Abstract
In this work, the behavior in tomato rhizosphere of Bacillus velezensis FZB42 was analyzed taking into account the surfactin production, the use of tomato roots exudate as substrates, and the biofilm formation. B. velezensis FZB42 and B. amyloliquefaciens S499 have a similar capability to colonize tomato rhizosphere. Little difference in this colonization was observed with surfactin non producing B. velezensis FZB42 mutant strains. B. velezensis is able to grow in the presence of root exudate and used preferentially sucrose, maltose, glutamic, and malic acids as carbon sources. A mutant enable to produce exopolysaccharide (EPS-) was constructed to demonstrate the main importance of biofilm formation on rhizosphere colonization. This mutant had completely lost its ability to form biofilm whatever the substrate present in the culture medium and was unable to efficiently colonize tomato rhizosphere.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Plant growth-promoting rhizobacteria (PGPR) provide beneficial effects to host plants and they contribute to increase yields of crops (Saharan and Nehra 2011). Direct mechanisms involved in the beneficial effects of these PGPR are biofertilization (increase of nutrient supply), stimulation of root development (production of phytohormones), and improvement of abiotic stress tolerance (Saharan and Nehra 2011; Glick 2012). PGPR can also indirectly favor plant growth and health by reducing the impact of diseases caused by phytopathogens via three main mechanisms that are competition for space and nutrients, antagonism toward infectious microbes, and elicitation of plant defense reactions (a phenomenon called “induced systemic resistance” (ISR)) (Van Loon and Bakker 2005; Choudhary and Johri 2009; Lugtenberg and Kamilova 2009; Kloepper et al. 2004). It is well established that to provide their beneficial effects, PGPR have to reach minimal population densities in the rhizosphere (Lugtenberg and Kamilova 2009; Das and Dkhar 2011). Thus, an efficient colonization of rhizosphere is a key step for providing both growth-promoting effect and disease control activity. The main hypothesis currently mentioned in literature to explain colonization efficiency is the strain abilities (i) to move toward the place to colonize, (ii) to use carbon and nitrogen sources (Bertin et al. 2003) provided by root exudates, (iii) to withstand plant response reaction (Budiharjo et al. 2014), and (iv) to form a biofilm at the root surface.
Bacillus amyloliquefaciens and Bacillus subtilis are well recognized PGPR. B. amyloliquefaciens strains are able to produce auxin (Chen et al. 2007; Idris et al. 2004) or can contribute to plant growth promotion under conditions of phosphate limitation by excreting phytase in phytate presence (Idris et al. 2002; Makarewicz et al. 2006).
In addition, some Bacillus amyloliquefaciens strains such as S499 or FZB42, now Bacillus velezensis FZB42 (Fan et al. 2017), have an impressive capacity to produce secondary metabolites having antimicrobial activities (Scholz et al. 2014; Molinatto et al. 2016). Besides B. velezensis, B. subtilis strains have been reported to synergistically increase plant nitrogen phosphate accumulation when co-inoculated with mycorrhiza Glomus intraradices (Kohler et al. 2007). Bacillus sp. strains are also able to synthesize a set of different secondary metabolites. Among the metabolites produced by both species B. subtilis and B. velezensis, cyclic lipopeptides (CLPs) belonging to the surfactin, iturin, and fengycin families have been well studied (Ongena and Jacques 2008). In vivo, fengycins and iturins display antifungal activities and inhibit the growth of several plant pathogens. Surfactins are poorly antifungal but may have some synergistic effects on biological activity of iturins (Deravel et al. 2014) and fengycins (Maget-Dana et al. 1992; Ongena and Jacques 2008). Surfactins, fengycins, and iturins are able to stimulate Induced Systemic Resistance (ISR), by playing a role of elicitor, in some plant species such as bean, tomato, melon, and grapewine (Ongena et al. 2007; Garcia-Gutierrez et al. 2013; Farace et al. 2015). Interestingly, the expression of the genes involved in the biosynthesis of some of these compounds is increased in the presence of root exudates (Fan et al. 2012).
Due to their surfactant and tension surface-lowering activities, it has been suggested that these lipopeptides, especially surfactin, may contribute to the root colonization process by the producing strains. This hypothesis is mainly supported by in vitro data (Leclere et al. 2006; Ongena and Jacques 2008), but also by studies demonstrating such a role in planta (Bais et al. 2004; Fan et al. 2011; Dietel et al. 2013). Using confocal laser microscopy and GFP-labeled strains, Fan et al. (2011 and 2012) have also demonstrated the ability of FZB42 strains to colonize roots of Lemna minor, Arabidopsis thaliana, and Zea mays.
In this work, we confirmed the role of surfactin in tomato root colonization, we characterized the influence of tomato root exudates on Bacillus velezensis growth and surfactin production, and we highlighted, for the first time, the main role played by the biofilm formation in the B. velezensis FZB42 colonization process. This last result was obtained by comparing the behavior of an exopolysaccharide non-producer mutant strain with the wild-type.
Materials and methods
Microorganisms and plants
Tomato seeds Solanum lycopersicum (Merveille des Marchés cultivar) and four bacterial strains (Table 1) were used in this study: B. amyloliquefaciens S499, B. velezensis FZB42, and two FZB42 derivatives: AK3 and CH1 (kindly provided by Dr. Rainer from Humboldt University, Berlin, Germany). The Bacillus strains were long-term preserved at − 80 °C in glycerol (40%) and routinely grown at 37 °C on Luria-Bertani (LB) broth.
Invasive growth assays
Nine centimeter diameter Petri dishes containing 25 mL of LB medium with 0.7% bacto agar (Julkowska et al. 2004) were prepared 90 min before inoculation and dried during 15 min lid open in a laminar flow hood. The center of the LB medium plate was inoculated with a drop of 3 μL of diluted culture in LB medium (OD600 = 0.1). The plates were incubated at 30 °C and the colonization was evaluated after 3 days. Each experiment was repeated at least three times.
