Abstract
Biosynthesis of gold nanoparticles (AuNPs) by microbes has received much attention as an efficient and eco-friendly process. However, the characteristics of AuNPs biosynthesized by different microbial cell-free extracts are rarely comparatively studied. In this study, three locally isolated strains, i.e., bacteria Labrys sp. WJW, yeast Trichosporon montevideense WIN, and filamentous fungus Aspergillus sp. WL-Au, were selected for AuNPs biosynthesis. UV-Vis absorption bands at 538, 539, and 543 nm confirmed the formation of AuNPs by these strains. Transmission electron microscopy and selected area electron diffraction analyses revealed that the as-synthesized AuNPs were crystalline with spherical or pseudo-spherical shapes. However, the average sizes of these AuNPs were diverse, which were 18.8, 22.2 and 9.5 nm, respectively. The biomolecules involved in nanoparticles stabilization were demonstrated by Fourier transform infrared spectroscopy analysis. Four common functional groups such as –N–H, –C=C, –N=O, and –S=O groups were detected in these AuNPs, while a distinct –C=O group was involved in WL-Au-AuNPs. The catalytic rate of WL-Au-AuNPs toward 4-nitrophenol reduction (0.37 min−1) was much higher than those of others (WJW-AuNPs 0.27 min−1 and WIN-AuNPs 0.23 min−1). This research would provide useful information for exploring efficient microbial candidates to synthesize AuNPs with excellent performances.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Gold nanoparticles (AuNPs) have immense applications in the fields of microbiology, physics, chemistry, medicine, and material science due to their distinctive electronic, optical, and catalytic properties (Arvizo et al. 2012; Guo and Wang 2007; He et al. 2014; Scari et al. 2012). Although wet-chemical procedure has been widely utilized for the synthesis of AuNPs due to its simplicity and high growth rate, the usage of toxic and hazardous organic reagents in the wet-chemical synthesis process has raised great concerns on the living organisms and the environment (Seo et al. 2015; Singh et al. 2016a, b). Therefore, it has been of increasing interest to develop an efficient and environmentally benign process to synthesize AuNPs. In recent years, green synthesis of AuNPs by microbes has gained much attention (Ahmad et al. 2013; Dhillon et al. 2012; Hulkoti and Taranath 2014; Park et al. 2016), which was considered as a safe, simple, nontoxic, biocompatible, and environment-friendly alternative to wet-chemical techniques because the microbes can serve as “green factory” for AuNP synthesis without addition of any reducing and capping agents (Kharissova et al. 2013; Moghaddam et al. 2015; Singh et al. 2016a, 2016b).
Many microbes, such as bacteria, actinomycetes, fungi, and yeasts, have been reported to synthesize AuNPs by reducing Au3+ (Kitching et al. 2015; Sugunan et al. 2007). For example, bacteria Geobacillus sp., Anabaena sp., and Lyngbya majuscula were reported to produce hexagonal and spherical AuNPs intracellularly by exposing cells to Au3+ (Brayner et al. 2007; Chakraborty et al. 2009; Correa-Llantén et al. 2013). Monodisperse AuNPs were extracellularly biosynthesized through reduction of aqueous chloroaurate ions by actinomycetes Thermomonospora sp. (Ahmad et al. 2003). Different fungal species, e.g., Aspergillus clavatus, Epicoccum nigrum, and Trichoderma sp. have been reported to synthesize AuNPs intracellularly or extracellularly (Qu et al. 2016; Sheikhloo et al. 2011; Verma et al. 2011). Yeasts like Pichia jadinii and Yarrowia lipolytica were also shown to have a good potential for AuNPs synthesis (Gericke and Pinches 2006; Pimprikar et al. 2009). In general, different microbes exhibit diverse ability to synthesize AuNPs, which is caused by varied oxidation-reduction enzymes and metabolites presented. As we know, fungi are able to secrete more proteins, produce more biomass, and show higher metal tolerance and bioaccumulation ability (Moghaddam et al. 2015), which may lead to possess superior ability to reduce soluble gold ions and synthesize insoluble nanocrystals (Kitching et al. 2015). However, the present researches were mostly focused on the characteristics of AuNPs synthesized by a single strain; there were few studies to comparing the characteristics of AuNPs biosynthesized by different microbes. Therefore, it is necessary to make some comparisons among different species of microbes to identify promising candidates for further study.
