Summary
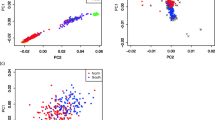
Genetic dissection of complex diseases is both important and challenging. The human major histocompatibility complex is involved in many human diseases and genetic mechanisms. This highly polymorphic chromosome region has been extensively studied in Caucasians but not as well in Asians. Thus, we compared genotypic distributions, linkage disequilibria and haplotype blocks between Caucasian and Taiwan’s Han Chinese populations. Moreover, we investigated the population admixture and phylogenetic system in Han Chinese residing in Taiwan. The results show that Taiwan’s Han Chinese differ drastically in genotypic information compared with Caucasians but are relatively homogeneous among the three major ethnic subgroups, Minnan, Hakka and Mainlanders. Differences in allele frequency (AF) between Taiwanese and Caucasians in some disease-associated loci may reveal clues to differences in disease prevalence. The results of ethnic heterogeneity imply that public databases should be used with caution in cases where the study population(s) differs from the population characterized in the database. The high homogeneity we observed among the Taiwanese subpopulations mitigates the possibility of spurious association caused by ignoring population stratification in Taiwanese disease gene association studies. These results are useful for understanding our genetic background and designing future disease gene mapping studies.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Wang W.Y.S., Barratt B.J., Clayton D.G. and Todd J.A. (2005). Genome-wide association studies: theoretical and practical concerns. Nat. Rev. Genet. 6: 109–118
Hirschhorn J.N. and Daly M.J. (2005). Genome-wide association studies for common diseases and complex traits. Nat. Rev. Genet. 6: 95–108
Maniatis N., Collins A. and Gibson J. (2004). Positional cloning by linkage disequilibrium. Am. J. Hum. Genet. 74: 846–855
Freedman M.L., Reich D., Penney K.L., McDonald G.J., Mignault A.A., Patterson N., Gabriel S.B., Topol E.J., Smoller J.W., Pato C.N., Pato M.T., Petryshen T.L., Kolonel L.N., Lander E.S., Sklar P., Henderson B., Hirschhorn J.N. and Altshuler D.(2004). Assessing the impact of population stratification on genetic association studies. Nat. Genet. 36: 388–393
Marchini J., Cardon L.R., Phillips M.S. and Donnelly P. (2004). The effects of human population structure on large genetic association studies. Nat. Genet. 36: 512–517
Helgason A., Yngvadóttir B., Hrafnkelsson B., Gulcher J. and Stefánsson K. (2005). An Icelandic example of the impact of population structure on association studies. Nat. Genet. 37: 90–95
Walsh E.C., Mather K.A., Schaffner S.F., Farwell L., Daly M.J., Patterson N., Cullen M., Carrington M., Bugawan T.L., Erlich H., Campbell J., Barrett J., Miller K., Thomson G., Lander E.S. and Rioux J.D. (2003). An integrated haplotype map of the human major histocompatibility complex. Am. J. Hum. Genet. 73: 580–590
Stenzel A., Lu T., Koch W.A., Hampe J., Guenther S.M., De La Vega F.M., Krawczak M. and Schreiber S. (2004). Patterns of linkage disequilibrium in the MHC region on human chromosome 6p. Hum. Genet. 114: 377–385
Horton R., Wilming L., Rand V., Lovering R.C., Bruford E.A., Khodiyar V.K., Lush M.J., Povey S., Talbot C.C., Wright M.W., Wain H.M., Trowsdale J., Ziegler A. and Beck S. (2004). Gene map of the extended human MHC. Nat. Rev. Genet. 5: 889–899
Miretti M.M., Walsh E.C., Ke X., Delgado M., Griffiths M., Hunt S., Morrison J., Whittaker P., Lander E.S., Cardon L.R., Bentley D.R., Rioux J.D., Beck S. and Deloukas P.(2005). A high-resolution linkage disequilibrium map of the human major histocompatibility complex and first generation of tag single-nucleotide polymorphisms. Am. J. Hum. Genet. 76: 634–646
Shiina T., Inoko H. and Kulski J.K. (2004). An update of the HLA genomic region, locus information and disease associations. Tissue Antigens 64: 631–649
Meacham W. (1981). Origins and development of the Yueh coastal Neolithic: a microcosm of culture change of the mainland of East Asia. In: Keightly D.K. (eds), The Origins of Chinese Civilization. University of California Press, California
Fang H.K. (eds) (1996). Deep Investigation of the Origin of Hakka. Woolin Publishing, Taipei, Taiwan
Lin M., Chu C.C., Chang S.L., Lee H.L., Loo J.H., Akaza T., Juji T., Ohashi J. and Tokunaga K. (2001). The origin of Minnan and Hakka, the so-called “Taiwanese”, inferred by HLA study. Tissue Antigens 57: 192–199
Zhao T.M. and Lee T.D. (1998). Gm and Km allotypes in 74 Chinese populations: a hypothesis of the origin of the Chinese Nation. Hum. Genet. 83: 101–110
Lo H.L. (eds) (1933). An Introduction to the Study of the Hakkas in its Ethnic, Historical, and Cultural Aspects. SMC Publishing, Taipei, Taiwan
Pan W.H., Fann C.S.J., Wu J.Y., Hung S.I., Hung Y.T., Chen Y.J., Hsu C.L., Liao C.J. and Chen, Y.T., Establishment of Taiwan Han Chinese cell and gene bank: comparing SNP profiles in MHC region with Caucasians. The American Society of Human Genetics 54th Annual Meeting, Toronto, Ontario, Canada, October 26–30, 2004.