Preparing tomato seeds for germination
Tomato seeds were surface sterilized with ethanol 75% during 2 min and in sodium hypochlorite 4.5% for 15 min and rinsed with sterile water. Then, they were put in Petri dish containing filter paper wetted with Hoagland solution and then left for germination during 4 days at 21 °C. After germination, they were used for the experiments of rhizosphere colonization, kinetic of bacterial growths, and surfactin production. They also were used for hydroponic experiments to collect root exudates.
Bacterial colonization of tomato rhizosphere
Bacterial strains were grown at 37 °C in LB medium and the bacterial cells were prepared for inocula by diluting them in the solution of 0.01 M of MgSO4 until a 1 × 105 CFU mL−1 concentration. Surface-sterilized and pregerminated tomato seeds were soaked for 10 min in such a diluted bacterial cell suspension and placed into a sterilized glass tube containing 2 g of perlite and 9 mL of Hoagland solution (final volume = 14 cm3). Tomato plantlets were grown at 21 °C in a culture room with a 16:8 (light/dark) hours of photoperiod. After 21 days of cultivation, three tubes were randomly chosen, aerial parts were removed, and 10 mL of trypton salt was added to each tube. These tubes were vortexed at 2500 rpm for 5 min and series of dilutions were released for bacterial plate count on LB agar. Results are expressed as total CFU per cubic centimeter of perlite.
Kinetic of rhizosphere colonization and surfactin production
B. velezensis FZB42 strain was used for kinetic study. Tomato seeds were prepared and grown as described earlier. Every 3 days, two samples of two treatments (without inoculum and inoculated with B. velezensis) were randomly taken. One sample was used for plate count and one for surfactin quantification. For plate count, aerial parts were removed and 10 mL of trypton salt was added to each tube, these tubes were vortexed for 5 min and series of dilutions in trypton salt were released for bacterial plate count on LB agar. Results are expressed as total CFU per cubic centimeter of perlite. The sample for surfactin extraction was also randomly selected and the aerial parts were removed. Nine milliliters of acetonitrile/formic acid 0.1% (V/V) and 2 g of glass beads were added to each tube. These tubes were first vortexed for 5 min and then incubated overnight at 30 °C under agitation (140 rpm). The tubes were centrifuged at 10,000 rpm during 10 min. Surfactin was recovered by loading the supernatant on Solid-Phase Extraction Cartridges C18 (Alltech Maxi-Clean). The cartridges were washed with water and the surfactin was eluted with a solution of acetonitrile. The solutions were vacuum dried (Speed Vac Plus, SC 110A, Savant, GMI, Ramsey, USA). Dried residues were suspended in 200 μL of acetonitrile/water/formic acid 80/20/0.1 (V/V/V) and analyzed by HPLC (Online Degaser, 717 Autosample, 660S Controller, 626 Pumps, 2996 PhotoDiode Array; Waters Corporation, Milford, MA, USA). The column used was a C-18 (5 μm, 250 × 3 mm, VYDAC 218 TP53; Grace-Davison, Deerfield, Illinois, USA). The liquid phase was a gradient of acetonitrile (0.1% trifluoroacetic acid) in double distilled water (0.1% trifluoroacetic acid) (Table 2), the volume of injection was 20 μL and the flow rate was 0.6 mL min−1.
Samples preparing for microscopic root observation
The microscopic observation of root was carried out on tomato plant roots after 21 days of cultivation. The dye used was acridine orange 0.01% (w/v) prepared in 0.1 M acetate buffer, pH 4 (36 mL of 0.1 M sodium acetate mixed with 164 mL of 0.1 M acetic acid). Several plants were randomly chosen, the aerial parts were removed, and the roots were submerged in acridine orange for 5 min. Samples were protected from light during this treatment. The roots were then fixed between slide and cover slip. The observation was performed under a fluorescence microscope, Nikon EFD-3, using oily lens with 100X magnification. The images were obtained by using a Nikon DS-1 Fi camera connected to a computer.
Root exudates collection
After 4 days of germination, sterilized seeds were put in sterile tubes containing Hoagland solution. The germinated seeds were left for growth at condition of 8:16 (dark/light) hours of photoperiod and at room temperature (21 °C). After 21 days, the root exudates were collected by recovering all the solution from the hydroponic experiment. Solution was sterilized by passing through a filter (0.22 μm) and then 50 times vacuum-concentrated and stored at − 20 °C.
Kinetic study for bacterial growth in root exudates and different carbon sources
A kinetic study was performed to elucidate the demeanor of bacterial growth during 72 h in root exudates and in different carbon sources. BioLector system was used as a simple and efficient high-throughput screening tool to follow microbial kinetics. By this tool, 48 samples can be studied in the same time under different conditions. The conditions of cultures, temperature, and agitation are controlled and pH, biomass, oxygen, and GFP are continuously measured by sensors supplied in BioLector system. One thousand two hundred microliters of root exudates and each carbon source in minimum medium were loaded in each well of BioLector microplates at 21 °C and 160 rpm for 72 h. Growth kinetics were followed by online measurement of optical density. The results were expressed as optical density (600 nm).
B. velezensis FZB42 growth and surfactin production on different carbon sources
Ten milliliters of concentrated root exudates and a set of five milliliters solutions containing an equivalent of 2 g of carbon were prepared with one of the following carbon sources: glucose, sucrose, fructose, maltose, or xylose as sugars and glutamic, malic, succinic, fumaric, citric, or oxalic acids, dissolved in 1 L of minimal medium composed of (Na2HPO4·2H2O 33.7 mM, KH2PO4 22.0 mM, NaCl 8.55 mM, MgSO4·7H2O 1 mM, CaCl2·H2O 0.3 mM, thiamin-HCl 0.003 μM, biotin 0.004 μM, EDTA 0.17 mM, FeCl3·6H2O 0.03 mM ZnCl2 0.0062 mM, CuCl2·2H2O 0.76 μM, CoCl2·2H2O 0.42 μM, H3BO3 1.62 μM, and MnCl2-4H2O 0.08 μM. pH was adjusted to 7 and the (C/N) ratio was (8:1). All these media and root exudates were inoculated by 1 × 105 CFU of B. velezensis FZB42 and were incubated at 21 °C under agitation (160 rpm) for 72 h. After 72 h, the population of bacteria was determined by using bacterial plate count. The bacterial suspensions were taken and centrifuged at 10,000 rpm through 10 min. The supernatants were prepurified and analyzed by HPLC as described above.
