Abstract
Introduction
Mass spectrometry and computational biology have advanced significantly in the past ten years, bringing the field of metabolomics a step closer to personalized medicine applications. Despite these analytical advancements, collection of blood samples for routine clinical analysis is still performed through traditional blood draws.
Objective
TAP capillary blood collection has been recently introduced for the rapid, painless draw of small volumes of blood (~ 100 μL), though little is known about the comparability of metabolic phenotypes of blood drawn via traditional venipuncture and TAP devices.
Methods
UHPLC-MS-targeted metabolomics analyses were performed on blood drawn traditionally or through TAP devices from 5 healthy volunteers. Absolute quantitation of 45 clinically-relevant metabolites was calculated against stable heavy isotope-labeled internal standards.
Results
Ranges for 39 out of 45 quantified metabolites overlapped between drawing methods. Pyruvate and succinate were over threefold higher in the TAP samples than in traditional blood draws. No significant changes were observed for other carboxylates, glucose or lactate. TAP samples were characterized by increases in reduced glutathione and decreases in urate and cystine, markers of oxidation of purines and cysteine—overall suggesting decreased oxidation during draws. The absolute levels of bile acids and acyl-carnitines, as well as almost all amino acids, perfectly correlated among groups (Spearman r ≥ 0.95).
Conclusion
Though further more extensive studies will be mandatory, this pilot suggests that TAP-derived blood may be a logistically-friendly source of blood for large scale metabolomics studies—especially those addressing amino acids, glycemia and lactatemia as well as bile acids, acyl-carnitine levels.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
The past decade has been characterized by significant technological advancements in the field of mass spectrometry (MS)-based metabolomics (Crutchfield et al. 2016; Dettmer et al. 2007) and computational methods to process data generated through metabolomics platforms (Uppal et al. 2016). Methods like flow-injection Time of Flight (Fuhrer et al. 2011) or isocratic/gradient-based high-throughput Ultra-High Pressure Liquid Chromatography (UHPLC)-MS-metabolomics (Lewis et al. 2016; Nemkov et al. 2017) have made it possible to process samples by the hundreds/thousands per day for routine screening of metabolites of clinical interest or discovery-mode research studies. This level of throughputness brought the field a step closer to personalized medicine applications, paving the way for the introduction of clinical metabolomics applications (D’Alessandro et al. 2012) and precision medicine (Beger et al. 2016). Despite these analytical advancements, collection of blood samples for routine clinical analysis is still performed through traditional blood draws. In the past few years, metabolomics analyses have informed on the impact of pre-analytical variables (Yin et al. 2015), such as drawing methods (Jobard et al. 2016; Townsend et al. 2016), collection tubes (López-Bascón et al. 2016), sample storage (refrigerated vs. frozen) on the reproducibility of metabolic phenotypes. In this view, notable was the introduction of dried blood spots as a logistically-friendly method to collect, store and ship blood samples for subsequent omics analyses, including metabolomics (Wang et al. 2016; Wilson 2011; Zukunft et al. 2013). Similarly, volumetric absorptive microsampling (VAMS) has been introduced over the past few years for the sampling of biological fluids and, particularly, whole blood, on a porous hydrophilic tip. A comprehensive overview of this technology and its applications has been recently provided by Kok and Fillet (2018).
More recently, the introduction of TAP (originally Touch-Activated Phlebotomy, now simply TAP) capillary blood collection device (Blicharz et al. 2018) has been welcomed as an innovative and user-friendly (i.e. no medical training required) strategy for the rapid and painless draw of small volumes of blood (~ 100 μL) for clinical analysis. This single-use sterile device is claimed to be superior to traditional blood draw by venipuncture or other methods such as fingerstick, owing to the totally painless experience they provide to the donor and the ease of use for untrained personnel. While the technology has quickly cleared Food and Drug Administration certification and attracted a significant degree of positive press, to the best of our knowledge no direct metabolomics comparison has been performed on blood drawn through traditional venipuncture methods and TAP devices on matched healthy volunteers. Though limited by the numbers of prototype available to perform this study, here we document for the first time that absolute quantitative profiles of whole blood drawn traditionally or through TAP are substantially comparable for most classes of small molecule compounds. The promising results reported here are based on the absolute quantitation of a defined set of targeted metabolites (45 total), including amino acids, glycolytic and Krebs cycle intermediates, acyl-carnitines and bile acids. As the panel of metabolites we selected here includes small molecules relevant to most of the basic science and clinical studies performed through the University of Colorado Denver Metabolomics Cores, we anticipate a potential widespread application of the TAP technology for blood-metabolomics related studies. Caveats are noted with respect to some carboxylates [e.g. succinate—relevant marker of mortality in the trauma population (DʼAlessandro et al. 2017; Lusczek et al. 2017)] and amino acids (e.g. arginine, valine and cystine), whose measurements in traditional blood draws differ significantly from the values determined in whole blood obtained through TAP. Finally, we noted decreased levels of oxidative markers (higher succinate and reduced glutathione—GSH, lower urate and cystine) in TAP-derived whole blood, potentially suggestive of decreased redox stress for blood drawn through this novel procedure.
