Abstract
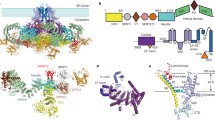
Ryanodine receptors are ion channels that allow for the release of Ca2+ from the endoplasmic or sarcoplasmic reticulum. They are expressed in many different cell types but are best known for their predominance in skeletal and cardiac myocytes, where they are directly involved in excitation-contraction coupling. With molecular weights exceeding 2 MDa, Ryanodine Receptors are the largest ion channels known to date and present major challenges for structural biology. Since their discovery in the 1980s, significant progress has been made in understanding their behaviour through multiple structural methods. Cryo-electron microscopy reconstructions of intact channels depict a mushroom-shaped structure with a large cytoplasmic region that presents many binding sites for regulatory molecules. This region undergoes significant motions during opening and closing of the channel, demonstrating that the Ryanodine Receptor is a bona fide allosteric protein. High-resolution structures through X-ray crystallography and NMR currently cover ∼11% of the entire protein. The combination of high- and low-resolution methods allows us to build pseudo-atomic models. Here we present an overview of the electron microscopy, NMR, and crystallographic analyses of this membrane protein giant.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Meissner G. Ryanodine activation and inhibition of the Ca2+ release channel of sarcoplasmic reticulum. J Biol Chem, 1986, 261: 6300–6306, 2422165, 1:CAS:528:DyaL28Xit1Sqs7g%3D
Takeshima H, Nishimura S, Matsumoto T, et al. Primary structure and expression from complementary DNA of skeletal muscle ryanodine receptor. Nature, 1989, 339: 439–445, 2725677, 1:CAS:528:DyaK3cXhsFertb8%3D
Otsu K, Willard H F, Khanna V K, et al. Molecular cloning of cDNA encoding the Ca2+ release channel (ryanodine receptor) of rabbit cardiac muscle sarcoplasmic reticulum. J Biol Chem, 1990, 265: 13472–13483, 2380170, 1:CAS:528:DyaK3cXmtVKis78%3D
Hakamata Y, Nakai J, Takeshima H, et al. Primary structure and distribution of a novel ryanodine receptor/calcium release channel from rabbit brain. FEBS Lett, 1992, 312: 229–235, 1330694, 1:CAS:528:DyaK3sXltFWmsL0%3D
Sorrentino V, Volpe P. Ryanodine receptors: How many, where and why? Trends Pharmacol Sci, 1993, 14: 98–103, 8387707, 1:CAS:528:DyaK3sXkt1Gjtbc%3D
Endo M. Calcium-induced calcium release in skeletal muscle. Physiol Rev, 2009, 89: 1153–1176, 19789379, 1:CAS:528:DC%2BD1MXhtlyqs73F
Rios E, Brum G. Involvement of dihydropyridine receptors in excitation-contraction coupling in skeletal muscle. Nature, 1987, 325: 717–720, 2434854, 1:CAS:528:DyaL2sXkt1Wht78%3D
Tanabe T, Beam K G, Powell J A, et al. Restoration of excitation-contraction coupling and slow calcium current in dysgenic muscle by dihydropyridine receptor complementary DNA. Nature, 1988, 336: 134–139, 2903448, 1:CAS:528:DyaL1MXltlGl
Block B A, Imagawa T, Campbell K P, et al. Structural evidence for direct interaction between the molecular components of the transverse tubule/sarcoplasmic reticulum junction in skeletal muscle. J Cell Biol, 1988, 107: 2587–2600, 2849609, 1:STN:280:DyaL1M%2FnvFOltA%3D%3D
Tanabe T, Beam K G, Adams B A, et al. Regions of the skeletal muscle dihydropyridine receptor critical for excitation-contraction coupling. Nature, 1990, 346: 567–569, 2165570, 1:CAS:528:DyaK3cXltFejt7w%3D
Saito A, Seiler S, Chu A, et al. Preparation and morphology of sarcoplasmic reticulum terminal cisternae from rabbit skeletal muscle. J Cell Biol, 1984, 99: 875–885, 6147356, 1:STN:280:DyaL2c3ovFKqsQ%3D%3D
Smith D S. The organization and function of the sarcoplasmic reticulum and T-system of muscle cells. Prog Biophys Mol Biol, 1966, 16: 107–142, 5338779, 1:STN:280:DyaF2s7nt1Gisw%3D%3D
Betzenhauser M J, Marks A R. Ryanodine receptor channelopathies. Pflugers Arch, 2010, 460: 467–480, 20179962, 1:CAS:528:DC%2BC3cXntlGnsrs%3D