Biofilm assay
To quantify B. velezensis FZB42 biofilm formation, the procedure described by Hsueh et al (2006) was used. The strains were grown in LB medium until mid-log phase, and the cells were collected by centrifugation at 10,000 rpm during 10 min and resuspended in minimum medium supplemented with the different carbon sources as described previously. The initial biomass of media containing different carbon sources and concentrated root exudates inoculated with B. velezensis FZB42 was 1 × 105 CFU mL−1. All cultures were incubated at 21 °C without shaking for 72 h. The contents of each well were then removed and the well was washed five times with PBS buffer and air-dried. Biofilm cells were stained with 1% crystal violet (CV) solution in 33% (v/v) acetic acid for 20 min. Excess CV was then removed with water for five times. The bound CV was solubilized in 200 μL of 33% acetic acid and the absorbance measured at 590 nm.
EPS− mutant construction
Escherichia coli JM109 and B. velezensis FZB42 were routinely cultured in LB liquid medium at 37 °C and 160 rpm or on LB agar plate at 37 °C. When appropriate, ampicillin (Ap; 100 μg mL−1 for E. coli) and erythromycin (Em; 20 μg mL−1 for B. velezensis FZB42) were added to the medium. The vectors used in this study were pGEM-T Easy and pMUTIN-GFP+. Firstly, an epsA amplicon was amplified using polymerase chain reaction procedure (denaturation temperature, 94 °C for 2 min, annealing temperature, 55 °C for 45 s, and elongation temperature, 72 °C for 2 min; during 35 cycles). The forward primer sequence was 5′GGTACCCTTTTCTTCTGCGG′3, whereas the reverse primer was 5′CGGCCGGCTTAAGAC′3. These primers were designed by both Primer3 (Version 4.0) and Amplifix programs. The PCR product was introduced in E. coli JM109 using pGEM-T Easy vector according to the instructions of the supplier (Promega Corp, Madison, WI, USA). The transformation mixture was spread onto LB medium containing the required antibiotic and the plates were incubated at 37 °C for 24 h. The pGEM-T Easy containing the epsA fragment was extracted using Gene Jet Plasmid Miniprep Kit (Thermo Scientific Fermentas, Vilnius, Lituania). A sufficient amount of epsA fragment was digested by selected restriction enzymes (Thermo Scientific Fermentas) and then transferred in E. coli JM109 after ligation within pMUTIN-GFP+ previously digested by the same enzymes. The resulting hybrid plasmid was transferred into B. velezensiss FZB42 using electroporation following two procedures (Zhang et al. 2011; Cao et al. 2011): an overnight LB culture of the FZB42 cells was diluted 100-fold in NCM fresh medium. When the optical density reached 0.5, the cell walls were weakened by adding 3.89% glycine and 1.06% DL-threonine. After 1 h of shaking, the cells were cooled on ice for 20 min and then collected by centrifugation at 4 °C and 8000×g for 5 min. Cells were washed four times with ice-cold ETM buffer (0.5 M sorbitol, 0.5 M mannitol, and 10% glycerol), containing KH2PO4, K2HPO4, and MgCl2 at 0.25, 0.25, and 0.5 mM, respectively (pH adjusted to 7.0). The electro-competent cells were resuspended in 1/100 volume of the original culture. One hundred microliter of this suspension was mixed with 100 ng of column-purified pMUTIN-GFP+ plasmid carrying the epsA amplicon. The mix was loaded into a prechilled 1 mm gap electroporation cuvette which was briefly incubated on ice and was shocked by a single 2.1 kV cm−1 pulse generated with resistance and capacitance set at 200 Ω and 25 μF, respectively. The cells were immediately diluted into 1 mL of recovery medium (growth medium containing 0.38 M mannitol and 0.5 sorbitol), following warming in a water bath at 46 °C for 6 min. Then, the cells were gently shaken for 3 h at 37 °C. Aliquots were spread onto LB medium agar plate supplemented with erythromycin (20 μg mL−1). After subculture in LB Em20, genomic DNAs were extracted from transformants using Wizard® Genomic DNA Purification Kit (Promega Corp.).
Interruption of epsA in B. velezensis FZB42 using fusion with GFP marker
A fragment from the eps operon was amplified by polymerase chain reaction (PCR) using the primers forward: 5′GGTACCCTTTTCTTCTGCGG′3 and reverse: 5′CGGCCGGCTTAAGAC′3 designed by both Primer3 (Version 4.0) and Amplifix programs, and chromosomal DNA from B. velezensis FZB42 as template. The PCR product was cloned in pGEM-T Easy and the ligation mixture was transformed into E. coli JM109 using a thermal shock procedure. Transformants were grown overnight in LB medium containing 100 μg mL−1 ampicillin. Then, the purified hybrid plasmid with the epsA-epsC fragment was extracted, purified, and cut using the restriction enzymes KpnI and XmaIII. The epsA-epsC amplicon was ligated to pMUTIN-GFP+ cut with the same enzymes. The ligation mixture served to transform E. coli JM109 as above, with a selection by resistance to 20 μg mL−1 erythromycin. After overnight growth of transformants in LB medium + Em20, the purified pMUTIN-GFP+::epsA-C was used to transform B. velezensis FZB42 using electroporation method with Em resistance selection. Transformants were grown overnight in LB medium and samples were analyzed by fluorescence microscopy. The results showed that the fusion gene eps-gfp was expressed in all cells as compared with the wild-type strain. To ensure that epsA was integrated within the corresponding chromosomal locus of FZB42, a fragment of 1972 bp was designed as above, with the primers forward: 5′ACTCATCTTCCGTGTCTCC′3 and reverse: 5′GTCTTGTAGTTCCCGTCATC′3. This fragment consisted of a part of slr, epsA, epsB and a part of gfp genes and was amplified using chromosomal DNA from both strain FZB42 and its Em-R fluorescent derivative. After agarose gel electrophoresis analysis, the 1972-bp amplicon was observed only in the Em-R transformant (data not shown).