2 Materials and methods
2.1 Blood collection
Commercial reagents were purchased from Sigma-Aldrich (Saint Louis, MO) unless otherwise noted. Blood was drawn from 5 fasting healthy donor volunteers (three females, 2 males, age 29 ± 4) at 9:00 a.m., in agreement with the Declaration of Helsinki and after signing of informed consent in conformity with the University of Colorado institutional review board regulations, both through traditional venipuncture (purple cap—EDTA tubes) or TAP (non-commercial investigational-use prototypes kindly provided by Seventh Sense Biosystems, containing EDTA anticoagulant—Seventh Sense Biosystems, Medford, MA, USA).
2.2 Quantitation and biofluids
Stable heavy isotope labeled standards (complete list, concentration and vendor information included in Supplementary Table 1) were diluted in 50 mL of a metabolite extraction solution consisting of methanol, acetonitrile and water (5:3:2, v/v/v). Whole blood aliquots (25 µL) were extracted in 225 µL (1:10 dilution) of ice cold extraction solution with described standards by vortexing for 30 min at 4 °C, as reported (D’Alessandro et al. 2017). Insoluble proteins were pelleted by centrifugation at 4 °C for 10 min at 18,213 g and supernatant extracts (100 µL) were collected for MS analysis as detailed below. Technical mixes were prepared by pooling extracts from each of the 5 venipuncture donors and 5 TAP device donors and injected every 5 runs with an additional two pre- and post-sample injections and blanks. Technical mixes were analyzed to ensure intraday technical variability, expressed as coefficients of variation (CV = standard deviation/mean) < 10%.
2.3 UHPLC-MS metabolomics and absolute quantitation against stable heavy-isotope labeled standards
Analysis was performed using a Vanquish UHPLC system (Thermo Fisher Scientific, San Jose, CA, USA) coupled to a Q Exactive mass spectrometer (Thermo Fisher Scientific, San Jose, CA, USA). Samples were resolved over a Kinetex C18 column, 2.1 × 150 mm, 1.7 µm particle size (Phenomenex, Torrance, CA, USA) equipped with a guard column (SecurityGuard™ Ultracartidge—UHPLC C18 for 2.1 mm ID Columns—AJO-8782—Phenomenex, Torrance, CA, USA) at 45 °C using a gradient elution over 5 min, flowing at 450 µL/min. The Q Exactive mass spectrometer was operated independently in positive or negative ion mode, scanning in Full MS mode (1 µscan) from 65 to 975 m/z at 70,000 resolution with 4 kV spray voltage, 45 sheath gas and 15 auxiliary gas. In addition, samples were resolved at 45 °C for 5 min using a gradient elution over 5 min, flowing at 450 µL/min. Mobile phase A was 0.1% formic acid in water; mobile phase B was 0.1% formic acid in acetonitrile. 95% A and 5% B was held from 0.00 to 0.50 min. From 0.5 to 1.10 min, A was decreased to 5% A and 95% B. This condition was held from 1.10 to 2.75 min. From 2.75 to 3.00 min, A was increased to the initial condition of 95% A and 5% B. This condition was held from 3.00 to 5.00 min. The Q Exactive mass spectrometer was operated in positive ion mode, scanning in Full MS mode (1 µscan) from 65 to 975 m/z at 70,000 resolution with 4 kV spray voltage, 45 sheath gas and 15 auxiliary gas. For negative ion mode, samples were resolved at 45 °C for 5 min using a gradient elution over 5 min, flowing at 450 µl/min. Mobile phase A was 5% acetonitrile and 95% water with 5 mM ammonium acetate; mobile phase B was 95% acetonitrile and 5% water with 5 mM ammonium acetate. 100% A and 0% B was held from 0.00 to 0.50 min. From 0.5 to 1.10 min, A was decreased to 0% A and 100% B. This condition was held from 1.10 to 2.75 min. From 2.75 to 3.00 min, A was increased to the initial condition of 100% A and 0% B. This condition was held from 3.00 to 5.00 min. The Q Exactive mass spectrometer was operated in negative ion mode using electrospray ionization, scanning in Full MS mode (1 µscan) from 65 to 975 m/z at 70,000 resolution with 4 kV spray voltage, 45 sheath gas and 15 auxiliary gas.
