Abstract
Purpose
Long-term intensive cultivation and heavy fertilization improve the nutritional conditions in acidic Ultisols, but also sharply accelerate soil acidification. However, the impact of such dramatic environmental changes on soil microorganisms is unclear. The aims of this work were to investigate the responses of microbial community composition and metabolic function to long-term fertilization, and to determine the key factors that primarily drive microbial changes in acidic Ultisols.
Materials and methods
A long-term fertilization experiment under a winter wheat–summer maize rotation was established in 1990 in acidic Ultisols of southern China. Soils were collected from four treatments in June 2014: (1) non-fertilization control (CK); (2) only N fertilization (N); (3) N, P, and K fertilization (NPK); and (4) NPK plus manure (NPKM; 70% of total N obtained from manure). The amount of N used in all N treatments was 300 kg N ha−1 year−1. The soil pH, cation exchange capacity (CEC), soil organic carbon (SOC), total nitrogen (TN), phosphorus (TP) and potassium (TK), available nitrogen (AN), phosphorus (AP), and potassium (AK) were measured. Soil microbial community composition and metabolic function were determined by phospholipid fatty acids analysis (PLFA) and community-level physiological profile (CLPP) method, respectively.
Results and discussion
Compared with CK, NPKM significantly increased total PLFA biomass and average well color development (AWCD); NPK increased total PLFA biomass by 2.2 times, but its AWCD was not significantly different from CK, indicating that microbial metabolic efficiency in NPK decreased. N decreased total PLFA biomass by 27.9%, while almost completely inhibiting metabolic activity. NPKM maintained microbial functional diversity indexes at similar levels as CK, while NPK and N significantly decreased microbial functional diversity indexes. Redundancy analysis (RDA) revealed that soil microbial community composition and metabolic pattern were more stably maintained by application of manure compared to chemical fertilizers. Soil pH showed the primary effect on microbial community composition, metabolic activity, and functional diversity indexes.
Conclusions
This research demonstrated that the negative effects of Ultisol acidification induced by long-term application of chemical N fertilizer on microorganisms overwhelmed the positive effects of soil nutrition improvement. The inhibiting effect of serious acidification on microbial metabolic function was stronger than that on community composition. Microorganisms live in a low active metabolic state to resist serious Ultisols acidification. Therefore, fertilization in acidic Ultisols should be based on the premise of preventing soil further acidification.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Fertilization is an important way to improve soil fertility and maximize crop yields (Dong et al. 2012; Li et al. 2017). In recent decades, intensive application of chemical fertilizers has replaced traditional organic fertilization in China (Cai et al. 2014). For instance, from 1980 to 2014, the application of chemical fertilizers increased by 2.12 times, and nitrogen (N) fertilizer contributed 55.3% to the total increase in chemical fertilizers (National Bureau of Statistics of China 2016). However, excessive chemical N fertilizer can lead to soil acidification, which was the main obstacle to modern agricultural production. The negative effects of soil acidification are closely related to soil type, land use pattern, crop cultivation, and fertilization regime (Guo et al. 2010; Rousk et al. 2010).
Ultisol is a typical soil in tropical and subtropical regions. In China, Ultisols comprise 21% of the country’s land area (about 2 million km2), and support approximately 40% of population (Huang et al. 2012). To meet the tremendous demand for food, high-intensity cultivation is frequently adopted in Ultisols of southern China, such as rotation of winter wheat and summer maize (Zhang et al. 2014). Thus, heavy fertilization is required to maintain soil productivity. Ultisols generally undergo a slow natural acidification process, exhibiting low soil pH, low organic matter content, and nutrient deficiencies, all of which make Ultisols more susceptible to anthropogenic acidification (Abdulahaal et al. 2017). Although application of chemical fertilizers significantly improves the fertility level of Ultisols, it also sharply accelerates the soil acidification process (Cai et al. 2014 and 2015). Guo et al. (2010) found that in some Ultisols with perennial rotation of southern China, anthropogenic acidification caused by application of chemical N fertilizers in recent decades has even surpassed natural acidification that typically occurs over hundreds to millions of years. The whole agro-ecosystem is inevitably affected by such rapid Ultisols acidification. The effects of Ultisols acidification on soil nutrient transformation and supply, crop growth, and yield have been studied (Huang et al. 2010; Cai et al. 2015). However, the response of soil microorganisms to serious Ultisols acidification is unclear.
Soil microorganisms play an essential role in soil structure formation, organic matter decomposition, and nutrient biogeochemical cycles (Kušlienė et al. 2014). They also have significant effects on crop health and yield (Berg 2009). Soil microorganisms are very sensitive to environmental changes. Nutrient deficiencies (e.g., C, N, P, and K) and environmental stresses (e.g., soil pH, moisture, and porosity) are the most important limiting factors for microbial growth and activity. A review (Allison and Martiny 2008) reported that 84% of the selected papers indicated that soil microbial characteristics were sensitive to the improved soil nutritional conditions caused by fertilization. Ultisols usually lack C, N, and P because of high weathering and leaching. Therefore, in many cases, soil nutritional status (e.g., SOC, TN, and AP) drive changes in soil microbial properties (Chen et al. 2015). However, some studies have demonstrated that pH was the key indicator affecting soil microbial community (Ma et al. 2015). Long-term fertilization triggered significant changes in Ultisols properties, so determining the dominant factors of microbial community shifts can help to adjust fertilization strategy in a time and effectively manner.
Phospholipid fatty acid (PLFA) analysis and community-level physiological profile (CLPP) methods have been widely employed to study soil microbial community composition and metabolic function, respectively (Chen et al. 2015; Ma et al. 2015). In combination with the PLFA and CLPP methods, the responses of microbial community to different fertilization regimes can be comprehensively assessed. In this work, a long-term fertilization field experiment (1990–2014) under a winter wheat–summer maize rotation in acidic Ultisols of southern China was investigated. The objectives of this research were (1) to study soil acidity and nutrients following long-term fertilizations, (2) to investigate the changes in soil microbial community composition and metabolic function, and (3) to determine the key factors that primarily drive soil microbial changes. The findings will help to maintain soil quality and achieve sustainable agriculture in acidic Ultisols of southern China.
