Abstract
Pathogenic Vibrio alginolyticus, a cause of severe infection in shellfish, as well as in humans, has been found at high frequency around all coastal areas of Korea. The aim of this study was to determine the occurrence of V. alginolyticus, to identify the strains isolated from oysters in West Sea, and to investigate their antimicrobial resistance profiles. Biochemical analyses of the 90 initially recovered presumptive V. alginolyticus colonies indicated that 16 isolates were V. alginolyticus. PCR analysis to detect the presence of the gyrB gene confirmed that 15 (93.8 %) of the 16 isolates were V. alginolyticus. These 15 isolates had the following profiles of resistance against 16 antibiotics: all isolates were resistant to ampicillin and vancomycin, and 26.7 % of the isolates exhibited resistance to cephalothin. A large number of isolates showed intermediate resistance to erythromycin (100 %) and rifampin (73.3 %). Five (33.3 %) of the V. alginolyticus isolates demonstrated multiple resistance to at least three antimicrobials.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Vibrio alginolyticus, as representative of halophilic vibrios, can be isolated from coastal waters and sediments all over the world (Holt et al. 1994), preferring temperatures between 17 and 35 °C and salinity between 5 and 25 ‰ (Hornstrup and Gahrn-Hansen 1993). V. alginolyticus is considered a marine fish and shellfish pathogen (Gomez-León et al. 2005). This bacterium is a common inhabitant of the marine environment in both temperate and tropical waters (Zanetti et al. 2000) and is associated with high mortality in aquaculture systems.
V. alginolyticus is associated with human infections related to consumption of raw or undercooked sea products (fishes and shellfishes) causing severe gastroenteritis and extraintestinal diseases (wounds, intracranial infection in immunocompromised and cirrhotic patients). These illnesses occur frequently during the summer related to an increase in seawater temperature (Croci et al. 2001; Cavallo and Stabili 2002; Thompson et al. 2004). This microorganism produces many extracellular proteases responsible for interactions between the bacterium and cell hosts (human and animals) and plays an important role in human infection and fish pathology (Ottaviani et al. 2001; Thompson et al. 2004). The mechanism of pathogenicity induced by Vibrio infections is still complex and related to several factors including cytotoxins, enterotoxins, and lytic enzymes (Ottaviani et al. 2001).
Antimicrobials and other chemotherapeutic agents are used in fish farms as feed additives and/or immersion baths to treat and prevent the spread of disease (Oh et al. 2011; Kesarcodi-Watson et al. 2008), and the presence of Vibrio species that are major fish and shellfish pathogens has led to the widespread use of such antimicrobials in the marine environment. The overuse of antimicrobials in veterinary therapy has led to the emergence of single- and multiple-drug resistant bacterial strains (Levy 2001) and to the selection of antibiotic-resistant bacteria (Son et al. 1997; Manjusha et al. 2005). Therefore, it is necessary to establish a monitoring system for the objective evaluation of local and national levels of antimicrobial resistance in aquaculture. Because Vibrio strains are common gut flora in farmed fish, they can be used as indicators of the development of resistance to antimicrobials. Since it was discovered in the 1950s in Japan, this antimicrobial resistance organism has been isolated from a variety of seafood, including fish and shellfish in USA, China, Korea, and many other countries (Fujino et al. 1953; Cook et al. 2002; Lee et al. 2008; Yamamoto et al. 2008; Zhao et al. 2011).
In this study, we gathered information about the potential bacterial pathogens in oysters and reported on the antimicrobial susceptibility patterns of V. alginolyticus isolated from oysters collected from the west coast of Korea.