In this study, AuNPs were biosynthesized using different microbial cell-free extracts, which were obtained from three locally isolated strains of bacterium Labrys sp. WJW, yeast Trichosporon montevideense WIN, and filamentous fungus Aspergillus sp. WL-Au. The characterizations of AuNPs were performed by UV-Vis spectroscopy and transmission electron microscopy (TEM) analyses. The interactions of biomolecules present in the extracts with surfaces of synthesized AuNPs were confirmed by Fourier transform infrared spectroscopy (FTIR). The catalytic activities of AuNPs were determined by reduction of 4-NP in the presence of excess NaBH4. In this study, the characteristics of AuNPs synthesized by different species of microbes were comparatively studied in detail. It will provide some valuable reference for exploring efficient microbial resources to biosynthesize AuNPs.
Materials and methods
Chemicals, media, and strains
Hydrogen tetrachloroaurate (III) hydrate (HAuCl4·3H2O) was purchased from J&K Scientific Ltd. (China). 4-NP and NaBH4 were obtained from Sinopharm Chemical Regent Beijing Co., Ltd. (China). All other reagents were of analytical grade.
Filamentous fungus Aspergillus WL-Au and yeast Trichosporon montevideense WIN were isolated from antibiotic-contaminated reactor in our laboratory, and bacteria Labrys WJW was isolated from graphene oxide degradation reactor in our laboratory. Strain WL-Au was aerobically cultured at 30 °C for 3 days in the modified martin broth (MMB, pH 7), containing 1 g/L (NH4)2SO4, 1 g/L KH2PO4, 0.5 g/L MgSO4·7H2O, and 2 g/L glucose, which was autoclaved at 115 °C for 15 min before use. Strains WJW and WIN were aerobically cultured at 30 °C for 2 days in lysogeny broth medium (LB, pH 7), containing 10 g/L NaCl, 5 g/L yeast extract and 10 g/L peptone, which was autoclaved at 121 °C for 20 min before use.
Biosynthesis of AuNPs by different microbial cell-free extracts
The microbial cell-free extracts used for AuNPs synthesis were prepared as follows: cells of strains WJW and WIN were firstly harvested after being cultivated for 2 days by centrifugation (10,000×g for 10 min at 4 °C), and cells of strain WL-Au was harvested by sieving through a qualitative filter paper. Then, the cells were washed with sterile deionized water to remove medium components before use. Subsequently, cells were re-suspended in phosphate sodium buffer (50 mM, pH 7) and lysed by ultrasonication (Ultrasonic Processor CPX 750, USA) for 40 min. After centrifugation of the broken cracking liquid samples at 10,000×g for 20 min (4 °C), the supernatants were then treated by filtering through 0.45-μm syringe Millipore filters and the filtrate samples were collected as microbial cell-free extracts. The protein concentrations of microbial cell-free extracts were regulated to 150 mg/L by the Bradford assays before use. The HAuCl4 solutions were added to the microbial cell-free extracts with final concentrations of 1 mM for AuNPs synthesis, and then the reaction mixtures were incubated in dark at 30 °C.
Characterization of AuNPs synthesized by different microbial cell-free extracts
The synthesis of AuNPs were determined by UV-Vis spectrophotometer (Metash UV-9000, China) recording the spectra between 400 and 800 nm at the resolution of 1 nm. The concentrations of synthesized AuNPs were measured by inductively coupled plasma optical emission spectrometer (ICP-OES, Perkin-Elmer Optima 2000 DV, USA). The shapes and sizes of biogenic AuNPs were characterized by TEM (FEI Tecnai G220 S-Twin, USA) with an accelerating voltage of 120 kV. FTIR spectra were recorded by a Shimadzu IRPrestige-21 spectrophotometer (Japan) with the wavelength ranging from 500 to 4000 cm−1 to investigate the conjugation of proteins on the nanoparticles surfaces.