2 Materials and methods
2.1 Site description
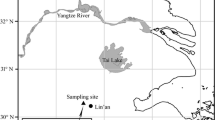
The long-term fertilization experiment was established in 1990 at the Red Soil Experimental Station (26° 45′ 12″N, 111° 52′ 32″E) of the Chinese Academy of Agricultural Sciences at Qiyang, Hunan Province, southern China. This region is a subtropical humid monsoon climate with annual mean temperature of 18.0 °C, mean precipitation of 1250–1400 mm, mean evaporation of 1470 mm, 300 frost-free days, and 1610 sunlight hours (Cai et al. 2015). The tested soil, derived from quaternary red clay, was wasteland before the experiment. The initial properties of the topsoil (0–20 cm) in 1990 were soil pH of 5.70 (water:soil = 2.5:1), soil organic carbon (SOC) of 6.06 g kg−1; total nitrogen (TN), phosphorus (TP), and potassium (TK) of 1.07, 0.52, and 13.7 g kg−1, respectively; available nitrogen (AN), phosphorus (AP), and potassium (AK) of 79.0, 13.9, and 104 mg kg−1, respectively; and soil cation exchange capacity (CEC) of 8.99 cmol(+) kg−1.
2.2 Experimental design and soil sampling
This work involved four treatments: (1) non-fertilization control (CK); (2) only N fertilization (N); (3) N, P, and K fertilization (NPK); and (4) NPK plus manure (NPKM). The application rates of N, P, and K were 300, 53, and 100 kg ha−1 year−1, respectively. In NPKM treatment, 70% of the total N obtained from pig manure and the remaining 30% came from urea. Between the two crops, 30% of the total chemical fertilizers were applied for winter wheat and the remaining 70% were applied for summer maize. However, all manure was applied before wheat sowing. Chemical N, P, and K fertilizers were urea, calcium superphosphate, and potassium chloride, respectively. The manure was completely composted pig manure with TC, TN, TP, and TK of 368.4, 18.2, 13.7, and 13.3 g kg−1, respectively. In order to simulate the actual field cultivation, each treatment was duplicated in relatively large experimental plots (10 × 20 m), which allowed common field management similar to that in local farmland (Cai et al. 2015; Chen et al. 2014). There was a 1-m deep concrete barrier between plots. More details of the experimental design and implementation were described by Cai et al. (2014, 2015).
Surface soil samples (0–20 cm) were collected in June 2014 (after wheat was harvested and before maize was planted). Twenty cores per plot were sampled randomly using a soil auger, and then were thoroughly mixed to form a composite sample. The composite sample was passed through a 2-mm sieve and then divided into three parts: one part was air-dried for further chemical analysis, the second part was preserved at 4 °C for CLPP analysis, and the remaining part was freeze-dried and kept at − 40 °C for PLFA analysis.
2.3 Analysis of soil chemical properties
Soil pH was determined with a soil-to-water ratio of 1:2.5 (w/v). SOC was determined by dichromate oxidation and titration with ferrous sulfate (Pansu and Gautheyrou 2006). TN, TP, and TK were determined using the Kjeldahl digestion-distillation method, molybdenum-blue colorimetry, and flame photometry, respectively (Pansu and Gautheyrou 2006). AN, AP, and AK were determined using alkali hydrolyzation, the Bray method, and flame photometry, respectively (Pansu and Gautheyrou 2006). CEC was determined by the ammonium acetate method (Pansu and Gautheyrou 2006).
2.4 PLFA analysis
Soil microbial community composition was determined based on PLFA analysis as described by Buyer et al. (2010). Phospholipid fatty acids were extracted from 3 g of freeze-dried soil with a mixture of chloroform, methanol, and phosphate buffer (pH 7.4) in a 1:2:0.8 volumetric ratio. Polar lipids in the extracted PLFAs were purified using a 500 mg sep-pak silica cartridge (Waters & Associates, Inc.) with sequential washes of chloroform, acetone, and methanol, and then dried using N2. The collections were converted to fatty acid methyl esters using methanolic KOH, and methyl nonadecanoate (19:0) was added as internal standard. Then, they were quantified using an Agilent 7890A gas chromatograph (Agilent Technologies, USA) with the MIDI Sherlock system (MIDI, Inc., USA).
The detected fatty acid methyl esters were categorized into various taxonomic microbial groups as follows: the fatty acid 18:2 ω6c was considered as fungi (Ma et al. 2016), the fatty acids with a methyl branch on the 10th C were chosen to indicate actinomycetes (Zelles 1997), the branched saturated fatty acids were selected to indicate gram-positive (G+) bacteria, and the monounsaturated and cyclopropyl fatty acids were selected to indicate gram-negative (G−) bacteria (Zelles 1997). The sum of biomass in G+ bacteria and G− bacteria was represented as the total bacterial biomass. The ratio of G+ bacterial to G− bacterial biomass (G+/G−) and the ratio of fungal to bacterial biomass (F/B) were also calculated, respectively. The ratio of total saturated fatty acids to total monounsaturated fatty acids (sat/mono) was calculated and used as a comprehensive indicator of environmental conditions (Fierer et al. 2003).
