Abstract
Purpose
The synthetic soil based bioremediation approach as reasonable and sustainable practice at the farming level where desired bioremediation could be established at lower cost.
Materials and methods
Metal-tolerant bacteria from different environmental field samples, (a) a municipal dump site, (b) an agricultural field and (c) sludge of electro-plating industries, were screened and characterized. Bioremediation of metal contaminants through isolated bacteria was compared under two different conditions, synthetic soil and basic minimal media containing copper, cobalt and nickel.
Results and discussion
The pollutants arising from industrial effluents are imparting a huge negative impact on agricultural land. Microbes are predominant in heavy metal-contaminated sites, which signifies as a potential opportunity for the researchers towards bioremediation. Three bacterial species showed high metal tolerance; 16S ribosomal DNA (rDNA) analysis revealed that the organisms were Proteus vulgaris strain, Stenotrophomonas sp. and Bacillus thuringiensis. Percentage removal of metals was also analysed under different concentrations and pH.
Conclusions
The current tested methods are helpful in streamlining the natural compliance of fragile elements and its uptake into the microbial system under in vitro and in situ conditions.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Over the past few decades, it has been observed that the pollutants arising from industrial effluents are imparting a huge negative impact on agricultural land and the water table underneath the existing soil profile. Spread of such elements and its consequent spatial variation in local contaminated sites causes variable loss at C and N level in the top 20 cm soil (Xu et al. 2008; Fromin et al. 2012). Common sources of metal pollutants include effluent discharge from electroplating, plastic manufacturing, fertilizer-producing industries as well as large-scale plants concerned with mining and metallurgical processes (Mudhoo et al. 2012). Whilst some of these metals are highly toxic with no physiological or cellular role (Appenroth 2010), others are essential for life at low concentrations. Copper, cobalt and nickel are heavy elements, with a long history of toxicity. Cobalt and copper are very close to nickel in the periodic table, and their close concentrations in soil persist for a longer time which allows them to exhibit different chemical forms (speciation) and for further bioavailability (McLaughlin et al. 2000). The major sources of these elements are usually weathered rock, mines, smelting operations, chemical industries (effluents), etc. Heavy metal exposure to humans causes chronic diseases and severe damage to the kidneys, nervous system, liver and brain (Tchounwou et al. 2012). Their exposure is becoming a great concern because of its toxic nature, wide spread occurrence and fate in biological system.
The exploitation of the metal-resistant microbial system in the bioremediation of metals from agricultural or cultivation lands holds a great biotechnological significance in health and environment sectors. Ozaki et al. (2003) claimed that heavy metals are difficult to remove from the environment and are ultimately indestructible, unlike many other pollutants (organic) that can be chemically or biologically degraded. Effective sequestration and immobilization of metals and minerals have been carried effectively through microbial interactions. Specific ion-exchange process influenced by microbes was noticed as the contact building approach towards their bioremediation (Gadd 2010). Microbial growth has been estimated by the redox potential (Eh) measurement in the specific fermentation-based growth curve (Maegan and Ilenys 2009). In addition, many minerals are biogenic in origin, and the formation of such biominerals is of global geological and industrial importance. Bioavailability of metals in different forms of compound has been understood as the metabolic precursors in plants (Hossain et al. 2012). Metal homeostasis in Arabidopsis model crop has been investigated in recent past; the redox nature of such precursors was found to be causative in functional loss of mitochondria due to oxidative stress, and similar consequences have been reported to a human context (Gadd 2010; Tan et al. 2010).
Researchers have indicated that microbes are predominant in heavy metal-contaminated sites, though its phenomenal act of survival is still a puzzle, which signifies a potential opportunity for the researchers towards bioremediation (Monachese et al. 2012; Girma 2015). Chatterjee et al. (2010) assessed the metal-binding capacity of Geobacillus thermodenitrificans in synthetic metal solutions and industrial waste water. They successfully determined biosorption of heavy metals like Fe, Cr, Co, Cu, Zn, Cd, Ag and Pb from industrial wastewater by G. thermodenitrificans. Similarly, Balaji et al. (2014) evaluated the biosorption effect of Zn and Ni under laboratory conditions which potentially influence the sustainability of Spirulina spp. in terms of doubling time, chlorophyll content and protein content.
