Abstract
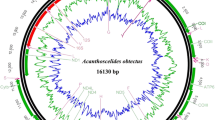
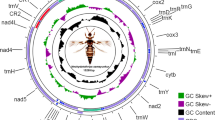
The complete mitochondrial genome of Chinese Bombyx mandarina (ChBm) was determined. The circular genome is 15682 bp long, and contains a typical gene complement, order, and arrangement identical to that of Bombyx mori (B. mori) and Japanese Bombyx mandarina (JaBm) except for two additional tRNA-like structures: tRNASer(TGA)-like and tRNAIle(TAT)-like. All protein-coding sequences are initiated with a typical ATN codon except for the COI gene, which has a 4-bp TTAG putative initiator codon. Eleven of 13 protein-coding genes (PCGs) have a complete termination codon (all TAA), but the remaining two genes terminate with incomplete codons. All tRNAs have the typical clover-leaf structures of mitochondrial tRNAs, with the exception of tRNASer(TGA)-like, with a four stem-and-loop structure. The length of the A+T-rich region of ChBm is 484 bp, shorter than those of JaBm (747 bp) and B. mori (494–499 bp). Phylogenetic analysis among B. mori, ChBm, JaBm, and Antheraea pernyi (Anpe) showed that B. mori is more closely related to ChBm than JaBm. The earliest divergence time estimate for B. mori-ChBm and B. mori-JaBm is about 1.08±0.18–1.41±0.24 and 1.53±0.20–2.01±0.26 Mya, respectively. ChBm and JaBm diverged around 1.11±0.16–1.45±0.21 Mya.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Wolstenholme D R. Animal mitochondrial DNA: Structure and evolution. Int Rev Cyt, 1992, 141: 173–216, 10.1016/S0074-7696(08)62066-5, 1:CAS:528:DyaK3sXkt1Wgs78%3D
Junqueira A C. The mitochondrial genome of the blowfly Chrysomya chloropyga (Diptera: Calliphoridae). Gene, 2004, 339: 7–15, 15363841, 10.1016/j.gene.2004.06.031, 1:CAS:528:DC%2BD2cXnsVKjtb8%3D
Cha S Y, Yoon H J, Lee E M, et al. The complete nucleotide sequence and gene organization of the mitochondrial genome of the bumblebee, Bombus ignites (Hymenoptera: Apidae). Gene, 2007, 392: 206–220, 17321076, 10.1016/j.gene.2006.12.031, 1:CAS:528:DC%2BD2sXjs1Cgsbg%3D
Fauron C M R, Wolstenholme D R. Extensive diversity among Drosophila species with respective to nucleotide sequences within the adenine + thymine rich region of mitochondrial DNA molecules. Nucleic Acids Res, 1980, 11: 2439–2453, 10.1093/nar/8.11.2439
Lewis D L, Farr C L, Kaguni L S. Drosophila melanogaster mitochondrial DNA, completion of the nucleotide sequence and evolutionary comparisons. Insect Mol Biol, 1995, 4: 263–278, 8825764, 10.1111/j.1365-2583.1995.tb00032.x, 1:CAS:528:DyaK2MXhtVSitLrM
Renfu S, Nick J H, Campbell H, et al. Numerous gene rearrangements in the mitochondrial genome of the wallaby louse, Heterodoxus macropus (Phthiraptera). Mol Biol Evol, 2001, 18: 858–865
Goldsmith M R, Shimada T, Abe H. The genetics and genomics of the silkworm, Bombyx mori. Annu Rev Entomol, 2005, 50: 71–100, 15355234, 10.1146/annurev.ento.50.071803.130456, 1:CAS:528:DC%2BD2MXhtFOqt7o%3D
Sasaki C. On the affinity of our wild and domestic silkworms. Ann Zool Jpn, 1889, 2: 2
Banno Y, Nakamura T, Nagashima E, et al. M chromosome of the wild silkworm, Bombyx mandarina (n=27), corresponds to two chromosomes in the domestication silkworm, Bomby mori (n=28). Genome, 2004, 47: 96–101, 15060606, 10.1139/g03-112, 1:CAS:528:DC%2BD2cXivFGqsb8%3D