2.5 CLPP analysis
Soil microbial metabolic function was evaluated using CLPP method with Biolog Eco Plates™ (Biolog Inc., USA). Fresh soil samples, which are equivalent to 5 g of dry soil, were added into 45 ml pre-sterilized 0.85% (w/v) NaCl solution. The mixture was shaken at 200 rpm for 30 min and then settled for 10 min. The suspension was further diluted 10-fold, and then, 150 μl dilution was inoculated into every well of the Biolog Eco Plates™. The plates were incubated at 25 °C for 10 days, and the value of optical density (OD) of each well was recorded every 24 h using a plate reader (Bio-Tek Instrument Inc., USA) at dual wavelengths of 590 and 750 nm. Finally, the 590-nm values minus the 750-nm values were used to denote the carbon source metabolic capacity (Peng et al. 2016). The data measured at 120 h were selected to calculate the average well color development (AWCD), substrate richness (S), Shannon index (H′), Simpson index (D), and Pielou index (E). The S index was simply measured by counting the total number of utilized substrates. The others were determined using the following formulas (Garland 1996):
where OD i is the optical density value of the ith well and p i is the ratio color development of the ith well on the total color development of all wells.
2.6 Statistical analysis
Significant differences among treatments were tested by one-way analysis of variance (ANOVA) with Tukey’s HSD post hoc test at P < 0.05. Relationships between soil chemical variables and microbial parameters were determined by stepwise regression analysis. Redundancy analysis (RDA) was used to study the changes in soil microbial community composition and metabolic pattern, and to determine the primary factors that drive these changes. Before RDA, the chord transformation was performed on each dataset of PLFA and CLPP (Legendre and Gallagher 2001), the soil chemical variables were standardized, and forward selection was conducted to retain variables that were significantly correlated to the dataset of PLFA or CLPP (Blanchet et al. 2008). Variance partitioning analysis (VPA) was applied to evaluate the effects of soil chemical variables on total variance of each RDA model. Relationships between individual carbon sources (or PLFAs) and ordination axes (RDA1 or 2) were determined by Pearson’s correlation analysis. One-way ANOVA, stepwise regression analysis, and Pearson’s correlation analysis were performed using PASW Statistics version 18.0 (IBM, USA). RDA and VPA were performed using R project (version 3.2.2) with the Vegan package.
3 Results
3.1 Soil acidity and nutrients
Soil chemical properties were significantly affected by long-term fertilization (Table 1). Compared with CK (pH 5.30), soil pH in N and NPK significantly decreased to 4.02 and 4.15, respectively, while pH in NPKM significantly increased to 5.99. Both NPK and NPKM improved soil nutrients with different extents. Compared with CK, NPK increased soil SOC, TN, TP, and AN by 54, 44, 150, and 70%, respectively, while those in NPKM were increased by 111, 118, 365, and 119%, respectively. Soil AP and AK responded much more sensitively: in NPK both increased by 22.4 and 3.71 times, respectively, and in NPKM, they increased by 55.5 and 9.00 times, respectively. Soil CEC in NPK and NPKM increased by 12.5 and 31.1%, respectively. However, N treatment only increased soil TN content by 0.33 times compared with CK, and significantly decreased the soil C/N ratio.
3.2 PLFA
A total of 30 PLFAs were used for data analysis (Table S1, Electronic Supplementary Material). PLFA results showed that NPKM and NPK significantly (P < 0.05) increased the biomass of total PLFA and all microbial groups (fungi, G+ and G− bacteria, and actinomycetes), while N treatment significantly decreased the biomass of total PLFA, G− bacteria, and actinomycetes (Fig. 1a). Compared with CK, NPKM significantly increased the relative abundance of G− bacteria (from 24.77 to 27.41%), but significantly decreased the relative abundance of actinomycetes (from 23.93 to 21.14%) (Fig. 1b). However, NPK significantly increased the relative abundance of G+ bacteria (from 50.19 to 54.25%), and significantly decreased the relative abundance of G− bacteria (from 24.77 to 19.93%). The relative abundance of fungi in fertilization treatments were all significantly improved as compared with CK.
The amounts (a) and relative abundance (b) of soil different microbial PLFAs under different treatments in a long-term (1990–2014) fertilization experiment in Qiyang, Hunan, China. CK non-fertilization control; N only chemical nitrogen fertilizer; NPK chemical nitrogen, phosphorous, and potassium fertilizers; NPKM chemical nitrogen, phosphorous, and potassium fertilizers plus manure. The error bar indicates standard deviation
Compared to CK, the G+/G− ratio was significantly increased in NPK, but significantly decreased in NPKM; the F/B ratio in all fertilization treatments and the sat/mono ratio in N and NPK treatments were significantly increased (Table 2). Stepwise regression analysis (Table 3) demonstrated that soil pH was the primary factor affecting the ratio of G+/G− and sat/mono, while SOC and TN were the primary factors affecting the total PLFA biomass and the ratio of F/B, respectively.
RDA results showed that the ordination axes 1 (RDA1) and 2 (RDA2) explained 79.44% (F = 56.480, P < 0.001) and 7.90% (F = 5.615, P < 0.001) of the total variance of the RDA model (F = 31.048, P < 0.001), respectively (Fig. 2a). The four treatments were distributed in four quadrants and separated from each other (P < 0.05), indicating that long-term different fertilization have led to different soil microbial community compositions. Soil pH and C/N ratio were the only two retained explanatory variables in the model through forward selection. Soil pH, accounted for 64.93% (F = 55.414, P < 0.001) of the total variance, was found to be the primary factor affecting the shifts in soil microbial community compositions and separating N and NPK from CK and NPKM along RDA1 (Fig. 3a). However, soil C/N ratio only explained 9.40% (F = 6.681, P = 0.012) of the total variance, and further distributed the treatments along RDA2 (Fig. 3b).
Redundancy analysis of PLFA (a) and CLPP (b) of soil microbial communities under different treatments in a long-term (1990–2014) fertilization experiment in Qiyang, Hunan, China. CK non-fertilization control; N only chemical nitrogen fertilizer; NPK chemical nitrogen, phosphorous, and potassium fertilizers; NPKM chemical nitrogen, phosphorous, and potassium fertilizers plus manure; C/N carbon to nitrogen ratio
Linear correlation between soil pH (a) and PLFA loading scores on RDA1, linear correlation between soil C/N (b) and PLFA loading scores on RDA2; linear correlation between soil pH (c) and CLPP loading scores on RDA2, linear correlation between soil C/N (d) and CLPP loading scores on RDA1 of different treatments in a long-term (1990–2014) fertilization experiment in Qiyang, Hunan, China. C/N carbon to nitrogen ratio
Pearson’s correlation analysis (Table S1, Electronic Supplementary Material) showed that twenty-two species of PLFAs (four actinomycetes, ten G− bacteria, and eight G+ bacteria) were significantly correlated to RDA1 data; thus these species induced the shifts among treatments along RDA1. However, only three species of PLFAs (two G− bacteria and one G+ bacteria) led to the differentiation among treatments along RDA2.
