Abstract
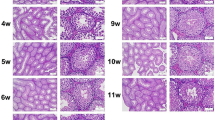
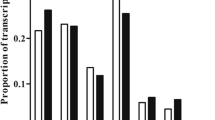
Mammalian testis development is a complex and highly sophisticated process. To study the dynamic change of normal testis development at the transcriptional level, we investigated mouse testes at three postnatal ages: 6 days postnatal, 4 weeks old, and 10 weeks old, representing infant (PN1), juvenile (PN2), and adult (PN3) stages, respectively. Using ultra high-throughput RNA sequencing (RNA-seq) technology, we obtained 211 million reads with a length of 35 bp. We identified 18837 genes that were expressed in mouse testes, and found that genes expressed at the highest level were involved in spermatogenesis. The gene expression pattern in PN1 was distinct from that in PN2 and PN3, which indicates that spermatogenesis has commenced in PN2. We analyzed a large number of genes related to spermatogenesis and somatic development of the testis, which play important roles at different developmental stages. We also found that the MAPK, Hedgehog, and Wnt signaling pathways were significantly involved at different developmental stages. These findings further our understanding of the molecular mechanisms that regulate testis development. Our study also demonstrates significant advantages of RNA-seq technology for studying transcriptome during development.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
McCarrey J R, O’Brien D A, Skinner M K. Construction and preliminary characterization of a series of mouse and rat testis cDNA libraries. J Androl, 1999, 20: 635–639
Su A I, Cooke M P, Ching K A, et al. Large-scale analysis of the human and mouse transcriptomes. Proc Natl Acad Sci USA, 2002, 99: 4465–4470
Tanaka K, Tamura H, Tanaka H, et al. Spermatogonia-dependent expression of testicular genes in mice. Dev Biol, 2002, 246: 466–479
Rockett J C, Christopher Luft J, Brian Garges J, et al. Development of a 950-gene DNA array for examining gene expression patterns in mouse testis. Genome Biol, 2001, 2: RESEARCH0014
Sha J, Zhou Z, Li J, et al. Identification of testis development and spermatogenesis-related genes in human and mouse testes using cDNA arrays. Mol Hum Reprod, 2002, 8: 511–517
Pang A L, Taylor H C, Johnson W, et al. Identification of differentially expressed genes in mouse spermatogenesis. J Androl, 2003, 24: 899–911
Pang A L, Johnson W, Ravindranath N, et al. Expression profiling of purified male germ cells: stage-specific expression patterns related to meiosis and postmeiotic development. Physiol Genomics, 2006, 24: 75–85
Yu Z, Guo R, Ge Y, et al. Gene expression profiles in different stages of mouse spermatogenic cells during spermatogenesis. Biol Reprod, 2003, 69: 37–47
Guo R, Yu Z, Guan J, et al. Stage-specific and tissue-specific expression characteristics of differentially expressed genes during mouse spermatogenesis. Mol Reprod Dev, 2004, 67: 264–272
Clemente E J, Furlong R A, Loveland K L, et al. Gene expression study in the juvenile mouse testis: identification of stage-specific molecular pathways during spermatogenesis. Mamm Genome, 2006, 17: 956–975
Schultz N, Hamra F K, Garbers D L. A multitude of genes expressed solely in meiotic or postmeiotic spermatogenic cells offers a myriad of contraceptive targets. Proc Natl Acad Sci USA, 2003, 100: 12201–12206
Shima J E, McLean D J, McCarrey J R, et al. The murine testicular transcriptome: characterizing gene expression in the testis during the progression of spermatogenesis. Biol Reprod, 2004, 71: 319–330
O’shaughnessy P J, Fleming L, Baker P J, et al. Identification of developmentally regulated genes in the somatic cells of the mouse testis using serial analysis of gene expression. Biol Reprod, 2003, 69: 797–808