Results and discussion
Rhizosphere colonization by different bacterial strains
B. amyloliquefaciens S499 and B. velezensis FZB42 and two derivatives (AK3 and CH1) were compared for their capacity to colonize tomato rhizosphere. The results obtained after 21 days of colonization are shown in Fig. 1. The results indicated similar colonization performance of the strains B. amyloliquefaciens S499, B. velezensis FZB42 and AK3 with respectively 3.8 × 107, 3.4 × 107, and 3.6 × 107 CFU cm−3 and a weaker result for the surfactin non-producing derivative CH1 with 2.2 × 107 CFU cm−3. This significant difference in colonization observed between the surfactin producers and the non-surfactin producer confirmed that surfactin production might promote the rhizosphere colonization. Nevertheless, this difference is weak (around 30% difference in biomass) indicating that other factors have to be considered.
Rhizosphere colonization by different strains of Bacillus spp.: B. velezensis S499, B. velezensis AK3, B. velezensis FZB42, and B. velezensis CH1. Tomato plants were planted in room culture at 21 °C, 16:8 h (light/dark) photoperiods during 21 days. Three replicates were used for bacterial count. S = strain producing surfactin
Kinetic of bacterial growth and surfactin production during rhizosphere colonization
The kinetic of bacterial growth and surfactin production in the tomato rhizosphere of B. velezensis FZB42 was followed during 21 days. The results are presented in Fig. 2.
The bacterial population of B. velezensis FZB42 in the rhizosphere continuously increased from an initial population of 1 × 105 to 2 × 108 CFU cm−3 at the end of experiment (21 days). The surfactin production followed the bacterial growth and final surfactin concentration was 6 μg cm−3. The surfactin production was harmonious with the biomass. Nevertheless, the specific production of surfactin was low compared to this obtained with a non-colonizing surfactin overproducing Bacillus strain (data not shown). This result confirmed that other parameters than surfactin are determinant for efficient root colonization.
Effect of root exudates on bacterial growth and surfactin production
In order to verify the role of root exudates on bacterial growth and surfactin production, root exudates from tomato roots were used as culture medium for B. velezensis FZB42.
B. velezensis FZB42 population reached value of 2.5 × 108 CFU mL−1 and surfactin production was 0.4 μg per 108 CFU. These findings are consistent with what has been obtained from the results of kinetic and this demonstrates the importance of root secretions to explain the rhizosphere colonization. Makarewicz et al. (2006) also found that the root exudates from tomato plant supported bacterial cell division and enhance the growth of B. amyloliquefaciens.
Root exudates include a diverse array of carbon sources like primary metabolites such as phenolic acids, organic acids, sugars and amino acids and secondary metabolites compounds (Badri and Vivanco 2009; Emmert and Handelsman 1999; Kohler et al. 2007). They provide the growth factors as well as nutrient sources for bacterial growth of B. velezensis FZB42.
The root exudates contain different organic compounds (Vancura and Hovadik 1965; Vancura and Hanzlikova 1972) and these compounds are indeed necessary to bacterial growth. Nevertheless, it is very complicated to get a good characterization of root exudates in terms of composition. Then, individual carbon sources (sugars and organic acids) were tested for bacterial growth and surfactin production.
Effect of different carbon sources on bacterial growth and surfactin production
As the root exudates contain many organic compounds that could affect growth and surfactin production, different sugars and organic acids were individually tested in this assay.
The effect of sugars and organic acids was tested using BioLector system. The results of growth are presented in Figs. 3 and 4. For sugars, the highest biomass was observed when maltose was used as a carbon source followed by sucrose, glucose, fructose, and xylose. These results indicated a non-negligible influence of the sugars used for bacterial growth of B. velezensis FZB42. Concerning organic acids, the highest biomass was obtained with glutamic acid, followed by malic, fumaric, succinic, citric, and oxalic acids. Oxalic acid had a negative role by inhibiting the bacterial growth; this observation was in agreement with the report of Rudrappa et al. (2007). The same observation occurred with Pseudomonas polymyxa SQR-21 (Ling et al. 2011).
The biomass was around two to three times higher with glutamic and malic acids than with citric and oxalic acids indicating also a great influence of the organic acids used as carbon sources for B. velezensis FZB42 growth. These results on different carbon sources generally present in root exudates indicate that root exudates composition has an influence on bacterial growth.
Sugars as carbon sources have also been tested for surfactin production by B. velezensis FZB42. The results are presented in Table 3. The production of surfactin depends on the sugar used by B. velezensis FZB42. Glucose and fructose are the best sugars and low production is observed with maltose and xylose.
When comparing the results of the specific production of surfactin, a difference between the different sugars was also clearly showed. For fructose and glucose, the values of specific production were almost the same but were reduced with sucrose, maltose, and xylose. The bigger contrast was observed with the maltose which allows the best biomass but with a weak production of surfactin. All these specific productions were higher than the one observed in the presence of the root exudates. However, surfactin had a very low critical micellar concentration (about 10 mg L−1) and its influence on the surface tension was still effective at low concentrations (Ongena and Jacques 2008).
Construction of a B. velezensis FZB42 EPS− mutant
Previous reports clearly indicated that mutants which were unable to synthesize exopolysaccharide were also unable to form biofilms, even though they may still form microcolonies and attach to the surfaces in limited scope (Allison and Sutherland 1987; Watnick and Kolter 1999; Sutherland 2001). As exopolysaccharides are known to be an important factor in biofilm formation, experiments were conducted for the purpose of interrupting a gene (eps) implied in exopolysaccharide synthesis in B. velezensis FZB42 strain.