3.3 CLPP
AWCD of carbon sources utilization could describe soil microbial metabolic activity. Compared with CK, NPKM significantly increased AWCD values and the metabolic activities of six categories of carbon sources (Figs. 4 and 5a). However, the AWCD values in N treatment stayed at a very low level (nearly 0), which indicated that soil microorganisms hardly metabolized any carbon source (Figs. 4 and 5a). The AWCD values in NPK maintained almost the same level as CK during the entire incubation period (Fig. 4); furthermore, NPK and CK only showed significant differences in the metabolic activities of amines (Fig. 5a).
Average well color development (AWCD) of soil microbial communities under different treatments in a long-term (1990–2014) fertilization experiment in Qiyang, Hunan, China. CK non-fertilization control; N only chemical nitrogen fertilizer; NPK chemical nitrogen, phosphorous, and potassium fertilizers; NPKM chemical nitrogen, phosphorous, and potassium fertilizers plus manure. The error bar indicates standard deviation
The metabolic activities (a) and relative abundance (b) of six categories of carbon substrate by soil microbial communities under different treatments in a long-term (1990–2014) fertilization experiment in Qiyang, Hunan, China. CK non-fertilization control; N only chemical nitrogen fertilizer; NPK chemical nitrogen, phosphorous, and potassium fertilizers; NPKM chemical nitrogen, phosphorous, and potassium fertilizers plus manure; AM amines; AA amino acids; CH carbohydrates; CA carboxylic acids; PC phenolic compounds; PM polymers. The error bar indicates standard deviation
Long-term fertilization changed the preference of soil microorganisms to metabolize different categories of carbon sources (Fig. 5b). The relative abundance of amines in N and NPK treatments were 1.20 and 0.91%, respectively, which were significantly lower than that in CK (7.96%). In contrast, the relative abundance of carbohydrates in N (44.64%) and NPK (37.29%) were significantly higher than that in CK (24.37%). In addition, the relative abundance of polymers in N (40.22%) was also significantly higher than that in CK (15.50%), while the relative abundance of amino acids, carboxylic acids, and phenolic compounds were significantly decreased. Compared with CK, NPKM significantly increased the metabolic activities of six categories of carbon sources by more than 100%, while the relative abundance of six categories of carbon sources showed little difference between NPKM and CK (P > 0.05). Similarly, there were minimal differences in all functional diversity indexes between NPKM and CK (Table 2). However, compared with CK, all diversity indexes in N and NPK treatments were significantly decreased, except the S index in NPK. Stepwise regression analysis indicated that the AWCD and all diversity indexes were significantly affected by soil pH (Table 3).
The RDA model (F = 7.368, P < 0.001) based on CLPP data is shown in Fig. 2b. RDA1 explained 47.90% (F = 11.370, P = 0.003) of the total variance and clearly differentiated the N treatment from the other treatments. In addition, RDA2 explained 14.18% (F = 3.365, P = 0.001) of the total variance and further separated NPK from NPKM and CK. NPKM and CK were located in the third quadrant of the RDA figure and were very close, indicating that they had similar metabolic patterns, and this result was in accordance with the above-mentioned results of diversity indexes (Table 2). Significant chemical variables (P < 0.05) were still soil pH and C/N ratio, and they accounted for 19.02% (F = 6.770, P < 0.001) and 33.56% (F = 7.965, P < 0.001) of the total variance, respectively. Soil pH and C/N ratio were primary factors that are causing differentiation among treatments along RDA2 and RDA1, respectively (Fig. 3c, d).
Twelve types of carbon sources (three amino acids, three carbohydrates, three carboxylic acids, one phenolic compound, and two polymers) had significant correlations with RDA1, and three types of carbon sources (two amines and one amino acid) had significant correlations with RDA2 (Table S1, Electronic Supplementary Material). Long-term different fertilization regimes changed the metabolic preferences of soil microorganisms on the above-mentioned carbon sources, resulting in metabolic pattern shifts.
4 Discussion
4.1 Effects of long-term fertilization on Ultisol acidity and nutrients
Excessive application of chemical N fertilizer has been the main reason of soil acidification in Chinese farmland in recent decades (Sun et al. 2015). However, the extent and trends of Ultisol acidification following fertilization were controversial. For example, Guo et al. (2010) reported that the decline of pH in Ultisols was less than that in other soil types, while many studies indicated that infertile Ultisols were more susceptible to acidification (Cai et al. 2014; Abdulahaal et al. 2017). Our results demonstrated that long-term chemical fertilizer application sharply accelerated the acidification process of acidic Ultisols. Compared with the soil pH of the original Ultisols (5.70), the pH in NPK and N treatments decreased by 1.55 and 1.68 units, respectively. Different soil acidification stages are accompanied by the weathering and liberation of different elements. In particular, when pH is below 4.2, the soil enters aluminum (Al) buffer range and a large amount of Al3+ releases from clay minerals (Ulrich 1986). Therefore, the negative impacts of soil acidification in NPK and N treatments were more pronounced because their aboveground plants and belowground microorganisms may have suffered from Al toxicity. In contrast, on the basis of the same N application rate, using organic N as a substitute for 70% of chemical N fertilizer could reverse Ultisols acidification in current cropping system. The alkalinity of organic manure, ammonification of organic N, and decarboxylation of organic anions were three primary causes for the decreases in soil acidity (Yuan et al. 2011; Rukshana et al. 2014). Furthermore, application of organic manure could also alleviate Al toxicity by forming Al-organic matter complexes (Gerke 2010).