Acquired data was then converted from .raw to .mzXML file format using Mass Matrix (Cleveland, OH, USA). Metabolite assignments and heavy isotopologue detection for absolute quantitation against internal standards were determined against in house standard libraries and KEGG database searches performed using MAVEN (Princeton, NJ, USA). Technical reproducibility (CVs) was assessed by monitoring internal heavy labeled standard mixes, as reported (Nemkov et al. 2017).
Calculation of absolute quantification for measured metabolites was performed using the following formula: [light] = (abundance light)/(abundance heavy) × [heavy corrected] [dilution factor], where [light] = concentration of non-isotopic metabolite, (abundance light) = total area abundance of non-isotopic metabolite, (abundance heavy) = total area abundance of isotopic metabolite, [heavy corrected] = known concentration of isotopic metabolite diluted according to amount of extraction solution used, as previously described (Nemkov et al. 2017).
Graphs, heat maps, statistical analyses (including t test, ANOVA, linear regression and Spearman correlations) and heat maps were prepared using GraphPad Prism 5.0 (GraphPad Software, Inc., La Jolla, CA, USA) and GENE-E (Broad Institute, MA, USA).
3 Results
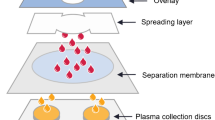
A total of 45 metabolites were quantified in whole blood collected from 5 healthy donor volunteers through paired draws (same subjects, minutes apart between draws) via traditional venipuncture or TAP device (Fig. 1a). A complete list and raw measurements and absolute quantification—as determined against spiked in heavy isotope-labeled internal standards (Fig. 1b)—are reported in Supplementary Table 1. None of the subjected reported any bruise (only a minor mark in the drawing area, visible for < 24 h) or adverse reaction to either procedure, though all five participants referred to the TAP draw as comparably painless. Amino acid analysis revealed comparable trends for both traditional and TAP blood draws (Fig. 2a–c), all within micromolar ranges consistent with those reported in the human metabolome database—HMDB (Wishart et al. 2018)—and our previous studies on whole blood and plasma (Gehrke et al. 2017). Median biological variability for all metabolites in blood drawn via traditional draws or TAP was 27.9 and 26.8%, respectively—showing substantial comparability across both drawing strategies (Supplementary Table 1). Significant changes were observed between drawing methods with respect to valine (fold-change TAP/traditional draw = 0.78; p = 0.01 paired T Test) and arginine (fold-change = 1.49; p = 0.02). Tryptophan and its metabolite kynurenine were extremely comparable between paired drawing methods (Fig. 2d), as were dimethylglycine and trimethylamine oxide (Fig. 2e). On the other hand, significantly lower levels of the oxidized cysteine dimer, cystine and higher levels of reduced glutathione (GSH) were observed in samples drawn through the TAP device (Fig. 2f).
Absolute quantification of amino acids (essential—a, conditionally essential—b and nonessential—c), tryptophan metabolites (d), metabolites involved in choline and microbiome metabolism (e), redox homeostasis mono- and tripeptides like reduced glutathione (GSH) (f) in traditionally drawn blood (red circles) or matched blood drawn via TAP (white). Paired T Test p-Value *<0.05; **<0.01
Absolute quantitative measurement ranges of glucose and lactate were extremely comparable between drawing methods, though blood drawn through the TAP device was characterized by higher levels of pyruvate (fold-change 2.9, p-Value < 0.001—Fig. 3a). Similarly, comparable levels of adenosine but lower levels of urate were determined in matched samples drawn from the TAP device when compared to the traditional venipuncture approach (Fig. 3b). Citrate and 2-oxoglutarate levels were not significantly different between drawing methods, while significantly higher levels of succinate (fold-change = 4, p < 0.01) were detected in the TAP samples (Fig. 3c). Blood obtained via TAP had higher biological CVs for alpha-ketoglutarate (131%) in comparison to traditionally drawn blood (58%—Supplementary Table 1).