Wu S M, Baxendale V, Chen Y, et al. Analysis of mouse germ-cell transcriptome at different stages of spermatogenesis by SAGE: Biological significance. Genomics, 2004, 84: 971–981
Yao J, Chiba T, Sakai J, et al. Mouse testis transcriptome revealed using serial analysis of gene expression. Mamm Genome, 2004, 15: 433–451
Almstrup K. Analysis of cell-type-specific gene expression during mouse spermatogenesis. Biol Reprod, 2004, 70: 1751–1761
Ellis P J, Furlong R A, Wilson A, et al. Modulation of the mouse testis transcriptome during postnatal development and in selected models of male infertility. Mol Hum Reprod, 2004, 10: 271–281
Divina P, Vlcek C, Strnad P, et al. Global transcriptome analysis of the C57BL/6J mouse testis by SAGE: evidence for nonrandom gene order. BMC Genomics, 2005, 6: 29
Chan W Y, Lee T L, Wu S M, et al. Transcriptome analyses of male germ cells with serial analysis of gene expression (SAGE). Mol Cell Endocrinol, 2006, 250: 8–19
Lee T L, Cheung H H, Claus J, et al. GermSAGE: a comprehensive SAGE database for transcript discovery on male germ cell development. Nucleic Acids Res, 2009, 37: D891–897
Ike A, Tokuhiro K, Hirose M, et al. Comprehensive analysis of gene expression in testes producing haploid germ cells using DNA microarray analysis. Int J Androl, 2007, 30: 462–475
Xiao P, Tang A, Yu Z, et al. Gene expression profile of 2058 spermatogenesis-related genes in mice. Biol Pharm Bull, 2008, 31: 201–206
Waldman Ben-Asher H, Shahar I, Yitzchak A, et al. Expression and chromosomal organization of mouse meiotic genes. Mol Reprod Dev, 2010, 77: 241–248
Mortazavi A, Williams B A, McCue K, et al. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods, 2008, 5: 621–628
Wang Z, Gerstein M, Snyder M. RNA-Seq: a revolutionary tool for transcriptomics. Nat Rev Genet, 2009, 10: 57–63
Cui P, Lin Q, Ding F, et al. A comparison between ribo-minus RNA-sequencing and polyA-selected RNA-sequencing. Genomics, 2010, 96: 259–265
Cui P, Lin Q, Xin C, et al. Hydroxyurea-induced global transcriptional suppression in mouse ES cells. Carcinogenesis, 2010, 31: 1661–1668
Gentleman R C, Carey V J, Bates D M, et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol, 2004, 5: R80
Ashburner M, Ball C A, Blake J A, et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet, 2000, 25: 25–29
Ye J, Fang L, Zheng H, et al. WEGO: a web tool for plotting GO annotations. Nucleic Acids Res, 2006, 34: W293–297
Wang L, Feng Z, Wang X, et al. DEGseq: an R package for identifying differentially expressed genes from RNA-seq data. Bioinformatics, 2010, 26: 136–138
Ogata H, Goto S, Sato K, et al. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res, 1999, 27: 29–34
Kanehisa M, Goto S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res, 2000, 28: 27–30
Kanehisa M, Goto S, Hattori M, et al. From genomics to chemical genomics: new developments in KEGG. Nucleic Acids Res, 2006, 34: D354–357
Kanehisa M, Goto S, Furumichi M, et al. KEGG for representation and analysis of molecular networks involving diseases and drugs. Nucleic Acids Res, 2010, 38: D355–360
Dahlquist K D, Salomonis N, Vranizan K, et al. GenMAPP, a new tool for viewing and analyzing microarray data on biological pathways. Nat Genet, 2002, 31: 19–20
Yeh J R, Nagano M C. Spermatogonial stem cell biomarkers: improved outcomes of spermatogonial transplantation in male fertility restoration? Expert Rev Mol Diagn, 2009, 9: 109–114
Kubota H, Avarbock M R, Brinster R L. Spermatogonial stem cells share some, but not all, phenotypic and functional characteristics with other stem cells. Proc Natl Acad Sci USA, 2003, 100: 6487–6492