Once this EPS− mutant obtained, preliminary experiments were performed to compare the behavior of the EPS− mutant to the wild-type FZB42. Growth kinetics of the two strains in LB medium showed no significant differences between these strains (same growth and same surfactin production), indicating that there was no effect of eps gene interruption on the bacterial growth and surfactin production of the EPS− mutant (data not shown).
In vitro biofilm comparative assays with B. velezensis FZB42, and its EPS− derivative
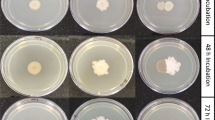
Thereafter, the two strains were grown in static cultures containing different carbon sources to investigate their ability to form a biofilm. The optical densities of biofilm stained with crystal violet were measured and the results are presented in Fig. 5.
A high difference was observed between these strains in forming a biofilm. With all tested carbon sources, the wild-type B. velezensis FZB42 was able to form a biofilm contrary to the EPS− mutant for which the biofilm formation was very weak. These results pointed out that the production of exopolysaccharides is necessary for biofilm formation by B. velezensis FZB42 and confirmed previous findings (Allison and Sutherland 1987; Watnick and Kolter 1999; Sutherland 2001).
Biofilm formation by B. velezensis FZB42 was also carbon sources dependent. The best sources being glucose, glutamic, succininc, and malic acids. Contrary to the results obtained on the effect of the substrates on the bacterial growth (Fig. 3), maltose and sucrose allowed a high biomass but not a high biofilm formation. Concerning the tested organic acids, a good correlation was observed between growth and biofilm formation.
Biofilm formation is an important process which represents the basis of root colonization and aggregate communities on soil particle surface by rhizobacteria (Davey et al. 2003; Tan et al. 2013). Biofilm formation was lower with concentrated root exudates than with other carbon sources, due to the facts that (i) the bacteria have a tendency to live in aggregate communities as a response to environmental stress and nutrient starvation (Donlan and Costerton 2002; Leclerc 2003; Swiecilo and Zych-Wezyk 2013) and (ii) the root exudates provide the essential elements for bacterial growth (Bertin et al. 2003; Vancura and Hanzlikova 1972; Vancura and Hovadik 1965). However, the low biofilm formation observed with the concentrated tomato root exudates compared to other carbon sources can be illustrated by the lack of both environmental harsh and nutrient deficiency and starvation in this concentrated tomato root exudates.
Microscopic observation and colonization assays
For more details, a microscopic observation was performed to compare the pattern colonization of both strains on the root of tomato plantlets using fluorescent microscopy (Fig. 6). Both strains colonized the rhizoplane of tomato following specific patterns.
These assays were realized depending on both the results of biofilm formation obtained under in vitro conditions and several reports which indicated the inability to form a biofilm in the absence of exopolysaccharide compounds (Allison and Sutherland 1987; Watnick and Kolter 1999; Sutherland 2001). Hence, the same strains were selected to inoculate germinated tomato seeds. They were left for growth in hydroponic system for 21 days. After this period, the aerial parts were removed and the roots were prepared for microscopic observation. B. velezensis FZB42 was treated with acridine orange, while the EPS− mutant carried the gfp marker. B. velezensis FZB42 was the best colonizer of roots as compared with its mutant (Fig. 6). A strong correlation was clearly demonstrated between the low biofilm formation and the weak colonization ability. Associated with the microscopic observation, samples of roots were used to count the number of bacteria that colonized these roots after 21 days. The results expressed as CFU per cubic centimeter were 17 × 107 for B. velezensis FZB42 and 0.60 × 107 for the EPS- mutant. Coupling the results obtained under in vitro conditions with the results of root colonization, the important role of biofilm in colonization was highlighted and we showed that biofilm formation plays a necessary role for roots and rhizosphere colonization.
As shown in previous reports, the rhizosphere colonization by plant growth-promoting bacteria is the most important step for the biocontrol agents (Weller et al. 2002; Vessey 2003; Pii et al. 2015). Our experiments shed light to the role of biofilm formation in rhizosphere colonization. Bacteria cells physically interact with plant by various means. These interactions commonly appear as the colonization of roots and/or rhizosphere. The bacteria adhere to the surface of plant tissues as individual and aggregated cells. The latter are defined as biofilms and they display various arrangements of dimensions, locations, and compositions (Nongkhlaw and Joshi 2014). The plant microenvironment has different characterizations such as saturation levels, nutrient availabilities, and surface chemistries, which strongly influence both the form and activity of biofilms (Ramey et al. 2004).
The amount of compounds within the biofilm depending on the carbon compounds availability and the equilibrium between carbon and other nutrients (Sutherland 2001; Fang et al. 2009), differences have been observed with different carbon sources. EPS- mutant presented a low biofilm formation and a low colonization of the root and the rhizosphere due to the fact that exopolysaccharide contribute directly to the properties of the biofilms and supply mechanical stability to the biofilms (Mayer et al. 1999; Flemming et al. 2007; Flemming and Wingender 2010).
Conclusion
Previous reports indicated that the rhizosphere colonization by plant growth-promoting bacteria is the most important step for the biocontrol agents (Weller et al. 2002; Vessey 2003; Pii et al. 2015). In this work, we investigated three main factors influencing the rhizosphere colonization by B. velezensis. Surfactin production, composition of root exudates, and capability of biofilm formation were evaluated. Surfactin-producer strains better colonize the rhizosphere but the difference is weaker (around 30% of difference of biomass) when compared with a non-surfactin producer strain. These results indicated that surfactin plays a role on the invasion of a Petri plate and on the rhizosphere but surfactin alone cannot explain this last phenomenon. Then, tomato root exudates and their components were evaluated by measuring the growth and the surfactin production by B. velezensis. Sugars and organic acids allowed a bacterial growth and a surfactin production but many differences were observed in the behavior of the strain. Depending on the substrates, biomass and surfactin productions differ but here also, root exudates associated to surfactin production cannot explain the capacity of a strain to be a good rhizosphere colonizer or not. Then, we investigated the role of biofilm formation in root colonization. A non-exopolysaccaride producer B. velezensis FZB42 mutant was constructed and compared to the wild type. By suppressing the capacity of exopolysaccharide production, the EPS− mutant also lost his capability to form a biofilm while the wild strain B. velezensis FZB42 showed a high biofilm formation. We also showed that the biofilm formation was influenced by the different substrates found in root exudates. Our results led that the rhizosphere colonization by B. velezensis FZB42 is dependent on surfactin production and on root exudates composition but the main factor influencing a good colonization is the capability of this strain to form a biofilm.