Bile acids—either unconjugated or taurine and glycine conjugated—were extremely comparable between blood draws (Fig. 4), with median Spearman correlation coefficient > 95%. Still, traditionally drawn blood showed biological CVs as high as 138% for taurocholate, which instead had 52.5% biological CVs in blood drawn via TAP. Similar considerations could be made for free carnitine and acyl-conjugated carnitines, (Fig. 5), as well as palmitate and sphingosine 1-phosphate, averaging median correlations of 0.94.
Finally, in Fig. 6a and Supplementary Table 1 we provide a correlation matrix summarizing the results from Spearman correlations of matching measurements of the same compound in each one of the five subjects between traditional and TAP device blood draws. Median correlations across blood draws for all the tested metabolites was > 0.7. Representative correlation curves are shown for metabolites showing the highest linear correlations between drawing methods, carnitine (r = 0.984—Fig. 6b) and taurodexycholate (r = 0.991—Fig. 6c), and the lowest, palmitate, histidine and aspartate—(Supplementary Figs. 1, 2, Supplementary Table 1). Of note, concentration ranges for metabolites showing poor correlations were comparable between the two drawing strategies and only in the case of aspartate were impacted by increased CVs in TAP draws owing to two major outliers among the tested samples (Supplementary Table 1).
Correlation matrix of metabolite levels from absolute quantification measurements performed in blood drawn traditionally or via TAP (a). High positive correlations were observed form most metabolites (average r > 0.7) amino acids, carnitines (Spearman correlation r = 0.984) (b) and bile acids (r = 0.991) (c). A vectorial version of a with metabolite names is provided in Supplementary Fig. 1. Raw data and analyses are included in Supplementary Table 1
4 Discussion
TAP devices afford an easy, painless, logistically-feasible alternative to routine blood draws through venipuncture for collection of blood for metabolomics studies. However, little is known about the comparability of whole blood metabolic phenotypes following either drawing strategy. Here we provide a preliminary characterization of paired blood samples from five healthy donor volunteers. The study was limited by the availability of non-commercial prototypes of the TAP device tested in here. Despite the exploratory nature of the present analysis, we decided to focus on metabolites of relevance to the majority of the studies performed in our facility. In this view, we focused on amino acids, energy-related metabolites (including glycolytic and Krebs cycle intermediates), redox-related metabolites (including reduced glutathione), acyl-carnitines and bile acids. We report that results from either drawing strategy were comparable, with overall median correlation coefficients > 0.7 for all the tested metabolites and, for bile acids and acyl-carnitines r > 0.95.
Blood amino acid analysis (Nemkov et al. 2015) is a critical research endeavor in many fields, from biomedical research—e.g. serine and glycine in cancer research (Maddocks et al. 2016), tryptophan/kynurenine and arginine in “inflammaging” (Mondanelli et al. 2017), glutamine/glutamate/alanine in hemorrhagic hypoxia (Reisz et al. 2017), etc—to foodomics analysis—e.g. to potentially predict the outcome of muscle to meat conversion for meat tenderization processes (Herrero et al. 2012; Lana et al. 2015). Of note, the levels of most amino acids, except for valine and arginine (still quantified within a reasonable micromolar range), were comparable between blood draws strategy, suggesting an overall suitability of TAP-derived whole blood for amino acid-determination strategy.
Of note, metabolites related to redox homeostasis, such as cystine and reduced glutathione, but also succinate and urate differed significantly between blood drawing strategies. Of note, lower urate and higher succinate are observed in ischemic/hemorrhagic hypoxia prior to reperfusion-induced oxidation (Chouchani et al. 2014; D’Alessandro et al. 2015), suggesting an overall decreased level of oxidation for TAP-drawn blood. However, this introduces caveats about the potential need to recalibrate predictive ranges for clinical outcomes [e.g. succinate and mortality in critically injured trauma patients (DʼAlessandro et al. 2017; Lusczek et al. 2017)] in case blood draws were to be routinely performed through TAP-device based approaches. On the other hand, comparable measurements were obtained for glycemia and lactatemia, two key biochemical parameters of clinical relevance, for example in the diabetic population [e.g. fasting and non-fasting levels of circulating glucose—(Avignon et al. 1997)] and, in like fashion to succinate, in the critically ill and septic patient populations (DʼAlessandro et al. 2017; Krishna et al. 2009).