Removal of metals from the soil and water or their remediation from the waste streams “at source” has been a long-term challenge (Ray and Ray 2009). The global environmental protection policy has raised the concerns on metallic pollutions in soil and supported the practice of microorganisms for remediation (Venosa 2004). Remediation of toxic metals from waste effluents as well as deployment of plants for landfill applications has taken the industrialized approach. Molecular demonstrations of microbial system have revealed the systematic ability to convert the metals onto acceptable forms of the plants existing in contaminated water effluent or sludge (Dixit et al. 2015).
With the modern advent, several bioremediation strategies have been proposed as an attractive alternative owing to the potential low cost, less hazardous nature and sustainability as compared to other remedial approaches (Cui et al. 2013). In this work, we isolated and characterized metal-resistant bacteria from the copper, cobalt and nickel containing environmental field samples which were confirmed through sample screening in order to determine microbial species capable of growing efficiently in metal-contaminated soils. The isolated microorganisms were further tested for its effectiveness to mitigate exposure to the metal contaminants in soil. The synthetic-contaminated soils were prepared for this study which could be a sustainable approach for laying bioremediation plant at lower cost. Current tested methods are helpful in streamlining the natural compliance of fragile elements and its uptake into the microbial system under in vitro and in situ conditions. In spite of its cost-effectiveness and environment friendliness, applications of this approach could be extrapolated to field conditions.
2 Materials and methods
2.1 Sampling and isolation of bacterial strains
Soil samples were collected in triplicate from three different sites: a municipal dump site, Dhapa, Kolkata, West Bengal (22° 53′ and 88° 41′); an agricultural field adjoining a rifle factory, Ichapore, Kolkata, West Bengal, eastern India (22° 47′ and 88° 5′) and sludge from a nearby electro-plating plant at Jalandhar, Punjab, northern India (30° 4′ and 75° 5′). Samples were sealed aseptically and brought to the laboratory in sterilized zip lock plastic bags, stored at 4 °C in the refrigerator during the study. The soil samples were properly homogenized, and sieving was done to remove particulate material and other impurities. The air-dried samples were analysed physiochemically with soil-testing parameters (Taylor 1987), and total concentration of heavy metals was determined according to ISO11466:1995 (Soil Testing Department, Punjab Agricultural University, Ludhiana, India). A total of 35, 47 and 29 bacterial isolates from municipal dump site, agricultural field and sludge waste-contaminated sites, respectively, were cultured in minimal medium conditions. Further, all the isolates from soil samples were cultured on copper (Cu2+)-, cobalt (Co2+)- and nickel (Ni2+)-enriched minimal nutrient media individually with six different concentrations (10, 50, 100, 150, 200 and 250 mg/L) at 37 °C for 24 and 48 h. Different concentrations of Cu, Co and Ni solutions were prepared from standard stock solutions of copper sulphate (CuSO4 · 7H2O), cobalt chloride (CoCl2 · 6H2O) and nickel sulphate (NiSO4 · 6H2O) (analytical grade, Sigma), respectively, and sterilized by autoclaving. The metal solutions were added to the medium after autoclaving at 45 to 50 °C. From the independently growing colonies, eight were randomly selected and those three bacteria with the highest resistance against targeted metals were screened and isolated.
2.2 Identification of the metal-resistant isolates by 16S rDNA sequencing
Genomic DNA was isolated and purified from the selected bacterial plates (identified biochemically and morphologically). Phenol chloroform extraction method was followed for genomic DNA isolation (Chomczynski and Sacchi 1987). Fragment of 16S ribosomal DNA (rDNA) gene was amplified by PCR from the above isolated DNA. Forward and reverse DNA sequencing reaction of PCR amplicon was carried out with 27F and 1492R primers (Mao et al. 2012).
2.3 Sequence analysis
Sequencing of 16S rDNA amplicon was performed using the Applied Biosystems 3130 × l Genetic Analyzer. The 16S rDNA sequenced products were submitted to GenBank databases for accession numbers.