Kawaguchi E. Zytologiche untersuchungen am seidenspinner und seinen verwandten. I. Gametogenes von Bombyx mori L. und B. mandarina M. und ihrer Bestarde. Z Zellforsch Mikrosk Anat, 1928, 7: 157–156, 10.1007/BF01264055
Astaurov B L, Golisheva M D, Roginskaya I S. Chromosome complex of Ussuri geographical race of Bombyx mandarina M. with spacial reference to the problem of the origin of the domesticated silkworm, Bombyx mori L. Cytology (in Russian), 1959, 1: 327–332
Yukuhiro K, Sezutsu H, Itoh M, et al. Significant levels of sequence divergence and gene rearrangements have occurred between the mitochondrial genomes of the wild mulberry silkmoth, Bombyx mandarina, and its close relative, the domesticated silkmoth, Bombyx mori. Mol Biol Evol, 2002, 19: 1385–1389, 12140251, 1:CAS:528:DC%2BD38XmtF2rs70%3D
Arunkumar K P, Metta M, Nagaraju J. Molecular phylogeny of silkworm reveals the origin of domesticated silkmoth, Bombyx mori from Chinese Bombyx mandarina and paternal inheritiance of Antheraea proylei mitochondrial DNA. Mol Phylogenet Evol, 2006, 40:419–427, 16644243, 10.1016/j.ympev.2006.02.023, 1:CAS:528:DC%2BD28XntFWqsLw%3D
Avise J C. Phylogeography: The History and Formation of Species. Cambridge: Harvard University Press, 2000
Moritz C, Dowling T E, Brown W M. Evolution of animal mitochondrial DNA: Relevance for population biology and systematics. Ann Rev Ecol Syst, 1987, 18: 269–292, 10.1146/annurev.es.18.110187.001413
Pesole G, Gissi C, Saccone C. Nucleotide substitution rate of mammalian mitochondrial genomes. J Mol Evol, 1999, 48: 427–434, 10079281, 10.1007/PL00006487, 1:CAS:528:DyaK1MXhvFyqu7w%3D
Sambrook J, Fritsch E F, Maniatis T. Molecular Cloning, A Laboratory Manual. 2nd ed. Cold Spring Harbor: Cold Spring Harbor Laboratory Press, 1989
Thomson J D, Gibson T J, Plewniak F, et al. The CLUSTAL X windows interface: Flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res, 1997, 24:173–216
Lowe T M, Eddy S R. tRNA-scan-SE: A program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res, 1997, 25: 955–964, 9023104, 10.1093/nar/25.5.955, 1:CAS:528:DyaK2sXhvVahtrk%3D
Kumar S, Tamura K, Jakobsen I B, et al. MEGA 2: Molecular evolution genetics analysis software. Bioinformatics, 2001, 17:1244–1245, 11751241, 10.1093/bioinformatics/17.12.1244, 1:CAS:528:DC%2BD38XmtVCktQ%3D%3D
Schmidt H A, Strimmer K, Vingron M, von Haseler A. TREE-PUZZLE: Maximum likelihood phylogenetic analysis using quartets and parallel computing. Bioinformatics, 2002, 18: 502–504, 11934758, 10.1093/bioinformatics/18.3.502, 1:CAS:528:DC%2BD38XivFKrsL0%3D
Adachi J, Hasegawa M. Model of amino acid substitution in proteins encoded by mitochondrial DNA. J Mol Evol, 1996, 42: 459–468, 8642615, 10.1007/BF02498640, 1:CAS:528:DyaK28XivFKnurw%3D
Kumar M. A simple method for estimating evolution rate of base substitution through comparative studies of nucleotide sequence. J Mol Evol, 1980, 16: 111–120, 10.1007/BF01731581
Zakharov E V, Caterino M S, Sperling F A. Molecular phylogeny, historical biogeography, and divergence time estimates for swallowtail butterflies of the genus Papilio (Lepidoptera: Papilionidae). Syst Biol, 2004, 53: 193–215, 15205049, 10.1080/10635150490423403
Fu Y X, Li W H. Estimating the age of the common ancestor of a sample of DNA sequences. Mol Biol Evol, 1997, 14: 195–199, 9029798, 1:CAS:528:DyaK2sXhtVKhurc%3D