Catalytic activities of different biogenic AuNPs for 4-NP reduction
The catalytic activities of different biogenic AuNPs were measured by reducing 4-NP in the presence of excess NaBH4. In a typical assay, 0.1 mL of 2 mM stock solution of 4-NP and 0.4 mL of 30 mM freshly prepared NaBH4 solution were mixed in a 3-mL standard quartz cuvette. The biogenic AuNPs with final concentrations of 1 mg/L quantified by ICP-OES were added to initiate the reactions. The final reaction volumes were adjusted to 2 mL with sterile water. To monitor the reduction of 4-NP, samples were taken at intervals and recorded by UV-Vis spectra in the range of 300–500 nm. The catalytic reduction rates of 4-NP by different biogenic AuNPs were calculated through measuring the decrease of UV-Vis absorbance at 401 nm. All the reactions were performed in triplicate at ambient temperature.
Results and discussion
UV-Vis spectra of AuNPs synthesized by different microbial cell-free extracts
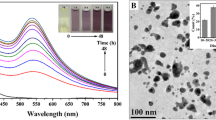
After incubation of the microbial cell-free extracts with HAuCl4 for 7 days, the formation of AuNPs was preliminary visually observed by colors, which changed from pale yellow to ruby red and purple as shown in Fig. 1 insets. Meanwhile, the UV-Vis spectra of different reaction solutions were recorded in the range of 400–800 nm, which could act as a reliable method to confirm the presence of AuNPs based on their optical transition and electronic band structure (Arumugam and Berchmans 2011). As shown in Fig. 1, the UV-Vis spectra of different reaction solutions respectively showed characteristic absorption peaks at 538 nm (WJW), 539 nm (WIN), and 543 nm (WL-Au) corresponding to the surface plasmon resonance of AuNPs. It is indicated that the AuNPs were biosynthesized by the cell-free extracts from these microbes (Das et al. 2012), named as WJW-AuNPs, WIN-AuNPs, and WL-Au-AuNPs, respectively. In addition, the stability of these biogenic AuNPs was also investigated. It was found that WJW-AuNPs, WIN-AuNPs, and WL-Au-AuNPs were stable even after being stored for 3 months at 4 °C. The UV-Vis spectra showed no evident changes in the wavelength and intensity of SPR bands, which indicated that the as-synthesized AuNPs were stable and well dispersed in solution without obvious aggregation. This suggested that the interactions between as-synthesized AuNPs and biomolecules in the microbial cell-free extracts take important roles in preventing aggregation and stabilization of the formed AuNPs (Das et al. 2012).
Shapes and particle sizes of AuNPs synthesized by different microbial cell-free extracts
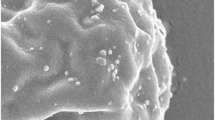
The morphologies of AuNPs formed by different microbial cell-free extracts were investigated through TEM analysis. As shown in Fig. 2, the major shapes of these biogenic AuNPs were common sphere or pseudo-sphere. However, the WL-Au-AuNPs were relatively uniform and nearly monodisperse. The selected area electron diffraction (SAED) patterns (Fig. 2 insets) of all the three biogenic AuNPs indicated that the biosynthesized AuNPs were highly crystalline in nature (Emmanuel et al. 2014). In addition, the size distributions of different AuNPs were calculated by analytical procedure. As shown in Fig. 3, the size distributions of WJW-AuNPs, WIN-AuNPs, and WL-Au-AuNPs were 4.3–52.1, 8.4–38.5, and 3.2–20.6 nm, respectively, which demonstrated the relatively uniform size of WL-Au-AuNPs compared with others. Meanwhile, the average sizes were calculated as 18.8 nm (WJW-AuNPs), 22.2 nm (WIN-AuNPs), and 9.5 nm (WL-Au-AuNPs), respectively. It was generally assumed that there were two main processes in the AuNP biosynthesis by microbes: (i) Au3+ reduction to Au0 and (ii) Au0 stabilized by biomolecules and formed AuNPs. In the process of Au3+ reduction, Au3+ were reduced by proteins, reducing sugars, and other metabolites, and therefore synthesized Au0 nucleus extracellularly or intracellularly. The Au0 nucleus would continuously grow and finally become aggregations if without addition of capping and stabilizing agents. In the process of AuNP biosynthesis by microbes, biomacromolecules were speculated to act as capping and stabilizing agents, and they can stabilize the growing Au0 nucleus and form AuNPs. The morphology of synthesized AuNPs was closely associated with the types of biomacromolecules (Kitching et al. 2015; Dhillon et al. 2015; Das et al. 2010). These results suggested that the filamentous fungus had a superior potential for synthesis of uniform and monodisperse AuNPs, which should provide insights into the green synthesis of metal nanoparticles.