References
Allison DG, Sutherland IW (1987) The role of exopolysaccharides in adhesion of freshwater bacteria. J Gen Microbiol 133:1319–1327. https://doi.org/10.1099/00221287-133-5-1319
Badri DV, Vivanco JM (2009) Regulation and function of root exudates. Plant Cell Environ 32:666–681. https://doi.org/10.1111/j.1365-3040.2009.01926.x
Bais HP, Fall R, Vivanco JM (2004) Biocontrol of Bacillus subtilis against infection of Arabidopsis roots by Pseudomonas syringae is facilitated by biofilm formation and surfactin production. Plant Physiol 134:307–319. https://doi.org/10.1104/pp.103.028712
Bertin C, Yang X, Weston LA (2003) The role of root exudates and allelochemicals in the rhizosphere. Plant Soil 256:67–83. https://doi.org/10.1023/A:1026290508166
Budiharjo A, Chowdhury SP, Dietel K, Beator B, Dolgova O, Fan B, Bleiss W, Ziegler J, Schmid M, Hartman A, Borriss R (2014) Transposon mutagenesis of the plant-associated Bacillus amyloliquefaciens ssp. plantarum FZB42 revealed that the nfrA and RBAM17410 genes are involved in plant- microbe-interactions. PLoS One 9(5):e98267. https://doi.org/10.1371/journal.pone.0098267
Cao G, Zhang X, Zhong L, Lu Z (2011) A modified electro-transformation method for Bacillus subtilis and its application in the production of antimicrobial lipopeptides. Biotechnol Lett 33:1047–1051. https://doi.org/10.1007/s10529-011-0531-x
Chen XH, Koumoutsi1 A, Scholz R, Eisenreich1 A, Schneider K, Heinemeyer I, Morgenstern B, Voss B, Hess WR, Reva O, Junge H, Voigt B, Jungblut PR, Vater J, Sussmuth R, Liesegang H, Strittmatter A, Gottschalk G, Borriss R (2007) Comparative analysis of the complete genome sequence of the plant growth-promoting bacterium Bacillus amyloliquefaciens FZB42. Nat Biotechnol (9):1007–1014. https://doi.org/10.1038/nbt1325
Choudhary DK, Johri BN (2009) Interactions of Bacillus spp. and plants with special reference to induced systemic resistance (ISR). Microbiol Res 164:493–513. https://doi.org/10.1016/j.micres.2008.08.007
Das BB, Dkhar MS (2011) Rhizosphere microbial populations and physico chemical properties as affected by organic and inorganic farming practices. Am Eur J Agric Environ Sci 10:140–150
Davey ME, Caiazza NC, O'Toole GA (2003) Rhamnolipid surfactant production affects biofilm architecture in Pseudomonas aeruginosa PAO1. J Bacteriol 185:1027–1036. https://doi.org/10.1128/JB.185.3.1027-1036.2003
Deravel J, Lemiere S, Coutte F, Krier F, Van Hese N, Bechet M, Jacques P (2014) Mycosubtilin and surfactin are efficient, low ecotoxicity molecules for the biocontrol of lettuce downy mildew. Appl Microbiol Biotechnol 98:6255–6264. https://doi.org/10.1007/s00253-014-5663-1
Dietel K, Beator B, Budiharjo A, Fan B, Borris R (2013) Bacterial traits involved in colonization of Arabidopsis thaliana roots by Bacillus amyloliquefaciens FZB42. Plant Pathol J 29:59–66. https://doi.org/10.5423/PPJ.OA.10.2012.0155
Donlan RM, Costerton JW (2002) Biofilms: survival mechanisms of clinically relevant microorganisms. Clin Microbiol Rev 15:167–193. https://doi.org/10.1128/CMR.15.2.167-193.2002
Emmert EA, Handelsman J (1999) Biocontrol of plant disease: a Gram-positive perspective. FEMS Microbiol Lett 171:1–9. https://doi.org/10.1111/j.1574-6968.1999.tb13405.x
Fan B, Chen XH, Budiharjo A, Bleiss W, Vater J, Borriss R (2011) Efficient colonization of plant roots by the plant growth promoting bacterium Bacillus amyloliquefaciens FZB42, engineered to express green fluorescent protein. J Biotechnol 151:303–311. https://doi.org/10.1016/j.jbiotec.2010.12.022
Fan B, Carvalhais LC, Becker A, Fedoseyenko D, von Wirén N, Borriss R (2012) Transcriptomic profiling of Bacillus amyloliquefaciens FZB42 in response to maize root exudates. BMC Microbiol 12:116. https://doi.org/10.1186/1471-2180-12-116
Fan B, Blom J, Klenk HP, Borriss R (2017) Bacillus amyloliquefaciens, Bacillus velezensis, and Bacillus siamensis form an “operational group B. amyloliquefaciens” within the B. subtilis species complex. Front Microbiol 8:22. https://doi.org/10.3389/fmicb.2017.00022
Fang W, Hu JY, Ong SL (2009) Influence of phosphorus on biofilm formation in model drinking water distribution systems. J Appl Microbiol 106:1328–1335. https://doi.org/10.1111/j.1365-2672.2008.04099.x
Farace G, Fernandez O, Jacquens L, Coutte F, Krier F, Jacques P, Clément C, Ait Barka E, Jacquard C, Dorey S (2015) Cyclic lipopeptides from Bacillus subtilis activate distinct patterns of defence responses in grapevine. Mol Plant Pathol 16(2):177–187. https://doi.org/10.1111/mpp.12170
Flemming HC, Wingender J (2010) The biofilm matrix. Nat Rev Microbiol 8:623–633. https://doi.org/10.1038/nrmicro2415
Flemming HC, Neu TR, Wozniak DJ (2007) The EPS matrix: the “house of biofilm cells”. J Bacteriol 189:7945–7947. https://doi.org/10.1128/JB.00858-07
