Abstract
The use of plants for the improvement of soils contaminated with hydrocarbons has been a primary research focus in phytoremediation studies. Obtaining insights regarding genes that are differentially induced by petroleum hydrocarbon stress and understanding plant response mechanisms against petroleum hydrocarbons at molecular level is essential for developing better phytoremediation strategies to remove these hazardous contaminants. The purpose of this study was to analyze the transcriptomal profile changes under hydrocarbon stress in maize plants and identify the genes associated with the phytoremediative capacity. Zea mays GeneChips were used to analyze the global transcriptome profiles of maize treated with different concentrations of petroleum hydrocarbons. In total, 883, 1281, and 2162 genes were differentially induced or suppressed in the comparisons of 0 (control) vs. 1% crude petroleum, 1 vs. 5% crude petroleum, and 0 vs. 5% crude petroleum, respectively. The differentially expressed genes were functionally associated with the osmotic stress response mechanism, likely preventing the uptake of water from the roots, and the phytoremediative capacity of plants, e.g., secretory pathway genes. The results presented here show the regulatory mechanisms in the response to petroleum hydrocarbon pollution in soil. Our study provides global gene expression data of Z. mays in response to petroleum hydrocarbon stress that could be useful for further studies investigating the biodegradation mechanism in maize and other plants.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Petroleum hydrocarbon contamination in soil fields is a serious problem, particularly in countries that produce, transport, and refine oil (Lu et al. 2014). Phytoremediation is an effective technique used for the long-term in situ rehabilitation of petroleum hydrocarbon contamination (Vidali 2001). The removal of pollutants from soil is a disruptive and expensive physical process (Trellua et al. 2016). Therefore, phytoremediation is an efficient and promising cleanup approach for extracting or inactivating organic and inorganic pollutants from soil.
Different plant groups have investigated to reveal specific mechanisms to reduce oil contamination in soil (Jones et al. 2004; Liste and Prutz 2006; Kechavarzi et al. 2007). Plant species vary considerably in their phytoremediation capacity (Liste and Alexander 1999; Liste and Prutz 2006). Wheat, rye, oat, maize, and other agricultural crops have been reported to tolerate crude oil pollutants in soil (Aprill and Sims 1990; Wild et al. 1992; Liste and Alexander 1999). The bioremediation capability of plant species has been established in different studies investigating oil-contaminated soil (Jones et al. 2004; Kaimi et al. 2006; White et al. 2006; Yergeau et al. 2012). Even though effective phytoremediation of hydrocarbon pollution has been demonstrated, the specific adaptations responsible for this ability and the mechanisms involved in managing the resistance of particular plants to pollutants are not fully understood.
The specific adaptation mechanisms regulating the resistance of specific plants to contaminants are very important for the efficient phytoremediation of organic compounds. Researchers are increasingly focused on this area, investigating factors such as pollutant-specific tolerance mechanisms and structural and functional changes in the rhizosphere under pollution. The identification of the most effective remediating species for particular compounds is a priority in phytoremediation experiments. During the phytoremediation of crude oil-contaminated soil, the effect of plant species on colonization pattern and metabolic activity of the inoculated endophytes was evaluated in different studies (Fatima et al. 2016).
Polycyclic-aromatic hydrocarbons (PAHs) are major components in petroleum-contaminated soils, are resistant to degradation processes, have strong carcinogenic effects on human health, and pose a serious risk to ecosystems (Totsche et al. 2006; Wehrer and Totsche 2009). PAH treatments in plants lead to different stresses. PAHs cause leaf deformations, accumulation of H2O2, oxidative stress, up-regulation of antioxidant systems, cell death, and reduced plant growth (Gao and Zhu 2004; Alkio et al. 2005; Burritt 2008; Liu et al. 2009). Phytotoxicity differs according to the specific PAH applied and the plant species (Baek et al. 2004). However, information regarding the mechanisms of PAH toxicity in plants, the genes responsible for PAH degradation and stress signaling mechanisms, is not fully understood.
cDNA microarrays have been reported to be effective tools for the identification of stress-responsive genes involved in tolerance and stress responses (Seki et al. 2004; Shinozaki et al. 2003). The present study aims to contribute fundamental information regarding the mechanisms underlying tolerance to the toxic effects of petroleum hydrocarbons on plant leaves and roots using the Affymetrix Zea mays GeneChip. The mechanism by which petroleum hydrocarbon exposure activates stress signaling pathways common to other abiotic or biotic stresses has not been well characterized, and whether the identified genes are restricted to petroleum hydrocarbon stress is unclear.
We report the common and distinct regulatory mechanisms involved in the response to petroleum hydrocarbon contaminated soil in leaf and root tissues of Z. mays. Molecular knowledge regarding the stress response and defense mechanisms is imperative for finding new approaches to promote plant tolerance to PAHs.
Materials and methods
Plant material and petroleum treatment
Zea mays L. Shemal seeds were surface sterilized for 5 min in 1% (w/v) sodium hypochlorite and washed with distilled water. After soaking the sterilized seeds in distilled water for 12 h, a pre-germination step was performed in Petri dishes for 10 days in the dark. Three independent experiments were performed for the effect of petroleum on Z. mays growth and each petroleum concentration was examined in triplicate. The germinated seeds were sown in pots containing a soil and sand mixture contaminated with 0 (control), 1, 2.5, and 5% Siberian Light (SBL) crude petroleum. The equal amount of ¼ Hoagland’s solution was applied twice a week to all the pots and grown under 25/22 ± 2 °C and 60 ± 5% % RH with the 8-h light/16-h dark cycle. After 30 days, the leaves and roots from the petroleum-treated (1, 2.5, and 5%) and untreated control plants were harvested and the roots were rinsed in double distilled water for 3 min to remove any surface contamination. Total fresh weights of leaves and roots were measured and frozen in liquid nitrogen for RNA isolation.
RNA isolation
Total RNA was isolated from the leaf and root tissues (100 mg) using the RNeasy Plant Kit (Qiagen, Germany). Each RNA sample was incubated with RNase-free DNaseI (Recombinant, Roche Applied Science GmbH, Germany) to eliminate the genomic DNA and purified according to a previously described method (Keskin et al. 2010). The RNA integrity and the quantity of the purified RNA samples were determined using a spectrophotometer (Nanodrop, ThermoFisher Scientific, USA) and 1.0% (w/v) formaldehyde agarose gel electrophoresis.
cDNA synthesis for quantitative real time PCR
The cDNA synthesis was performed using MMLV reverse transcriptase (Roche Transcriptor High Fidelity cDNA Synthesis Kit), according to the manufacturer’s instructions. The efficiencies of the cDNA synthesis reactions were assessed by performing conventional PCR using a housekeeping gene-specific primer designed from Z. mays β-actin. The QRT-PCR experiments were performed in triplicate of each sample for three biological replicates.