FTIR analyses of different biogenic AuNPs
The capping effects of biomolecules in different microbial cell-free extracts for AuNP synthesis were investigated by FTIR analysis (Fig. 4). The bands at 3442 cm−1 (WJW-AuNPs), 3329 cm−1 (WIN-AuNPs), and 3300 cm−1 (WL-Au-AuNPs) were due to –N–H stretching vibrations arising from the peptide linkages (Mukherjee et al. 2008). The bands at 2798 cm−1, 2924 cm−1 (WL-Au-AuNPs), and 2920 cm−1 (WIN-AuNPs) could have resulted from stretching vibrations of –C–H groups of aliphatic acids (Kora et al. 2012). The bands at 1635~1653, 1500~1540, and 1026~1041 cm−1 were attributed to the stretching vibrations of –C=C, –N=O, and –S=O groups, respectively (Ashour et al. 2015). The additional band at 1751 cm−1 in the WL-Au-AuNPs spectrum indicated the presence of –C=O group (Gajbhiye et al. 2009). This suggested that the microbial-synthesized AuNPs had a strong binding ability with many functional biomolecules, such as proteins, esters, and carboxylic acids, which could serve as capping and stabilizing agents, thereby offering stability to particles by preventing agglomeration and growth of nanocrystals (Au0) (Ashour et al. 2015). Researchers have made many efforts to investigate the chemicals involved in the AuNP stabilization. FTIR analysis is one of the most commonly used methods, which could detect the functional groups bonded to the synthesized AuNP surfaces. For example, Faramarzi and Forootanfar found that –OH/–NH functional groups and carbonyl group were used in bioconjugation and immobilization of the AuNPs synthesized by laccase from Paraconiothyrium variabile (Faramarzi and Forootanfar 2011). In addition, Binupriya et al. detected the amide I and II of proteins on the synthesized AuNP surfaces using FTIR analysis. They pointed out that proteins could bind to AuNPs through either free amine groups or crysteine residues in the proteins, and therefore, stabilization of the AuNPs by surface-bound proteins is a possibility (Binupriya et al. 2010). In this study, many functional groups including –N–H, –C–H, –C=C, –N=O, –S=O, and –C=O groups were detected in the synthesized AuNP surfaces, indicating these groups possibly participated in the AuNP stabilization. In addition, it was also found that the surfaces of WL-Au-AuNPs attached more biological groups. Filamentous fungi had advantages over bacteria and yeasts for AuNP biosynthesis due to the large amounts of secreted proteins with diverse functions, which could lead to the formation of homogeneous nanoparticles with enhanced stability (Kitching et al. 2015).
