Abstract
Biodegradable polymer was used as carbon source and biofilm support for nitrate removal from aqueous solution as an attractive alternative for biological denitrification. The objective of this paper was to investigate the denitrification performance and microbial community of a packed-bed bioreactor using poly (butanediol succinate) (PBS), a biodegradable polymer, as carbon source and biofilm support. NO3–N concentration was determined by UV spectrophotometer. NO2–N concentration was assayed by hydrochloric acid naphthyl ethylenediamine spectrophotometry method. Total organic carbon (TOC) was measured using a TOC analyzer. The morphology of the samples was observed using an environmental scanning electron microscope (ESEM). The microbial community was analyzed by pyrosequencing method. The experimental results showed that an average removal efficiency of nitrate was 95 %. ESEM observation and FTIR analysis indicated the changes of PBS granules before and after microbial utilization. Pyrosequencing results showed that Betaproteobacteria predominated, and most of PBS-degrading denitrifying bacteria were assigned to the family Comamonadaceae. Denitrifying bacteria accounted for 13.02 % in total population. The PBS granules were suitable support and carbon source for denitrifying bacteria.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
In recent years, nitrate pollution of water body has become more and more serious and caused a series of environmental problems such as eutrophication in China. The removal of nitrate from contaminated body of water and municipal and industrial wastewater is a topic of intense research (Chu and Wang 2011a, b; Shen and Wang 2011). Nitrogen removal is one of the most important issues in wastewater treatment. In decades, researchers put much effort into developing efficient denitrification system, among which is the solid phase denitrification using biodegradable polymer (BDP) as carbon source (Boley et al. 2000; Hille et al. 2009; Walters et al. 2009; Wang and Wang 2009). While conventional process with methanol and other liquid organic has the risk of overdosing and increased dissolved organic carbon (DOC) in effluent, solid phase denitrification with BDP can avoid such concerns since it is insoluble (Zhao et al. 2009). Acting as both energy source and a biofilm carrier, biodegradable polymers make ease for operation. Polyhydroxyalkanoates including poly(3-hydroxybutyrate) (PHB) and poly (3-droxybutyrate-co-hyroxyvelate) (PHBV) are easily degraded and have shown high rate of nitrogen removal. Yet, the prohibitive cost prevents their use in practice (Hiraishi and Khan 2003). In light of this, poly (butanediol succinate) (PBS) is economically more attractive.
PBS is widely used in package, tableware, agricultural film, and medical field due to its excellent comprehensive property and cost-effectiveness (Zhou et al. 2009). Moreover, it can be modified with additives and polymerized with other materials easily. As a potential carbon source, the study of its application in solid denitrification is of great importance. To have a deep understanding of the process utilizing PBS, functional and structural analyses of the microbial communities are necessary. Yet, little information is available on PBS-degrading denitrifying bacteria.
This study aims to reveal the microbial composition and diversity of the PBS-acclimated reactor. Pyrosequencing is chosen to achieve this purpose giving the limitation of other molecular techniques. Unlike other approaches, pyrosequencing does not require subcloning or handling of individual clones (Margulies et al. 2005). Thus, it provides comprehensive information without bias. With advantages over traditional DNA approaches, pyrosequencing gives a more in-depth understanding of microbial diversity (Huse et al. 2007; Manter et al. 2010; Uroz et al. 2010). The pyrosequencing technology has been used to explore microbes in human gut, soil, and oceans (Roesch et al. 2007; Andersson et al. 2008; Biddle et al. 2008; Kirchman et al. 2010; Wu et al. 2010). Yet, few papers have reported its application in bioreactors.
The objectives of this study were to investigate the denitrification performance and material characterization in a packed-bed bioreactor. The microbial community was analyzed by pyrosequencing.
Materials and methods
Material
PBS was kindly supplied by Shenzhen Guanghuaweiye Co. Ltd. The average molecular weight of the polymer is ca. 40,000. PBS is in shape of pellet with a size of 2.5–3 mm.
Apparatus
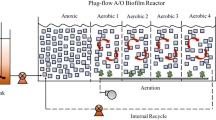
The flow chart was shown in the following. The packing height of the reactor is 25 cm, and the whole volume is 0.69 L (Fig. 1).