Phillips B T, Gassei K, Orwig K E. Spermatogonial stem cell regulation and spermatogenesis. Philos Trans R Soc Lond B Biol Sci, 2010, 365: 1663–1678
Suter L, Koch E, Bechter R, et al. Three-parameter flow cytometric analysis of rat spermatogenesis. Cytometry, 1997, 27: 161–168
Franke F E, Pauls K, Rey R, et al. Differentiation markers of Sertoli cells and germ cells in fetal and early postnatal human testis. Anat Embryol (Berl), 2004, 209: 169–177
O’shaughnessy P J, Hu L, Baker P J. Effect of germ cell depletion on levels of specific mRNA transcripts in mouse Sertoli cells and Leydig cells. Reproduction, 2008, 135: 839–850
Haider S G. Cell biology of Leydig cells in the testis. Int Rev Cytol, 2004, 233: 181–241
O’shaughnessy P J, Willerton L, Baker P J. Changes in Leydig cell gene expression during development in the mouse. Biol Reprod, 2002, 66: 966–975
Zhang F P, Pakarainen T, Zhu F, et al. Molecular characterization of postnatal development of testicular steroidogenesis in luteinizing hormone receptor knockout mice. Endocrinology, 2004, 145: 1453–1463
Kanamori M, Konno H, Osato N, et al. A genome-wide and nonredundant mouse transcription factor database. Biochem Biophys Res Commun, 2004, 322: 787–793
Ravasi T, Suzuki H, Cannistraci C V, et al. An atlas of combinatorial transcriptional regulation in mouse and man. Cell, 2010, 140: 744–752
Zhou Q, Griswold M D. Regulation of spermatogonia. StemBook. Cambridge (MA): Harvard Stem Cell Institute, 2008
Sekido R. SRY: A transcriptional activator of mammalian testis determination. Int J Biochem Cell Biol, 2010, 42: 417–420
Barrionuevo F, Scherer G. SOX E genes: SOX9 and SOX8 in mammalian testis development. Int J Biochem Cell Biol, 2010, 42: 433–436
Tanaka H, Baba T. Gene expression in spermiogenesis. Cell Mol Life Sci, 2005, 62: 344–354
Meistrich M L, Mohapatra B, Shirley C R, et al. Roles of transition nuclear proteins in spermiogenesis. Chromosoma, 2003, 111: 483–488
Nayernia K, Adham I M, Burkhardt-Gottges E, et al. Asthenozoospermia in mice with targeted deletion of the sperm mitochondrion-associated cysteine-rich protein (Smcp) gene. Mol Cell Biol, 2002, 22: 3046–3052
Anway M D, Ravindranath N, Dym M, et al. Identification of a murine testis complementary DNA encoding a homolog to human A-kinase anchoring protein-associated sperm protein. Biol Reprod, 2002, 66: 1755–1761
Kierszenbaum A L, Tres L L. Structural and transcriptional features of the mouse spermatid genome. J Cell Biol, 1975, 65: 258–270
Heidaran M A, Kistler W S. Transcriptional and translational control of the message for transition protein 1, a major chromosomal protein of mammalian spermatids. J Biol Chem, 1987, 262: 13309–13315
Kleene K C. Poly(A) shortening accompanies the activation of translation of five mRNAs during spermiogenesis in the mouse. Development, 1989, 106: 367–373
Kleene K C, Bagarova J. Comparative genomics reveals gene-specific and shared regulatory sequences in the spermatid-expressed mammalian Odf1, Prm1, Prm2, Tnp1, and Tnp2 genes. Genomics, 2008, 92: 101–106
Sozubir S, Barber T, Wang Y, et al. Loss of Insl3: a potential predisposing factor for testicular torsion. J Urol, 2010, 183: 2373–2379
Nakamura N, Mori C, Eddy E M. Molecular complex of three testis-specific isozymes associated with the mouse sperm fibrous sheath: hexokinase 1, phosphofructokinase M, and glutathione S-transferase mu class 5. Biol Reprod, 2010, 82: 504–515
Valentin M, Balvers M, Pusch W, et al. Structure and expression of the mouse gene encoding the endozepine-like peptide from haploid male germ cells. Eur J Biochem, 2000, 267: 5438–5449