Catalytic activities of different biogenic AuNPs
To evaluate the catalytic activities of different biogenic AuNPs, 4-NP reduction assays were performed by AuNPs with excess NaBH4, which was studied as a model reaction. Though the reduction of 4-NP to 4-AP by NaBH4 was thermodynamically favorable, the presence of the kinetic barrier due to large potential difference between donor and acceptor molecules reduced the feasibility. AuNPs could help transfer the electrons from BH4− ions to the nitro group of 4-NP, reducing it to 4-AP (Gangula et al. 2011). For all experiments, the initial concentrations of 4-NP, NaBH4, and AuNPs were kept at 0.1, 10, and 1 mg/L, respectively. Since the absorption peak of 4-NP shifted from 317 to 401 nm in the presence of NaBH4, the change in the absorption spectra at 401 nm was monitored to track the reaction process (Gangula et al. 2011). Figure 5a showed the UV-Vis spectra of 4-NP reduction by WL-Au-AuNPs, which was chosen as a representative to analyze the catalytic reduction of 4-NP by the biogenic AuNPs. The SPR absorption band at 401 nm decreased rapidly after addition of the biogenic AuNPs, corresponding to the disappearance of the p-nitrophenolate ion. Meanwhile, a new absorption peak at 310 nm appeared, indicating the formation of 4-AP (Panigrahi et al. 2007). The reduction reactions were also repeated by WJW-AuNPs and WIN-AuNPs. Figure 5b showed the catalytic reduction rates of 4-NP by different biogenic AuNPs. The reactions could be completed within 16, 16, and 10 min, respectively when using WJW-AuNPs, WIN-AuNPs, and WL-Au-AuNPs as the catalysts. However, in the control experiment without addition of AuNP catalyst, the absorbance at 401 nm decreased slightly even after 2 days.
Since the concentrations of NaBH4 were much higher than those of 4-NP, the catalytic reductions should follow pseudo-first-order kinetics (Das et al. 2012). The apparent rate constants (k app ) of different biogenic AuNPs were calculated as shown in Fig. 5c. The k app values of WJW-AuNPs, WIN-AuNPs, and WL-Au-AuNPs were 0.27, 0.23, and 0.37 min−1, respectively, which were comparable to those of reported biogenetic AuNPs. For example, the biogenic AuNPs synthesized by Cylindrocladium floridanum and stem extract of Breynia rhamnoides could function as efficient catalysts for 4-NP reduction with reaction rate constants of 0.03 min−1 (Narayanan and Sakthivel 2011) and 0.55 min−1 (Gangula et al. 2011), respectively. The AuNPs synthesized by Prunus domestica fruit extract had reaction rate constants of 0.11 to 0.31 min−1 for 4-NP reduction when diverse volume of AuNPs were used (Dauthal and Mukhopadhyay 2012). The WL-Au-AuNPs showed the highest catalytic rate of 0.37 min−1 with a smallest average size of 9.5 nm, which may be due to the higher ratio of surface area to volume. The result was also determined by Das et al. (2012), who found that the surface area to volume ratio of AuNPs could affect their catalytic activities. This research demonstrated that the filamentous fungi showed superior ability to produce well-dispersed and uniform AuNPs with high catalytic activity (Dhillon et al. 2012; Kitching et al. 2015). It might provide a well-founded pathway for exploring efficient microbes to green synthesize AuNPs with high catalytic performances.
Conclusions
In this study, three different microbes, containing one bacterium, one yeast, and one filamentous fungus were employed to green synthesize AuNPs. The formation of AuNPs were determined by color change and UV-Vis spectra of reaction solutions. TEM and SAED analyses showed that crystalline AuNPs of similar sphere or pseudo-sphere were formed by different microbial cell-free extracts but the sizes of different AuNPs were diverse (WL-Au-AuNPs 9.5 nm, WJW-AuNPs 18.8 nm, and WIN-AuNPs 22.2 nm). FTIR analyses demonstrated that the biomolecules were involved in stabilizing the synthesized AuNPs. The as-synthesized AuNPs exhibited high catalytic activities for 4-NP reduction with catalytic rates of 0.37 min−1 (WL-Au-AuNPs), 0.27 min−1 (WJW-AuNPs), and 0.23 min−1 (WIN-AuNPs). In conclusion, filamentous fungus WL-Au-mediated AuNPs were more uniform and better dispersed and exhibited superior catalytic activity than other biogenic AuNPs.