Analytical methods
Samples were taken and filtered through 0.45 μm membrane before analysis. NO3–N was determined by UV spectrophotometer (Shimadzu UV-3100) at 220 and 275 nm. NO2–N was assayed by hydrochloric acid naphthyl ethylenediamine spectrophotometry method. Total organic carbon (TOC) was measured using a TOC analyzer (HACH, IL530 TOC-TN). The morphology of the samples was observed using an environmental scanning electron microscope ESEM (Quanta 200F), which is the ultimate low-vacuum SEM with extended low-vacuum capabilities for the really challenging samples and dynamic experiments. Fourier transform infrared spectroscopy (FTIR) spectra were measured using an attenuated total reflection method with Spectrum GX FTIR system (Perkin Elmer).
Pyrosequencing
DNA extraction and purification
Biofilm was collected from the reactor at day 120 when the performance of reactor is stable. Genomic DNA was extracted using E.Z.N.A. Soil DNA Kit (Omega).
PCR amplification
Polymerase chain reaction (PCR) technique was used to amplify DNA. For each sample, we amplified V1–V3 region of bacterial 16S rRNA genes using a broadly conserved primer set (27F and 533R). The forward primer (5′-GCC TTG CCA GCC CGC TCA GAG AGT TTG ATC CTG GCT CAG-3′) contained the 454 Life Sciences primer B sequence and the broadly conserved bacterial primer 27F. The reverse primer (5′-GCC TCC CTC GCG CCA TCA GNN NNN NNN NNT TAC CGC GGC TGC TGG CAC-3′) contained the 454 Life Sciences primer A sequence, a unique 10-nt barcode used to tag each PCR product (designated by NNNNNNNNNN), and the broad-range bacterial primer 533R. PCR reactions were carried out in triplicate 20-μL reactions with 0.4 μM forward and reverse primers, 1-μL template DNA, 250 nM dNTP and 1× FastPfu Buffer. All dilutions were carried out using certified DNA-free PCR water. Thermal cycling consisted of initial denaturation at 95°C for 2 min followed by 25 cycles of denaturation at 95°C for 30 s, annealing at 55°C for 30 s, and extension at 72°C for 30 s, with a final extension of 5 min at 72°C. Replicate amplicons were pooled and visualized on 2.0 % agarose gels using SYBR Safe DNA gel stain in 0.5× TBE. Amplicons were purified using AxyPrep™ DNA Gel Extraction Kit (Axygen) according to the manufacturer’s instructions.
Amplicon quantitation, pooling, and pyrosequencing
Amplicon DNA concentrations were measured using the Quant-iT PicoGreen dsDNA reagent and kit (Invitrogen). DNA samples were diluted in 30 uL 1× TE, and an equal volume of 2× PicoGreen working solution was added in a total reaction volume of 60 μL in minicell cuvette. Fluorescence was measured on a Turner Biosystems TBS-380 Fluorometer using the 465–485/515–575-nm excitation/emission filter pair. Following quantitation, cleaned amplicons were combined in equimolar ratios into a single tube. Pyrosequencing was carried out on a 454 Life Sciences Genome Sequencer FLX Titanium instrument (Roche) by Shanghai Majorbio Bio-pharm Biotechnology Co., Ltd. (Shanghai, China).
Results and discussion
Reactor performance
The denitrification performance of the reactor was studied. The reactor was operated at 25°C with a hydraulic retention time of 0.5 h at first 70 days. The experimental results indicated that the average influent concentration of nitrate was about 15 mg/L. An average nitrate removal efficiency of 94.87 % was achieved under such conditions, which was higher than that using PLA as carbon source. When the temperature dropped to 15°C at day 71, the denitrification performance was significantly influenced, though HRT and influent concentration remained the same. During this period, about 70 % nitrate was removed on average.
In the first phase, from day 1 to day 70, there was little nitrite accumulation. The nitrite concentration was almost all below 0.1 mg/L. The average concentration of nitrite in effluent was 0.05 mg/L. Yet, from day 71, when the denitrification rate slowed down due to the drop of temperature, the nitrite concentration showed a sharp increase. Ranging from 0.30 to 0.50 mg/L, the average concentration of nitrite was increased to 0.40 mg/L, nearly eight times of previous effluent.
At low temperature, the relative slow denitrification rate might cause nitrite accumulation. In turn, the high concentration of nitrite deteriorated the denitrification process. Yet, no matter what the temperature was, the nitrate removal performance of the reactor was quite considerable. PBS made itself a suitable carbon source for denitrification even when the temperature is not quite suitable (Fig. 2).
The denitrification performance at different initial pH (5.0–10.0) was investigated. Compared with an uncontrolled initial pH condition, at both acid and basic condition, the nitrate removal efficiency decreased, indicating that when pH was beyond the optimal range, denitrification enzymatic activity was inhibited. It was similar to the optimal pH range reported for denitrification (Tang et al. 2011).