Microarray analysis
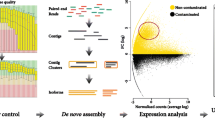
GeneChip Maize Genome Arrays were used to identify the differentially expressed genes. The arrays contain approximately 14,850 Zea mays transcripts that correspond to 13,339 genes. One microgram DNA-free RNA was used to generate biotin-labeled cDNA. The leaves and roots of Z. mays seedlings that were grown for 30 days in different petroleum hydrocarbon contaminated soils (control, 1, and 5%) were individually sampled, and each sample included three independent biological replicates for the microarray analysis. The hybridization, washing, staining, and scanning steps were performed according to the Affymetrix GeneChip standard protocol, and all protocols were performed at AY-KA Ltd. (Ankara, Turkey). Three biological replicates of each sample were performed to assess the quality and reproducibility of the chip hybridization. GeneChip suite 4.0 (Affymetrix) was used for the data normalization, and the fold changes were determined using GENESPRING 5.0. The Affymetrix CEL data analysis was performed using R Studio Version 0.98.1091 (http://www.bioconductor.org/biocLite.R). The data pre-processing included a background correction performed using the Robust Multiarray Average (RMA) algorithm. The normalization and PM correction were performed using the LOESS method and MAS5 algorithm, respectively, and the median polish method was applied for the summary.
Quantitative real-time PCR analysis
To independently confirm the microarray data, qRT-PCR experiments were performed using an IQ5 System (BioRad Laboratories, Hercules, USA). The qRT-PCR reactions were performed in 25 μl total volumes in 96 well polypropylene plates containing 200 ng of the cDNA samples, 10 pmole of each primer, and 12.5 μl SYBR Green Mix (Roche FastStart Universal SYBR Green Master), as described by Cevher-Keskin et al. (2012). The IQ5 System (BioRad Laboratories, Hercules, USA) was used for all qRT-PCR reactions in cycles of 3 min at 95 °C, followed by 50 cycles at 95 °C for 30 s, 55 °C for 30 s, and 72 °C for 40 s. To rule out possible contamination, we subjected a negative control containing no template cDNA to the same procedure in all experiments. To accurately quantify the transcript level, we performed three technical replicates for each experiment. The relative abundance levels of Cys protease (NM_001112009.1), CBL Interacting Protein Kinase (CIPK) (NM_001115006.2), 12-oxophytodienoic acid reductase (OPR) (NM_001112429.1), polyamine oxidase1 (pao1) (NM_001111636.1), early nodulin 93 (Enod93) (NM_001153581.1), and glutamine synthetase2 (GS2) (NM_001111973.1) were detected. The primer sequences of these genes are shown in Table 1. The comparative CT method was used to calculate the fold changes in expression (Schmittgen et al. 2000). The ΔCT values of all mRNA transcripts were averaged across all treatments and experimental replicates. The mRNA expression levels of the genes were normalized to the levels of β-actin in all qRT-PCR experiments.
Results
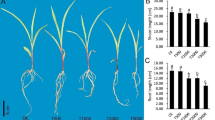
To determine the differences in the transcript levels among the petroleum hydrocarbon-treated Z. mays, we performed microarray experiments using whole plants grown for 30 days in pots containing 0, 1, 2.5, and 5% petroleum contaminated soil. Petroleum hydrocarbons in soils negatively impact plant growth and development (Fig. 1). In all tested concentrations, chlorosis symptoms were detected in the leaves of Z. mays. Significant decreases were observed in the shoot and root lengths and the number of leaves, nodes, and branches (Fig. 2).
The GeneChip Maize Genome Array (Affymetrix, California, USA) was used to characterize the petroleum hydrocarbon responsive genes in maize. Figures 3 and 4 show the distribution of the different expression levels of the genes in leaf and root tissues treated with two different petroleum hydrocarbon percentages (1 and 5%). The Venn diagrams show the number of genes that are responsive to petroleum hydrocarbon in the leaf (Fig. 3) and root (Fig. 4) tissues in Z. mays.
Venn diagram demonstrating the number of up- and down-regulated genes in leaf tissues under petroleum hydrocarbon stress. Genes whose expression was significantly changed (p < 0.05) by twofold are shown. The maximum gene expression difference (2668 genes) was observed in the leaf tissues of the plants treated with 5% petroleum hydrocarbon. a Differentially expressed genes in leaf tissues exposed to 1% oil are shown in pink. b Differentially expressed genes in leaf tissues exposed to 5% oil are shown in green. c Differentially expressed genes regulated in leaf tissue by 1 and 5% oil are shown in blue
Number of differentially expressed probe sets under petroleum hydrocarbon stress. Venn diagrams of up-regulated and down-regulated genes in root tissue. Genes that changed their expression significantly (p < 0.05) by twofold are given. The maximum gene expression difference (2162 genes) was observed in root tissues of 5% petroleum hydrocarbon treated plants. a Genes that change their expression in root tissue exposed to 1% oil are shown in pink. b Genes that change their expression in root tissue exposed to 5% oil are shown in green. c Genes that are commonly regulated by 1 and 5% oil are shown in blue
The distribution of the expression levels of the genes in the leaf tissues was more variable than that in the root tissues (Fig. 3). Expectedly, the maximum gene expression difference in both the root and leaf tissues was observed in the 5% petroleum-treated Z. mays plants (Figs. 3 and 4). In the 5% petroleum hydrocarbon-treated plants, 2668 and 2162 up- and down-regulated genes, respectively, with greater than a twofold change were identified in the leaf and root tissues (Suppl. Data 1 and 2). After the pre-processing step, the farthest neighbor algorithm was applied to select ten highly differentially expressed genes and the 20 genes clustered in the diagrams (Suppl. Data 3 and 4). Table 2 shows the gene IDs and functional descriptions of these 20 highly differentially expressed genes.