Thomas N L, Maxwell C, Mukherjee S, et al. Ryanodine receptor mutations in arrhythmia: The continuing mystery of channel dysfunction. FEBS Lett, 2010, 584: 2153–2160, 20132818, 1:CAS:528:DC%2BC3cXlvVaqs74%3D
Priori S G, Chen S R. Inherited dysfunction of sarcoplasmic reticulum Ca2+ handling and arrhythmogenesis. Circ Res, 2011, 108: 871–883, 21454795, 1:CAS:528:DC%2BC3MXktFeks7Y%3D
Kushnir A, Marks A R. The ryanodine receptor in cardiac physiology and disease. Adv Pharmacol, 2010, 59: 1–30, 20933197, 1:CAS:528:DC%2BC3cXhs1Smtr%2FF
Bers D M. Macromolecular complexes regulating cardiac ryanodine receptor function. J Mol Cell Cardiol, 2004, 37: 417–429, 15276012, 1:CAS:528:DC%2BD2cXmtVaisr0%3D
Zalk R, Lehnart S E, Marks A R. Modulation of the ryanodine receptor and intracellular calcium. Annu Rev Biochem, 2007, 76: 367–385, 17506640, 1:CAS:528:DC%2BD2sXhtVehtb%2FI
Lanner J T, Georgiou D K, Joshi A D, et al. Ryanodine receptors: structure, expression, molecular details, and function in calcium release. Cold Spring Harb Perspect Biol, 2010, 2: a003996, 20961976, 1:CAS:528:DC%2BC3cXhsFagtrbM
Marx S. Ion channel macromolecular complexes in the heart. J Mol Cell Cardiol, 2003, 35: 37–44, 12623298, 1:CAS:528:DC%2BD3sXhs1Cgsbg%3D
Fill M, Copello J A. Ryanodine receptor calcium release channels. Physiol Rev, 2002, 82: 893–922, 12270947, 1:CAS:528:DC%2BD38Xot1Ggurk%3D
Revel J P. The sarcoplasmic reticulum of the bat cricothroid muscle. J Cell Biol, 1962, 12: 571–588, 14491637, 1:STN:280:DyaF38%2FptlWqtw%3D%3D
Franzini-Armstrong C. Studies of the triad: I. Structure of the junction in frog twitch fibers. J Cell Biol, 1970, 47: 488–499, 19866746, 1:STN:280:DC%2BC3cnisFWhug%3D%3D
Bonilla E. Staining of transverse tubular system of skeletal muscle by tannic acid-glutaraldehyde fixation. J Ultrastruct Res, 1977, 162–165
Somlyo A V. Bridging structures spanning the junctioning gap at the triad of skeletal muscle. J Cell Biol, 1979, 80: 743–750, 313399, 1:CAS:528:DyaE1MXhsV2gtbc%3D
Campbell K P, Franzini-Armstrong C, Shamoo A E. Further characterization of light and heavy sarcoplasmic reticulum vesicles. Identification of the ’sarcoplasmic reticulum feet’ associated with heavy sarcoplasmic reticulum vesicles. Biochim Biophys Acta, 1980, 602: 97–116, 6448074, 1:CAS:528:DyaL3cXmtlensr4%3D
Franzini-Armstrong C. Membrane particles and transmission at the triad. Fed Proc, 1975, 34: 1382–1389, 1079008, 1:STN:280:DyaE2M7jtFCltQ%3D%3D
Franzini-Armstrong C. Structure of sarcoplasmic reticulum. Fed Proc, 1980, 39: 2403–2409, 7371874, 1:CAS:528:DyaL3cXkt1Srsbg%3D
Ferguson D G, Schwartz H W, Franzini-Armstrong C. Subunit structure of junctional feet in triads of skeletal muscle: A freeze-drying, rotary-shadowing study. J Cell Biol, 1984, 99: 1735–1742, 6386826, 1:STN:280:DyaL2M%2FjtVGrsg%3D%3D
Inui M, Saito A, Fleischer S. Purification of the ryanodine receptor and identity with feet structures of junctional terminal cisternae of sarcoplasmic reticulum from fast skeletal muscle. J Biol Chem, 1987, 262: 1740–1747, 3805051, 1:CAS:528:DyaL2sXpvVahsw%3D%3D
Saito A, Inui M, Radermacher M, et al. Ultrastructure of the calcium release channel of sarcoplasmic reticulum. J Cell Biol, 1988, 107: 211–219, 2455723, 1:CAS:528:DyaL1cXltVKltro%3D
Yin C C, Lai F A. Intrinsic lattice formation by the ryanodine receptor calcium-release channel. Nat Cell Biol, 2000, 2: 669–671, 10980710, 1:CAS:528:DC%2BD3cXmslGks7Y%3D
Yin C C, Han H, Wei R, et al. Two-dimensional crystallization of the ryanodine receptor Ca2+ release channel on lipid membranes. J Struct Biol, 2005, 149: 219–224, 15681238, 1:CAS:528:DC%2BD2MXos1OhtQ%3D%3D
Cheng H, Lederer W J. Calcium sparks. Physiol Rev, 2008, 88: 1491–1545, 18923188, 1:CAS:528:DC%2BD1cXhtlGgtbnF
Wagenknecht T, Grassucci R, Frank J, et al. Three-dimensional architecture of the calcium channel/foot structure of sarcoplasmic reticulum. Nature, 1989, 338: 167–170, 2537473, 1:STN:280:DyaL1M7ktlOqsA%3D%3D