At acidic pH, the average denitrification rates were also decreased. In addition, the effluent nitrite and DOC concentrations at pH of 5.0 were higher than that at pH of 7.0. It was interesting to note that the average denitrification rate increased when pH increased from 7.0 to 10.0, which might be due to the higher DOC release and accumulation at pH of 10.0. At basic condition, more organic acid products were produced to neutralize the alkalinity, which is confirmed by pH change. These excess release of dissolved organic carbon should be exceeded the need of denitrifying bacteria for denitrification, so a higher DOC accumulation was observed at basic condition. Effluent pH tended to be neutral both at acidic and basic initial conditions, which would be a net result of acidity by acidic degradation products and alkalinity derived from denitrification.
Characterization of PBS
ESEM observation
To elucidate the biological utilization of PBS, environmental scanning electron microscope (ESEM) observation was conducted. The surface of raw PBS shown in Fig. 3a was smooth. In contrast with it, the surface of PBS after 50 days of biological utilization changed a lot. As shown in Fig. 3b, it was full of bumps and hollows. A few bacilli living in the pit of the material failed to be removed in the process of biofilm detachment. These bacilli were firmly attached to the material, able to take full advantage of the carbon source for denitrification. As a result of biodegradation, pores and hollows were formed. The pores in turn provide more area for bacterial attachment as well as an environment for anoxic processes.
Biofilms in different period were also scanned by ESEM. After a week of operation, a thin film of bacilli could be observed (Fig. 3c). The biofilm which developed in a month was much thicker and richer. Besides bacilli, we also found Coccus and filamentous bacteria. Filamentous organisms twined between bacilli and Coccus, increasing the stability of biofilm. An anoxic inner microenvironment was also formed. Comparing the biofilms formed at different depths, we could deduce the developing process of biofilm.
FTIR analysis
FTIR spectroscopy of the material was carried out before and after the biofilm colonization. Yet, the spectra in Fig. 4a, b showed little differences, indicating that biodegradation does not significantly change the chemical structure. The comparison of peak before and after microbial utilization was presented in Table 1.
Microbial community analysis
Pyrosequencing analysis was performed to reveal the bacterial community of biofilm developed on the surface of PBS. The results showed that it yielded a total number of 9,722 sequences after removal of short and low-quality reads. The average read length was 452.5 bp. The numbers of OTUs represented by the pyrosequencing reads were determined at three dissimilarity levels—3, 5, and 10 %. The shape of rarefaction curve indicated that bacterial richness was complete. In this paper, we choose 3 % level for further analysis and discussion. The Chao index, estimating the richness of total bacterial community, ranged from 4,532 to 5,160 with a median of 4,823.
The sequences represented 24 known phyla or candidate divisions. The vast majority of sequences (96.09 %) belonged to three phyla (Fig. 5), among which Proteobacteria was predominant, taking up 88.35 %. The other two were Bacteroidetes and Chloroflexi with percentage of 3.23 and 4.51 %, respectively. Within Proteobacteria phylum, Betaproteobacteria was the most abundant, followed by Gammaproteobacteria and Alphaproteobacteria. Also, there was a little part occupied by Deltaproteobacteria. As to phylum Bacteroidetes, Sphingobacteria and Flavobacteria were relatively abundant representatives.
However, in genus level, a very large fraction of the diversity represented a hitherto unidentified bacterial population. Nearly 80 % of the sequences represented unidentified non-culturable bacteria, underlining the vast untapped diversity. Despite of that, representative denitrifying bacteria still manifested themselves with a considerable percentage of 13.02 %, which accounts for more than one half of the identified genus. Typically, the abundance of denitrifiers among total population in soils was 3–8 % (Henry et al. 2006). Yet, few papers have reported the actual percentage of denitrifying bacteria in reactors. Our study herein showed a relatively high proportion of denitrifying genus.
In the identified genera, Diaphorobacter has been reported to be capable of degrading PHB and PHBV (Khan and Hiraishi 2002). Previous study showed that genera Dechloromonas and Thauera were the dominant denitrifiers in methanol-acclimated (Blackall et al. 2004) and acetate-acclimated (Blackall et al. 2005) activated sludge, respectively. Alicycliphilus was reported denitrifiers belonging to Comamonadaceae (Mechichi et al. 2003; Khardenavis et al. 2007). Besides, denitrifying genera (Khan et al. 2002; Lu et al. 2007; Heylen et al. 2008) such as Simplicispira, Acidovorax, Azospira, Hydrogenophaga, and Desulfovibrio were observed. In denitrifying clusters, Betaproteobacteria also predominate. This was in consistent with previous studies that Betaproteobacteria play a primary role in nitrogen removal with biodegradable polymer as a carbon source (Khan et al. 2002).