Molecular functional categories of the differentially regulated genes in the root and leaf tissues in Z. mays
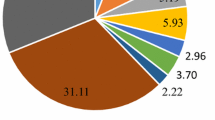
The differentially expressed genes in the root and leaf tissues of the petroleum hydrocarbon-treated Z. mays were categorized according to their biological and molecular functions and cellular location using the agriGO (http://bioinfo.cau.edu.cn/ agriGO/) program. To identify the genes that were regulated by petroleum hydrocarbon, we classified the up- and down-regulated genes according to their functions, including binding, catalytic activity, electron carrier, transporter, structural molecule, transcription regulator activity, and molecular transducer activity. Notably, most differentially expressed genes in the leaf tissues belonged to the structural molecular activity group (Fig. 5). This group includes basic vital functional genes, such as new ribosome biogenesis genes. This result was consistent with the observation that stress-related gene expression activation induces protein synthesis in leaf tissues. Because petroleum hydrocarbon treatments induce responses similar to those induced by drought stress, increased mRNA levels of osmotic stress- and antioxidant enzyme-related genes were observed in the root tissues (Fig. 6).
In leaf tissues, the differentially expressed genes with greater than twofold changes were classified according to their molecular functions. Most differentially expressed genes in the leaf tissues belonged to the structural molecular activity group. This group includes basic vital functional genes, such as new ribosome biogenesis genes. This result was consistent with the observation that stress-related gene expression activation induces protein synthesis in leaf tissues
The expression levels of catalytic activator genes in the root tissues were up-regulated but were not higher than those in the leaf tissues (Fig. 7). The expression level of catalytic activator genes, such as hormone biosynthesis-related genes (abscisic acid and gibberellic acid), was increased in the petroleum hydrocarbon-treated leaf tissues of Z. mays (Fig. 7). The structural molecular activity in the leaf tissues was also up-regulated by petroleum hydrocarbons (Fig. 7).
Differentially expressed genes with greater than twofold changes in three leaf tissue groups (0–1%, 0–5%, and 1–5%) were classified according to their molecular functions. The expression levels of catalytic activator genes, such as hormone biosynthesis-related genes (abscisic acid and gibberellic acid), and the structural molecular activity were increased in the petroleum hydrocarbon-treated leaf tissues of Z. mays. The antioxidant and electron carrier activity was up-regulated in the root tissues of the 1 and 5% petroleum hydrocarbon-treated Z. mays (Fig. 8)
Regarding the biological categories of genes, the expression of cell component biogenesis-related genes in the leaf tissues was higher than that in the root tissues of the petroleum hydrocarbon-treated plants (Figs. 9 and 10). Biological regulation and the regulation of biological processes were suppressed in the root and leaf tissues of the petroleum-treated Z. mays (Figs. 9 and 10). The response to stimulus, multi-organism, and developmental process was up-regulated in the root tissues of the petroleum-treated plants (Fig. 10).
In the leaf tissues, the differentially expressed genes with greater than twofold changes were classified according to their biological function. The expression of cell component biogenesis-related genes in the leaf tissues was higher than that in the root tissues of petroleum hydrocarbon-treated plants
Quantitative RT-PCR
To confirm the microarray data, differentially expressed six genes that might be implicated in different stress responses were selected for qRT-PCR experiments (Table 1). qRT-PCR analyses of these transcripts were performed in Z. mays roots and leaves grown in soils polluted with different concentrations of petroleum (0, 1, 2.5, and 5%).
Cys proteases are involved in cellular regulation (Wagstaff et al. 2002), and the maximal Cys expression was observed in the 2.5% petroleum hydrocarbon-treated leaf tissues. In the 1% petroleum hydrocarbon-treated leaf tissues, the mRNA expression of Cys was higher than that in the control tissue. The maximal Cys expression was observed in the 2.5% treated-plants, and the Cys expression was lower in the 5% petroleum-treated plant leaves (Fig. 11a). The mRNA expression levels of the Cys gene were decreased by petroleum in the root tissues of Z. mays (Fig. 11b).
Total RNA was isolated from 30-day-old Z. mays grown on 0, 1, 2.5, and 5% petroleum hydrocarbon-contaminated soil and used for qRT-PCR experiments. The mRNA regulation of Cys protease, CIPK, and Opr in root and leaf tissues of Z. mays are shown. Error bars represent the standard deviation of the qRT-PCRs performed in triplicate. The gene expression was normalized using β-actin as a housekeeping gene. a Maximal CIPK expression was observed in the 2.5% petroleum hydrocarbon-treated plant leaves. b In the root tissues, the expression levels decreased as the petroleum hydrocarbon concentration increased. c Maximal CIPK expression was observed in the 2.5% petroleum hydrocarbon-treated plant leaves. d In the root tissues, the expression levels were decreased by the petroleum hydrocarbon treatment. e Opr mRNA from plants grown in soil with 2.5 and 5% petroleum hydrocarbon showed 25- and 30-fold changes in expression level relative to the control tissues, respectively. f Opr expression in the root tissues was not dramatically affected by the different petroleum concentrations
The CBLs constitute a family of Ca2+ sensors that are only found in plants. The responses of CIPK (CBL-interacting protein kinase) to the petroleum hydrocarbon treatment were different between the root and leaf tissues (Fig. 11c, d). The maximal CIPK expression was observed in the 2.5% petroleum hydrocarbon-treated plant leaves (fourfold). In contrast, the expression levels were decreased by the petroleum hydrocarbon treatment in the root tissues (Fig. 11d).
The 12-oxo-phytodienoic acid reductases (OPRs) are the enzymes that are important for the detoxification process. In leaves, Opr mRNAs from plants grown in soil contaminated by 2.5 and 5% petroleum hydrocarbon showed 25- and 30-fold expression level changes relative to the control tissues, respectively (Fig. 11e, f). The Opr expression in the root tissues was not dramatically affected by the different petroleum concentrations (Fig. 11f).
Pao is involved in ABA-induced cytosolic antioxidant defense through H2O2 in maize leaves. H2O2 is derived from polyamine oxidation due to cell death, the hypersensitivity response, and the expression of defense genes (Yoda et al. 2006; Xue et al. 2009). According to our qRT-PCR analysis, the highest Pao1 gene expression was observed in the 1% petroleum hydrocarbon-treated leaf tissues. Decreasing expression patterns were observed in the leaf tissues as the concentration of petroleum hydrocarbon increased (Fig. 12a). In the root tissues, significant differences were observed among the 1, 2.5 and 5% petroleum hydrocarbon-treated Z. mays, but there was no correlation between the Pao gene expression and the concentration of petroleum hydrocarbon (Fig. 12b).