Our mechanistic understanding about the interconnection between host and microbiome metabolism has recently expanded (Palau-Rodriguez et al. 2015). Emerging concepts in this research endeavor highlighted the relevance in health and disease (e.g. inflammation) of blood metabolites like the choline-catabolite dimethylglycine, TMAO and bile acids, which can only be deconjugated by the gut microbiome and play a key role in inflammation and cholestasis (Kurbegov et al. 2003). Notably, these metabolites were comparably quantified in either blood drawing strategies, with correlation coefficients > 95%. Similar considerations can be made for circulating acyl-conjugated carnitines, key metabolites in the study of athletic performances and obesity (McCurdy et al. 2016). It is indeed in sports (e.g. outdoor) scenarios that a technology as versatile and easy to use as the TAP device would likely find its best application.
This study holds several limitations. First, a limited number of replicates were tested (five, limited by the availability of the hitherto not-commercially available prototype device—kindly and gratuitously offered by Seventh Sense). Further follow-up studies will be necessary to expand the metabolic coverage to other metabolites than those tested in this study, as well as to further verify the findings reported herein with orthogonal quantitative technology than the UHPLC-MS approach employed here. Discrepancies between some of the metabolites quantified (6 out of 45) may be due to biological rather than technical issues, such as the site of blood draw and potential stress on the drawn blood cells when forced through the TAP device microcapillaries. Furthermore, even though we carefully selected blood drawing areas as close as possible (spatially and temporally—i.e. paired blood draws on the same subjects were performed only minutes apart and in very close spots), it is likely that some of the observed differences were more attributable to blood source (e.g. vein vs. capillary blood). Finally, since the main focus of this study was to compare metabolic phenotypes of whole blood samples for potential streamlined clinical application, we did not sort plasma from cellular blood components. It is indeed worth noting that most of the metabolites quantified here are usually dosed in plasma/serum in most common databases (e.g. HMDB), which limits the comparability of the present findings with published repositories. In keeping with this consideration, we recently commented on the caveats associated with comparability between the metabolic phenotypes of plasma, whole blood and cellular components (e.g. red blood cells) (Gehrke et al. 2017). On the other hand, whole blood is a “richer” matrix that also informs on the levels of metabolites of cellular origin, such as oxidative stress markers like GSH.
5 Conclusion
Overall, with all the caveats and limitations pointed out above, our results indicate that the TAP device may be a viable strategy to perform routine blood draws in a painless, easy and logistically-friendly fashion. Blood obtained through the TAP device appears to be suitable for routine analysis of acyl-carnitines and bile acids, as well as glycemia, lactatemia and measurements of most amino acids of basic science or clinical relevance. For current clinical tests based on the analysis of small molecule compounds, large scale studies may be necessary to cross-validate existing ranges calibrated on traditional venipuncture-drawn blood or to prospectively validate new ranges calibrated directly on TAP device-drawn blood.
References
Avignon, A., Radauceanu, A., & Monnier, L. (1997). Nonfasting plasma glucose is a better marker of diabetic control than fasting plasma glucose in type 2 diabetes. Diabetes Care, 20(12), 1822–1826.
Beger, R. D., Dunn, W., Schmidt, M. A., Gross, S. S., Kirwan, J. A., Cascante, M., et al. (2016). Metabolomics enables precision medicine: “A white paper, community perspective”. Metabolomics, 12(9), 149. https://doi.org/10.1007/s11306-016-1094-6.
Blicharz, T. M., Gong, P., Bunner, B. M., Chu, L. L., Leonard, K. M., Wakefield, J. A., et al. (2018). Microneedle-based device for the one-step painless collection of capillary blood samples. Nature Biomedical Engineering, 2(3), 151–157. https://doi.org/10.1038/s41551-018-0194-1.
Chouchani, E. T., Pell, V. R., Gaude, E., Aksentijević, D., Sundier, S. Y., Robb, E. L., et al. (2014). Ischaemic accumulation of succinate controls reperfusion injury through mitochondrial ROS. Nature, 515(7527), 431–435. https://doi.org/10.1038/nature13909.
Crutchfield, C. A., Thomas, S. N., Sokoll, L. J., & Chan, D. W. (2016). Advances in mass spectrometry-based clinical biomarker discovery. Clinical Proteomics. https://doi.org/10.1186/s12014-015-9102-9.
D’Alessandro, A., Giardina, B., Gevi, F., Timperio, A. M., & Zolla, L. (2012). Clinical metabolomics: The next stage of clinical biochemistry. Blood Transfusion = Trasfusione Del Sangue, 10(Suppl 2), s19–24. https://doi.org/10.2450/2012.005S.