Radermacher M, Rao V, Grassucci R, et al. Cryo-electron micro-scopy and three-dimensional reconstruction of the calcium release channel/ryanodine receptor from skeletal muscle. J Cell Biol, 1994, 127: 411–423, 7929585, 1:CAS:528:DyaK2cXmt1Clsr8%3D
Radermacher M, Wagenknecht T, Grassucci R, et al. Cryo-EM of the native structure of the calcium release channel/ryanodine receptor from sarcoplasmic reticulum. Biophys J, 1992, 61: 936–940, 1316182, 1:CAS:528:DyaK38XisFWgtb8%3D
Serysheva, II, Orlova E V, Chiu W, et al. Electron cryomicroscopy and angular reconstitution used to visualize the skeletal muscle calcium release channel. Nat Struct Biol, 1995, 2: 18–24, 7719847, 1:CAS:528:DyaK2MXivF2ls7s%3D
Sharma M R, Penczek P, Grassucci R, et al. Cryoelectron microscopy and image analysis of the cardiac ryanodine receptor. J Biol Chem, 1998, 273: 18429–18434, 9660811, 1:CAS:528:DyaK1cXkvVCgsb4%3D
Jeyakumar L H, Copello J A, O’Malley A M, et al. Purification and characterization of ryanodine receptor 3 from mammalian tissue. J Biol Chem, 1998, 273: 16011–16020, 9632651, 1:CAS:528:DyaK1cXkt1Giu70%3D
Murayama T, Oba T, Katayama E, et al. Further characterization of the type 3 ryanodine receptor (RyR3) purified from rabbit diaphragm. J Biol Chem, 1999, 274: 17297–17308, 10358090, 1:CAS:528:DyaK1MXjvVGrsrY%3D
Sharma M R, Jeyakumar L H, Fleischer S, et al. Three-dimensional structure of ryanodine receptor isoform three in two conformational states as visualized by cryo-electron microscopy. J Biol Chem, 2000, 275: 9485–9491, 10734096, 1:CAS:528:DC%2BD3cXitlCru70%3D
Ludtke S J, Serysheva II, Hamilton S L, et al. The pore structure of the closed RyR1 channel. Structure, 2005, 13: 1203–1211, 16084392, 1:CAS:528:DC%2BD2MXntVOhsrs%3D
Serysheva II, Ludtke S J, Baker M L, et al. Subnanometer-resolution electron cryomicroscopy-based domain models for the cytoplasmic region of skeletal muscle RyR channel. Proc Natl Acad Sci USA, 2008, 105: 9610–9615, 18621707, 1:CAS:528:DC%2BD1cXovVOjt70%3D
Samso M, Wagenknecht T, Allen P D. Internal structure and visualization of transmembrane domains of the RyR1 calcium release channel by cryo-EM. Nat Struct Mol Biol, 2005, 12: 539–544, 15908964, 1:CAS:528:DC%2BD2MXks12rtr0%3D
Samso M, Feng W, Pessah I N, et al. Coordinated movement of cytoplasmic and transmembrane domains of RyR1 upon gating. PLoS Biol, 2009, 7: e85, 19402748
Minor D L Jr. Bend to open? Structure, 2005, 13: 1094–1095, 16084381, 1:CAS:528:DC%2BD2MXntVOiu7o%3D
Frank J. Single-particle imaging of macromolecules by cryo-electron microscopy. Annu Rev Biophys Biomol Struct, 2002, 31: 303–319, 11988472, 1:CAS:528:DC%2BD38XltVynu78%3D
van Heel M, Schatz M. Fourier shell correlation threshold criteria. J Struct Biol, 2005, 151: 250–262, 16125414
Du G G, Avila G, Sharma P, et al. Role of the sequence surrounding predicted transmembrane helix M4 in membrane association and function of the Ca(2+) release channel of skeletal muscle sarcoplasmic reticulum (ryanodine receptor isoform 1). J Biol Chem, 2004, 279: 37566–37574, 15226293, 1:CAS:528:DC%2BD2cXntFSnsrk%3D
Doyle D A, Morais Cabral J, Pfuetzner R A, et al. The structure of the potassium channel: molecular basis of K+ conduction and selectivity. Science, 1998, 280: 69–77, 9525859, 1:CAS:528:DyaK1cXitlWksrY%3D
Jiang Y, Lee A, Chen J, et al. The open pore conformation of potassium channels. Nature, 2002, 417: 523–526, 12037560, 1:CAS:528:DC%2BD38XktVeht7k%3D
D’Avanzo N, Cho H C, Tolokh I, et al. Conduction through the inward rectifier potassium channel, Kir2.1, is increased by negatively charged extracellular residues. J Gen Physiol, 2005, 125: 493–503, 15824191
Imoto K, Busch C, Sakmann B, et al. Rings of negatively charged amino acids determine the acetylcholine receptor channel conductance. Nature, 1988, 335: 645–648, 2459620, 1:CAS:528:DyaL1cXmt1Cqs7w%3D
Andersen C, Koronakis E, Hughes C, et al. An aspartate ring at the TolC tunnel entrance determines ion selectivity and presents a target for blocking by large cations. Mol Microbiol, 2002, 44: 1131–1139, 12068802, 1:CAS:528:DC%2BD38XksFGqtbo%3D