2.4 Preparation of synthetic soil media
Stock solutions of Cu, Co and Ni salts having concentration of 10 mg/L were prepared in distilled water. All the solutions were sterilized by membrane filters of pore size 0.22 μm and stored at 4 °C until use. Sterilized field soil weighed 100 g (wet weight) for each flask. The metals from stock solution were incorporated into soil to achieve the desired metal concentration along with minimal nutrient media. The concentration was equivalent to both liquid and semisolid media (10, 20, 50, 100 and 200 mg/L for Ni and Cu and 10, 20, 50 and 100 mg/L for Co) and then set to pH 7.3. For storage purpose, soil samples were dried at 140 °C for 3 h using soil-specific hot air oven.
2.5 Antibiotic sensitivity
Commercially prepared discs (antibiotic discs) of ampicillin (10 mcg), tetracycline (30 mcg), chloramphenicol (30 mcg), kanamycin (30 mcg) and erythromycin (15 mcg) (Himedia, India) were kept in Muller-Hinton agar medium swabbed with a faintly opalescent bacterial culture and were incubated at 37 °C for 24 h media and also maintained the negative control for the comparison (Bharagava et al. 2014). After 24 h, the inhibition zones were measured and isolates were classified as resistant or susceptible on the basis of zone size. Bacterial strains were susceptible when inhibition zone was 3 mm or more in diameter (Gales et al. 2001), and, henceforth, susceptibility test was carried out in replicates.
2.6 Growth of isolates on different concentrations of metals
Each isolate grown in log phase on metal-supplemented nutrient agar media was inoculated into the test tubes containing 10 mL of nutrient broth with increasing concentrations of respective metals, ranging from 10 to 200 mg/L (Karbasizaed et al. 2004). After 48 h, the growth was observed and analysed spectrophotometrically by measuring the optical density measurements at 600 nm (Shimadzu UV-1800 UV/Vis Spectrophotometer). Nutrient broth (100 mL) was poured into the flasks containing synthetic soil media prepared earlier having the concentrations and pH value described in Section 2.4 (Buers et al. 1997). At subsequent time intervals (36, 48 and 60 h), soil sample was collected and stored at 4 °C. The above samples were analysed for the presence of metals at respective concentrations by Inductively Coupled Plasma-Atomic Emission Spectroscopy (ICP-AES) Model Labtum-8440 Australia Model Labtum-8440 Australia by the technique of Jones (1977).
2.7 Estimation of percentage removal of metals
Percentage removal, defined as a percentage reduction of metal ion present in solution, was analysed using the equation:
M0 = metal initial concentration (mg/L); M1 = metal final concentration (mg/L).
The input percentage removals were provided by the Soil Testing Department, Punjab Agricultural University, Ludhiana, India.
2.8 Statistical analysis
Two-way ANOVA with repeated measures was performed using statistical software Graph Pad Prism Version 5.0 to observe the significance in the variation of treatment against control. Triplicate samples were used amongst treatments, and statistical differences were determined (P < 0.05). In this respect also, guidance has been followed to non-linear calibration lines (EPA method 6010B).
3 Results
3.1 Physic-chemical characterization of soil samples
Environmental field samples were collected from three different industrially polluted cities of India (North and East): (i) waste disposal region of an agricultural field adjacent to rifle factory, West Bengal, (ii) municipal dumping yard, West Bengal and (iii) plating industry, Punjab. Physic-chemical analysis of the soil samples from the targeted regions showed statistical differences (P < 0.05) in data based on variable parameters including pH, percentage moisture content, percentage organic carbon, available N, P, K, and trace elements Zn, Mn and Fe (mg/kg) (Table 1). Samples collected from these sites showed high alkalinity. The maximum alkalinity was observed in the contaminated soil collected from the waste disposal region of an agricultural field just nearby to rifle factory, West Bengal (pH 8.9) followed by the sample from the plating industry, Punjab (pH 8.5) and the samples from the municipal dumping yard, West Bengal (pH 8.0).