Hung S.I., Chung W.H., Liou L.B., Chu C.C., Lin M., Huang H.P., Lin Y.L., Lan J.L., Yang L.C., Hong H.S., Chen M.J., Lai P.C., Wu M.S., Chu C.Y., Wang K.H., Chen C.H., Fann C.S.J., Wu J.Y. and Chen Y.T. (2005). HLA-B*5801 allele as a genetic marker for severe cutaneous adverse reactions caused by allopurinol. Proc. Natl Acad. Sci. USA 102: 4134–4139
Benjamini Y. and Hochberg Y. (1995). Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B Met. 57: 289–300
Barrett J.C., Fry B., Maller J. and Daly M.J. (2005). Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21: 263–265
Lewontin R.C. (1964). The interaction of selection and linkage. I. General considerations; heterotic models. Genetics 49: 49–67
Gabriel S.B., Schaffner S.F., Nguyen H., Moore J.M., Roy J., Blumenstiel B., Higgins J., DeFelice M., Lochner A., Faggart M., Liu-Cordero S.N., Rotimi C., Adeyemo A., Cooper R., Ward R., Lander E.S., Daly M.J. and Altshuler D.(2002). The structure of haplotype blocks in the human genome. Science 296: 2225–2229
Pritchard J.K., Stephens M. and Donnelly P.(2000). Inference of population structure using multilocus genotype data. Genetics 155: 945–959
Felsenstein J.(1995). Phylip: Phylogeny inference package, ver. 3.6, Software User Manual. University of Washington, Washington
Nei M.(1972). Genetic distance between populations. Am. Nat. 106: 283–292
Woo K.S., Chook P., Chan W.B., So W.Y., Cockram C.S. and Celermajer D.S. (2001). Type 1 diabetes and arterial dysfunction in asymptomatic Chinese adults. Diabetes Care 24: 173
Lench N., Iles M.M., Mackay I., Patel R., Sagoo G.S., Ward S.J., Dechairo B., Olavesen M., Carey A., Duff G.W., Cork M.J. and Tazi-Ahnini R. (2005). Single-point haplotype scores telomeric to human leukocyte antigen-C give a high susceptibility major histocompatability complex haplotype for psoriasis in a Caucasian population. J. Invest. Dermatol. 124: 545–552
Hung S.I., Chung W.H. and Chen Y.T. (2005). HLA-B genotyping to detect carbamazipine-induced Stevens-Johnson syndrome: implications for personalizing medicine. Pers. Med. 2: 225–237
Bhimma R. and Coovadia H.M. (2004). Hepatitis B virus associated nephropathy. Am. J. Nephrol. 24: 198–211
Lin J.C., Cherng J.M., Lin H.J., Tsang C.W., Liu Y.X. and Lee S.P. (2004). Amino acid changes in functional domains of latent membrane protein 1 of Epstein-Barr virus in nasopharyngeal carcinoma of southern China and Taiwan: prevalence of an HLA A2-restricted ‚epitope-loss variant’. J. Gen. Virol. 85: 2023–2034
Lu C.C., Chen J.C., Tsai S.T., Jin Y.T., Tsai J.C., Chan S.H. and Su I.J. (2005). Nasopharyngeal carcinoma–susceptibility locus is localized to a 132 kb segment containing HLA-A using high-resolution microsatellite mapping. Int. J. Cancer 115: 742–746
Odom D.T., Zizlsperger N., Gordon D.B., Bell G.W., Rinaldi N.J., Murray H.L., Volkert T.L., Schreiber J., Rolfe P.A., Gifford D.K., Fraenkel E., Bell G.I. and Young R.A. (2004). Control of pancreas and liver gene expression by HNF transcription factors. Science 303: 1378–1381
Zouari B.K., Chouchane L., Jellouli K., Cherif S., Haddad S., Gabbouj S. and Danguir J. (2004). Polymorphism of stress protein HSP70–2 gene in Tunisians: susceptibility implications in type 2 diabetes and obesity. Diabetes Metab. 30: 175–180
Zimmet P., Alberti K.G. and Shaw J. (2001). Global and societal implications of the diabetes epidemic. Nature 414: 782–787
Pociot F., Ronningen K.S. and Nerup J. (1993). Polymorphic analysis of the human MHC-linked heat shock protein 70 (HSP70–2) and HSP70-Hom genes in insulin-dependent diabetes mellitus (IDDM). Scand. J. Immunol. 38: 491–495
Acknowledgments
This project was funded by grants from the Taiwan National Science Council (NSC 93-2320-B-001-0.26 and NSC 94-311-B001-007-Y) and the Genomics and Proteomics Program of Academia Sinica (94F003-2).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
11373_2006_9077_MOESM1_ESM.pdf
Supplementary material is available for this article at http://dx.doi.org/10.1007/s11373-006-9077-7 and is accessible for authorized users.
Rights and permissions
About this article
Cite this article
Yang, HC., Lin, CH., Hsu, CL. et al. A comparison of major histocompatibility complex SNPs in Han Chinese residing in Taiwan and Caucasians. J Biomed Sci 13, 489–498 (2006). https://doi.org/10.1007/s11373-006-9077-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11373-006-9077-7