mRNA expression patterns of Pao, GS2, and Enod93 in the root and leaf tissues of Z. mays grown in soil contaminated with 0, 1, 2.5, and 5% petroleum hydrocarbon. Gene expression was normalized to β-actin, which was used as a housekeeping gene. Error bars represent the standard deviation of the qRT-PCRs performed in triplicate. a The highest Pao gene expression was observed in the 1% petroleum hydrocarbon-treated leaf tissue. Decreasing expression patterns were observed in the leaf tissues as the concentration of petroleum hydrocarbon increased. b In the root tissues, no correlation was observed between the Pao gene expression and the concentration of petroleum hydrocarbon. c Maximal GS2 gene expression was observed in the 1% petroleum hydrocarbon-treated leaf tissues (20-fold). Decreasing expression pattern was observed as the concentration of petroleum hydrocarbon increased. d However, no significant difference was observed in the root tissues. f Enod93 mRNA expression in the root tissues increased as the concentration of petroleum hydrocarbons increased. Maximal expression was observed in the 5% petroleum hydrocarbon-treated root tissues. e In contrast, the opposite Enod93 expression profile was observed in the leaf tissue
In plant cells, GS2 plays an important role in the flow of nitrogen into nitrogenous organic compounds and is involved in glutamate synthesis. Maximal GS2 gene expression was observed in the 1% petroleum hydrocarbon-treated leaf tissue (20-fold). Decreasing expression patterns were observed in the leaf tissues as the concentration of petroleum hydrocarbon increased. In contrast, no significant difference was observed in the root tissues (Fig. 12c, d).
The ENOD93 proteins are assumed to play roles in cell structure, the control of nodule ontogeny, and carbon metabolism (Okubara et al. 2000). According to our qRT-PCR experiments, Enod93 mRNA expression increased in the root tissues as the concentration of petroleum hydrocarbon increased. Maximal expression was observed in the 5% petroleum hydrocarbon-treated root tissues. However, the opposite Enod93 expression profile was observed in the leaf tissues (Fig. 12e, f).
Discussion
The main goal of this study was to identify the global expression profile in response to petroleum hydrocarbon stress in Z. mays. The reduced growth of Z. mays plants due to the petroleum-contaminated soil might be the cause of the reduction in leaf growth and/or delay in cell expansion (Agbogidi et al. 2007; Athar et al. 2016). Organic chemical pollution in soil and water has been demonstrated to extensively affect microbial populations (Vilchez-Vargas et al. 2013; Gonzalez et al. 2015). According to the microarray analysis, the expression of genes related to general stress response mechanisms (such as GST) was increased as the petroleum hydrocarbon concentration was increased. The increases in the percentage of the oil prevented plant water intake and, therefore, incurred transcriptional changes that are observed following drought stress. The increased expression levels of certain genes, such as those in the Derlin general secretion pathway, may influence the provision of microorganisms in the rhizosphere. In addition, the suppression of certain root meristematic stem cell differentiation genes, such as Ocl4, may retard the growth of plant root tissue.
Cys
Cys protease expression was up-regulated in the root tissues as the concentrations of petroleum hydrocarbon increased, closely correlating with the microarray data (Fig. 11b). Cys proteases are important for cellular regulation and several developmental events associated with programmed cell death (Elbaz et al. 2002; Estelle 2001; Matarasso et al. 2005). Cys protease has also been shown to be expressed during drought, senescence ripening, and wounding (Alonso and Granell 1995; Harrak et al. 2001).
Cysteine proteases are involved in signaling pathways and the response to biotic and abiotic stresses. Four different cysteine protease probes were induced in the leaf tissues of Z. mays plants.
CBL–CIPK complexes regulate abiotic stress responses, such as responses to drought, cold, and salt and ABA signaling (Li et al. 2006; Xu et al. 2006). According to the qRT-PCR analysis, the responses of CIPK to the petroleum hydrocarbon treatment differed between the root and leaf tissues (Fig. 11c, d). Decreasing expression was observed with increasing concentrations of petroleum in the root tissues of Z. mays (Fig. 11d).
According to the qRT-PCR experiments, the Opr expression levels in the root tissues increased with increasing concentrations of petroleum (Fig. 11f). The OPRs catalyze the decrease in the double bonds adjacent to an oxo group in a, b-unsaturated aldehydes or ketones. Therefore, these enzymes are very important for the detoxification process. Previous reports (Zhang et al. 2005) suggest that different Opr genes are also differentially regulated in response to stress hormones, wounding, and biotic stress, such as pathogen infection (Sobajima et al. 2007; Zhang et al. 2005).
In plants, polyamine oxidation-derived hydrogen peroxide (H2O2) production has been associated with cell wall development, ABA-induced antioxidant mechanism, wound healing, cell death, and biotic stress (Angelini et al. 2008; Yoda et al. 2006; Xue et al. 2009). Abiotic stress, such as drought, heat, and salt induced Pao gene expression in plants and bacteria (Aziz et al. 1998; Suzuki et al. 2004). Pao expression has also been reported in the ABA-related antioxidant mechanism in maize leaves (Cona et al. 2006). According to our microarray data, Pao expression in the leaf tissues increased as the oil concentration increased. The qRT-PCR results confirmed the petroleum hydrocarbon stress responsive behavior of Pao (Fig. 12a, b).
GS is an important enzyme for nitrogen metabolism, which converts toxic ammonium to nontoxic glutamine (Gln) in the presence of glutamate and ATP. Genetic transformations of GS genes have been performed in different systems, such as transgenic lotus plants, tobacco, wheat, and poplar trees, to enhance nitrogen assimilation (Fuentes et al. 2001; Habash et al. 2001; Oliveira et al. 2002; Pascual et al. 2008). Maximal GS2 gene expression was observed in the 1% petroleum hydrocarbon-treated leaf tissues (20-fold) according to the qRT-PCR analysis (Fig. 12c, d). Decreased expression patterns were observed in leaf tissues as the petroleum hydrocarbon concentrations increased.
According to the qRT-PCR analysis, Enod93 mRNA expression was up-regulated in the root tissues and down-regulated in the leaf tissues with increasing petroleum hydrocarbon concentrations (Fig. 12e, f) “Nodulation (nod)” genes play an important role in the induction of nodules in leguminous plant roots. The development of the nodule depends on the coordinated expression of plant and bacterial genes. The nodulation process is initiated by the stimulation of root cortical cell division (Tam et al. 1997). Enod40 is rapidly triggered by rhizobia in the root pericycle and dividing cortical cells of the nodule primordium (Yang et al. 1993; Crespi 1994).
Antioxidant enzymes
Under biotic or abiotic stress, major reactive oxygen species (ROS) (superoxide oxidase and H2O2) are produced in plant cells (Moller et al. 2007), and antioxidant and detoxification enzymes are up-regulated under these stresses. In maize, CAT (Zm.6357.3.S1_a_at), APX Zm.5762.1.A1_at), SOD (Zm.241.1.S1_at), and GR (Zm.7168.1.S1_at) were up-regulated under petroleum hydrocarbon pollution. Ten microarray probes representing peroxidase were up-regulated in our microarray data, particularly in the root tissues. The APX1 gene transcript was highly abundant in willow trees grown in petroleum-contaminated soil (Gonzalez et al. 2015).