Selvaraj V, Asano A, Page J L, et al. Mice lacking FABP9/PERF15 develop sperm head abnormalities but are fertile. Dev Biol, 2010, 348: 177–189
Dakhova O, O’Day D, Kinet N, et al. Dickkopf-like1 regulates postpubertal spermatocyte apoptosis and testosterone production. Endocrinology, 2009, 150: 404–412
Choi H S, Lee S H, Kim H, et al. Germ cell-specific gene 1 targets testis-specific poly(A) polymerase to the endoplasmic reticulum through protein-protein interactions. FEBS Lett, 2008, 582: 1203–1209
Bellve A R, Millette C F, Bhatnagar Y M, et al. Dissociation of the mouse testis and characterization of isolated spermatogenic cells. J Histochem Cytochem, 1977, 25: 480–494
Tegelenbosch R A, de Rooij D G. A quantitative study of spermatogonial multiplication and stem cell renewal in the C3H/101 F1 hybrid mouse. Mutat Res, 1993, 290: 193–200
Gassei K, Ehmcke J, Wood M A, et al. Immature rat seminiferous tubules reconstructed in vitro express markers of Sertoli cell maturation after xenografting into nude mouse hosts. Mol Hum Reprod, 2010, 16: 97–110
Steger K, Rey R, Louis F, et al. Reversion of the differentiated phenotype and maturation block in Sertoli cells in pathological human testis. Hum Reprod, 1999, 14: 136–143
Dahia C L, Rao A J. Regulation of FSH receptor, PKIbeta, IL-6 and calcium mobilization: Possible mediators of differential action of FSH. Mol Cell Endocrinol, 2006, 247: 73–81
Heckert L, Griswold M D. Expression of the FSH receptor in the testis. Recent Prog Horm Res, 1993, 48: 61–77
Li M W, Mruk D D, Cheng C Y. Mitogen-activated protein kinases in male reproductive function. Trends Mol Med, 2009, 15: 159–168
Gnessi L, Emidi A, Jannini E A, et al. Testicular development involves the spatiotemporal control of PDGFs and PDGF receptors gene expression and action. J Cell Biol, 1995, 131: 1105–1121
Basciani S, Mariani S, Spera G, et al. Role of platelet-derived growth factors in the testis. Endocr Rev, 2010, 31: 916–939
Szczepny A, Hime G R, Loveland K L. Expression of hedgehog signalling components in adult mouse testis. Dev Dyn, 2006, 235: 3063–3070
Bitgood M J, Shen L, McMahon A P. Sertoli cell signaling by Desert hedgehog regulates the male germline. Curr Biol, 1996, 6: 298–304
Golestaneh N, Beauchamp E, Fallen S, et al. Wnt signaling promotes proliferation and stemness regulation of spermatogonial stem/progenitor cells. Reproduction, 2009, 138: 151–162
Tanwar P S, Kaneko-Tarui T, Zhang L, et al. Constitutive WNT/betacatenin signaling in murine Sertoli cells disrupts their differentiation and ability to support spermatogenesis. Biol Reprod, 2010, 82: 422–432
Ma P, Wang H, Guo R, et al. Stage-dependent Dishevelled-1 expression during mouse spermatogenesis suggests a role in regulating spermatid morphological changes. Mol Reprod Dev, 2006, 73: 774–783
Hacker A, Capel B, Goodfellow P, et al. Expression of Sry, the mouse sex determining gene. Development, 1995, 121: 1603–1614
De Cesare D, Fimia G M, Sassone-Corsi P. CREM, a master-switch of the transcriptional cascade in male germ cells. J Endocrinol Invest, 2000, 23: 592–596
Blendy J A, Kaestner K H, Weinbauer G F, et al. Severe impairment of spermatogenesis in mice lacking the CREM gene. Nature, 1996, 380: 162–165
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is published with open access at Springerlink.com
Electronic supplementary material
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Gong, W., Pan, L., Lin, Q. et al. Transcriptome profiling of the developing postnatal mouse testis using next-generation sequencing. Sci. China Life Sci. 56, 1–12 (2013). https://doi.org/10.1007/s11427-012-4411-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-012-4411-y