Combination of chemical fertilizers and organic manure is an effective measure to improve soil fertility (Sun et al. 2015). In this study, chemical fertilizer balanced application (NPK) also improved soil nutrient contents, but the effects were much weaker than with NPKM. SOC in NPK was also significantly increased; the reason was that balanced nutrition improvement increased crop biomass, which in turn increased the amounts of litter, root residue, and root exudates return back to soil (Xiang et al. 2017). However, long-term N application alone only increased soil TN content, which was not conducive to improve soil fertility. The differences between C and N inputs and outputs caused by long-term different fertilization regimes also significantly changed the soil C/N.
4.2 Effects of long-term fertilization on PLFA
This study showed that NPKM and NPK significantly increased the biomass of total PLFA and each microbial group (i.e., bacteria and fungi), possibly because the significant increase in SOC (Tables 1 and 3) provided a large number of substrates for different microbial growth. Many studies based on high-throughput sequencing and quantitative PCR techniques have also obtained similar conclusion; that is, NPK balanced application or manure incorporation could increase the abundance of soil bacteria and fungi (Sun et al. 2015; Hu et al. 2017). In this study, the microbial relative abundance in different treatments also showed large changes. NPKM significantly improved the relative abundance of G− bacteria and was marked by an increase in fatty acid 16:1 ω7c (Table S1, Electronic Supplementary Material). Waldrop and Firestone (2004) found that G− bacteria are the most sensitive microbial group to labile organic substrates, and 16:1 ω7c has typically been identified as a prominent biomarker for the response of soil microorganisms to manure amendment. The significant decrease in the G+/G− ratio in NPKM indicated that soil condition changed from oligotrophic to copiotrophic following long-term fertilization (Fanin et al. 2014). However, the G+/G− ratio in NPK significantly increased, despite the fact that application of chemical fertilizers also improved soil nutritional status. The reason was that long-term chemical fertilizers application exacerbated soil acidification, and thus, the bacterial evolution tended to be dominated by G+ bacteria, which were more protected because of their thicker cell walls (Hammesfahr et al. 2008). As for N treatment, the soil pH was the lowest and the nutritional condition was very poor, neither suitable for G+ bacteria nor G− bacteria, so the G+/G− ratio showed no significant change.
Fungal relative abundance in the NPKM treatment increased significantly relative to CK, which supports the previous conclusion that manure application promoted soil fungal growth, and the fungal abundance was proportional to the application rate of manure (Ma et al. 2016). However, manure incorporation increased fungal proportion at the expense of actinomycetes, which is shown by an obvious decrease in the proportion of fatty acid 17:1 ω7c 10-methyl in NPKM (Table S1, Electronic Supplementary Material). The reason was that improvement of soil nutritional condition caused by manure amendment could exert survival pressure on actinomycetes (Ma et al. 2016). The increase in the F/B ratio was beneficial for stabilizing and protecting soil organic carbon and promoting the formation of soil aggregates (Six et al. 2006). Thus, the highest F/B value in NPKM indicated that manure incorporation was an effective measure to regulate microbial community structure and improve soil quality. Because the range of optimum pH for bacterial growth is narrow, while that for fungi is wider (Rousk et al. 2010), the significant decrease in soil pH also increased fungal abundance and F/B ratio in NPK and N treatments.
The higher sat/mono ratio indicated the stronger environmental stress and vice versa (Fierer et al. 2003). Application of chemical fertilizers, especially in the NPK treatment, obviously alleviated soil nutrient insufficiency or unavailability, but seriously exacerbated soil acidification, resulting in a significant increase in the sat/mono ratio (Table 2). Therefore, we infer that the negative impact of soil acidification on microbial community was much greater than the positive impact of nutrition improvement. The stepwise regression also identified soil pH as the primary factor in determining the sat/mono ratio, followed by nutrient conditions (i.e., SOC and AP) (Table 3). On the contrary, NPKM not only increased soil nutrients but also reversed soil acidification; thus, the ratio of sat/mono showed a slight decrease (Table 2).
RDA analysis further demonstrated that the composition of microbial community was mainly affected by soil pH (Figs. 2a and 3a). This result was consistent with some fertilization studies (Ma et al. 2015; Sun et al. 2015). However, many previous studies suggested that the availability of soil nutrients drove the shifts in microbial community composition (Zhong et al. 2010). The reason for this inconsistency depended on the degree of soil pH change. Xun et al. (2015) proposed a “rugby model” to explain this phenomenon. When soil pH showed a little change, soil nutrient conditions were the main factors affecting the microbial community composition; on the contrary, when soil acidification or alkalization was exacerbated, the effects of soil pH gradually increased, while the effects of other factors were strictly limited. In this study, CK and NPKM had similar soil pH, which might induce the clustering of microbial community composition between them (Fig. 2a). Besides pH, soil C/N ratio acted as a comprehensive indicator of soil nutrient status following long-term fertilization, and also had impacts on soil microbial community compositions (Zhang et al. 2015).
4.3 Effects of long-term fertilization on CLPP
Manure application increased the carbon metabolic activity of soil microorganisms (Ma et al. 2015). However, there were minimal differences in carbon metabolic preference and in functional diversity (S, H′, D, and E) between NPKM and CK (Fig. 5b and Table 2). This suggested that NPKM and CK had similar carbon metabolic patterns, which was further demonstrated by the RDA of CLPP (Fig. 2b). Sun et al. (2015) used high-throughput sequencing technique and found that long-term manure incorporation did not change the bacterial genetic diversity. Many studies repeatedly verified that bacterial genetic diversity was mainly affected by soil pH, but less affected by soil nutrient status (Hartman et al. 2008; Lauber et al. 2009; Shen et al. 2013). Our results demonstrated that soil pH is also the primary factor affecting the functional diversity of soil microorganisms (Table 3). Compared to long-term application of chemical fertilizer, manure application is an effective measure to maintain soil pH, so NPKM had high level of microbial functional diversity similar to CK.