Materials and methods
Sample collection and isolation
During July to October 2014, we tested oysters from shellfish farms on the west coast of Korea. All oyster samples were collected once a month for 4 months, and the position in which the oyster samples were collected is as follows: 37○ 09′ 20″ N and 126○ 20′ 00″ E, 37○ 10′ 04″ N and 126○ 15′ 50″ E, 37○ 11′ 10″ N and 126○ 13′ 27″ E, and 37○ 13′ 00″ N (latitude) and 126○ 10′ 55″ E (longitude), respectively. All samples were collected in ice box kept on ice and transported to the laboratory to be processed within 6 h. Oyster meat (200 g) was transferred to an autoclaved beaker and mixed with 200 mL of phosphate-buffered saline (PBS, 2.5 mM KH2PO4; pH 7.2). Next, 100 μL of oyster homogenates and serial tenfold dilutions (in 0.1 % peptone water) of the homogenates were spread on tryptic soy agar plates (TSA; Difco, MI, USA) containing 3 % NaCl and incubated at 35 °C for 24 h.
Sampling sites were located at shellfish growing areas in the west coast of Korea, where abiotic parameter measurements were done once a month from July to October 2014. Water column depth and surface water salinity, temperature, dissolved oxygen, conductivity, and pH were measured on every sampling date and at each location with a YSI 556 Multiprobe System (YSI Incorporated, Yellow spring, OH, USA) multiparameter display system in accordance with the manufacturer’s instructions.
Biochemical testing
After incubation of the TSA plates, three to five colonies suspected to be of the Vibrio spp. were picked and streaked on thiosulfate citrate bile salt sucrose agar (TCBS; Difco, MI, USA) plates to obtain pure colonies. Yellow colonies on the TCBS agar plates considered presumptive V. alginolyticus were transferred onto CHROMagar™ Vibrio (CHROMagar, ASANPHARM.CO., LTD., Korea). Colorless colonies on CHROMagar™ Vibrio were streaked onto 3 % NaCl TSA. These were selected and screened for the production of oxidase and for fermentation activity on triple sugar iron (TSI; Oxoid, UK) agar for 18 to 24 h at 35 °C. Presumptive colonies were confirmed as V. alginolyticus with the API 20E kit (BioMerieux, France).
DNA extraction
DNA from presumptive V. alginolyticus colonies was extracted using a boiled cell lysate method (Han et al. 2012). Colonies from semisolid nutrient agar were revived in TSB with 3 % w/v NaCl and incubated in a shaker incubator at 200 rpm at 35 °C. Then, 1 mL of overnight culture suspension was pipetted into a sterile 1.5-mL Eppendorf tube and centrifuged at 10,000 rpm for 5 min. The supernatant was carefully discarded, 1 mL of sterile ultrapure water was added, and the tube was vortexed and then boiled at 100 °C for 7 min. The tubes were immediately immersed in ice for 5 min and then centrifuged at 13,000 rpm for 1 min to separate the debris from the DNA-containing supernatant. The supernatant was transferred to a new tube and used as a template for PCR analysis.
Identification of V. alginolyticus using gyrB-based PCR assay
Identification of isolated Vibrio strains was carried out by PCR amplification and DNA sequence determination of the gyrB gene (Luo and Hu 2008). PCR amplification was performed under the following conditions: initial denaturation at 94 °C for 4 min, 32 cycles of 94 °C for 30 s, 64 °C for 30 s, and 72 °C for 1 min, followed by a final extension at 72 °C for 8 min. All amplified products were separated using a 2 % agarose gel containing 0.5 % ethidium bromide. An isolate was considered positive if amplification of a product of the expected size was observed in this simplex PCR.