Terlau H, Heinemann S H, Stuhmer W, et al. Mapping the site of block by tetrodotoxin and saxitoxin of sodium channel II. FEBS Lett, 1991, 293: 93–96, 1660007, 1:CAS:528:DyaK38XltlaktA%3D%3D
Long S B, Campbell E B, Mackinnon R. Voltage sensor of Kv1.2: structural basis of electromechanical coupling. Science, 2005, 309: 903–908, 16002579, 1:CAS:528:DC%2BD2MXmvFSks7s%3D
Kuo A, Gulbis J M, Antcliff J F, et al. Crystal structure of the potassium channel KirBac1.1 in the closed state. Science, 2003, 300: 1922–1926, 12738871, 1:CAS:528:DC%2BD3sXks1Kmurk%3D
Baker M L, Ju T, Chiu W. Identification of secondary structure elements in intermediate-resolution density maps. Structure, 2007, 15: 7–19, 17223528, 1:CAS:528:DC%2BD2sXmtVentw%3D%3D
Nishida M, Cadene M, Chait B T, et al. Crystal structure of a Kir3.1-prokaryotic Kir channel chimera. EMBO J, 2007, 26: 4005–4015, 17703190, 1:CAS:528:DC%2BD2sXpvFWhsbw%3D
Nishida M, MacKinnon R. Structural basis of inward rectification: cytoplasmic pore of the G protein-gated inward rectifier GIRK1 at 1.8 A resolution. Cell, 2002, 111: 957–965, 12507423, 1:CAS:528:DC%2BD3sXis1Kisw%3D%3D
Pegan S, Arrabit C, Zhou W, et al. Cytoplasmic domain structures of Kir2.1 and Kir3.1 show sites for modulating gating and rectification. Nat Neurosci, 2005, 8: 279–287, 15723059, 1:CAS:528:DC%2BD2MXhslyqsbo%3D
Zagotta W N, Olivier N B, Black K D, et al. Structural basis for modulation and agonist specificity of HCN pacemaker channels. Nature, 2003, 425: 200–205, 12968185, 1:CAS:528:DC%2BD3sXntFCisbo%3D
Orlova E V, Serysheva II, van Heel M, et al. Two structural configurations of the skeletal muscle calcium release channel. Nat Struct Biol, 1996, 3: 547–552, 8646541, 1:CAS:528:DyaK28Xjtlylsr8%3D
Serysheva II, Schatz M, van Heel M, et al. Structure of the skeletal muscle calcium release channel activated with Ca2+ and AMP-PCP. Biophys J, 1999, 77: 1936–1944, 10512814, 1:CAS:528:DyaK1MXmtl2msLY%3D
Liu Z, Zhang J, Sharma M R, et al. Three-dimensional reconstruction of the recombinant type 3 ryanodine receptor and localization of its amino terminus. Proc Natl Acad Sci USA, 2001, 98: 6104–6109, 11353864, 1:STN:280:DC%2BD3MzhtlChtA%3D%3D
Wang R, Chen W, Cai S, et al. Localization of an NH(2)-terminal disease-causing mutation hot spot to the “clamp” region in the three-dimensional structure of the cardiac ryanodine receptor. J Biol Chem, 2007, 282: 17785–17793, 17452324, 1:CAS:528:DC%2BD2sXmt1ersb8%3D
Paul-Pletzer K, Yamamoto T, Bhat M B, et al. Identification of a dantrolene-binding sequence on the skeletal muscle ryanodine receptor. J Biol Chem, 2002, 277: 34918–34923, 12167662, 1:CAS:528:DC%2BD38Xnt1Whurk%3D
Paul-Pletzer K, Yamamoto T, Ikemoto N, et al. Probing a putative dantrolene-binding site on the cardiac ryanodine receptor. Biochem J, 2005, 387: 905–909, 15656791, 1:CAS:528:DC%2BD2MXjsFyqs78%3D
Wang R, Zhong X, Meng X, et al. Localization of the dantrolene-binding sequence near the FK506-binding protein-binding site in the three-dimensional structure of the ryanodine receptor. J Biol Chem, 2011, 286: 12202–12212, 21262961, 1:CAS:528:DC%2BC3MXktVKqtbg%3D
Liu Z, Wang R, Zhang J, et al. Localization of a disease-associated mutation site in the three-dimensional structure of the cardiac muscle ryanodine receptor. J Biol Chem, 2005, 280: 37941–37947, 16157601, 1:CAS:528:DC%2BD2MXhtF2is7%2FM
Ikemoto N, Yamamoto T. Regulation of calcium release by interdomain interaction within ryanodine receptors. Front Biosci, 2002, 7: d671–683, 11861212, 1:CAS:528:DC%2BD38Xit12qu7k%3D
Du G G, Khanna V K, MacLennan D H. Mutation of divergent region 1 alters caffeine and Ca2+ sensitivity of the skeletal muscle Ca2+ release channel (ryanodine receptor). J Biol Chem, 2000, 275: 11778–11783, 10766801, 1:CAS:528:DC%2BD3cXisl2js7s%3D
Du G G, MacLennan D H. Ca2+ inactivation sites are located in the COOH-terminal quarter of recombinant rabbit skeletal muscle Ca2+ release channels (ryanodine receptors). J Biol Chem, 1999, 274: 26120–26126, 10473562, 1:CAS:528:DyaK1MXmtVKmtLg%3D