Conclusions
An average removal efficiency of nitrate could reach 95 % at stable stage when using PBS as the carbon source and biofilm support in a packed-bed bioreactor. ESEM observation and FTIR analysis confirmed the changes of PBS granules before and after microbial utilization. Pyrosequencing results showed that Betaproteobacteria predominated, and most of PBS-degrading denitrifying bacteria assigned to the family of Comamonadaceae. Denitrifying bacteria accounted for 13.02 % in total population. PBS granules were suitable supporter and carbon source for denitrifying bacteria.
References
Andersson AF, Lindberg M, Jakobsson H, Backhed F, Nyren P, Engstrand L (2008) Comparative analysis of human gut microbiota by barcoded pyrosequencing. PLoS One 3(7):e2836
Biddle JF, Fitz-Gibbon S, Schuster SC, Brenchley JE, House CH (2008) Metagenomic signatures of the Peru Margin subseafloor biosphere show a genetically distinct environment. Proc Natl Acad Sci U S A 105(30):10583–10588
Blackall LL, Ginige MP, Hugenholtz P, Daims H, Wagner M, Keller J (2004) Use of stable-isotope probing, full-cycle rRNA analysis, and fluorescence in situ hybridization-microautoradiography to study a methanol-fed denitrifying microbial community. Appl Environ Microbiol 70(1):588–596
Blackall LL, Ginige MP, Keller J (2005) Investigation of an acetate-fed denitrifying microbial community by stable isotope probing, full-cycle rRNA analysis, and fluorescent in situ hybridization-microautoradiography. Appl Environ Microbiol 71(12):8683–8691
Boley A, Muller WR, Haider G (2000) Biodegradable polymers as solid substrate and biofilm carrier for denitrification in recirculated aquaculture systems. Aquac Eng 22(1–2):75–85
Chu LB, Wang JL (2011a) Comparison of polyurethane foam and biodegradable polymer as carriers in moving bed biofilm reactor for treating wastewater with a low C/N ratio. Chemosphere 83:63–68
Chu LB, Wang JL (2011b) Nitrogen removal using biodegradable polymers as carbon source and biofilm carriers in a moving bed biofilm reactor. Chem Eng J 170:220–225
Henry S, Bru D, Stres B, Hallet S, Philippot L (2006) Quantitative detection of the nosZ gene, encoding nitrous oxide reductase, and comparison of the abundances of 16S rRNA, narG, nirK, and nosZ genes in soils. Appl Environ Microbiol 72(8):5181–5189
Heylen K, Lebbe L, De Vos P (2008) Acidovorax caeni sp. nov., a denitrifying species with genetically diverse isolates from activated sludge. Int J Syst Evol Microbiol 58:73–77
Hille A, He M, Ochmann C, Neu T, Horn H (2009) Application of two component biodegradable carriers in a particle-fixed biofilm airlift suspension reactor: development and structure of biofilms. Bioproc Biosyst Eng 32(1):31–39
Hiraishi A, Khan ST (2003) Application of polyhydroxyalkanoates for denitrification in water and wastewater treatment. Appl Microbiol Biotechnol 61(2):103–109
Huse SM, Huber JA, Morrison HG, Sogin ML, Mark Welch D (2007) Accuracy and quality of massively parallel DNA pyrosequencing. Genome Biol 8(7):R143.1–R143.9
Khan ST, Hiraishi A (2002) Diaphorobacter nitroreducens gen. nov., sp, nov., a poly(3-hydroxybutyrate)-degrading denitrifying bacterium isolated from activated sludge. J Gen Appl Microbiol 48(6):299–308
Khan ST, Horiba Y, Yamamoto M, Hiraishi A (2002) Members of the family Comamonadaceae as primary poly(3-hydroxybutyrate-co-3-hydroxyvalerate)-degrading denitrifiers in activated sludge as revealed by a polyphasic approach. Appl Environ Microbiol 68(7):3206–3214
Khardenavis AA, Kapley A, Purohit HJ (2007) Simultaneous nitrification and denitrification by diverse Diaphorobacter sp. Appl Microbiol Biotechnol 77(2):403–409