References
Ahmad A, Senapati S, Khan MI, Kumar R, Sastry M (2003) Extracellular biosynthesis of monodisperse gold nanoparticles by a novel extremophilic actinomycete, Thermomonospora sp. Langmuir 19(8):3550–3553. https://doi.org/10.1021/la026772l
Ahmad T, Wani IA, Manzoor N, Ahmed J, Asiri AM (2013) Biosynthesis, structural characterization and antimicrobial activity of gold and silver nanoparticles. Colloids Surf B Biointerfaces 107:227–234. https://doi.org/10.1016/j.colsurfb.2013.02.004
Arumugam P, Berchmans S (2011) Synthesis of gold nanoparticles: an ecofriendly approach using Hansenula anomala. ACS Appl Mater Interfaces 3:1418–1425
Arvizo RR, Bhattacharyya S, Kudgus RA, Giri K, Bhattacharya R, Mukherjee P (2012) Intrinsic therapeutic applications of noble metal nanoparticles: past, present and future. Chem Soc Rev 41(7):2943–2970. https://doi.org/10.1039/c2cs15355f
Ashour AA, Raafat D, El-Gowelli HM, El-Kamel AH (2015) Green synthesis of silver nanoparticles using cranberry powder aqueous extract: characterization and antimicrobial properties. Int J Nanomedicine 10:7207–7221. https://doi.org/10.2147/IJN.S87268
Binupriya AR, Sathishkumar M, Vijayaraghavan K, Yun SI (2010) Bioreduction of trivalent aurum to nano-crystalline gold particles by active and inactive cells and cell-free extract of Aspergillus oryzae var. viridis. J Hazard Mater 177(1-3):539–545. https://doi.org/10.1016/j.jhazmat.2009.12.066
Brayner R, Barberousse H, Hemadi M, Djedjat C, Yéprémian C, Coradin T, Livage J, Fiévet F, Couté A (2007) Cyanobacteria as bioreactors for the synthesis of Au, Ag, Pd, and Pt nanoparticles via an enzyme-mediated route. J Nanosci Nanotechnol 7(8):2696–2708. https://doi.org/10.1166/jnn.2007.600
Chakraborty N, Banerjee A, Lahiri S, Panda A, Ghosh AN, Pal R (2009) Biorecovery of gold using cyanobacteria and a eukaryotic alga with special reference to nanogold formation-a novel phenomenon. J Appl Phycol 21(1):145–152. https://doi.org/10.1007/s10811-008-9343-3
Correa-Llantén DN, Muñoz-Ibacache SA, Castro ME, Muñoz PA, Blamey JM (2013) Gold nanoparticles synthesized by Geobacillus sp. strain ID17 a thermophilic bacterium isolated from Deception Island, Antarctica. Microb Cell Factories 12:1
Das SK, Das AR, Guha AK (2010) Microbial synthesis of multishaped gold nanostructures. Small 6(9):1012–1021. https://doi.org/10.1002/smll.200902011
Das SK, Dickinson C, Lafir F, Brougham DF, Marsili E (2012) Synthesis, characterization and catalytic activity of gold nanoparticles biosynthesized with Rhizopus oryzae protein extract. Green Chem 14(5):1322–1334. https://doi.org/10.1039/c2gc16676c
Dauthal P, Mukhopadhyay M (2012) Prunus domestica fruit extract-mediated synthesis of gold nanoparticles and its catalytic activity for 4-nitrophenol reduction. Ind Eng Chem Res 51(40):13014–13020. https://doi.org/10.1021/ie300369g
Dhillon GS, Brar SK, Kaur S, Verma M (2012) Green approach for nanoparticle biosynthesis by fungi: current trends and applications. Crit Rev Biotechnol 32(1):49–73. https://doi.org/10.3109/07388551.2010.550568
Dhillon M, Ramani M, Marsili E (2015) Fungal biosynthesis of gold nanoparticles: mechanism and scale up. Microb Biotechnol 8:904–917
