Abstract
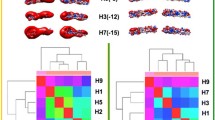
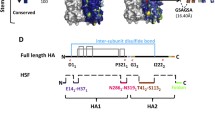
The 2009 swine-origin influenza virus (S-OIV, H1N1 subtype) has developed into a new pandemic influenza as announced by the World Health Organization. In order to uncover clues about the determinants for virulence and pathogenicity of the virus, we characterized the functional modules of the surface glycoprotein hemagglutinin (HA), the most important protein in molecular epidemiology and pathogenesis of influenza viruses. We analyzed receptor binding sites, basic patch, neutralization antibody epitopes and T cell epitopes in the HA protein of the current S-OIV according to the corresponding functional and structural modules previously characterized in other H1 HA molecules or HA molecules of other subtypes. We compared their differences and similarities systematically. Based on the amino acids defined as the functional and structural modules, the HA protein of 2009 S-OIV should specifically bind to the human 2,6-receptor. The D225G/E mutation in HA, which is found in some isolates, may confer dual binding specificity to the 2,3- and 2,6-receptor based on previously reported work. This HA variant contains two basic patches, one of which results in increased basicity, suggesting enhanced membrane fusion function. The 2009 S-OIV HA also has an extra glycosylation site at position 276. Four of the five antibody neutralization epitopes identified in A/RP/8/34(H1N1) were exposed, but the other was hidden by a glycosylation site. The previously identified cytotoxic T cell epitopes in various HA molecules were summarized and their corresponding sequences in 2009 S-OIV HA were defined. These results are critical for understanding the pathogenicity of the virus and host immune response against the virus.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Smith G J, Vijaykrishna D, Bahl J, et al. Origins and evolutionary genomics of the 2009 swine-origin H1N1 influenza A epidemic. Nature, 2009, 459: 1122–1125 10.1038/nature08182, 19516283, 1:CAS:528:DC%2BD1MXnslKqtL0%3D
Gao G F, Sun Y P. It is not just AIV: From avian to swine-origin influenza virus. Sci China Life Sci, 2010, 53: 151–153 10.1007/s11427-010-0017-4
Trifonov V, Khiabanian H, Rabadan R. Geographic dependence, surveillance, and origins of the 2009 influenza A (H1N1) virus. N Engl J Med, 2009, 361: 115–119 10.1056/NEJMp0904572, 19474418, 1:CAS:528:DC%2BD1MXosVKkt7w%3D
Itoh Y, Shinya K, Kiso M, et al. In vitro and in vivo characterization of new swine-origin H1N1 influenza viruses. Nature, 2009, 460: 1021–1025 19672242, 1:CAS:528:DC%2BD1MXhtVWhsbvJ
Yang Y, Sugimoto J D, Halloran M E, et al. The transmissibility and control of pandemic influenza A (H1N1) virus. Science, 2009, 326: 729–733 10.1126/science.1177373, 19745114, 1:CAS:528:DC%2BD1MXhtlagtL3F
Louie J K, Acosta M, Jamieson D J, et al. Severe 2009 H1N1 Influenza in pregnant and postpartum women in California. N Engl J Med, 2010, 362: 27–35 10.1056/NEJMoa0910444, 20032319, 1:CAS:528:DC%2BC3cXjvFGnug%3D%3D
Wilson I A, Cox N J. Structural basis of immune recognition of influenza virus hemagglutinin. Annu Rev Immunol, 1990, 8: 737–771 10.1146/annurev.iy.08.040190.003513, 2188678, 1:CAS:528:DyaK3cXkslGqsb8%3D
Gao G F, Shaw P C. The challenges of avian influenza virus: Mechanism, epidemiology and control. Sci China Ser C-Life Sci, 2009, 52: 405–406 10.1007/s11427-009-0058-8
Liu D, Liu X, Yan J, et al. Interspecies transmission and host restriction of avian H5N1 influenza virus. Sci China Ser C-Life Sci, 2009, 52: 428–438 10.1007/s11427-009-0062-z