3.2 Metal analysis
This study showed Cu, Co and Ni bioremediation in two comparative environments, i.e. nutrient broth (in vitro) and synthetic soil (in situ) under manifested contact time, metal concentration and pH conditions. ICP-AES was used for the elemental analysis as shown in Table 2. Concentrations of Co in soils from the plating industry place were remarkably higher (P < 0.05) values than found in municipal dumping site and Rifle factory, whilst Ni and As concentrations were much higher in municipal dumping sites compared to the other sites (Table 2). Similarly, other existing elements (Pb, Cd, Cr and As) in the tested samples from the sites also exhibited higher concentrations compared to the threshold values as suggested by the Food and Fertilizer Technology Center, Taiwan. The analysis on the metal concentration reveals the fact that Co, Ni and Cu were predominantly higher when compared to the other heavy metals (Table 2).
3.3 Identification of resistant bacteria
During this study, 35, 47 and 29 bacterial isolates were municipal dump site, agricultural field and sludge waste-contaminated sites, respectively. The resistant bacteria strain Ni200 (Proteus vulgaris), Co100 (Stenotrophomonas sp.) and Cu200 (Bacillus thuringiensis) were predominantly identified in Ni-, Co- and Cu-spiked liquid media and artificial soil, respectively, at different concentrations and pH. NCBI accession number has been obtained for each strain, and their molecular phylogenetic analysis was carried out using 16S rDNA gene sequence analysis as mentioned in Table 3.
3.4 Antibiotic sensitivity of metal-resistant strains
Bacterial strains identified as P. vulgaris, Stenotrophomonas sp. and B. thuringiensis for bioremediation of Ni2+, Cu2+ and Co2+ were subjected to the antibiotic sensitivity test. Antibiogram has been raised for its resistance and susceptibility towards the specific antibiotics (Table 4). The zone more than 3 mm and less than 3 mm (mean) was considered to be susceptible and resistant, respectively. Ni200 strain (mutant) was found to be susceptible to all the antibiotics used in the experiment except ampicillin. Testing control strain (untreated) and Ni200 mutant were found to be similar in response to each antibiotic treatment.
Cu200 strain and control were both extremely susceptible to all antibiotics. Co100 strain shows resistance against ampicillin whereas control strain showed susceptibility. Above analysis indicates the presence and absence of metal resistance gene in the corresponding plasmids of strains.
3.5 Comparative study of Ni, Cu and Co removal
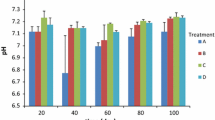
Increasing the contact between microbes and heavy metals in either liquid or soil in media resulted in increased rate of heavy metal removal. For instance, at least 20 % of 10 mg/L of Ni was removed after 12-h contact time but as the contact time increased to 48 h, a significantly higher (>80 %) amount of 10 mg/L Ni was removed (P < 0.05). However, as the concentration of Ni was increased, lower amount of Ni was removed (%) by the bacterial inoculum. At 150 and 200 mg/L Ni in liquid media, less than 20 % was removed in 12 h but as the time was increased to 48 h, 60 and 50 % of Ni were removed, respectively (Fig. 1a). Similarly in the soil medium, 40 % of 50 mg/L Ni was removed after 36-h contact time, but as contact time increased to 48 h, 55 % was totally removed by the P. vulgaris strain Ni200 (Fig. 1b). A specific range of pH (8 and 9) screened in both conditions was found to be effective for Ni removal (≥75 %) in liquid media and (≥50 %) in artificial soil containing 100 mg/kg, of Ni, respectively, as described in Fig. 2a, b.
In the case of Cu, observations were quite different, at lower concentration in liquid media; higher amount of Cu removal (%) (P < 0.05) by the Stenotrophomonas sp. Cu200 was observed after 24 h; and at 100 mg/kg concentration in soil media, higher amount of Cu removal (%) (P < 0.05) was noticed. For instance, at least 50 % of 10 mg/L of Cu was removed after 36-h contact time, which further increased to 55 % at 48 and 60 h, respectively (P < 0.05). However, at initial condition, higher concentration of Cu (200 mg/L) was shown to be removed (%) effectively at 12-h contact time by the bacterial inoculums (Fig. 3a). In the soil medium, for lower concentrations of Cu metal (50 and 100 mg/kg), higher removal (%) (P < 0.05) was observed by the bacterial inoculum. Thirty and 34 % removal after 48-h contact time, further as contact time increased to 60 h, quite less i.e. 29 and 32 % was totally removed by the Stenotrophomonas sp. Cu200 (Fig. 3b). There was a persistent removal rate of Cu (80 %) in liquid media and (23 %) in artificial soil containing 100 mg/kg, of Cu, respectively, under pH factor 8 to 9 at 48 and 60 h, respectively (Fig. 4a, b).