Antimicrobial susceptibility test
Susceptibility to several antimicrobial agents was determined using a disk diffusion technique (NCCLS 2003). Bacterial suspensions were spread onto Muller-Hinton agar (MHA; Difco, MI, USA) plates onto which antibiotic disks (Oxoid, UK) were then placed. The plates were incubated at 35 °C for 18–24 h under aerobic conditions. The diameter of the zone of inhibition around each disk was measured and recorded. Each bacterial species was classified as resistant (R), intermediately resistant (I), or sensitive (S) according to the guidelines of the Clinical and Laboratory Standards Institute (NCCLS 2003). The following antibiotics, with their concentrations given in parentheses, were tested: ampicillin (AM; 10 μg), cefotaxime (CTX; 30 μg), cefotetan (CTT; 30 μg), cephalothin (CF; 30 μg), chloramphenicol (C; 30 μg), ciprofloxacin (CIP; 5 μg), cefepime (CEP; 30 μg), erythromycin (E; 15 μg), gentamicin (GM; 10 μg), kanamycin (K; 30 μg), nalidixic acid (NA; 30 μg), rifampicin (RA; 5 μg), streptomycin (S; 10 μg), tetracycline (TE; 30 μg), trimethoprim/sulfamethoxazole (SXT; 1.25 μg and 23.75 μg, respectively), and vancomycin (VA; 30 μg). The multiple antibiotic resistance (MAR) index of isolates was defined as x/y, where x represents the number of antimicrobials to which a particular isolate was resistant and y represents the number of antimicrobials to which the isolate was exposed (Krumperman 1983).
Results and discussion
Physical and chemical water quality
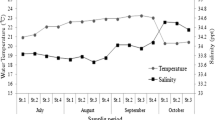
Water temperature, salinity, pH, and dissolved oxygen (DO) were uniform across the four sampling locations. Water temperature at the sampling sites varied from 20.35 to 23.27 °C (average = 22.07 ± 1.06 °C), and salinity ranged from 33.67 to 34.52 ‰ (average = 33.98 ± 0.27 ‰). Dissolved oxygen and pH values varied from 5.88 to 7.32 mg/L (average = 6.59 ± 0.45 mg/L) and from 7.99 to 8.14 (average = 8.07 ± 0.04), respectively (Table 1).
Isolation and identification
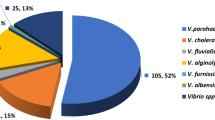
In the present study, conventional cultural methods based on colony appearance (yellow colonies cultivated on TCBS agar) and microscopic examination (gram negative, curved rod shape, non-spore-forming) could detect presumptive V. alginolyticus in 90 out of the 200 (45.0 %) Vibrio species. These 90 presumptive V. alginolyticus strains consisted of 25 (27.8 %), 25 (27.8 %), 25 (27.8 %), and 15 (16.6 %) isolates collected in July, August, September, and October, respectively (Fig. 1). Further biochemical tests of the presumptive V. alginolyticus isolates, however, indicated that seven (7.8 %), six (6.7 %), two (2.2 %), and one (1.1 %) of the isolates collected in July, August, September, and October, respectively, were V. alginolyticus. Variation in the prevalence of V. alginolyticus among the examined samples was likely attributable to the different geographical locations of the study area, sampling sites, water salinity, seasons of the year, species of fish, postharvest practices, and hygienic standards applied during the handling, transport, and storage of seafood products, as well as the methods used for the isolation and identification of the organism (Abd-Elghany and Sallam 2013). In Korea, raw oysters are the most important source of vibrios, the numbers of which increase in summer (Jun et al. 2012), peaking from July to September (Lee et al. 2008). The occurrence of V. alginolyticus is influenced by seasonal variations in water temperature (Kelly 1982; Cavallo and Stabili 2002), and according to our study and other reports (Vezzulli et al. 2013), Vibrio concentrations are generally higher in warmer waters.