Nakai J, Gao L, Xu L, et al. Evidence for a role of C-terminus in Ca2+ inactivation of skeletal muscle Ca2+ release channel (ryanodine receptor). FEBS Lett, 1999, 459: 154–158, 10518010, 1:CAS:528:DyaK1MXmsVWnu7g%3D
Benacquista B L, Sharma M R, Samso M, et al. Amino acid residues 4425–4621 localized on the three-dimensional structure of the skeletal muscle ryanodine receptor. Biophys J, 2000, 78: 1349–1358, 10692321, 1:CAS:528:DC%2BD3cXhsF2jt7w%3D
Liu Z, Zhang J, Li P, et al. Three-dimensional reconstruction of the recombinant type 2 ryanodine receptor and localization of its divergent region 1. J Biol Chem, 2002, 277: 46712–46719, 12324472, 1:CAS:528:DC%2BD38XovFKgt70%3D
Yamazawa T, Takeshima H, Shimuta M, et al. A region of the ryanodine receptor critical for excitation-contraction coupling in skeletal muscle. J Biol Chem, 1997, 272: 8161–8164, 9079632, 1:CAS:528:DyaK2sXitF2hsLY%3D
Perez C F, Mukherjee S, Allen P D. Amino acids 1-1,680 of ryanodine receptor type 1 hold critical determinants of skeletal type for excitation-contraction coupling. Role of divergence domain D2. J Biol Chem, 2003, 278: 39644–39652, 12900411, 1:CAS:528:DC%2BD3sXnvFantbY%3D
Liu Z, Zhang J, Wang R, et al. Location of divergent region 2 on the three-dimensional structure of cardiac muscle ryanodine receptor/calcium release channel. J Mol Biol, 2004, 338: 533–545, 15081811, 1:CAS:528:DC%2BD2cXjtVegtbo%3D
Hayek S M, Zhao J, Bhat M, et al. A negatively charged region of the skeletal muscle ryanodine receptor is involved in Ca2+-dependent regulation of the Ca2+ release channel. FEBS Lett, 1999, 461: 157–164, 10567689, 1:CAS:528:DyaK1MXntlels7k%3D
Hayek S M, Zhu X, Bhat M B, et al. Characterization of a calcium-regulation domain of the skeletal-muscle ryanodine receptor. Biochem J, 2000, 351: 57–65, 10998347, 1:CAS:528:DC%2BD3cXnslWltro%3D
Proenza C, O’Brien J, Nakai J, et al. Identification of a region of RyR1 that participates in allosteric coupling with the alpha(1S) (Ca(V)1.1) II-III loop. J Biol Chem, 2002, 277: 6530–6535, 11726651, 1:CAS:528:DC%2BD38XhvFyqsL0%3D
Koop A, Goldmann P, Chen S R, et al. ARVC-related mutations in divergent region 3 alter functional properties of the cardiac ryanodine receptor. Biophys J, 2008, 94: 4668–4677, 18326664, 1:CAS:528:DC%2BD1cXmvFKgtro%3D
Zhang J, Liu Z, Masumiya H, et al. Three-dimensional localization of divergent region 3 of the ryanodine receptor to the clamp-shaped structures adjacent to the FKBP binding sites. J Biol Chem, 2003, 278: 14211–14218, 12576471, 1:CAS:528:DC%2BD3sXjtVShsrY%3D
Danila C I, Hamilton S L. Phosphorylation of ryanodine receptors. Biol Res, 2004, 37: 521–525, 15709678
Petrovic M M, Vales K, Putnikovic B, et al. Ryanodine receptors, voltage-gated calcium channels and their relationship with protein kinase A in the myocardium. Physiol Res, 2008, 57: 141–149, 17465689, 1:CAS:528:DC%2BD1cXptVWhu7c%3D
Wehrens X H, Lehnart S E, Reiken S, et al. Ryanodine receptor/calcium release channel PKA phosphorylation: A critical mediator of heart failure progression. Proc Natl Acad Sci USA, 2006, 103: 511–518, 16407108, 1:CAS:528:DC%2BD28XhtVOitLk%3D
Xiao B, Jiang M T, Zhao M, et al. Characterization of a novel PKA phosphorylation site, serine-2030, reveals no PKA hyperpho-sphorylation of the cardiac ryanodine receptor in canine heart failure. Circ Res, 2005, 96: 847–855, 15790957, 1:CAS:528:DC%2BD2MXjsFagsLg%3D
Xiao B, Zhong G, Obayashi M, et al. Ser-2030, but not Ser-2808, is the major phosphorylation site in cardiac ryanodine receptors responding to protein kinase A activation upon beta-adrenergic stimulation in normal and failing hearts. Biochem J, 2006, 396: 7–16, 16483256, 1:CAS:528:DC%2BD28XjvFSmu7c%3D
Jones P P, Meng X, Xiao B, et al. Localization of PKA phosphorylation site, Ser(2030), in the three-dimensional structure of cardiac ryanodine receptor. Biochem J, 2008, 410: 261–270, 17967164, 1:CAS:528:DC%2BD1cXhvVymsbk%3D