Fouchier R A, Munster V, Wallensten A, et al. Characterization of a novel influenza A virus hemagglutinin subtype (H16) obtained from black-headed gulls. J Virol, 2005, 79: 2814–2822 10.1128/JVI.79.5.2814-2822.2005, 15709000, 1:CAS:528:DC%2BD2MXitlaqurs%3D
Stevens J, Corper A L, Basler C F, et al. Structure of the uncleaved human H1 hemagglutinin from the extinct 1918 influenza virus. Science, 2004, 303: 1866–1870 10.1126/science.1093373, 14764887, 1:CAS:528:DC%2BD2cXitFehuro%3D
Lin T, Wang G, Li A, et al. The hemagglutinin structure of an avian H1N1 influenza A virus. Virology, 2009, 392: 73–81 10.1016/j.virol.2009.06.028, 19628241, 1:CAS:528:DC%2BD1MXhtVOis7bM
Russell R J, Gamblin S J, Haire L F, et al. H1 and H7 influenza haemagglutinin structures extend a structural classification of haemagglutinin subtypes. Virology, 2004, 325: 287–296 10.1016/j.virol.2004.04.040, 15246268, 1:CAS:528:DC%2BD2cXls1WksL4%3D
Gamblin S J, Haire L F, Russell R J, et al. The structure and receptor binding properties of the 1918 influenza hemagglutinin. Science, 2004, 303: 1838–1842 10.1126/science.1093155, 14764886
Liu J, Stevens D J, Haire L F, et al. Structures of receptor complexes formed by hemagglutinins from the Asian Influenza pandemic of 1957. Proc Natl Acad Sci USA, 2009, 106: 17175–17180 10.1073/pnas.0906849106, 19805083
Fleury D, Wharton S A, Skehel J J, et al. Antigen distortion allows influenza virus to escape neutralization. Nat Struct Biol, 1998, 5: 119–123 10.1038/nsb0298-119, 9461077, 1:CAS:528:DyaK1cXovV2ktA%3D%3D
Chen J, Lee K H, Steinhauer D A, et al. Structure of the hemagglutinin precursor cleavage site, a determinant of influenza pathogenicity and the origin of the labile conformation. Cell, 1998, 95: 409–417 10.1016/S0092-8674(00)81771-7, 9814710, 1:CAS:528:DyaK1cXntlCqt70%3D
Ha Y, Stevens D J, Skehel J J, et al. X-ray structure of the hemagglutinin of a potential H3 avian progenitor of the 1968 Hong Kong pandemic influenza virus. Virology, 2003, 309: 209–218 10.1016/S0042-6822(03)00068-0, 12758169, 1:CAS:528:DC%2BD3sXjvFeqsL4%3D
Russell R J, Kerry P S, Stevens D J, et al. Structure of influenza hemagglutinin in complex with an inhibitor of membrane fusion. Proc Natl Acad Sci USA, 2008, 105: 17736–17741 10.1073/pnas.0807142105, 19004788
Fleury D, Daniels R S, Skehel J J, et al. Structural evidence for recognition of a single epitope by two distinct antibodies. Proteins, 2000, 40: 572–578 10.1002/1097-0134(20000901)40:4<572::AID-PROT30>3.0.CO;2-N, 10899782, 1:CAS:528:DC%2BD3cXmtVWnsbs%3D
Fleury D, Barrere B, Bizebard T, et al. A complex of influenza hemagglutinin with a neutralizing antibody that binds outside the virus receptor binding site. Nat Struct Biol, 1999, 6: 530–534 10.1038/9299, 10360354, 1:CAS:528:DyaK1MXjs1ykt7o%3D
Barbey-Martin C, Gigant B, Bizebard T, et al. An antibody that prevents the hemagglutinin low pH fusogenic transition. Virology, 2002, 294: 70–74 10.1006/viro.2001.1320, 11886266, 1:CAS:528:DC%2BD38Xhslemtrg%3D
Bullough P A, Hughson F M, Skehel J J, et al. Structure of influenza haemagglutinin at the pH of membrane fusion. Nature, 1994, 371: 37–43 10.1038/371037a0, 8072525, 1:CAS:528:DyaK2cXmt1Cmsbw%3D
Ekiert D C, Bhabha G, Elsliger M A, et al. Antibody recognition of a highly conserved influenza virus epitope. Science, 2009, 324: 246–251 10.1126/science.1171491, 19251591, 1:CAS:528:DC%2BD1MXktFals7w%3D
Sui J, Hwang W C, Perez S, et al. Structural and functional bases for broad-spectrum neutralization of avian and human influenza A viruses. Nat Struct Mol Biol, 2009, 16: 265–273 10.1038/nsmb.1566, 19234466, 1:CAS:528:DC%2BD1MXitFKks7s%3D