Versicolorin reductase
Oxidoreductase activity was increased in the leaf tissues. In the presence of oxygen and water, aldehyde oxidase produces carboxylic acids from aldehydes and catalyzes the conversion of an aldehyde to an acid and hydrogen peroxide. According to our microarray data, two probes representing aldehyde oxidase, four probes representing 12-oxo-phytodienoic acid reductase, and four probes representing peroxidase were induced in the leaf tissues by petroleum hydrocarbon.
Lipoxygenase gene expression is involved in different stress responses, such as wounding, abiotic (water deficiency), or biotic (pathogen attack) stress responses (Melan et al. 1993; Porta et al. 1999). The LOX5 and LOX8 probes, representing lipoxygenase, were up-regulated in the leaf tissues. Our comparative microarray approach found that Z. mays MAP kinase4 (MPK4) mRNA was up-regulated by 2.6- and 1.5-fold in the leaf tissue grown in the 1 and 5% petroleum hydrocarbon-contaminated soils, respectively. According to a DDRT-PCR analysis in T. aestivum performed by Keskin et al. (2010), the expression of MPK4 mRNA was induced by ABA treatment. Water stress induced MPK4 mRNA expression in T. aestivum, according to an RNAseq analysis (Cevher-Keskin et al. 2015).
Pyruvate decarboxylase catalyzes the oxidation of decarboxylation of pyruvate to acetaldehyde. We observed that pyruvate decarboxylase (pcd2) mRNA was up-regulated in our microarray experiments (twofold by 1% petroleum and 1.9-fold by 5% petroleum). Pyruvate decarboxylase (pcd3) was also up-regulated in the leaf tissues (2.3-fold by 1% and 4.1-fold by 5% petroleum). In contrast, in the root tissue, Pcd1 was down-regulated (25-fold by 1% and 11-fold by 5% petroleum). In the absence of oxygen, an increase in the activity of pyruvate decarboxylase was observed in wheat and maize root tips (Waters et al. 1991; Drew et al. 1994). Thus, increased pyruvate decarboxylase activity is likely a marker of tissue hypoxia or anoxia (Mohanty and Ong 2003).
Carbonyl reductase 1 (cbr1) has oxidoreductase activity and was induced 28.5-fold and 9.8-fold in the leaf tissues of maize grown in 1 and 5% petroleum hydrocarbon-contaminated soil, respectively. Versicolor in reductase, which is another oxidoreductase, was up-regulated in the leaf (18.3-fold by 1% and 11.0-fold by 5% petroleum) and root (2.2-fold by 1% and 2.0-fold by 5% petroleum) tissues.
Transport- and cell wall-related genes
In cellular transport processes, the ion transporters support homeostasis in the plant cytoplasm. The “ion transporter genes” are up-regulated by high salinity stress (Hasegawa et al. 2000). The cell wall protein expansin is involved in cell wall loosening, enlargement, and other developmental processes (Sampedro and Cosgrove 2005). Beta expansins (exB4, exB5, exB7, and exB8) and α-expansins (exA3 and exA5) were up-regulated in the root tissues of Z. mays by petroleum hydrocarbon according to the microarray data. β-Expansin mRNA is induced by ABA treatment in leaf tissues of T. aestivum (Keskin et al. 2010), and six different probes representing glutathione S-transferase12 (gst6, 9, 11, 12, 13, 14) were increased by petroleum in the Z. mays root tissues. Twenty-two probes representing glutathione S-transferase 2 and three different glutamine synthetase2 (gln2) probes were up-regulated in the leaf tissues. The gene expression of gln2 was induced by petroleum hydrocarbon, but no significant difference was found in the root tissues (Fig. 12c, d). The microarray data are consistent with the data obtained from the qRT-PCR experiments.
Abiotic stresses, such as salt and water, stimulate proteolytic activity and decrease protein synthesis, leading to free amino acid accumulation in the cells. Excessive amino acids reduce the need for ammonium incorporation to form ammonium acids and indirectly cause ammonium accumulation. Higher GS1 and GS2 mRNA expression levels induce more ammonium absorption in roots, causing ammonium accumulation and eventually inducing cell senescence.
Tonoplast-membrane-integral protein ZmTIP2-2 (NM_001111561) was induced (threefold) in the root tissues by 1% petroleum hydrocarbon. TIP1 (TaAQP6) participates in internal water redistribution by regulating water transfer under drought stress (Zhang et al. 2008). The overexpression of plasma membrane aquaporin BnPIP1 in Nicotiana tabacum caused an enhanced tolerance to drought stress (Yu et al. 2005). In contrast, ABA-induced TIP1 mRNA expression was observed in the leaf tissues of T. aestivum (Keskin et al. 2010). TIP1 expression is differentially regulated in leaf and root tissues under drought conditions in drought tolerant and non-tolerant T. aestivum cultivars (Cevher-Keskin et al. 2015).
Hormone signaling network
ABA-induced plasma membrane protein PM19 and ABA-induced protein were up-regulated in response to the petroleum hydrocarbon treatment in the root tissues of Z. mays. ABA-treated T. aestivum immature embryo cells were closely related to the considerable accumulation of PM19 (Koike et al. 1997). However, in leaf tissues, auxin inducible protein gene expression was up-regulated approximately 2.3- and 3.5-fold by the 1 and 5% petroleum contaminated-soils, respectively.
Transporters of ion or sugar function by adjusting the osmotic potential between two sides of the membrane. According to our microarray results, inorganic phosphate transporter 3 and polyol transporter protein 4 were down- and up-regulated, respectively, in the root tissues (Suppl. Data 5 and 6). However, inorganic phosphate transporter 3 and plastidic 2-oxoglutarate/malate transporters were down-regulated in the leaf tissues, and zinc transporter 4 was up-regulated (Suppl. Data 7 and 8). Bray (2004) demonstrated that genes related to cell wall metabolism were down-regulated during osmotic stress in Arabidopsis.
Two different probes representing transglutaminase (TGase) were up-regulated by petroleum hydrocarbon. Calcium-dependent TGase activity is regulated by senescence, salt stress, programmed cell death, differentiation, and light in different plants (Serafini-Fracassini and Del Duca 2002; Dondini et al. 2003).