Meng X, Xiao B, Cai S, et al. Three-dimensional localization of serine 2808, a phosphorylation site in cardiac ryanodine receptor. J Biol Chem, 2007, 282: 25929–25939, 17606610, 1:CAS:528:DC%2BD2sXpsFWksbw%3D
Ahern G P, Junankar P R, Dulhunty A F. Subconductance states in single-channel activity of skeletal muscle ryanodine receptors after removal of FKBP12. Biophys J, 1997, 72: 146–162, 8994600, 1:CAS:528:DyaK2sXmtFKjsw%3D%3D
Chelu M G, Danila C I, Gilman C P, et al. Regulation of ryanodine receptors by FK506 binding proteins. Trends Cardiovasc Med, 2004, 14: 227–234, 15451514, 1:CAS:528:DC%2BD2cXnvV2msbw%3D
Marx S O, Reiken S, Hisamatsu Y, et al. PKA phosphorylation dissociates FKBP12.6 from the calcium release channel (ryanodine receptor): defective regulation in failing hearts. Cell, 2000, 101: 365–376, 10830164, 1:CAS:528:DC%2BD3cXjsFekurY%3D
Jayaraman T, Brillantes A M, Timerman A P, et al. FK506 binding protein associated with the calcium release channel (ryanodine receptor). J Biol Chem, 1992, 267: 9474–9477, 1374404, 1:CAS:528:DyaK38Xkt1Sitr8%3D
Timerman A P, Onoue H, Xin H B, et al. Selective binding of FKBP12.6 by the cardiac ryanodine receptor. J Biol Chem, 1996, 271: 20385–20391, 8702774, 1:CAS:528:DyaK28Xlt1ygtbg%3D
Wagenknecht T, Grassucci R, Berkowitz J, et al. Cryoelectron microscopy resolves FK506-binding protein sites on the skeletal muscle ryanodine receptor. Biophys J, 1996, 70: 1709–1715, 8785329, 1:CAS:528:DyaK28XhvVGjsr4%3D
Wagenknecht T, Radermacher M, Grassucci R, et al. Locations of calmodulin and FK506-binding protein on the three-dimensional architecture of the skeletal muscle ryanodine receptor. J Biol Chem, 1997, 272: 32463–32471, 9405457, 1:CAS:528:DyaK1cXislWktg%3D%3D
Burkhard P, Taylor P, Walkinshaw M D. X-ray structures of small ligand-FKBP complexes provide an estimate for hydrophobic interaction energies. J Mol Biol, 2000, 295: 953–962, 10656803, 1:CAS:528:DC%2BD3cXot12qtQ%3D%3D
Samso M, Shen X, Allen P D. Structural characterization of the RyR1-FKBP12 interaction. J Mol Biol, 2006, 356: 917–927, 16405911, 1:CAS:528:DC%2BD28XotVOitw%3D%3D
Sharma M R, Jeyakumar L H, Fleischer S, et al. Three-dimensional visualization of FKBP12.6 binding to an open conformation of cardiac ryanodine receptor. Biophys J, 2006, 90: 164–172, 16214874, 1:CAS:528:DC%2BD2MXhtlertrbP
Wagenknecht T, Berkowitz J, Grassucci R, et al. Localization of calmodulin binding sites on the ryanodine receptor from skeletal muscle by electron microscopy. Biophys J, 1994, 67: 2286–2295, 7696469, 1:CAS:528:DyaK2MXisVajtbg%3D
Samso M, Wagenknecht T. Apocalmodulin and Ca2+-calmodulin bind to neighboring locations on the ryanodine receptor. J Biol Chem, 2002, 277: 1349–1353, 11694536, 1:CAS:528:DC%2BD38XnvFOrtA%3D%3D
Ikemoto T, Iino M, Endo M. Enhancing effect of calmodulin on Ca2+-induced Ca2+ release in the sarcoplasmic reticulum of rabbit skeletal muscle fibres. J Physiol, 1995, 487( Pt 3): 573–582, 8544122, 1:CAS:528:DyaK2MXosVGkt7k%3D
Tripathy A, Xu L, Mann G, et al. Calmodulin activation and inhibition of skeletal muscle Ca2+ release channel (ryanodine receptor). Biophys J, 1995, 69: 106–119, 7669888, 1:CAS:528:DyaK2MXmsFWqsrg%3D
Buratti R, Prestipino G, Menegazzi P, et al. Calcium dependent activation of skeletal muscle Ca2+ release channel (ryanodine receptor) by calmodulin. Biochem Biophys Res Commun, 1995, 213: 1082–1090, 7544580, 1:CAS:528:DyaK2MXnslWktbo%3D
Board P G, Coggan M, Watson S, et al. CLIC-2 modulates cardiac ryanodine receptor Ca2+ release channels. Int J Biochem Cell Biol, 2004, 36: 1599–1612, 15147738, 1:CAS:528:DC%2BD2cXktVelsbY%3D
Littler D R, Harrop S J, Goodchild S C, et al. The enigma of the CLIC proteins: Ion channels, redox proteins, enzymes, scaffolding proteins? FEBS Lett, 2010, 584: 2093–2101, 20085760, 1:CAS:528:DC%2BC3cXlvVaqsr0%3D
Meng X, Wang G, Viero C, et al. CLIC2-RyR1 interaction and structural characterization by cryo-electron microscopy. J Mol Biol, 2009, 387: 320–334, 19356589, 1:CAS:528:DC%2BD1MXjt1SmurY%3D