Stevens J, Blixt O, Tumpey TM, et al. Structure and receptor specificity of the hemagglutinin from an H5N1 influenza virus. Science, 2006, 312: 404–410 10.1126/science.1124513, 16543414, 1:CAS:528:DC%2BD28XjslSktbY%3D
Ha Y, Stevens D J, Skehel J J, et al. H5 avian and H9 swine influenza virus haemagglutinin structures: possible origin of influenza subtypes. EMBO J, 2002, 21: 865–875 10.1093/emboj/21.5.865, 11867515, 1:CAS:528:DC%2BD38XitFylsLw%3D
Shen J, Ma J, Wang Q. Evolutionary trends of A (H1N1) influenza virus hemagglutinin since 1918. PLoS One, 2009, 4: e7789 10.1371/journal.pone.0007789, 19924230, 1:CAS:528:DC%2BD1MXhsVGhsrfK
Kornfeld R, Kornfeld S. Assembly of asparagine-linked oligosaccharides. Annu Rev Biochem, 1985, 54: 631–664 10.1146/annurev.bi.54.070185.003215, 3896128, 1:STN:280:DyaL2M3nvVSjsA%3D%3D
Schwede T, Kopp J, Guex N, et al. SWISS-MODEL: An automated protein homology-modeling server. Nucleic Acids Res, 2003, 31: 3381–3385 10.1093/nar/gkg520, 12824332, 1:CAS:528:DC%2BD3sXltVWjsLg%3D
Pettersen E F, Goddard T D, Huang C C, et al. UCSF Chimera—a visualization system for exploratory research and analysis. J Comput Chem, 2004, 25: 1605–1612 10.1002/jcc.20084, 15264254, 1:CAS:528:DC%2BD2cXmvVOhsbs%3D
Wilson I A, Skehel J J, Wiley D C. Structure of the haemagglutinin membrane glycoprotein of influenza virus at 3 Å resolution. Nature, 1981, 289: 366–373 10.1038/289366a0, 7464906, 1:CAS:528:DyaL3MXhvFSjtb8%3D
Connor R J, Kawaoka Y, Webster R G, et al. Receptor specificity in human, avian, and equine H2 and H3 influenza virus isolates. Virology, 1994, 205: 17–23 10.1006/viro.1994.1615, 7975212, 1:CAS:528:DyaK2cXmvFKktbk%3D
Rogers G N, Paulson J C, Daniels R S, et al. Single amino acid substitutions in influenza haemagglutinin change receptor binding specificity. Nature, 1983, 304: 76–78 10.1038/304076a0, 6191220, 1:CAS:528:DyaL3sXkvVKnsLs%3D
Yamada S, Suzuki Y, Suzuki T, et al. Haemagglutinin mutations responsible for the binding of H5N1 influenza A viruses to human-type receptors. Nature, 2006, 444: 378–382 10.1038/nature05264, 17108965, 1:CAS:528:DC%2BD28Xht1Sgtb7J
Rogers G N, D’souza B L. Receptor binding properties of human and animal H1 influenza virus isolates. Virology, 1989, 173: 317–322 10.1016/0042-6822(89)90249-3, 2815586, 1:CAS:528:DyaK3cXhvFOrsQ%3D%3D
Matrosovich M N, Gambaryan A S, Teneberg S, et al. Avian influenza A viruses differ from human viruses by recognition of sialyloligosaccharides and gangliosides and by a higher conservation of the HA receptor-binding site. Virology, 1997, 233: 224–234 10.1006/viro.1997.8580, 9201232, 1:CAS:528:DyaK2sXktlajtrY%3D
Reid A H, Janczewski T A, Lourens R M, et al. 1918 influenza pandemic caused by highly conserved viruses with two receptor-binding variants. Emerg Infect Dis, 2003, 9: 1249–1253 14609459
Skehel J J, Wiley D C. Receptor binding and membrane fusion in virus entry: The influenza hemagglutinin. Annu Rev Biochem, 2000, 69: 531–569 10.1146/annurev.biochem.69.1.531, 10966468, 1:CAS:528:DC%2BD3cXnt1ajtbY%3D
Tumpey T M, Maines T R, Van Hoeven N, et al. A two-amino acid change in the hemagglutinin of the 1918 influenza virus abolishes transmission. Science, 2007, 315: 655–659 10.1126/science.1136212, 17272724, 1:CAS:528:DC%2BD2sXhtVyisL0%3D
Srinivasan A, Viswanathan K, Raman R, et al. Quantitative biochemical rationale for differences in transmissibility of 1918 pandemic influenza A viruses. Proc Natl Acad Sci USA, 2008, 105: 2800–2805 10.1073/pnas.0711963105, 18287068
Thomas P G, Keating R, Hulse-Post D J, et al. Cell-mediated protection in influenza infection. Emerg Infect Dis, 2006, 12: 48–54 16494717, 1:CAS:528:DC%2BD28XosV2jsg%3D%3D