A significant response to oil contamination was observed in the gene expression of chitinase. In the leaf and root tissues, the functional genes encoding chitinase were dramatically down-regulated as the concentration of petroleum hydrocarbon increased. Regarding the nitrogen fixation-related functional genes, the early nodulin 93 genes were significantly down-regulated in the root tissue by the 1 and 5% petroleum hydrocarbon contamination. Nodulin-like protein, which is responsible for transmembrane transport, was up-regulated in the leaf tissues grown in the 1 and 5% petroleum hydrocarbon-contaminated soil.
Because of the hydrophobicity of petroleum-polluted soils, petroleum contamination induces drought stress (Li et al. 1997). Our comparative transcriptome analyses in both root and leaf tissues of Z. mays show that the metabolism of water uptake and osmotic homeostasis-related gene expression is predominantly influenced by the petroleum hydrocarbon treatment.
Conclusion
In conclusion, transcriptome studies performed in maize provided data regarding the mechanism of tolerance to petroleum hydrocarbon. Plants grown in petroleum-contaminated soils showed increased protein and gene expression that induced microorganisms. Because of the toxic crude oil, related genes are activated in plants. Since plant water uptake is affected, the expression levels of genes associated with osmotic balance are also affected. The present results are highly valuable for breeding and engineering plants for the phytoremediation of petroleum hydrocarbons.
Abbreviations
- PAH:
-
Polycyclic-aromatic hydrocarbon
- CIPK:
-
CBL-interacting protein kinase
- OPR:
-
12-oxophytodienoic acid reductases
- pao1:
-
Polyamine oxidase1
- enod93:
-
Early nodulin 93
- GS2:
-
Glutamine synthetase 2
References
Agbogidi OM, Eruotor PG, Akparobi SO (2007) Effects of crude oil levels on the growth of maize (Zea mays L.). Am J Food Technol 2:529–535
Alkio M, Tabuchi TM, Wang X, Colon-Carmona A (2005) Stress responses to polycyclic aromatic hydrocarbons in Arabidopsis include growth inhibition and hypersensitive response-like symptoms. J Exp Botany 56:2983–2994
Alonso JM, Granell A (1995) A putative vacuolar processing protease is regulated by ethylene and also during fruit ripening in citrus-fruit. Plant Physiol 109:541–547
Angelini R, Tisi A, Rea G, Chen MM, Botta M, Federico R et al (2008) Involvement of polyamine oxidase in wound healing. Plant Physiol 146:162–177
Aprill W, Sims RC (1990) Evaluation of the use of prairie grasses for stimulating polycyclic aromatic hydrocarbon treatment in soil. Chemosphere 20:253–265
Athar HR, Ambreen S, Javed M, Hina M, Rasul S, Ullah Z, Hamid Z, Manzoor Ogbaga CC, Afzal M, Al-Qurainy F, Ashraf M (2016) Influence of sub-lethal crude oil concentration on growth, water relations and photosynthetic capacity of maize (Zea mays L.) plants. Environ Sci Pollut Res 23(18):18320–18331
Aziz A, Martin-Tanguy J, Larher F (1998) Stress-induced changes in polyamine and tyramine levels can regulate proline accumulation in tomato leaf discs treated with sodium chloride. Physiol Plant 104:195–202
Baek KH, Kim HS, Oh HM, Yoon BD, Kim J, Lee IS (2004) Effects of crude oil, oil components, and bioremediation on plant growth. J Environ Sci Health A39:2473–2484
Bray EA (2004) Genes commonly regulated by water-deficit stress in Arabidopsis thaliana. J Exp Bot 55:2331–2341
Burritt DJ (2008) The polycyclic aromatic hydrocarbon phenanthrene causes oxidative stress and alters polyamine metabolism in the aquatic liverwort Riccia fluitans L. Plant Cell Environ 31:1416–1431
Cevher-Keskin B, Yuca E, Ertekin O, Yuksel B, Memon AR (2012) The effect of developmental stages and light conditions on the expression characteristics of ARF1 and SAR1. Plant Biol 14:24–32
Cevher-Keskin B, Yıldızhan Y, Kulen O, Onarıcı S (2015) Quantitative expression analysis of TaMPK4 and TaTIP1 genes in drought tolerant and non-tolerant wheat (Triticum aestivum L.) cultivars. Plant Omics J 8:270–277
Cona A, Rea G, Angelini R, Federico R, Tavladoraki P (2006) Functions of amine oxidases in plant development and defense. Trends Plant Sci 11:80–88
Crespi MD (1994) ENOD40, a gene expressed during nodule organogenesis, codes for a nontranslatable RNA involved in plant-growth. EMBO J 13(21):5099–5112
Dondini L, Del Duca S, Dall'Agata L, Bassi R, Gastaldelli M, Della Mea M, Di Sandro A, Claparols I, Serafini-Fracassini D (2003) Suborganellar localisation and effect of light on Helianthus tuberosus chloroplast transglutaminases and their substrates. Planta 217:84–95
Drew MC, Cobb BG, Johnson JR, Andrews D, Morgan PW, Jordan W, He CJ (1994) Metabolic acclimation of root-tips to oxygen deficiency. Ann Bot 74:281–286
Elbaz M, Avni A, Weil M (2002) Constitutive caspase-like machinery executes programmed cell death in plant cells. Cell Death Differ 9:726–733
Estelle M (2001) Proteases and cellular regulation in plants. Curr Opin Plant Biol 4:254–260
Fatima K, Imran A, Amin I, Khan QM, Afzal M (2016) Plant species affect colonization patterns and metabolic activity of associated endophytes during phytoremediation of crude oil-contaminated soil. Environ Sci Pollut Res 23(7):6188–6196
Fuentes SI, Allen DJ, Ortiz-Lopez A, Hernandez G (2001) Over-expression of cytosolic glutamine synthetase increases photosynthesis and growth at low nitrogen concentrations. J Exp Bot 52:1071–1081
Gao YZ, Zhu LZ (2004) Plant uptake, accumulation and translocation of phenanthrene and pyrene in soils. Chemosphere 55:1169–1178
Gonzalez E, Brereton NJB, Marleau J, Nissim WG, Labrecque M, Pitre FE, Joly S (2015) Meta-transcriptomics indicates biotic crosstolerance in willow trees cultivated on petroleum hydrocarbon contaminated soil. BMC Plant Biol 15:246. https://doi.org/10.1186/s12870-015-0636-9