Confirmation of the V. alginolyticus isolates by PCR
Phenotypic characterization of the isolates using the API 20E kit confirmed that V. alginolyticus was the only isolated Vibrio species. Sixteen bacterial strains were screened using PCR primers specific for V. alginolyticus. The gyrB gene has been shown to be highly useful for this level of species differentiation (Tacão et al. 2005; Kumar et al. 2006) and has also been used for PCR identification of other Vibrio species (Kumar et al. 2006; Thaithongnum et al. 2006). Fifteen of the isolated strains produced the predicted 568-bp PCR fragments (93.8 % occurrence rate), indicating that they were likely to be V. alginolyticus (Fig. 2). The gyrB-based PCR identification of V. alginolyticus in the present study, and the PCR methods for the identification of other Vibrio species, validates the usefulness of gyrB as a potential candidate for the molecular identification of the Vibrio species. Not all of the selected 90 isolates recovered by colonial identification were confirmed by the biochemical methods to be V. alginolyticus, as some of the yellow colonies on selective media were not V. alginolyticus. The limited selectivity of the medium allows some bacteria, particularly V. alginolyticus, to overgrow and mask the presence of V. parahaemolyticus colonies. In addition, the morphological and growth characteristics of V. alginolyticus on TCBS are similar to those of other Vibrio spp. (Vibrio vulnificus, Vibrio mimicus, Vibrio fluvialis), Photobacterium spp., Chryseomonas spp., or Shewanella spp., which further complicates its identification (Hara-Kudo et al. 2003; Martinez-Urtaza et al. 2008). Due to the possibility of both false-positive and false-negative results in all the biochemical identification methods used, some authors (O’Hara et al. 2003) have suggested caution in the interpretation of such identifications and recommend the use of additional confirmatory tests, such as PCR.
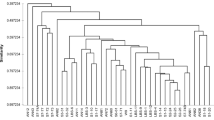
Representative agarose gel electrophoresis for PCR analysis of gyrB gene in V. alginolyticus isolated from oyster. Amplified bands of the expected molecular size of 568 bp for gyrB gene were visualized under UV light. M: DNA marker used as a reference for fragment size; lanes with the numbers from 17 to 85 are the 16 selected strains
Antibiotic resistance
In this study, 16 antimicrobials with various modes of action were chosen for susceptibility testing, and the results are shown in Table 2. Among the 15 isolates tested, all isolates were resistant to AM and VA, and 26.7 % of the isolates exhibited resistance to CF. Prevalence of AM resistance in Vibrio isolates from marine environments is generally high (Ferrini et al. 2008; Han et al. 2007; Stabili et al. 2010). In addition, more than 90.0 % of the V. alginolyticus isolates are resistant to AM (Lee et al. 1998; Son et al. 2005). In this study, 100 % of the V. alginolyticus isolates were resistant to AM, even though this drug is not commonly used in Korean fisheries. According to a previous report, AM resistance is mostly due to the production of β-lactamase (Zanetti et al. 2001).
Relatively small numbers of the isolates were resistant to K (6.7 %), S (6.7 %), and RA (6.7 %), whereas a large number of isolates showed intermediate to E (100 %), RA (73.3 %), CIP (46.6 %), and CF (46.6 %). Evidence of complete resistance to E was not found in the V. alginolyticus isolates, even though E is a representative macrolide in Korean fisheries. Quinolones have been used as an alternative therapy to combat TE-resistant bacteria in Asia (Aoki 1988). Selvin and Lipton (2003) showed that V. alginolyticus isolated from shrimp was sensitive to NA, and we also found that all the V. alginolyticus isolates were sensitive to NA. As our results, along with other reports, demonstrate that V. alginolyticus is sensitive to quinolones, they are likely to be effective antibiotics against V. alginolyticus.
All isolates were susceptible to CTT, NA, SXT, TE, and C, and 93.3 % of the isolates exhibited susceptibility to CEP and S. TE has long been the antimicrobial most commonly used in Korean fisheries, particularly for the treatment of heavy infection by Vibrio species (Morris and Tenny 1985; Liu et al. 2006). Low rates of tetracycline-resistant Vibrio isolates have also been reported in other studies. These results are understandable, as TE has long been the most commonly used antimicrobial in Korean livestock, and the prevalence of antimicrobial resistance was correlated with the amount of antibiotic consumed by the animals.