Many studies reported that the shifts in microbial community composition will trigger the changes in microbial metabolic function (Zhong et al. 2010; Fanin et al. 2014). However, due to the inherent diversity, potential adaptability, and functional redundancy of microorganisms (Birgander et al. 2013; Yue et al. 2016), the response of microbial community composition and metabolic function to external disturbance may be asymmetric. In this study, long-term application of NPK increased the total PLFA biomass by 2.2 times, but its AWCD value was almost the same as CK (Fig. 4). The result indicated that the carbon metabolic efficiency of soil microorganisms decreased in NPK. In general, microorganisms enter a low active metabolic state when they encounter unfavorable environmental conditions, such as high temperature, drought, or hypoxia; if the environmental conditions continue to deteriorate, they will take a bet-hedging strategy, such as metabolic dormancy, while microbial biomass might be relatively less affected (Lennon and Jones 2011; Rittershaus et al. 2013). This phenomenon was particularly prominent in the N treatment. As Compared to CK, the total PLFA in the N treatment only decreased by 27.9%, but the carbon metabolic ability was almost completely inhibited (Fig. 4). This extreme situation has rarely been reported in previous fertilization studies. Stepwise regression analysis indicated that soil pH was the primary limiting factor for the AWCD (Table 3). We hypothesized that this might be related to soil acidification into the Al buffer range, and speculated that the inhibitory effect of Al toxicity on microbial metabolic activity was stronger than that on microbial biomass (Dorea and Clarke 2008). In addition, soil acidification also led to the significant decreases in microbial functional diversity of NPK and N treatments (Tables 2 and 3), which was detrimental to crop health and ecosystem stability (Sun et al. 2015). Treseder (2008) reported that excessive amounts of N react with soil C to produce polymers. In this study, the relative abundance of polymers used in the N treatment was increased from 15.50 to 40.22%, which might be a response of soil microorganisms to environmental change.
According to the RDA analysis, soil C/N ratio became the deterministic factor affecting the metabolic patterns of microbial community (Figs. 2b and 3d). Some studies reported that soil microbial metabolism is regulated by the soil C/N ratio, because this ratio is a comprehensive reflection of the quality and quantity of soil C and N (Chen et al. 2015). When the variation in soil C/N is small, soil microorganisms could adjust their macromolecular compositions and alter their intrinsic biochemical stoichiometry to adapt to soil C/N changes; however, when the variation exceeds the microbial self-regulating ability, the microbial community composition will eventually be dominated by groups with different metabolic patterns (Sistla and Schimel 2012). Unlike PLFA profiles, soil pH became the second important driving factor affecting microbial metabolic pattern (Figs. 2b and 3c). Although soil microbial community composition could be dramatically reshaped by pH, due to the characteristics of functional redundancy, the different microbial communities could metabolize similar carbon source (Ma et al. 2015). In addition, the CLPP method has the disadvantage of being biased by incubation conditions and not incorporating the contributions of fungi and slow-growing bacteria, so the effects of pH on soil microbial metabolic patterns may not be completely reflected in current results.
5 Conclusions
Our results demonstrated that different long-term fertilizations triggered asymmetric effects on microbial community compositions and metabolic functions in acidic Ultisols of southern China. The negative effects of Ultisol acidification induced by long-term application of chemical N fertilizer on microorganisms overwhelmed the positive effects of soil nutrition improvement. The inhibiting effect of serious Ultisol acidification on microbial metabolic function was stronger than that on community composition. In this study, NPK increased the total PLFA biomass, while its AWCD was not significantly different from CK; N slightly decreased the total PLFA biomass, but almost completely inhibited metabolic activity. In summary, we speculated that in situations of serious Ultisol acidification, microorganisms live in a low active metabolic state to resist severe soil acidity. In contrast, manure incorporation was an effective measure to alleviate soil acidity, increase total PLFA biomass and metabolic activity, and maintain the stability of microbial community composition, metabolic pattern, and functional diversity. Therefore, fertilization in acidic Ultisols should be based on the premise of preventing soil further acidification. This work helps us to better understand how long-term fertilization affects microbial community composition and metabolic function in acidic Ultisol agro-ecosystems.