Habash DZ, Massiah AJ, Rong HL, Wallsgrove RM, Leigh RA (2001) The role of cytosolic glutamine synthetase in wheat. Ann Appl Biol 138:83–89
Harrak H, Azelmat S, Baker EN, Tabaeizadeh Z (2001) Isolation and characterization of a gene encoding a drought-induced cysteine protease in tomato (Lycopersicon esculentum). Genome 44:368–374
Hasegawa PM, Bressan RA, Zhu JK, Bohnert HJ (2000) Plant cellular and molecular responses to high salinity. Ann Rev Plant Physiol Plant Mol Biol 51:463–499
Jones DL, Hodge A, Kuzyakov Y (2004) Plant and mycorrhizal regulation of rhizodeposition. New Phytol 163:459–480
Kaimi E, Mukaidani T, Miyoshi S, Tamaki M (2006) Ryegrass enhancement of biodegradation in diesel-contaminated soil. Environ Exp Bot 55:110–119
Kechavarzi C, Pettersson K, Leeds-Harrison P, Ritchie L, Ledin S (2007) Root establishment of perennial ryegrass (L-perenne) in diesel contaminated subsurface soil layers. Environ Pollut 145:68–74
Keskin BC, Sarikaya AT, Yüksel B, Memon AR (2010) Abscisic acid regulated gene expression in bread wheat (Triticum aestivum L.). Aust J Crop Sci 4:617–625
Koike M, Takezawa D, Arakawa K, Yoshida S (1997) Accumulation of 19-kDa plasma membrane polypeptide during induction of freezing tolerance in wheat suspension-cultured cells by abscisic acid. Plant Cell Physiol 38:707–716
Li X, Feng Y, Sawatsky N (1997) Importance of soil-water relations in assessing the endpoint of bioremediated soils. Plant Growth Plant Soil 192:219–226
Li L, Kim BG, Cheong YH, Pandey GK, Luan S (2006) A Ca2+ signaling pathway regulates a K+ channel for low-K response in Arabidopsis. Proc Natl Acad Sci U S A 103:12625–12630
Liste HH, Alexander M (1999) Rapid screening of plants promoting phenanthrene degradation. J Environ Qual 28:1376–1377
Liste HH, Prutz I (2006) Plant performance, dioxygenase-expressing rhizosphere bacteria, and biodegradation of weathered hydrocarbons in contaminated soil. Chemosphere 62:1411–1420
Liu H, Weisman D, Ye Y-b, Cui B, Huang Y-h, Colon-Carmona A, Wang Z-h (2009) An oxidative stress response to polycyclic aromatic hydrocarbon exposure is rapid and complex in Arabidopsis thaliana. Plant Sci 176:375–382
Lu L, Huggins T, Jin S, Zuo Y, Ren ZJ (2014) Microbial metabolism and community structure in response to bioelectrochemically enhanced remediation of petroleum hydrocarbon-contaminated soil environ. Sci Technol 48:4021–4029
Matarasso N, Schuster S, Avni A (2005) A novel plant cysteine protease has a dual function as a regulator of 1-aminocyclopropane-1-carboxylic acid synthase gene expression. Plant Cell 17:1205–1216
Melan MA, Dong XN, Endara ME, Davis KR, Ausubel FM, Peterman TK (1993) An Arabidopsis thaliana lipoxygenase gene can be induced by pathogens, abscisic-acid, and methyl jasmonate. Plant Physiol 101:441–450
Mohanty B, Ong BL (2003) Contrasting effects of submergence in light and dark on pyruvate decarboxylase activity in roots of rice lines differing in submergence tolerance. Ann Bot 91:291–300
Moller IM, Jensen PE, Hansson A (2007) Oxidative modifications to cellular components in plants. Ann Rev Plant Biol 58:459–481
Okubara PA, Fujishige NA, Hirsch AM, Berry AM (2000) Dg93, a nodule-abundant mRNA of Datisca glomerata with homology to a soybean early nodulin gene. Plant Physiol 122:1073–1079
Oliveira IC, Brears T, Knight TJ, Clark A, Coruzzi GM (2002) Overexpression of cytosolic glutamine synthetase. Relation to nitrogen, light, and photorespiration. Plant Physiol 129:1170–1180
Pascual MB, Ping Jing Z, Kirby EG, Canovas FM, Gallardo F (2008) Response of transgenic poplar overexpressing cytosolic glutamine synthetase to phosphinothricin. Phytochemistry 69:382–389
Porta H, Rueda-Benitez P, Campos F, Colmenero-Flores JM, Colorado JM, Carmona MJ, Covarrubias AA, Rocha-Sosa M (1999) Analysis of lipoxygenase mRNA accumulation in the common bean (Phaseolus vulgaris L.) during development and under stress conditions. Plant Cell Physiol 40:850–858
Sampedro J, Cosgrove DJ (2005) The expansin superfamily. Genome Biol 6:242
Seki M, Satou M, Sakurai T, Akiyama K, Iida K, Ishida J, Nakajima M, Enju A, Narusaka M, Fujita M, Oono Y, Kamei A, Yamaguchi-Shinozaki K, Shinozaki K (2004) RIKEN Arabidopsis full-length (RAFL) cDNA and its applications for expression profiling under abiotic stress conditions. J Exp Bot 55:213–223
Schmittgen TD, Zakrajsek BA, Mills AG, Gorn V, Singer MJ, Reed MW.(2000) Quantitative Reverse Transcription–Polymerase Chain Reaction to Study mRNA Decay: Comparison of Endpoint and Real-Time Methods Analytical Biochemistry 285 ( 2):194–204
Shinozaki K, Yamaguchi-Shinozaki K, Seki M (2003) Regulatory network of gene expression in the drought and cold stress responses. Curr Opin Plant Biol 6(5):410–417
Sobajima H, Tani T, Chujo T, Okada K, Suzuki K, Mori S, Minami E, Nishiyama M, Nojiri H, Yamane H (2007) Identification of a jasmonic acid-responsive region in the promoter of the rice 12-oxophytodienoic acid reductase 1 gene OsOPR1. Biosci Biotech Biochem 71:3110–3115
Serafini-Fracassini D, Del Duca S (2002) Biochemistry and function of plant transglutaminases. Minerva Biotecnol 14: 135–141
Suzuki H, Higashi Y, Asano M, Suguro M, Kigawa M, Maeda M, Katayama S, Mukouyama EB, Uchiyama K (2004) Sequencing and expression of the L-phenylalanine oxidase gene from Pseudomonas sp P-501. Proteolytic activation of the proenzyme. J Biochem 136:617–627
Tam W, BenYehuda D, Hayward WS (1997) bic, a novel gene activated by proviral insertions in avian leukosis virus-induced lymphomas, is likely to function through its noncoding RNA. Mol Cell Biol 17:1490–1502
Totsche KU, Jann S, Kogel-Knabner I (2006) Release of polycyclic aromatic hydrocarbons, dissolved organic carbon, and suspended matter from disturbed NAPL-contaminated gravelly soil material. Vadose Zone J 5:469–479