Garcia-Gutierrez L, Zeriouh H, Romero D, Cubero J, Vicente A, Perez-Garcia A (2013) The antagonistic strain Bacillus subtilis UMAF6639 also confers protection to melon plants against cucurbit powdery mildew by activation of jasmonate-and salicylic acid-dependent defence responses. Microb Biotechnol 6:264–274. https://doi.org/10.1111/1751-7915.12028
Glick BR (2012) Plant growth-promoting bacteria: mechanisms and applications. Scientifica:1–15. https://doi.org/10.6064/2012/963401
Hsueh YH, Somers EB, Lereclus D, Wong ACL (2006) Biofilm formation by Bacillus cereus is influenced by PlcR, a pleiotropic regulator. Appl Environ Microbiol 72:5089–5092. https://doi.org/10.1128/AEM.00573-06
Idris EE, Makarewicz O, Farouk A, Rosner K, Greiner R, Bochow H, Borriss R (2002) Extracellular phytase activity of Bacillus amyloliquefaciens FZB45 contributes to its plant-growth-promoting effect. Microbiology 148:2097–2109. https://doi.org/10.1099/00221287-148-7-2097
Idris EE, Bochow H, Ross H, Borriss R (2004) Use of Bacillus subtilis as biocontrol agent. VI. Phytohormone-like action of culture filtrates prepared from plant growth-promoting Bacillus amyloliquefaciens FZB24, FZB42, FZB45 and Bacillus. Z Pflanzenkrankh Pflanzenschutz 111:583–597. https://doi.org/10.1111/j.1574-6968.1999.tb13405.x
Jacques P, Hbid C, Destain J, Razafindralambo H, Paquot M, De Pauw E and Thonart P (1999) Optimization of biosurfactant lipopeptide production from Bacillus subtilis S499 by Plackett-Burman design. In Twentieth Symposium on Biotechnology for Fuels and Chemicals. Humana Press, pp 223–233
Julkowska D, Obuchowski M, Holland IB, Séror SJ (2004) Branched swarming patterns on a synthetic medium formed by wild-type Bacillus subtilis strain 3610: detection of different cellular morphologies and constellations of cells as the complex architecture develops. Microbiology 150:1839–1849. https://doi.org/10.1099/mic.0.27061-0
Kloepper JW, Ryu CM, Zhang S (2004) Induced systemic resistance and promotion of plant growth by Bacillus spp. Phytopathology 94:1259–1266. https://doi.org/10.1094/PHYTO.2004.94.11.1259
Kohler J, Caravaca F, Carrasco L, Roldan A (2007) Interactions between a plant growth-promoting rhizobacterium, an AM fungus and a phosphate-solubilizing fungus in the rhizosphere of Lactuca sativa. Appl Soil Ecol 35:480–487. https://doi.org/10.1016/j.apsoil.2006.10.006
Koumoutsi A, Chen XH, Henne A, Liesegang H, Hitzeroth G, Franke P, Vater J, Borris R (2004) Structural and functional characterization of gene clusters directing nonribosomal synthesis of bioactive cyclic lipopeptides in Bacillus amyloliquefaciens strain FZB42. J Bacteriol 186:1084–1096. https://doi.org/10.1128/JB.186.4.1084-1096.2004
Leclerc H (2003) Relationships between common water bacteria and pathogens in drinking-water. Heterotrophic Plate Counts and Drinking-water Safety. IWA Publishing, London, pp 80–118
Leclere V, Marti R, Béchet M, Fickers P, Jacques P (2006) The lipopeptides mycosubtilin and surfactin enhance spreading of Bacillus subtilis strains by their surface active properties. Arch Microbiol 186:475–483. https://doi.org/10.1007/s00203-006-0163-z
Ling N, Raza W, Ma J, Huang Q, Shen Q (2011) Identification and role of organic acids in watermelon root exudates for recruiting Paenibacillus polymyxa SQR-21 in the rhizosphere. Europ J Soil Biol 47:374–379. https://doi.org/10.1016/j.ejsobi.2011.08.009
Lugtenberg B, Kamilova F (2009) Plant-growth-promoting rhizobacteria. Annu Rev Microbiol 63:541–556. https://doi.org/10.1146/annurev.micro.62.081307.162918
Maget-Dana R, Thimon L, Peypoux F, Ptak M (1992) Surfactin/iturin A interactions may explain the synergistic effect of surfactin on the biological properties of iturin A. Biochimie 74:1047–1051. https://doi.org/10.1016/0300-9084(92)90002-V
Makarewicz O, Dubrac S, Msadek T, Borriss R (2006) Dual role of the PhoP∼P response regulator: Bacillus amyloliquefaciens FZB45 phytase gene transcription is directed by positive and negative interactions with the phyC promoter. J Bacteriol 188:6953–6965. https://doi.org/10.1128/JB.00681-06
Mayer C, Moritz R, Kirschner C, Borchard W, Maibaum R, Wingender J, Flemming HC (1999) The role of intermolecular interactions: studies on model systems for bacterial biofilms. Int J Biol Macromol 26:3–16. https://doi.org/10.1016/S0141-8130(99)00057-4
Molinatto G, Puopoloa G, Sonego P, Moretto M, Engelen K, Viti C, Ongena M, Pertota I (2016) Complete genome sequence of Bacillus amyloliquefaciens subsp. plantarum S499, a rhizobacterium that triggers plant defences and inhibits fungal phytopathogens. J Biotechnol 238:56–59. https://doi.org/10.1016/j.jbiotec.2016.09.013
Nongkhlaw FMW, Joshi SR (2014) Distribution pattern analysis of epiphytic bacteria on ethnomedicinal plant surfaces: a micrographical and molecular approach. J Microscopy Ultrastructure 2:34–40. https://doi.org/10.1016/j.jmau.2014.02.003