References
Abdulahaal BM, Li JY, Xu CY, Mehmood K, Xu RK (2017) Determination of critical pH and Al concentration of acidic Ultisols for wheat and canola crops. Solid Earth 8:149–159
Allison SD, Martiny JB (2008) Resistance, resilience, and redundancy in microbial communities. Proc Natl Acad Sci U S A 105(Supplement 1):11512–11519. https://doi.org/10.1073/pnas.0801925105
Berg G (2009) Plant-microbe interactions promoting plant growth and health: perspectives for controlled use of microorganisms in agriculture. Appl Microbiol Biot 84(1):11–18. https://doi.org/10.1007/s00253-009-2092-7
Birgander J, Reischke S, Jones DL, Rousk J (2013) Temperature adaptation of bacterial growth and 14C-glucose mineralisation in a laboratory study. Soil Biol Biochem 65:294–303. https://doi.org/10.1016/j.soilbio.2013.06.006
Blanchet FG, Legendre P, Borcard D (2008) Forward selection of explanatory variables. Ecology 89:2623–2632
Buyer JS, Teasdale JR, Roberts DP, Zasada IA, Maul JE (2010) Factors affecting soil microbial community structure in tomato cropping systems. Soil Biol Biochem 42(5):831–841. https://doi.org/10.1016/j.soilbio.2010.01.020
Cai ZJ, Wang BR, Xu MG, Zhang HM, He XH, Zhang L, Gao SD (2015) Intensified soil acidification from chemical N fertilization and prevention by manure in an 18-year field experiment in the red soil of southern China. J Soils Sediments 15(2):260–270. https://doi.org/10.1007/s11368-014-0989-y
Cai ZJ, Wang BR, Xu MG, Zhang HM, Zhang L, Gao SD (2014) Nitrification and acidification from urea application in red soil (Ferralic Cambisol) after different long-term fertilization treatments. J Soils Sediments 14(9):1526–1536. https://doi.org/10.1007/s11368-014-0906-4
Chen L, Xun WB, Sun L, Zhang N, Shen QR, Zhang RF (2014) Effect of different long-term fertilization regimes on the viral community in an agricultural soil of southern China. Eur Soil Bio 62:121–126. https://doi.org/10.1016/j.ejsobi.2014.03.006
Chen XF, Li ZP, Liu M, Jiang CY, Che YP (2015) Microbial community and functional diversity associated with different aggregate fractions of a paddy soil fertilized with organic manure and/or NPK fertilizer for 20 years. J Soils Sediments 15(2):292–301. https://doi.org/10.1007/s11368-014-0981-6
Dong WY, Zhang XY, Wang HM, Dai XQ, Sun XM, Qiu WW, Yang FT (2012) Effect of different fertilizer application on the soil fertility of paddy soils in red soil region of southern China. PLoS One 7(9):e44504. https://doi.org/10.1371/journal.pone.0044504
Dorea CC, Clarke BA (2008) Effect of aluminium on microbial respiration. Water Air Soil Pollut 189(1-4):353–358. https://doi.org/10.1007/s11270-007-9553-3
Fanin N, Hättenschwiler S, Fromin N (2014) Litter fingerprint on microbial biomass, activity, and community structure in the underlying soil. Plant Soil 379:79–91
Fierer N, Schimel JP, Holden PA (2003) Variations in microbial community composition through two soil depth profiles. Soil Biol Biochem 35(1):167–176. https://doi.org/10.1016/S0038-0717(02)00251-1
Garland JL (1996) Analytical approaches to the characterization of samples of microbial communities using patterns of potential C source utilization. Soil Biol Biochem 28(2):213–221. https://doi.org/10.1016/0038-0717(95)00112-3
Gerke J (2010) Humic (organic matter)-Al (Fe)-phosphate complexes: an underestimated phosphate form in soils and source of plant-available phosphate. Soil Sci 175(9):417–425. https://doi.org/10.1097/SS.0b013e3181f1b4dd
Guo JH, Liu XJ, Zhang Y, Shen JL, Han WX, Zhang WF, Christie P, Goulding KW, Vitousek PM, Zhang FS (2010) Significant acidification in major Chinese croplands. Science 327(5968):1008–1010. https://doi.org/10.1126/science.1182570
Hammesfahr U, Heuer H, Manzke B, Smalla K, Thielebruhn S (2008) Impact of the antibiotic sulfadiazine and pig manure on the microbial community structure in agricultural soils. Soil Biol Biochem 40:1583–1591
Hartman WH, Richardson CJ, Vilgalys R, Bruland GL (2008) Environmental and anthropogenic controls over bacterial communities in wetland soils. Proc Natl Acad Sci U S A 105(46):17842–17847. https://doi.org/10.1073/pnas.0808254105
Hu XJ, Liu JJ, Wei D, Zhu P, Cui XA, Zhou BK, Chen XL, Jin J, Liu XB, Wang GH (2017) Effects of over 30-year of different fertilization regimes on fungal community compositions in the black soils of northeast China. Agric Ecosyst Environ 248:113–122. https://doi.org/10.1016/j.agee.2017.07.031
Huang R, Wu YC, Zhang JB, Zhong WH, Jia ZJ, Cai ZC (2012) Nitrification activity and putative ammonia-oxidizing archaea in acidic red soils. J Soils Sediments 12(3):420–428. https://doi.org/10.1007/s11368-011-0450-4
Huang S, Zhang W, Yu X, Huang QR (2010) Effects of long-term fertilization on corn productivity and its sustainability in an Ultisol of southern China. Agric Ecosyst Environ 138(1-2):44–50. https://doi.org/10.1016/j.agee.2010.03.015
Kušlienė G, Rasmussen J, Kuzyakov Y, Eriksen J (2014) Medium-term response of microbial community to rhizodeposits of white clover and ryegrass and tracing of active processes induced by 13C and 15N labelled exudates. Soil Biol Biochem 76:22–33. https://doi.org/10.1016/j.soilbio.2014.05.003
Lauber CL, Hamady M, Knight R, Fierer NP (2009) Pyrosequencing-based assessment of soil pH as a predictor of soil bacterial community structure at the continental scale. Appl Environ Microb 75(15):5111–5120. https://doi.org/10.1128/AEM.00335-09
Legendre P, Gallagher E (2001) Ecologically meaningful transformations for ordination of species data. Oecologia 129(2):271–280. https://doi.org/10.1007/s004420100716
Lennon JT, Jones SE (2011) Microbial seed banks: the ecological and evolutionary implications of dormancy. Nat Rev Microbiol 9(2):119–130. https://doi.org/10.1038/nrmicro2504
Li WT, Liu M, Jiang CY, Wu M, Chen XF, Ma XY, Li ZP (2017) Changes in soil aggregate- associated enzyme activities and nutrients under long-term chemical fertilizer applications in a phosphorus-limited paddy soil. Soil Use Manage 33(1):25–33. https://doi.org/10.1111/sum.12322