Kirchman DL, Cottrell MT, Lovejoy C (2010) The structure of bacterial communities in the western Arctic Ocean as revealed by pyrosequencing of 16S rRNA genes. Environ Microbiol 12(5):1132–1143
Lu SP, Ryu SH, Chung BS, Chung YR, Park W, Jeon CO (2007) Simplicispira limi sp. nov., isolated from activated sludge. Int J Syst Evol Microbiol 57:31–34
Manter DK, Delgado JA, Holm DG, Stong RA (2010) Pyrosequencing reveals a highly diverse and cultivar-specific bacterial endophyte community in potato roots. Microb Ecol 60(1):157–166
Margulies M, Egholm M, Altman WE, Attiya S, Bader JS, Bemben LA, Berka J, Braverman MS, Chen YJ, Chen ZT, Dewell SB, Du L, Fierro JM, Gomes XV, Godwin BC, He W, Helgesen S, Ho CH, Irzyk GP, Jando SC, Alenquer MLI, Jarvie TP, Jirage KB, Kim JB, Knight JR, Lanza JR, Leamon JH, Lefkowitz SM, Lei M, Li J, Lohman KL, Lu H, Makhijani VB, McDade KE, McKenna MP, Myers EW, Nickerson E, Nobile JR, Plant R, Puc BP, Ronan MT, Roth GT, Sarkis GJ, Simons JF, Simpson JW, Srinivasan M, Tartaro KR, Tomasz A, Vogt KA, Volkmer GA, Wang SH, Wang Y, Weiner MP, Yu PG, Begley RF, Rothberg JM (2005) Genome sequencing in microfabricated high-density picolitre reactors. Nature 437(7057):376–380
Mechichi T, Stackebrandt E, Fuchs G (2003) Alicycliphilus denitrificans gen. nov., sp nov., a cyclohexanol-degrading, nitrate-reducing beta-proteobacterium. Int J Syst Evol Microbiol 53:147–152
Roesch LF, Fulthorpe RR, Riva A, Casella G, Hadwin AKM, Kent AD, Daroub SH, Camargo FAO, Farmerie WG, Triplett EW (2007) Pyrosequencing enumerates and contrasts soil microbial diversity. ISME J 1(4):283–290
Shen ZQ, Wang JL (2011) Biological denitrification using cross-linked starch/PCL blends as solid carbon source and biofilm carrier. Bioresour Technol 102:8835–8838
Tang YN, Zhou C, Ziv-El M, Rittmann BE (2011) A pH-control model for heterotrophic and hydrogen-based autotrophic denitrification. Water Res 45:232–240
Uroz S, Buee M, Murat C, Frey-Klett P, Martin F (2010) Pyrosequencing reveals a contrasted bacterial diversity between oak rhizosphere and surrounding soil. Environ Microbiol Rep 2(2):281–288
Walters E, Hille A, He M, Ochmann C, Horn H (2009) Simultaneous nitrification/denitrification in a biofilm airlift suspension (BAS) reactor with biodegradable carrier material. Water Res 43(18):4461–4468
Wang XM, Wang JL (2009) Removal of nitrate from groundwater by heterotrophic denitrification using the solid carbon source. Sci China Ser B Chem 52(2):236–240
Wu GD, Lewis JD, Hoffmann C, Chen YY, Knight R, Bittinger K, Hwang J, Chen J, Berkowsky R, Nessel L, Li HZ, Bushman FD (2010) Sampling and pyrosequencing methods for characterizing bacterial communities in the human gut using 16S sequence tags. BMC Microbiol 10:206
Zhao X, Meng XL, Wang JL (2009) Biological denitrification of drinking water using biodegradable polymer. Int J Environ Pollut 38(3):328–338
Zhou HH, Zhao X, Wang JL (2009) Nitrate removal from groundwater using biodegradable polymers as carbon source and biofilm support. Int J Environ Pollut 38:339–348
Acknowledgments
The authors are grateful to the National Natural Science Foundation of China (grant no. 50978001) and the National S&T Major Project (grant no. 2008ZX07102-003) for their financial support.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible editor: Vinod Kumar Gupta
Rights and permissions
About this article
Cite this article
Wu, W., Yang, L. & Wang, J. Denitrification using PBS as carbon source and biofilm support in a packed-bed bioreactor. Environ Sci Pollut Res 20, 333–339 (2013). https://doi.org/10.1007/s11356-012-0926-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-012-0926-9