Five strains (33.3 %) of V. alginolyticus demonstrated multiple antimicrobial resistance (MAR) to at least three antimicrobials (Table 3). The MAR value of most of the isolates was 0.13, but it ranged up to 0.31 in one isolate that was resistant to four antimicrobial types. The MAR index, first proposed by Krumperman (1983), reflects the extent of antimicrobial contamination and gives an indication of the potential risk to human health. MAR values higher than 0.2 suggest that seafood may have originated from a high-risk contaminated source in which antimicrobials were used. In this study, the MAR values of most of the isolates were 0.13 and 0.19, suggesting that most of oysters were not extensively exposed to antimicrobials. However, the oysters, collected from Deokjeokdo (37° 13′ 00″ N and 126° 10′ 55″ E), containing isolates with a MAR value of 0.31, were likely exposed to a high-risk contaminated environment. Several authors have observed a close relationship between the use of antimicrobials in aquaculture and the selection of resistant strains, with the potential risk of resistance and its transfer into the aquatic environment (Balebona et al. 1998). The results of the antimicrobial susceptibility of V. alginolyticus from farmed fish in Korea (Oh et al. 2011) showed a pattern of resistance that was similar to our results.
Conclusions
In Asian countries including Korea, where consumption of raw seafood is common, the Vibrio species tend to be increasingly common pathogens causing human morbidity and mortality. As raw oysters constitute the most important source of V. alginolyticus, and the number of V. alginolyticus increases in summer, purchasing oysters from retail outlets in the summer is prohibited in Korea. In the present study, 90 presumptive V. alginolyticus colonies were isolated from July to October. Subsequently, 15 isolates were identified as V. alginolyticus based on biochemical and PCR analyses. These 15 isolates were characterized by profiling their resistance against 16 antibiotics. The monitoring of the advent and prevalence of antimicrobial resistance in vibrios is necessary, given the trend in antimicrobial usage in fisheries, to identify and implement proper public health measures for the control and prevention of diseases caused by pathogenic vibrios.
References
Abd-Elghany SM, Sallam KI (2013) Occurrence and molecular identification of Vibrio parahaemolyticus in retail shellfish in Mansoura, Egypt. Food Control 33:399–405
Aoki T (1988) Drug-resistant plasmids from fish pathogens. Microbiol Sci 5:219–223
Balebona MC, Zorrilla I, Morinigo MA, Borrego JJ (1998) Survey of bacterial pathologies affecting farmed gilthead sea bream (Sparus aurata L.) in southwestern Spain from 1990 to 1996. Aquaculture 166:19–35
Cavallo RA, Stabili L (2002) Presence of vibrios in seawater and Mytilus galloprovincialis (lam.) from the Mar Piccolo of Taranto (Ionian Sea). Water Res 36:3719–3726
Cook DW, O’Leary P, Hunsucker JC, Sloan EM, Bowers JC, Blodgett RJ, DePaola A (2002) Vibrio vulnificus and Vibrio parahaemolyticus in US retail shell oysters: a national survey from June 1998 to July 1999. J Food Prot 65:79–87
Croci L, Serratore P, Cozzi L, Stacchini A, Milandri S, Suffredini E, Toti L (2001) Detection of Vibrionaceae in mussels and in their seawater growing area. Lett Appl Microbiol 32:57–61
Ferrini AM, Mannoni V, Suffredini E, Cozzi L, Croci L (2008) Evaluation of antibacterial resistance in Vibrio strains isolated from imported seafood and Italian aquaculture settings. Food Anal Methods 1:164–170