Trellua C, Mousset E, Pechaud Y, Huguenot D, van Hullebuscha ED, Esposito G, Oturan MA (2016) Removal of hydrophobic organic pollutants from soil washing/flushing solutions: A critical review. J Hazard Mater 306:149–174
Vidali M (2001) Bioremediation, an overview. Pure Appl Chem 73(7):1163–1172
Vilchez-Vargas R, Geffers R, Suárez-Diez M, Conte I, Waliczek A, Kaser VS, Kralova M, Junca H, Pieper DH (2013) Analysis of the microbial gene landscape and transcriptome for aromatic pollutants and alkane degradation using a novel internally calibrated microarray system. Environ Microbiol 15(4):1016–1039
Wagstaff C, Leverentz MK, Griffiths G, Thomas B, Chanasut U, Stead AD, Rogers H (2002) Cys proteases are involved in cellular regulation. J Exp Bot 53:233–240
Waters I, Morrell S, Greenway H, Colmer TD (1991) Effects of anoxia on wheat seedlings: II. Influence of O2 supply prior to anoxia on tolerance to anoxia, alcoholic fermentation, and sugar levels. J Exp Bot 42:1437–1447
Wehrer M, Totsche KU (2009) Difference in PAH release processes from tar-oil contaminated soil materials with similar contamination history. Chem Erde-Geochem 69:109–124
White PM, Wolf DC, Thoma GJ, Reynolds CM (2006) Phytoremediation of alkylated polycyclic aromatic hydrocarbons in a crude oil-contaminated soil. Water Air Soil Pollut 169:207–220
Wild SR, Jones KC, Johnston AE (1992) The polynuclear aromatic hydrocarbon (PAH) content of herbage from a long-term grassland experiment. Atmos Environ Part A 26:1299–1307
Xu J, Li HD, Chen LQ, Wang Y, Liu LL, He L, Wu WH (2006) A protein kinase, interacting with two calcineurin B-like proteins, regulates K+ transporter AKT1 in Arabidopsis. Cell 125:1347–1360
Xue B, Zhang A, Jiang M (2009) Involvement of polyamine oxidase in abscisic acid induced cytosolic antioxidant defense in leaves of maize. J Integr Plant Biol 51:225–234
Yang WC, Katinakis P, Hendriks P, Smolders A, Devries F, Spee J, Vankammen A, Bisseling T, Franssen H (1993) Characterization of GmENOD40, a gene showing novel patterns of cell-specific expression during soybean nodule development. Plant J 3:573–585
Yergeau E, Sanschagrin S, Beaumier D, Greer CW (2012) Metagenomic analysis of the bioremediation of diesel contaminated Canadian high arctic soils. PloSone 7:1–10
Yoda H, Hiroi Y, Sano H (2006) Polyamine oxidase is one of the key elements for oxidative burst to induce programmed cell death in tobacco cultured cells. Plant Physiol 142:193–206
Yu Q, Hu Y, Li J, Wu Q, Lin Z (2005) Sense and antisense expression of plasma membrane aquaporin BnPIP1 from Brassica napus in tobacco and its effects on plant drought resistance. Plant Science 169 (4): 647–656. https://doi.org/10.1016/j.plantsci.2005.04.013
Zhang JL, Simmons C, Yalpani N, Crane V, Wilkinson H, Kolomiets M (2005) Genomic analysis of the 12-oxo-phytodienoic acid reductase gene family of Zea mays. Plant Mol Biol 59:323–343
Zhang JF, Deng ZY, Cao SH, Wang XP, Zhang AM, Zhang XQ (2008) Isolation of six novel aquaporin genes from Triticum aestivum L. and functional analysis of TaAQP6 in water redistribution. Plant Mol Biol Rep 26:32–45
Acknowledgements
This project was supported by The Scientific and Technological Council of Turkey (TUBITAK) TARAL 1007 program “The Bioremediation of the Hydrocarbon Polluted Areas by use of Plants, Algae and Microorganisms” (105G079).
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible editor: Yi-ping Chen
Electronic supplementary material
Suppl. Data 1
Expression levels of significantly up (−) and down (+) regulated to petroleum hydrocarbon-stress responsive transcripts in Z. mays. Differentially expressed genes with greater than twofold changes in three root tissue groups (0–1, 0–5, and 1–5%) were compared. (DOC 285 kb)
Suppl. Data 2
Expression levels of significantly up (−) and down (+) regulated to petroleum hydrocarbon-stress responsive transcripts in Z. mays. Differentially expressed genes with greater than twofold changes in three leaf tissue groups (0–1, 0–5, and 1–5%) were compared. (DOC 711 kb)
Suppl. Data 3
The pre-processing step and farthest neighbor algorithm were performed to select ten highly differentially expressed genes. (PNG 52 kb)
Suppl. Data 4
The farthest neighbor algorithm was applied to select 20 highly differentially expressed genes. (PNG 106 kb)
Suppl. Data 5
Petroleum hydrocarbon up-regulated genes in root tissues (0, 1, and 5%) as obtained by microarray analyses. Rows and columns of the cluster diagram represent genes and treatments, respectively. Bar represents the colors corresponding to expression values. (PNG 118 kb)
Suppl. Data 6
Petroleum hydrocarbon down-regulated genes in root tissues (0, 1, and 5%) as obtained by microarray analyses. Rows and columns of the cluster diagram represent genes and treatments, respectively. Bar represents the colors corresponding to expression values. (PNG 124 kb)
Suppl. Data 7
Petroleum hydrocarbon up-regulated genes in leaf tissues (0, 1, and 5%) as obtained by microarray analyses. Rows and columns of the cluster diagram represent genes and treatments, respectively. Bar represents the colors corresponding to expression values. (PNG 132 kb)
Suppl. Data 8
Petroleum hydrocarbon down-regulated genes in leaf tissues (0, 1, and 5%) as obtained by microarray analyses. Rows and columns of the cluster diagram represent genes and treatments, respectively. Bar represents the colors corresponding to expression values. (PNG 100 kb)
Rights and permissions
About this article
Cite this article
Cevher-Keskin, B., Selçukcan-Erol, Ç., Yüksel, B. et al. Comparative transcriptome analysis of Zea mays in response to petroleum hydrocarbon stress. Environ Sci Pollut Res 25, 32660–32674 (2018). https://doi.org/10.1007/s11356-018-3078-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-018-3078-8