Tripathy A, Resch W, Xu L, et al. Imperatoxin A induces subconductance states in Ca2+ release channels (ryanodine receptors) of cardiac and skeletal muscle. J Gen Physiol, 1998, 111: 679–690, 9565405, 1:CAS:528:DyaK1cXjtFWgsbY%3D
Gurrola G B, Arevalo C, Sreekumar R, et al. Activation of ryanodine receptors by imperatoxin A and a peptide segment of the II–III loop of the dihydropyridine receptor. J Biol Chem, 1999, 274: 7879–7886, 10075681, 1:CAS:528:DyaK1MXitFSktbg%3D
Samso M, Trujillo R, Gurrola G B, et al. Three-dimensional location of the imperatoxin A binding site on the ryanodine receptor. J Cell Biol, 1999, 146: 493–499, 10427100, 1:CAS:528:DyaK1MXkvVGmtLk%3D
Zhou Q, Wang Q L, Meng X, et al. Structural and functional characterization of ryanodine receptor-natrin toxin interaction. Biophys J, 2008, 95: 4289–4299, 18658224, 1:CAS:528:DC%2BD1cXhtlagu7nL
Maximciuc A A, Putkey J A, Shamoo Y, et al. Complex of calmodulin with a ryanodine receptor target reveals a novel, flexible binding mode. Structure, 2006, 14: 1547–1556, 17027503, 1:CAS:528:DC%2BD28XhtVCrtrnF
Wright N T, Prosser B L, Varney K M, et al. S100A1 and calmodulin compete for the same binding site on ryanodine receptor. J Biol Chem, 2008, 283: 26676–26683, 18650434, 1:CAS:528:DC%2BD1cXhtFejtrzL
Balshaw D M, Xu L, Yamaguchi N, et al. Calmodulin binding and inhibition of cardiac muscle calcium release channel (ryanodine receptor). J Biol Chem, 2001, 276: 20144–20153, 11274202, 1:CAS:528:DC%2BD3MXktlKnsLo%3D
Yamaguchi N, Xu L, Pasek D A, et al. Molecular basis of calmodulin binding to cardiac muscle Ca2+ release channel (ryanodine receptor). J Biol Chem, 2003, 278: 23480–23486, 12707260, 1:CAS:528:DC%2BD3sXkslygsb0%3D
Xiong L W, Newman R A, Rodney G G, et al. Lobe-dependent regulation of ryanodine receptor type 1 by calmodulin. J Biol Chem, 2002, 277: 40862–40870, 12185083, 1:CAS:528:DC%2BD38XnvFynurc%3D
Zhang H, Zhang J Z, Danila C I, et al. A noncontiguous, intersubunit binding site for calmodulin on the skeletal muscle Ca2+ release channel. J Biol Chem, 2003, 278: 8348–8355, 12509414, 1:CAS:528:DC%2BD3sXhsF2hsLs%3D
Menegazzi P, Larini F, Treves S, et al. Identification and characterization of three calmodulin binding sites of the skeletal muscle ryanodine receptor. Biochemistry, 1994, 33: 9078–9084, 8049209, 1:CAS:528:DyaK2cXlvFyktL8%3D
Zorzato F, Fujii J, Otsu K, et al. Molecular cloning of cDNA encoding human and rabbit forms of the Ca2+ release channel (ryanodine receptor) of skeletal muscle sarcoplasmic reticulum. J Biol Chem, 1990, 265: 2244–2256, 2298749, 1:CAS:528:DyaK3cXksFegu74%3D
Chen S R, MacLennan D H. Identification of calmodulin-, Ca2+-, and ruthenium red-binding domains in the Ca2+ release channel (ryanodine receptor) of rabbit skeletal muscle sarcoplasmic reticulum. J Biol Chem, 1994, 269: 22698–22704, 7521330, 1:CAS:528:DyaK2cXlvFyqsbo%3D
Most P, Remppis A, Pleger S T, et al. Transgenic overexpression of the Ca2+-binding protein S100A1 in the heart leads to increased in vivo myocardial contractile performance. J Biol Chem, 2003, 278: 33809–33817, 12777394, 1:CAS:528:DC%2BD3sXmvVCrtLg%3D
Treves S, Scutari E, Robert M, et al. Interaction of S100A1 with the Ca2+ release channel (ryanodine receptor) of skeletal muscle. Biochemistry, 1997, 36: 11496–11503, 9298970, 1:CAS:528:DyaK2sXlslOitrg%3D
Prosser B L, Wright N T, Hernandez-Ochoa E O, et al. S100A1 binds to the calmodulin-binding site of ryanodine receptor and modulates skeletal muscle excitation-contraction coupling. J Biol Chem, 2008, 283: 5046–5057, 18089560, 1:CAS:528:DC%2BD1cXhvFeksbw%3D
Du X J, Cole T J, Tenis N, et al. Impaired cardiac contractility response to hemodynamic stress in S100A1-deficient mice. Mol Cell Biol, 2002, 22: 2821–2829, 11909974, 1:CAS:528:DC%2BD38XislWnt7w%3D
McCarthy T V, Quane K A, Lynch P J. Ryanodine receptor mutations in malignant hyperthermia and central core disease. Hum Mutat, 2000, 15: 410–417, 10790202, 1:CAS:528:DC%2BD3cXjtlylsb4%3D