Ongena M, Jacques P (2008) Bacillus lipopeptides: versatile weapons for plant disease biocontrol. Trends Microbiol 16:115–125. https://doi.org/10.1016/j.tim.2007.12.009
Ongena M, Jourdan E, Adam A, Paquot M, Brans A, Joris B, Thonart P (2007) Surfactin and fengycin lipopeptides of Bacillus subtilis as elicitors of induced systemic resistance in plants. Environ Microbiol 9:1084–1090. https://doi.org/10.1111/j.1462-2920.2006.01202.x
Pii Y, Mimmo T, Tomasi N, Terzano R, Cesco S, Crecchio C (2015) Microbial interactions in the rhizosphere: beneficial influences of plant growth-promoting rhizobacteria on nutrient acquisition process. A review. Biol Fertil Soils 51:403–415. https://doi.org/10.1007/s00374-015-0996-1
Ramey BE, Koutsoudi M, von Bodman SB, Fuqua C (2004) Biofilm formation in plant–microbe associations. Curr Opin Microbiol 7:602–609. https://doi.org/10.1016/j.mib.2004.10.014
Rudrappa T, Quinn WJ, Stanley-Wall NR, Bais HP (2007) A degradation product of the salicylic acid pathway triggers oxidative stress resulting in down-regulation of Bacillus subtilis biofilm formation on Arabidopsis thaliana roots. Planta 226:283–297. https://doi.org/10.1007/s00425-007-0480-8
Saharan BS, Nehra V (2011) Plant growth promoting rhizobacteria: a critical review. Life Sci Med Res 21:1–30
Scholz R, Vater J, Budiharjo A, Wang Z, He Y, Dietel K, Borriss R (2014) Amylocyclicin, a novel circular bacteriocin produced by Bacillus amyloliquefaciens FZB42. J Bacteriol 196:1842–1852. https://doi.org/10.1128/JB.01474-14
Sutherland IW (2001) Biofilm exopolysaccharides: a strong and sticky framework. Microbiology 147:3–9. https://doi.org/10.1099/00221287-147-1-3
Swiecilo A, Zych-Wezyk I (2013) Bacterial stress response as an adaptation to life in a soil environment. Pol J Environ Stud 6:1577–1587
Tan S, Yang C, Mei X, Shen S, Raza W, Shen Q, Xu Y (2013) The effect of organic acids from tomato root exudates on rhizosphere colonization of Bacillus amyloliquefaciens T-5. Appl Soil Ecol 64:15–22. https://doi.org/10.1016/j.apsoil.2012.10.011
Van Loon LC, Bakker PAHM (2005) Induced systemic resistance as a mechanism of disease suppression by rhizobacteria. In: PGPR: biocontrol and biofertilization. Springer, Netherlands, pp 39–66
Vancura V, Hanzlikova A (1972) Root exudates of plants: IV. Differences in chemical composition of seed and seedlings exudates. Plant Soil 36:271–282
Vancura V, Hovadik A (1965) Root exudates of plants: II. Composition of root exudates of some vegetables. Plant Soil 22:21–32
Vessey JK (2003) Plant growth promoting rhizobacteria as biofertilizers. Plant Soil 255:571–586. https://doi.org/10.1023/A:1026037216893
Watnick PI, Kolter R (1999) Steps in the development of a Vibrio cholerae El Tor biofilm. Mol Microbiol 34:586–595. https://doi.org/10.1046/j.1365-2958.1999.01624.x
Weller DM, Raaijmakers JM, Gardener BBM, Thomashow LS (2002) Microbial populations responsible for specific soil suppressiveness to plant pathogens 1. Annu Rev Phytopathol 40:309–348. https://doi.org/10.1146/annurev.phyto.40.030402.110010
Zhang GQ, Bao P, Zhang Y, Deng AH, Chen N, Wen TY (2011) Enhancing electro-transformation competency of recalcitrant Bacillus amyloliquefaciens by combining cell-wall weakening and cell-membrane fluidity disturbing. Anal Biochem 409:130–137. https://doi.org/10.1016/j.ab.2010.10.013
Acknowledgements
The authors thank Dr. Rainer Borriss for kindly providing the Bacillus velezensis strains.
Funding
This work was supported by the University of Lille 1 Sciences and Technologies, the European Funds of INTERREG IV PhytoBio Project and of INTERREG V Smartbiocontrol portfolio, BioProd project and the CPER FEDER project ALIBIOTECH. The authors thank the REALCAT platform for the use of BioLector in this work. The REALCAT platform is benefiting from a state subsidy administrated by the French National Research Agency (ANR) within the frame of the ‘Future Investments’ program (PIA), with the contractual reference ‘ANR-11-EQPX-0037’. The European Union, through the ERDF funding administered by the Hauts-de-France Region, has co-financed the platform. Centrale Lille, the CNRS, and Lille 1 University as well as the Centrale Initiatives Foundation, are thanked for their financial contributions to the acquisition and implementation of the equipment of the REALCAT platform. Ameen Al-Ali was a recipient of PhD scholarship awarded by Campus France through joint French-Iraqi governments program.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible editor: Philippe Garrigues
Rights and permissions
About this article
Cite this article
Al-Ali, A., Deravel, J., Krier, F. et al. Biofilm formation is determinant in tomato rhizosphere colonization by Bacillus velezensis FZB42. Environ Sci Pollut Res 25, 29910–29920 (2018). https://doi.org/10.1007/s11356-017-0469-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-017-0469-1