For Co, the same pattern as obtained for Ni was observed. For lower concentrations of metals in either liquid or soil in media, higher amount of Co removal (%) (P < 0.05) was observed by the B. thuringiensis. For instance, at least 12 % of 10 mg/L of Co was removed after 12-h contact time but as the contact time increased to 48 h, a significantly higher (>50 %) amount of 10 mg/L Co was removed (P < 0.05). However, as the concentration of Co was increased, lower amount of Co was removed (%) by the bacterial inoculum. At 50 and 100 mg/L Co in liquid media, less than 12 % was removed in 12 h but as the time was increased to 48 h, 40 and 22 % of Co were removed, respectively (Fig. 5a). Similarly in the soil medium, 22 % of 50 mg/L Co was removed after 36-h contact time, but as contact time increased to 48 h, 28 % was totally removed by the bacterial inoculum (Fig. 5b). There was a persistent removal rate of Co (40 %) in liquid media and (30 %) in artificial soil containing 100 mg/kg, of Co, respectively, under pH factor 8 to 9 at 48 h, respectively (Fig. 6a, b).
4 Discussion
Soil effluents were collected from three different cities of North and Northeast India virtually known as industrial, and agriculture belt coincides with the highest alkalinity range (pH ≥8.0) as reported in this study. The alkaline nature of effluents indicates the existence of metals and its derivatives in the dump yard, changes the soil parameters to an adverse level (Singh et al. 1997; Noorjahan 2014; Pantano et al. 2014). Variations in contaminated soil at pH level have already been reported by several researchers in the recent past (Abskharon et al. 2009; Kermani et al. 2009). Changes in the chemical nature of soil due to the contamination are the sign of microbial response, which brings change in capacity of neutralization, pH, redox potential and electric conductivity (Kermani et al. 2009; Choi et al. 2012)Abrupt pH variation creates inductive atmosphere, accelerating the resistance in the pre-existing microbial population. In this study, metal bioremediation was assessed under two comparative conditions, i.e. nutrient broth (in vitro) and synthetic soil (in situ) under different metal concentrations and pH. The observations of both conditions were found to be similar, which resulted in remediation of these heavy metals. Several researchers have reported remediation of individual metal (Cu/Ni/Fe) with microbial treatment under in vitro condition alone, whereas in situ analysis was poorly understood as an effective model of heavy metal removal (Abskharon et al. 2009; Perpetuo et al. 2011; Mishra and Malik 2013).
In this study, all the selected metals were present at a concentration much greater than the threshold acceptable values. The reference values of Co, Cu and Ni agriculture soils are 50, 100 and 75 mg/kg, respectively (Kabata-Pendias and Pendias 2001). In order to determine the fate of contaminants at degradation level, researchers employed the percentage removal formula which calculates the absorption efficiency of microbes. It is presumed that the rates of ionic conversions are least affected by conditions imposed here in the study as the depth of the chemical in synthetic soil, adsorption, humidity and pH which had the common interest.
Microorganisms play a role of the active/passive carrier of contaminants as resulted in bioaccumulation into the plant tissues, further found to be phyto-remediated under symbiotic condition (Srinath et al. 2002; Sriprang and Murooka 2007; Umysova et al. 2009). Several studies related to environmental factors for metal uptake in the plants have described the fate of plant-microbe interaction (Weber et al. 2006; Gonzalez-Guerrero et al. 2007; Hedrich et al. 2011; Mishra and Malik 2013).
