts are derived from late endosomes and mature lysosomes. J Exp Med, 1993, 178: 1845–1856
Introne W, Boissy RE, Gahl WA. Clinical, molecular, and cell biological aspects of Chediak-Higashi syndrome. Mol Genet Metab, 1999, 68: 283–303
Barbosa MD, Nguyen QA, Tchernev VT, Ashley JA, Detter JC, Blaydes SM, Brandt SJ, Chotai D, Hodgman C, Solari RC, Lovett M, Kingsmore SF. Identification of the homologous beige and Chediak-Higashi syndrome genes. Nature, 1996, 382: 262–265
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is published with open access at springerlink.bibliotecabuap.elogim.com
Electronic supplementary material
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 2.0 International License (https://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Peng, J., Zhang, R., Cui, Y. et al. Atg5 regulates late endosome and lysosome biogenesis. Sci. China Life Sci. 57, 59–68 (2014). https://doi.org/10.1007/s11427-013-4588-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-013-4588-8