Doherty P C, Brown L E, Kelso A, et al. Immunity to avian influenza A viruses. Rev Sci Tech, 2009, 28: 175–185 19618625, 1:STN:280:DC%2BD1Mvps1CrtA%3D%3D
Cao W, Myers-Powell B A, Braciale T J. The weak CD8+ CTL response to an influenza hemagglutinin epitope reflects limited T cell availability. J Immunol, 1996, 157: 505–511 8752895, 1:CAS:528:DyaK28XktVynsrg%3D
Gould KG, Scotney H, Brownlee GG. Characterization of two distinct major histocompatibility complex class I Kk-restricted T-cell epitopes within the influenza A/PR/8/34 virus hemagglutinin. J Virol, 1991, 65: 5401–5409 1716691, 1:CAS:528:DyaK3MXmslOkt7Y%3D
Chen W, Anton L C, Bennink J R, et al. Dissecting the multifactorial causes of immunodominance in class I-restricted T cell responses to viruses. Immunity, 2000, 12: 83–93 10.1016/S1074-7613(00)80161-2, 10661408, 1:CAS:528:DC%2BD3cXhtVWjtbo%3D
Zhong W, Reche P A, Lai C C, et al. Genome-wide characterization of a viral cytotoxic T lymphocyte epitope repertoire. J Biol Chem, 2003, 278: 45135–45144 10.1074/jbc.M307417200, 12960169, 1:CAS:528:DC%2BD3sXoslyqur8%3D
Gianfrani C, Oseroff C, Sidney J, et al. Human memory CTL response specific for influenza A virus is broad and multispecific. Hum Immunol, 2000, 61: 438–452 10.1016/S0198-8859(00)00105-1, 10773346, 1:CAS:528:DC%2BD3cXisFGnt7w%3D
Alexander J, Oseroff C, Sidney J, et al. Derivation of HLA-A11/Kb transgenic mice: Functional CTL repertoire and recognition of human A11-restricted CTL epitopes. J Immunol, 1997, 159: 4753–4761 9366399, 1:CAS:528:DyaK2sXntlKnt74%3D
Sun Y, Liu J, Yang M, et al. Identification and structural definition of H5-specific CTL epitopes restricted by HLA-A*0201 derived from H5N1 subtype of influenza A viruses. J Gen Virol, 2010, 91: 919–930 10.1099/vir.0.016766-0, 19955560, 1:CAS:528:DC%2BC3cXkvFKitL8%3D
Enserink M, Cohen J. Virus of the year. The novel H1N1 influenza. Science, 2009, 326: 1607 1:CAS:528:DC%2BD1MXhs1agt7nF
Shinya K, Ebina M, Yamada S, et al. Avian flu: Influenza virus receptors in the human airway. Nature, 2006, 440: 435–436 10.1038/440435a, 16554799, 1:CAS:528:DC%2BD28Xis1Omu7s%3D
Kilbourne E D. Influenza pandemics of the 20th century. Emerg Infect Dis, 2006, 12: 9–14 16494710
Lee L Y, Ha do L A, Simmons C, et al. Memory T cells established by seasonal human influenza A infection cross-react with avian influenza A (H5N1) in healthy individuals. J Clin Invest, 2008, 118: 3478–3490 18802496, 1:CAS:528:DC%2BD1cXht1ChsrnO
Kreijtz J H, de Mutsert G, van Baalen C A, et al. Cross-recognition of avian H5N1 influenza virus by human cytotoxic T-lymphocyte populations directed to human influenza A virus. J Virol, 2008, 82: 5161–5166 10.1128/JVI.02694-07, 18353950, 1:CAS:528:DC%2BD1cXmt1Smu78%3D
Greenbaum J A, Kotturi M F, Kim Y, et al. Pre-existing immunity against swine-origin H1N1 influenza viruses in the general human population. Proc Natl Acad Sci USA, 2009, 106: 20365–20370 10.1073/pnas.0911580106, 19918065
Skehel J J, Stevens D J, Daniels R S, et al. A carbohydrate side chain on hemagglutinins of Hong Kong influenza viruses inhibits recognition by a monoclonal antibody. Proc Natl Acad Sci USA, 1984, 81: 1779–1783 10.1073/pnas.81.6.1779, 6584912, 1:CAS:528:DyaL2cXhvFSjsr4%3D
Author information
Authors and Affiliations
Corresponding author
Additional information
Contributed equally to this work
Rights and permissions
About this article
Cite this article
Sun, Y., Shi, Y., Zhang, W. et al. In silico characterization of the functional and structural modules of the hemagglutinin protein from the swine-origin influenza virus A (H1N1)-2009. Sci. China Life Sci. 53, 633–642 (2010). https://doi.org/10.1007/s11427-010-4010-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-010-4010-8