Our study suggests that the microbial community in soils compromises culturable organisms which can be very important in bioremediation. Nieto et al. (1989) and Deshpande et al. (1993) isolated several Acinetobacter sp. strains commonly occurring at metal-contaminated sites and confirmed Acinetobacter species to be highly resistant to metals. Synthetic soil and in vitro medium were evaluated as complementary models and possibly explained the fate of metals under the defined parameters. Researchers have proven that bacterial cells have the inherent potential to scavenge the sludge effluents once they get exposed to; as a result, a considerable intake machinery is established for the ionic conversion of metals under a particular environment (Roane and Kellogg 1996; Wireman et al. 1997; Eze et al. 2009). Though the current study does not address any changes in cellular or biochemical pathway level, moreover, it is assumed that resistance gain is possible only because of the changes at genetic constitution level. McArthur and Tuckfield (2000) have addressed in a broad manner that the metal and antibiotic resistances in bacteria are correlative action at genome or plasmid level. Kumar et al. (2013) have already reported and characterized the resistance strains of Bacillus and Stenotrophomonas spp. from chromium, cadmium and arsenic-affected zone from North India.
5 Conclusions
In the present work, the synthetic-contaminated soils were prepared for the study could be a sustainable approach for efficient bioremediation of copper, cobalt and nickel contaminants, respectively. Current tested methods are helpful in streamlining the natural compliance of fragile elements and its uptake into the microbial system under in vitro and in situ conditions. In spite of its cost-effectiveness and environment friendliness, applications of this approach could be translated in field conditions.
References
Abskharon RNN, Gad El-Rab SMF, Hassan SHA, Shoreit AAM (2009) Reduction of toxic hexavalent chromium by E. coli. J Biotechnol Biochem 4:98–103
Appenroth KJ (2010) Definition of “heavy metals” and their role in biological systems. In: Sherameti I, Varma A (eds) Soil heavy metals, soil biology, vol 19. Springer, Berlin, pp 19–29
Balaji S, Kalaivani T, Rajasekaran C (2014) Biosorption of zinc and nickel and its effect on growth of different Spirulina strains. Clean Soil Air Water 42:507–512
Bharagava RN, Yadav S, Chandra R (2014) Antibiotic and heavy metal resistance properties of bacteria isolated from the aeration lagoons of common effluent treatment plant (CETP) of tannery industries (Unnao, India). Ind J Biotechnol 13:514–519
Buers KLM, Prince EL, Knowles CJ (1997) The ability of selected bacterial isolates to utilise components of synthetic metal-working fluids as sole sources of carbon and nitrogen for growth. Biotechnol Lett 19:791–794
Chatterjee SK, Bhattacharya I, Chandra G (2010) Biosorption of heavy metals from industrial wastewater by Geobacillus thermodenitrificans. J Hazard Mater 175:117–125
Choi J, Yang JS, Tae Park Y, Kim JO, Kim KJ, Shim YS, Kwon HH, Khan HA, Park JW, Um JG, Jeon BH (2012) Comparison of As, Ni, Zn, Cd, and Pb removals using treatment agents. Environ Technol 33:445–454
Chomczynski P, Sacchi N (1987) Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal Biochem 162:156–159
Cui H, Zhou J, Zhao Q, Si Y, Mao J, Fang G, Liang J (2013) Fractions of Cu, Cd, and enzyme activities in a contaminated soil as affected by applications of micro- and nanohydroxyapatite. J Soils Sediments 13:742–752
Deshpande LM, Kapadnis BP, Chopade BA (1993) Metal resistance in Acinetobacter and its relation to beta-lactamase production. Biometals 6:55–59
Dixit R, Malaviya D, Pandiyan K, Singh UB, Sahu A, Shukla R, Singh BP, Rai JP, Sharma PK, Lade H, Paul D (2015) Bioremediation of heavy metals from soil and aquatic environment: an overview of principles and criteria of fundamental processes. Sustainability 7:2189–2212