Amador F J, Liu S, Ishiyama N, et al. Crystal structure of type I ryanodine receptor amino-terminal beta-trefoil domain reveals a disease-associated mutation “hot spot” loop. Proc Natl Acad Sci USA, 2009, 106: 11040–11044, 19541610, 1:CAS:528:DC%2BD1MXptVSnu78%3D
Lobo P A, Van Petegem F. Crystal structures of the N-terminal domains of cardiac and skeletal muscle ryanodine receptors: insights into disease mutations. Structure, 2009, 17: 1505–1514, 19913485, 1:CAS:528:DC%2BD1MXhsVajsrnI
Tung C C, Lobo P A, Kimlicka L, et al. The amino-terminal disease hotspot of ryanodine receptors forms a cytoplasmic vestibule. Nature, 2010, 468: 585–588, 21048710, 1:CAS:528:DC%2BC3cXhtlOht7nE
Lobo P A, Kimlicka L, Tung C C, et al. The deletion of exon 3 in the cardiac ryanodine receptor is rescued by beta strand switching. Structure, 2011, 19: 790–798, 21645850, 1:CAS:528:DC%2BC3MXntFGgu70%3D
Wriggers W, Birmanns S. Using situs for flexible and rigid-body fitting of multiresolution single-molecule data. J Struct Biol, 2001, 133: 193–202, 11472090, 1:CAS:528:DC%2BD3MXlsVSjs78%3D
Wriggers W, Chacon P. Modeling tricks and fitting techniques for multiresolution structures. Structure, 2001, 9: 779–788, 11566128, 1:CAS:528:DC%2BD3MXntVajsrk%3D
Baker M L, Serysheva II, Sencer S, et al. The skeletal muscle Ca2+ release channel has an oxidoreductase-like domain. Proc Natl Acad Sci USA, 2002, 99: 12155–12160, 12218169, 1:CAS:528:DC%2BD38XntlClu7k%3D
Wolfram F, Morris E, Taylor C W. Three-dimensional structure of recombinant type 1 inositol 1,4,5-trisphosphate receptor. Biochem J, 2010, 428: 483–489, 20377523, 1:CAS:528:DC%2BC3cXmslOgsbo%3D
Sato C, Hamada K, Ogura T, et al. Inositol 1,4,5-trisphosphate receptor contains multiple cavities and L-shaped ligand-binding domains. J Mol Biol, 2004, 336: 155–164, 14741211, 1:CAS:528:DC%2BD2cXkvFOguw%3D%3D
Serysheva II, Bare D J, Ludtke S J, et al. Structure of the type 1 inositol 1,4,5-trisphosphate receptor revealed by electron cryomicroscopy. J Biol Chem, 2003, 278: 21319–21322, 12714606, 1:CAS:528:DC%2BD3sXksVenurw%3D
da Fonseca P C, Morris S A, Nerou E P, et al. Domain organization of the type 1 inositol 1,4,5-trisphosphate receptor as revealed by single-particle analysis. Proc Natl Acad Sci USA, 2003, 100: 3936–3941, 12651956
Hamada K, Terauchi A, Mikoshiba K. Three-dimensional rearrangements within inositol 1,4,5-trisphosphate receptor by calcium. J Biol Chem, 2003, 278: 52881–52889, 14593123, 1:CAS:528:DC%2BD3sXpvFGjt7Y%3D
Jiang Q X, Thrower E C, Chester D W, et al. Three-dimensional structure of the type 1 inositol 1,4,5-trisphosphate receptor at 24 A resolution. EMBO J, 2002, 21: 3575–3581, 12110570, 1:CAS:528:DC%2BD38XlslOkt7k%3D
Bosanac I, Yamazaki H, Matsu-Ura T, et al. Crystal structure of the ligand binding suppressor domain of type 1 inositol 1,4,5-trisphosphate receptor. Mol Cell, 2005, 17: 193–203, 15664189, 1:CAS:528:DC%2BD2MXht12ltrk%3D
Chan J, Yamazaki H, Ishiyama N, et al. Structural studies of inositol 1,4,5-trisphosphate receptor: coupling ligand binding to channel gating. J Biol Chem, 2010, 285: 36092–36099, 20843799, 1:CAS:528:DC%2BC3cXhtlyntb%2FF
Bosanac I, Alattia J R, Mal T K, et al. Structure of the inositol 1,4,5-trisphosphate receptor binding core in complex with its ligand. Nature, 2002, 420: 696–700, 12442173, 1:CAS:528:DC%2BD38XpsVSiurs%3D
Yuchi Z, Van Petegem F. Common allosteric mechanisms between ryanodine and inositol-1,4,5-trisphosphate receptors. Channels (Austin), 2011, 5: 120–123, 1:CAS:528:DC%2BC3MXls1Crtb0%3D
Pettersen E F, Goddard T D, Huang C C, et al. UCSF Chimera—a visualization system for exploratory research and analysis. J Comput Chem, 2004, 25: 1605–1612, 15264254, 1:CAS:528:DC%2BD2cXmvVOhsbs%3D
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is published with open access at Springerlink.com
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Kimlicka, L., Van Petegem, F. The structural biology of ryanodine receptors. Sci. China Life Sci. 54, 712–724 (2011). https://doi.org/10.1007/s11427-011-4198-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-011-4198-2