Fujino T, Okuno Y, Nakada D, Aoyama A, Fukai K, Mukai T, Ueho T (1953) On the bacteriological examination of shirasu food poisoning. Med J Osaka Univ 4:299–304
Gomez-León J, Villamil L, Lemos ML, Novoa B, Figueras A (2005) Isolation of Vibrio alginolyticus and Vibrio splendidus from aquacultured carpet shell clam (Ruditapes decussatus) larvae associated with mass mortalities. Appl Environ Microbiol 71:98–104
Han AR, Yoon YJ, Kim JW (2012) Antibiotic resistance and plasmid profile of Vibrio parahaemolyticus strains isolated from Kyunggi-Incheon coastal area. Korean J Microbiol 48:22–28
Han F, Walker RD, Janes ME, Prinyawiwatkul W, Ge B (2007) Antimicrobial susceptibilities of Vibrio parahaemolyticus and Vibrio vulnificus isolated from Louisiana Gulf and retail raw oysters. Appl Environ Microbiol 73:7096–7098
Hara-Kudo Y, Sugiyama K, Nishibuchi M, Chowdhury A, Yatsuyanagi J, Ohtomo Y (2003) Prevalence of pandemic thermostable direct hemolysin-producing Vibrio parahaemolyticus O3:K6 in seafood and the coastal environment in Japan. Appl Environ Microbiol 69:3883–3891
Holt JG, Krieg NR, Sneath PHA, Staley JT, Williams ST (1994) Bergey’s manual of determinative bacteriology, 9th edn. Williams & Wilkins, PA, pp. 260–274
Hornstrup MK, Gahrn-Hansen B (1993) Extraintestinal infections caused by Vibrio alginolyticus in a Danish county, 1987–1992. Scand J Infect Dis 25:735–740
Jun JW, Kim JH, Choresca CH, Shin SP, Han JE, Han SY, Chai JY, Park SC (2012) Isolation, molecular characterization, and antibiotic susceptibility of Vibrio parahaemolyticus in Korean seafood. Foodborne Pathog Dis 9:224–231
Kesarcodi-Watson A, Kaspar H, Lategan JM, Gibson L (2008) Probiotics in aquaculture: the need, principles and mechanisms of action and screening processes. Aquaculture 274:1–14
Kelly MT (1982) Effect of temperature and salinity on Vibrio (Beneckea) vulnificus occurrence in a Gulf Coast environment. Am Soc Microbiol 44:820–824
Krumperman PH (1983) Multiple antibiotic resistance indexing of Escherichia coli to identify high-risk sources of fecal contamination of foods. Appl Environ Microbiol 46:165–170
Kumar HS, Parvathi A, Karunasagar I, Karunasagar I (2006) A gyrB-based PCR for the detection of Vibrio vulnificus and its application for direct detection of this pathogen in oyster enrichment broths. Int J Food Microbiol 111:216–220
Lee HK, Yoon YH, Lee SS, Ha KH (1998) Biochemical characteristics of Vibrios isolated from cultured shellfish, Ruditapes philippinarum, and some species of wild shellfish. J Korean Soc Microbiol 33:567–574
Lee JK, Jung DW, Eom SY (2008) Occurrence of Vibrio parahaemolyticus in oysters from Korean retail outlets. Food Control 19:990–994
Levy SB (2001) Antibiotic resistance: consequences of inaction. Clin Infect Dis 33:124–129
Liu JW, Lee IK, Tang HJ, Ko WC, Lee HC, Liu YC, Hsueh PR, Chuang YC (2006) Prognostic factors and antibiotics in Vibrio vulnificus septicemia. Arch Intern Med 166:2117–2123
Luo P, Hu C (2008) Vibrio alginolyticus gyrB sequence analysis and gyrB-targeted PCR identification in environmental isolates. Dis Aquat Org 82:209–216
Manjusha S, Sarita GB, Elyas KK, Chandrasekaran M (2005) Multiple antibiotic resistances of Vibrio isolated from coastal and brackish water areas. Am J Biochem Biotechnol 1:201–206
Martinez-Urtaza J, Huapaya B, Gavilan RG, Blanc-Abad V, Ansede-Bermejo J, Cadarso-Suarez C (2008) Emergence of Asiatic vibrio disease in South America in phase with El Nino. Epidemiology 19:829–837
Morris JG, Tenny J (1985) Antibiotic therapy for Vibrio vulnificus infection. J Am Med Assoc 253:1121–1122