Eze E, Eze U, Eze C, Ugwu K (2009) Association of metal tolerance with multidrug resistance among bacteria isolated from sewage. J Rural Trop Public Health 8:25–29
Fromin N, Porte B, Lensi R, Hamelin J, Domenach AM, Buatois B, Roggy JC (2012) Spatial variability of the functional stability of microbial respiration process: a microcosm study using tropical forest soil. J Soils Sediments 12:1030–1039
Gadd GM (2010) Metals, minerals and microbes: geomicrobiology and bioremediation. Microbiol 156:609–643
Gales AC, Reis AO, Jones RN (2001) Contemporary assessment of antimicrobial susceptibility testing methods for polymyxin B and colistin: review of available interpretative criteria and quality control guidelines. J Clin Microbiol 39:183–190
Girma G (2015) Microbial bioremediation of some heavy metals in soils: an updated review. Ind J Sci Res 6:147–161
Gonzalez-Guerrero M, Cano C, Azcon-Aguilar C, Ferrol N (2007) GintMT1 encodes a functional metallothionein in Glomus intraradices that responds to oxidative stress. Mycorrhiza 17:327–335
Hedrich S, Schlomann M, Johnson DB (2011) The iron-oxidizing proteobacteria. Microbiology 157:1551–1564
Hossain MA, Piyatida P, Silva JAT, Fujita M (2012) Molecular mechanism of heavy metal toxicity and tolerance in plants: central role of glutathione in detoxification of reactive oxygen species and methylglyoxal and in heavy metal chelation. J Bot 2012:1–37
Jones JB Jr (1977) Elemental analysis of soil extracts and plant tissue ash by plasma emission spectroscopy. Comm Soil Sci Plant Anal 8:349–365
Kabata-Pendias A, Pendias H (2001) Trace elements in soils and plants, 3rd edn. CRC Press, Boca Raton, p 413
Karbasizaed V, Badami N, Emtiazi G (2004) Antimicrobial, heavy metal resistance and plasmid profile of coliforms isolated from nosocomial infections in a hospital in Isfahan, Iran. Afr J Biotechnol 2(10):379–383
Kermani JN, Ghasemi MF, Khosravan A, Farahmand A, Shakibaie MR (2009) Cadmium bioremediation by metal-resistant mutated bacteria isolated from active sludge of industrial effluents. Iran J Environ Health Sci Eng 7:279–286
Kumar M, Kaur N, Kamini G, Pathak R, Khasa YP, Raj L (2013) Reporting heavy metal resistance bacterial strains from industrially polluted sites of northern India using fatty acid methyl ester (FAME) analysis and plasma-atomic emission spectroscopy (ICP-AES). Adv Sci Lett 19:3311–3314
Maegan JO, Ilenys MPD (2009) Influence of microbial growth on the redox potential of fermented cucumbers. J Food Sci 74:M149–M153
Mao DP, Zhou Q, Chen CY, Quan ZX (2012) Coverage evaluation of universal bacterial primers using the metagenomic datasets. BMC Microbiol 12:66
McArthur JV, Tuckfield RC (2000) Spatial patterns in antibiotic resistance among steam bacteria: effects of industrial pollution. Appl Environ Microbiol 66:3722–3726
McLaughlin MJ, Zarcinas BA, Stevens DP, Cook N (2000) Soil testing for heavy metals. Commun Soil Sci Plant Anal 31:1661–1700
Mishra A, Malik A (2013) Recent advances in microbial metal bioaccumulation. Crit Rev Environ Sci Technol 43:1162–1222
Monachese M, Burton JP, Reid G (2012) Bioremediation and tolerance of humans to heavy metals through microbial processes: a potential role for probiotics? Appl Environ Microbiol 78:6397–6404
Mudhoo A, Garg VK, Wang S (2012) Removal of heavy metals by biosorption. Environ Chem Lett 10:109–117
Nieto JJ, Fernandez-Castillo R, Marquez MC, Ventosa A, Quesada E, Ruiz-Berraquero F (1989) Survey of metal tolerance in moderately halophilic eubacteria. Appl Environ Microbiol 55:2385–2390
Noorjahan CM (2014) Physicochemical characteristics, identification of bacteria and biodegradation of industrial effluent. J Bioremed Biodeg 5:219
Ozaki T, Kimura T, Ohnuki T, Yoshida Z, Francis A (2003) Association mechanisms of europium (III) and curium (III) with Chlorella vulgaris. Environ Toxicol Chem 22:2800–2805