Ma XY, Liu M, Li ZP (2015) Changes in microbial properties and community composition in acid soils receiving wastewater from concentrated animal farming operations. Appl Soil Ecol 90:11–17
Ma XY, Liu M, Li ZP (2016) Shifts in microbial biomass and community composition in subtropical paddy soils under a gradient of manure amendment. Biol Fert. Soils 52:775–787
National Bureau of Statistics of China (2016) China statistical yearbook. China Statistics Press, Beijing
Pansu M, Gautheyrou J (2006) Handbook of soil analysis: mineralogical, organic and inorganic methods. Springer, Berlin Heidelberg, Berlin. https://doi.org/10.1007/978-3-540-31211-6
Peng CJ, Lai SS, Luo XS, Lu JW, Huang QY, Chen WL (2016) Effects of long term rice straw application on the microbial communities of rapeseed rhizosphere in a paddy-upland rotation system. Sci Total Environ 557-558:231–239. https://doi.org/10.1016/j.scitotenv.2016.02.184
Rittershaus ESC, Baek SH, Sassetti CM (2013) The normalcy of dormancy: common themes in microbial quiescence. Cell Host Microbe 13(6):643–651. https://doi.org/10.1016/j.chom.2013.05.012
Rousk J, Bååth E, Brookes PC, Lauber CL, Lozupone C, Caporaso JG, Knight R, Fierer N (2010) Soil bacterial and fungal communities across a pH gradient in an arable soil. ISME J 4(10):1340–1351. https://doi.org/10.1038/ismej.2010.58
Rukshana F, Butterly CR, Xu JM, Baldock JA, Tang CX (2014) Organic anion-to-acid ratio influences pH change of soils differing in initial pH. J Soils Sediments 14(2):407–414. https://doi.org/10.1007/s11368-013-0682-6
Shen CC, Xiong JB, Zhang HY, Feng YZ, Lin XG, Li XY, Liang WJ, Chu HY (2013) Soil pH drives the spatial distribution of bacterial communities along elevation on Changbai Mountain. Soil Biol Biochem 57:204–211. https://doi.org/10.1016/j.soilbio.2012.07.013
Sistla SA, Schimel JP (2012) Stoichiometric flexibility as a regulator of carbon and nutrient cycling in terrestrial ecosystems under change. New Phytol 196(1):68–78. https://doi.org/10.1111/j.1469-8137.2012.04234.x
Six J, Frey SD, Thiet RK, Batten KM (2006) Bacterial and fungal contributions to carbon sequestration in agroecosystems. Soil Sci Soc Am J 70(2):555–569. https://doi.org/10.2136/sssaj2004.0347
Sun RB, Zhang XX, Guo XS, Wang DZ, Chu HY (2015) Bacterial diversity in soils subjected to long-term chemical fertilization can be more stably maintained with the addition of livestock manure than wheat straw. Soil Biol Biochem 88:9–18. https://doi.org/10.1016/j.soilbio.2015.05.007
Treseder KK (2008) Nitrogen additions and microbial biomass: a meta-analysis of ecosystem studies. Ecol Lett 11(10):1111–1120. https://doi.org/10.1111/j.1461-0248.2008.01230.x
Ulrich B (1986) Natural and anthropogenic components of soil acidification. J Plant Nutr Soil Sci 149:702–717
Waldrop M, Firestone M (2004) Microbial community utilization of recalcitrant and simple carbon compounds: impact of oak-woodland plant communities. Oecologia 138(2):275–284. https://doi.org/10.1007/s00442-003-1419-9
Xiang XJ, He D, He JS, Myrold DD, Chu HY (2017) Ammonia-oxidizing bacteria rather than archaea respond to short-term urea amendment in an alpine grassland. Soil Biol Biochem 107:218–225. https://doi.org/10.1016/j.soilbio.2017.01.012
Xun WB, Huang T, Zhao J, Ran W, Wang BR, Shen QR, Zhang RF (2015) Environmental conditions rather than microbial inoculum composition determine the bacterial composition, microbial biomass and enzymatic activity of reconstructed soil microbial communities. Soil Biol Biochem 90:10–18. https://doi.org/10.1016/j.soilbio.2015.07.018
Yuan JH, Xu RK, Qian W, Wang RH (2011) Comparison of the ameliorating effects on an acidic ultisol between four crop straws and their biochars. J Soils Sediments 11(5):741–750. https://doi.org/10.1007/s11368-011-0365-0
Yue XL, Zhang JG, Shi AD, Yao SH, Zhang B (2016) Manure substitution of mineral fertilizers increased functional stability through changing structure and physiology of microbial communities. Eur J Soil Biol 77:34–43. https://doi.org/10.1016/j.ejsobi.2016.10.002
Zelles L (1997) Phospholipid fatty acid profiles in selected members of soil microbial communities. Chemosphere 35(1-2):275–294. https://doi.org/10.1016/S0045-6535(97)00155-0
Zhang Q, Zhou W, Liang GQ, Sun JW, Wang XB, He P (2015) Distribution of soil nutrients, extracellular enzyme activities and microbial communities across particle-size fractions in a long-term fertilizer experiment. Appl Soil Ecol 94:59–71. https://doi.org/10.1016/j.apsoil.2015.05.005
Zhang XB, Wu LH, Sun N, Ding XS, Li JW, Wang BR, Li DC (2014) Soil CO2 and N2O emissions in maize growing season under different fertilizer regimes in an upland red soil region of South China. J Integr Agr 13(3):604–614. https://doi.org/10.1016/S2095-3119(13)60718-2
Zhong WH, Gu T, Wei W, Zhang B, Lin XG, Huang QR, Shen WS (2010) The effects of mineral fertilizer and organic manure on soil microbial community and diversity. Plant Soil 326(1-2):511–522. https://doi.org/10.1007/s11104-009-9988-y
Acknowledgements
This study is supported by the National Basic Research Program (973 Program) of China (No. 2014CB441003) and the National Natural Science Foundation of China (Nos. 41661052 and 31660599). In addition, we thank the anonymous reviewers and editors for their helpful comments regarding the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible editor: Huaiying Yao
Electronic supplementary material
ESM 1
(DOC 62 kb)
Rights and permissions
About this article
Cite this article
Liu, J., Liu, M., Wu, M. et al. Soil pH rather than nutrients drive changes in microbial community following long-term fertilization in acidic Ultisols of southern China. J Soils Sediments 18, 1853–1864 (2018). https://doi.org/10.1007/s11368-018-1934-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11368-018-1934-2