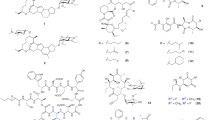
Abstract
Chemomics is an interdisciplinary study using approaches from chemoinformatics, bioinformatics, synthetic chemistry, and other related disciplines. Biological systems make natural products from endogenous small molecules (natural product building blocks) through a sequence of enzyme catalytic reactions. For each reaction, the natural product building blocks may contribute a group of atoms to the target natural product. We describe this group of atoms as a chemoyl. A chemome is the complete set of chemoyls in an organism. Chemomics studies chemomes and the principles of natural product syntheses and evolutions. Driven by survival and reproductive demands, biological systems have developed effective protocols to synthesize natural products in order to respond to environmental changes; this results in biological and chemical diversity. In recent years, it has been realized that one of the bottlenecks in drug discovery is the lack of chemical resources for drug screening. Chemomics may solve this problem by revealing the rules governing the creation of chemical diversity in biological systems, and by developing biomimetic synthesis approaches to make quasi natural product libraries for drug screening. This treatise introduces chemomics and outlines its contents and potential applications in the fields of drug innovation.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Collins FS, Morgan M, Patrinos A. The human genome project: Lessons from large-scale biology. Science, 2003, 300: 286–290
Luscombe NM, Greenbaum D, Gerstein M. What is Bioinformatics? A proposed definition and overview of the field. Methods Inf Med, 2001, 40: 346–358
Dhar PK. The next step in biology: A periodic table. J Biosci, 2007, 32: 1005–1008
Lander ES. The new genomics: Global views of biology. Science, 1996, 274: 536–539
Biro JC, Benyó B, Sansom C, Szlávecz Á, Fördös G, Micsik T. A common periodic table of codons and amino acids. Biochem Biophys Res Commun, 2003, 306: 408–415
Van Drie JH. Pharmacophore discovery-Lessons learned. Curr Pharm Design, 2003, 9: 1649–1664
Evans BE, Rittle KE, Bock MG, Dipardo RM, Freidinger RM, Whitter WL. Methods for drug discovery: Development of potent, selective, orally effective cholecystokinin antagonists. J Med Chem, 1988, 31: 2235–2246
DeSimone RW, Currie KS, Mitchell SA, Darrow JW, Pippin DA. Privileged structures: Applications in drug discovery. Comb Chem & High T Scr, 2004, 7: 473–493
Lewell XQ, Judd DB, Watson SP, Hann MM. RECAP—retrosynthetic combinatorial analysis procedure: A powerful new technique for identifying privileged molecular fragments with useful applications in combinatorial chemistry. J Chem Inf Comput Sci, 1998, 38: 511–522
Corey EJ. General methods for the construction of complex molecules. Pure Appl Chem, 1967, 14: 19–38
Burke MD, Schreiber SL. A planning strategy for diversity-oriented synthesis. Angew Chem Int Ed, 2004, 43: 46–58
Böhm HJ. Site-directed structure generation by fragment-joining. Perspect Drug Discovery Des, 1995, 3: 21–33
Rees DC, Congreve M, Murray CW, Carr R. Fragment-based lead discovery. Nat Rev Drug Discov, 2004, 3: 660–672
Miflin BJ, Lea PJ. Amino acid metabolism. Ann Rev Plant Physiol, 1977, 28: 299–329
Dewick PM. Medicinal Natural Products: A Biosynthetic Approach. Wiley, 2009
Munos B. Lessons from 60 years of pharmaceutical innovation. Nat Rev Drug Discov, 2009, 8: 959–968
Michal G. Biochemical Pathways: An Atlas of Biochemistry and Molecular Biology. Wiley, 1999
Kotera M, Hirakawa M, Tokimatsu T, Goto S, Kanehisa M. The KEGG databases and tools facilitating omics analysis: Latest developments involving human diseases and pharmaceuticals, next generation microarray bioinformatics. Methods Mol Biol, 2012, 802: 19–39
Reitz M, Sacher O, Tarkhov A, Trümbach D, Gasteiger J. Enabling the exploration of biochemical pathways. Org Biomol Chem, 2004, 2: 3226–3237
Johnson RE, Washington MT, Prakash S, Prakash L. Fidelity of human DNA polymerase η. J Biol Chem, 2000, 275: 7447–7450
Pray L A. DNA replication and causes of mutation. Nat Educ, 2008, 1: 1
Brown JR, Gentry D, Becker JA, Ingraham K, Holmes DJ, Stanhope MJ. Horizontal transfer of drug-resistant, aminoacyl-transfer-RNA synthetases of anthrax and Gram-positive pathogens. EMBO reports, 2003, 4: 692–698
Strieker M, Tanović A, Marahiel MA. Nonribosomal peptide synthetases: Structures and dynamics. Curr Opin Struct Biol, 2010, 20: 234–240
Wiermann R. Secondary plant products and cell and tissue differentiation. In: Conn EE, Ed. The Biochemistry of Plants. New York: Academic Press, 1980. 85–137
Nicolaou KC, Pfefferkorn JA, Roecker AJ, Cao GQ, Barluenga S, Mitchell HJ. Natural product-like combinatorial libraries based on privileged structures. 1. General principles and solid-phase synthesis of benzopyrans. J Am Chem Soc, 2000, 122: 9939–9953
Bu XZ, Wu XM, Ng NLJ, Mak CK, Qin CG, Guo ZH. Synthesis of gramicidin S and its analogues via an on-resin macrolactamization assisted by a predisposed conformation of the linear precursors. J Org Chem, 2004, 69: 2681–2685
Zhou J, Xie G, Yan X. Encyclopedia of Traditional Chinese Medicines—Molecular Structures, Pharmacological Activities, Natural Sources and Applications. Berlin: Springer-Verlag, 2011
Janga SC, Tzakos A. Structure and organization of drug-target networks: Insights from genomic approaches for drug discovery. Mol Bio Syst, 2009, 5: 1536–1548
Hopkins AL. Network pharmacology: The next paradigm in drug discovery. Nat Chem Biol, 2008, 4: 682–690
Berger SI, Iyengar R. Network analyses in systems pharmacology. Bioinformatics, 2009, 25: 2466–2472
Purnick PEM, Weiss R. The second wave of synthetic biology: From modules to systems. Nat Rev Mol Cell Biol, 2009, 10: 410–422
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Xu, J., Gu, Q., Liu, H. et al. Chemomics and drug innovation. Sci. China Chem. 56, 71–85 (2013). https://doi.org/10.1007/s11426-012-4761-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11426-012-4761-0