Emmanuel R, Karuppiah C, Chen SM, Palanisamy S, Padmavathy S, Prakash P (2014) Green synthesis of gold nanoparticles for trace level detection of a hazardous pollutant (nitrobenzene) causing Methemoglobinaemia. J Hazard Mater 279:117–124. https://doi.org/10.1016/j.jhazmat.2014.06.066
Faramarzi MA, Forootanfar H (2011) Biosynthesis and characterization of gold nanoparticles produced by laccase from Paraconiothyrium variabile. Colloids Surf B Biointerfaces 87(1):23–27. https://doi.org/10.1016/j.colsurfb.2011.04.022
Gajbhiye M, Kesharwani J, Ingle A, Gade A, Rai M (2009) Fungus-mediated synthesis of silver nanoparticles and their activity against pathogenic fungi in combination with fluconazole. Nanomedicine 5(4):382–386. https://doi.org/10.1016/j.nano.2009.06.005
Gangula A, Podila R, Ramakrishna M, Karanam L, Janardhana C, Rao AM (2011) Catalytic reduction of 4-nitrophenol using biogenic gold and silver nanoparticles derived from Breynia rhamnoides. Langmuir 27(24):15268–15274. https://doi.org/10.1021/la2034559
Gericke M, Pinches A (2006) Biological synthesis of metal nanoparticles. Hydrometallurgy 83(1-4):132–140. https://doi.org/10.1016/j.hydromet.2006.03.019
Guo S, Wang E (2007) Synthesis and electrochemical applications of gold nanoparticles. Anal Chim Acta 598(2):181–192. https://doi.org/10.1016/j.aca.2007.07.054
He R, Wang YC, Wang X, Wang Z, Liu G, Zhou W, Wen L, Li Q, Wang X, Chen X, Zeng J, Hou JG (2014) Facile synthesis of pentacle gold–copper alloy nanocrystals and their plasmonic and catalytic properties. Nat Commun 5:4327
Hulkoti NI, Taranath TC (2014) Biosynthesis of nanoparticles using microbes—a review. Colloids Surf B Biointerfaces 121:474–483. https://doi.org/10.1016/j.colsurfb.2014.05.027
Kharissova OV, Dias HVR, Kharisov BI, Pérez BO, Jiménez Pérez VM (2013) The greener synthesis of nanoparticles. Trends Biotechnol 31(4):240–248. https://doi.org/10.1016/j.tibtech.2013.01.003
Kitching M, Ramani M, Marsili E (2015) Fungal biosynthesis of gold nanoparticles: mechanism and scale up. Microb Biotechnol 8(6):904–917
Kora AJ, Sashidhar RB, Arunachalam J (2012) Aqueous extract of gum olibanum (Boswellia serrata): a reductant and stabilizer for the biosynthesis of antibacterial silver nanoparticles. Process Biochem 47(10):1516–1520. https://doi.org/10.1016/j.procbio.2012.06.004
Moghaddam AB, Namvar F, Moniri M, Tahir PM, Azizi S, Mohamad R (2015) Nanoparticles biosynthesized by fungi and yeast: a review of their preparation, properties, and medical applications. Molecules 20(9):16540–16565. https://doi.org/10.3390/molecules200916540
Mukherjee P, Roy M, Mandal BP, Dey GK, Mukherjee PK, Ghatak J, Tyagi AK, Kale SP (2008) Green synthesis of highly stabilized nanocrystalline silver particles by a non-pathogenic and agriculturally important fungus T. asperellum. Nanotechnology 19(7):075103. https://doi.org/10.1088/0957-4484/19/7/075103
Narayanan KB, Sakthivel N (2011) Synthesis and characterization of nano-gold composite using Cylindrocladium floridanum and its heterogeneous catalysis in the degradation of 4-nitrophenol. J Hazard Mater 189(1-2):519–525. https://doi.org/10.1016/j.jhazmat.2011.02.069