Shao R, Barker S C. The highly rearranged mitochondrial genome of the plague thrips, Thrips imaginis (Insecta: Thysanoptera): Convergence of two novel gene boundaries and an extraordinary arrangement of rRNA genes. Mol Biol Evol, 2003, 20: 362–370, 12644556, 10.1093/molbev/msg045, 1:CAS:528:DC%2BD3sXisFaqs78%3D
Thao M L, Baumann L, Baumann P. Organization of the mitochondrial genomes of whiteflies, aphids, and psyllids (Hemiptera, Sternorrhyncha). BMC Evol Biol, 2004, 4: 25, 15291971, 10.1186/1471-2148-4-25, 1:CAS:528:DC%2BD2cXnsF2qsLw%3D
Zhang D X, Hewitt G M. Insect mitochondrial control region: A review of its structure, evolution and usefulness in evolutionary studies. BiochemSyst Ecol, 1997, 25: 99–120, 10.1016/S0305-1978(96)00042-7
Lu C, Liu Y Q, Liao S Y, et al. Complete sequence dternination and analysis of Bombyx mori mitochondrial genome. J Agric Biotechnol (in Chinese), 2002, 10: 163–170
Yamauchi Y, Hoeffer C, Yamamoto A, et al. cDNA and deduced amino acid sequences of apolipophorin-IIIs from Bombyx mori and Bombyx mandarina. Arch Insect Biochem Physiol, 2000, 43: 16–21, 10613959, 10.1002/(SICI)1520-6327(200001)43:1<16::AID-ARCH3>3.0.CO;2-W, 1:CAS:528:DC%2BD3cXkt1SltQ%3D%3D
Li A, Zhao Q, Tang S, et al. Molecular phylogeny of the domesticated silkworm, Bombyx mori, based on the sequences of mitochondrial cytochrome b genes. J Genet, 2005, 84: 137–142, 16131713, 10.1007/BF02715839, 1:CAS:528:DC%2BD2MXhtFSkt77E
Maekawa H, Takada N, Mikitani K, et al. Nucleolus organizers in the wild silkworm Bombyx mandarina and domesticated silkworm B. mori. Chromosoma, 1988, 96: 263–269, 10.1007/BF00286912, 1:CAS:528:DyaL1cXktVWktLk%3D
Nakamura T, Banno Y, Nakada T, et al. Geographic dimorphism of the wild silkworm, Bombyx mandarina, in the chromosome number and the occurrence of a retroposon-like insertion in the arylphorin gene. Genome, 1999, 42: 1117–1120, 10659778, 10.1139/gen-42-6-1117, 1:CAS:528:DC%2BD3cXpt12rsA%3D%3D
Shimada T, Kurimoto Y, Kobayashi M. Phylogenetic relationship of silkmoths inferred from sequence data of the arylphorin gene. Mol Phylogenet Evol, 1995, 4: 223–234, 8845960, 10.1006/mpev.1995.1021, 1:CAS:528:DyaK2MXptFShtbg%3D
Zhou Y. General Entomology (in Chinese). 2nd ed. Beijing: High Education Publication House, 1958
Oshima K. The history of straits around the Japanese Islands in the late-quaternary. Quaternary Res, 1990, 29: 193–208
Matsui H, Tada R, Oba T. Low-salinity isolation event in the Japan Sea in response to eustatic sea-level drop during LGM: Reconstruction based on salinity-balance model. Quaternary Res, 1998, 37:221–233, 10.4116/jaqua.37.221
Author information
Authors and Affiliations
Corresponding author
Additional information
Supported by the National Basic Research Program of China (Grant No. 2005cb121000), and National Natural Science Foundation of China (Grant No. 30771630)
Rights and permissions
About this article
Cite this article
Pan, M., Yu, Q., Xia, Y. et al. Characterization of mitochondrial genome of Chinese wild mulberry silkworm, Bomyx mandarina (Lepidoptera: Bombycidae). SCI CHINA SER C 51, 693–701 (2008). https://doi.org/10.1007/s11427-008-0097-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-008-0097-6