NCCLS (2003) Performance standards for antimicrobial disc susceptibility tests: approved standard, National Committee for Clinical Laboratory Standard of Antimicrobial Susceptibility, document M2-AS. Pennsylvania, USA
O’Hara CM, Sowers EG, Bopp CA, Duda SB, Strockbine NA (2003) Accuracy of six commercially available systems for identification of members of the family Vibrionaceae. J Clin Microbiol 41:5654–5659
Oh EG, Son KT, Yu H, Lee TS, Lee HJ, Shin SB, Kwon JY, Park KBW, Kim JH (2011) Antimicrobial resistance of Vibrio parahaemolyticus and Vibrio alginolyticus strains isolated from farmed fish in Korea from 2005 through 2007. J Food Prot 74:380–386
Ottaviani D, Bacchiocchi I, Masini L, Leoni F, Carraturo A, Giammarioli M, Giovanni S (2001) Antimicrobial susceptibility of potentially pathogenic halophilic vibrios isolated from seafood. Int J Antimicrob Agents 18:135–140
Selvin J, Lipton AP (2003) Vibrio alginolyticus associated with white spot disease of Penaeus monodon. Dis Aquat Org 57:147–150
Son KT, Oh EG, Lee TS, Lee HJ, Kim PH, Kim JH (2005) Antimicrobial susceptibility of Vibrio parahaemolyticus and Vibrio alginolyticus from fish farms on the southern coast of Korea. J Kor Fish Soc 38:365–371
Son R, Rusul G, Sahilah AM, Zainuri A, Raha AR, Salmah I (1997) Antibiotic resistance and plasmid profile of Aeromonas hydrophila isolates from cultured fish, tilapia (Tilapia mossambica). Lett Appl Microbiol 24:479–482
Stabili L, Gravili C, Boero F, Tredici SM, Alifano P (2010) Susceptibility to antibiotics of Vibrio sp AO1 growing in pure culture or in association with its hydroid host Aglaophenia octodonta (Cnidaria, hydrozoa). Microb Ecol 59:555–562
Tacão M, Moura A, Henriques I, Savedra MJ, Correia A (2005) Evaluation of 16S rDNA- and gyrB-DGGE for typing members of the genus Aeromonas. FEMS Microbiol Lett 246:11–18
Thaithongnum S, Ratanama P, Weeradechapol K, Sukhoom A, Vuddhakul V (2006) Detection of V. harveyi in shrimp postlarvae and hatchery tank water by the most probable number technique with PCR. Aquaculture 261:1–9
Thompson FL, Iida T, Swings J (2004) Biodiversity of vibrios. Microbiol Mol Biol Rev 68:3403–3431
Vezzulli L, Colwell RR, Pruzzo C (2013) Ocean warming and spread of pathogenic vibrios in the aquatic environment. Microb Ecol 65:817–825
Yamamoto A, Iwahori J, Vuddhakul V, Charernjiratragul W, Vose D, Osaka K, Shigematsu M, Toyofuku H (2008) Quantitative modeling for risk assessment of Vibrio parahaemolyticus in bloody clams in southern Thailand. Int J Food Microbiol 124:70–78
Zanetti S, Deriu A, Volterra L, Falchi MP, Molicotti P, Fadda G, Sechi L (2000) Virulence factors in Vibrio alginolyticus strains isolated from aquatic environments. Ann Ig 12:487–491
Zanetti S, Spanu T, Deriu A, Romano L, Sechi LA, Fadda G (2001) In vitro susceptibility of Vibrio spp. isolated from the environment. Int J Antimicrob Agents 17:407–409
Zhao F, Zhou D, Cao H, Ma L, Jiang Y (2011) Distribution, serological and molecular characterization of Vibrio parahaemolyticus from shellfish in the eastern coast of China. Food Control 22:1095–1100
Acknowledgments
This work was supported by an Inha University Research Grant.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible editor: Philippe Garrigues
Rights and permissions
About this article
Cite this article
Kang, CH., Shin, Y., Jang, S. et al. Antimicrobial susceptibility of Vibrio alginolyticus isolated from oyster in Korea. Environ Sci Pollut Res 23, 21106–21112 (2016). https://doi.org/10.1007/s11356-016-7426-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-016-7426-2