Panigrahi S, Basu S, Praharaj S, Pande S, Jana S, Pal A, Ghosh SK, Pal T (2007) Synthesis and size-selective catalysis by supported gold nanoparticles: study on heterogeneous and homogeneous catalytic process. J Phys Chem C 111(12):4596–4605. https://doi.org/10.1021/jp067554u
Park TJ, Lee KG, Lee SY (2016) Advances in microbial biosynthesis of metal nanoparticles. Appl Microbiol Biotechnol 100(2):521–534. https://doi.org/10.1007/s00253-015-6904-7
Pimprikar PS, Joshi SS, Kumar AR, Zinjarde SS, Kulkarni SK (2009) Influence of biomass and gold salt concentration on nanoparticle synthesis by the tropical marine yeast Yarrowia lipolytica CIM 3589. Colloids Surf B Biointerfaces 74(1):309–316. https://doi.org/10.1016/j.colsurfb.2009.07.040
Qu YY, Shen WL, Pei XF, Ma F, You SN, Li SZ, Wang JW, Zhou JT (2016) Biosynthesis of gold nanoparticles by Trichoderma sp. WL-Go for azo dyes decolorization. J Environ Sci 56:79–86
Scari G, Porta F, Fascio U, Avvakumova S, Santo VD, Simone MD, Saviano M, Leone M, Gatto AD, Pedone C, Zaccaro L (2012) Gold nanoparticles capped by a GC-containing peptide functionalized with an RGD motif for integrin targeting. Bioconjug Chem 23(3):340–349. https://doi.org/10.1021/bc200143d
Seo JM, Kim EB, Hyun MS, Kim BB, Park TJ (2015) Self-assembly of biogenic gold nanoparticles and their use to enhance drug delivery into cells. Colloids Surf B Biointerfaces 135:27–34. https://doi.org/10.1016/j.colsurfb.2015.07.022
Sheikhloo Z, Salouti M, Katiraee F (2011) Biological synthesis of gold nanoparticles by fungus Epicoccum nigrum. J Clust Sci 22(4):661–665. https://doi.org/10.1007/s10876-011-0412-4
Singh P, Kim YJ, Zhang D, Yang DC (2016a) Biological synthesis of nanoparticles from plants and microorganisms. Trends Biotechnol 34(7):588–599. https://doi.org/10.1016/j.tibtech.2016.02.006
Singh P, Singh H, Kim YJ, Mathiyalagan R, Wang C, Yang DC (2016b) Extracellular synthesis of silver and gold nanoparticles by Sporosarcina koreensis DC4 and their biological applications. Enzym Microb Technol 86:75–83. https://doi.org/10.1016/j.enzmictec.2016.02.005
Sugunan A, Melin P, Schnürer J, Hilborn JG, Dutta J (2007) Nutrition-driven assembly of colloidal nanoparticles: growing fungi assemble gold nanoparticles as microwires. Adv Mater 19(1):77–81. https://doi.org/10.1002/adma.200600911
Verma VC, Singh SK, Solanki R, Prakash S (2011) Biofabrication of anisotropic gold nanotriangles using extract of endophytic Aspergillus clavatus as a dual functional reductant and stabilizer. Nanoscale Res Lett 6:16–22
Acknowledgements
This work was supported by the Program for New Century Excellent Talents in University (No. NCET-13-0077), the Fundamental Research Funds for the Central Universities (No. DUT14YQ107), and the Open Project of State Key Laboratory of Urban Water Resource and Environment, Harbin Institute of Technology (No. ESK201529).
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible editor: Santiago V. Luis
Rights and permissions
About this article
Cite this article
Shen, W., Qu, Y., Li, X. et al. Comparison of gold nanoparticles biosynthesized by cell-free extracts of Labrys, Trichosporon montevideense, and Aspergillus. Environ Sci Pollut Res 25, 13626–13632 (2018). https://doi.org/10.1007/s11356-017-1050-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-017-1050-7