Pantano G, Campanha MB, Moreira AB, Bisinoti MC (2014) Occurrence of Cu and Cr in the sedimentary humic substances and pore water from a typical sugar cane cultivation area in São Paulo, Brazil. J Soils Sediments 14:377–384
Perpetuo EA, Souza CB, Nascimento CAO (2011) Engineering bacteria for bioremediation. In: Carpi A (ed) Progress in molecular and environmental bioengineering: from analysis and modeling to technology applications. InTech, Rijeka, pp 605–632
Ray SA, Ray MK (2009) Bioremediation of heavy metal toxicity-with special reference to chromium. Al Ameen J Med Sci 2:57–63
Roane TM, Kellogg ST (1996) Characterization of bacterial communities in heavy metal contaminated soils. Can J Microbiol 42:593–603
Singh M, Ansari AA, Müller G, Singh IB (1997) Heavy metals in freshly deposited sediments of the Gomati River (a tributary of the Ganga River): effects of human activities. Environ Geol 29:246–252
Srinath T, Verma T, Ramteke PW, Garg SK (2002) Chromium (VI) biosorption and bioaccumulation by chromate resistant bacteria. Chemosphere 48:427–435
Sriprang R, Murooka Y (2007) Accumulation and detoxification of metals by plants and microbes. In: Singh SN, Tripathi RD (eds) Environmental bioremediation technologies. Springer, Berlin, pp 77–100
Tan YF, O’Toole N, Taylor NL, Millar AH (2010) Divalent metal ions in plant mitochondria and their role in interactions with proteins and oxidative stress-induced damage to respiratory function. Plant Physiol 152:747–761
Taylor JK (1987) Quality assurance of chemical measurements. Lewis Publishers, Inc, Chelsea
Tchounwou PB, Yedjou CG, Patlolla AK, Sutton DJ (2012) Heavy metal toxicity and the environment. In: Luch A (ed) Molecular, clinical and environmental toxicology. Springer, Basel, pp 133–164
Umysova D, Vitova M, Douskova I, Bisova K, Hlavova M, Cizkova M, Machat J, Doucha J, Zachleder V (2009) Bioaccumulation and toxicity of selenium compounds in the green alga Scenedesmus quadricauda. BMC Plant Biol 9:58
Venosa AD (2004) Literature review on the use of commercial bioremediation agents for cleanup of oil-contaminated estuarine environments. National risk management research laboratory, office of research and development, U.S. environmental protection agency, Cincinnati, OH, pp 1–56
Weber KA, Achenbach LA, Coates JD (2006) Microorganisms pumping iron: anaerobic microbial iron oxidation and reduction. Nat Rev Microbiol 4:752–764
Wireman J, Liebert CA, Smith T, Summers AD (1997) Association of mercury resistance with antibiotic resistance in gram-negative fecal bacteria of primates. Appl Environ Microbiol 63:4494–4503
Xu ZH, Ward S, Chen CR, Blumfield T, Prasolova NV, Liu JX (2008) Soil carbon and nutrient pools, microbial properties and gross nitrogen transformations in adjacent natural forest and hoop pine plantations of subtropical Australia. J Soils Sediments 8:99–105
Acknowledgments
This research was supported by the Amity University Uttar Pradesh and Lovely Professional University, Punjab, India. The authors are grateful to the University for the support.
Author contributions
MK and JS both are having joint corresponding authorship based on the equal contribution. MK, JS, VK, AV, AP, AKS and AA were involved in manuscript designing and data interpretation. AP was involved in sample collection and analysis. JS and RP were involved in the statistical analysis. All authors read and approved the final manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interests.
Additional information
Responsible editor: Kirk T. Semple
Manoj Kumar and Vivek Kumar contributed equally to this article.
Rights and permissions
About this article
Cite this article
Kumar, M., Kumar, V., Varma, A. et al. An efficient approach towards the bioremediation of copper, cobalt and nickel contaminated field samples. J Soils Sediments 16, 2118–2127 (2016). https://doi.org/10.1007/s11368-016-1398-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11368-016-1398-1