D’Alessandro, A., Moore, H. B., Moore, E. E., Wither, M., Nemkov, T., Gonzalez, E., et al. (2015). Early hemorrhage triggers metabolic responses that build up during prolonged shock. American Journal of Physiology. Regulatory, Integrative and Comparative Physiology, 308(12), R1034–R1044. https://doi.org/10.1152/ajpregu.00030.2015.
D’Alessandro, A., Nemkov, T., Yoshida, T., Bordbar, A., Palsson, B. O., & Hansen, K. C. (2017). Citrate metabolism in red blood cells stored in additive solution-3. Transfusion, 57(2), 325–336. https://doi.org/10.1111/trf.13892.
DʼAlessandro, A., Moore, H. B., Moore, E. E., Reisz, J. A., Wither, M. J., Ghasasbyan, A., et al. (2017). Plasma succinate is a predictor of mortality in critically injured patients. The Journal of Trauma and Acute Care Surgery, 83(3), 491–495. https://doi.org/10.1097/TA.0000000000001565.
Dettmer, K., Aronov, P. A., & Hammock, B. D. (2007). Mass spectrometry-based metabolomics. Mass Spectrometry Reviews, 26(1), 51–78. https://doi.org/10.1002/mas.20108.
Fuhrer, T., Heer, D., Begemann, B., & Zamboni, N. (2011). High-throughput, accurate mass metabolome profiling of cellular extracts by flow injection-time-of-flight mass spectrometry. Analytical Chemistry, 83(18), 7074–7080. https://doi.org/10.1021/ac201267k.
Gehrke, S., Reisz, J. A., Nemkov, T., Hansen, K. C., & D’Alessandro, A. (2017). Characterization of rapid extraction protocols for high-throughput metabolomics. Rapid Communications in Mass Spectrometry: RCM, 31(17), 1445–1452. https://doi.org/10.1002/rcm.7916.
Herrero, M., Simó, C., García-Cañas, V., Ibáñez, E., & Cifuentes, A. (2012). Foodomics: MS-based strategies in modern food science and nutrition. Mass Spectrometry Reviews, 31(1), 49–69. https://doi.org/10.1002/mas.20335.
Jobard, E., Trédan, O., Postoly, D., André, F., Martin, A.-L., Elena-Herrmann, B., & Boyault, S. (2016). A systematic evaluation of blood serum and plasma pre-analytics for metabolomics cohort studies. International Journal of Molecular Sciences. https://doi.org/10.3390/ijms17122035.
Kok, M. G. M., & Fillet, M. (2018). Volumetric absorptive microsampling: Current advances and applications. Journal of Pharmaceutical and Biomedical Analysis, 147, 288–296. https://doi.org/10.1016/j.jpba.2017.07.029.
Krishna, U., Joshi, S. P., & Modh, M. (2009). An evaluation of serial blood lactate measurement as an early predictor of shock and its outcome in patients of trauma or sepsis. Indian Journal of Critical Care Medicine, 13(2), 66–73. https://doi.org/10.4103/0972-5229.56051.
Kurbegov, A. C., Setchell, K. D. R., Haas, J. E., Mierau, G. W., Narkewicz, M., Bancroft, J. D., et al. (2003). Biliary diversion for progressive familial intrahepatic cholestasis: Improved liver morphology and bile acid profile. Gastroenterology, 125(4), 1227–1234.
Lana, A., Longo, V., Dalmasso, A., D’Alessandro, A., Bottero, M. T., & Zolla, L. (2015). Omics integrating physical techniques: Aged Piedmontese meat analysis. Food Chemistry, 172, 731–741. https://doi.org/10.1016/j.foodchem.2014.09.146.
Lewis, M. R., Pearce, J. T. M., Spagou, K., Green, M., Dona, A. C., Yuen, A. H. Y., et al. (2016). Development and application of ultra-performance liquid chromatography-TOF MS for precision large scale urinary metabolic phenotyping. Analytical Chemistry, 88(18), 9004–9013. https://doi.org/10.1021/acs.analchem.6b01481.
López-Bascón, M. A., Priego-Capote, F., Peralbo-Molina, A., Calderón-Santiago, M., & Luque de Castro, M. D. (2016). Influence of the collection tube on metabolomic changes in serum and plasma. Talanta, 150, 681–689. https://doi.org/10.1016/j.talanta.2015.12.079.
Lusczek, E. R., Muratore, S. L., Dubick, M. A., & Beilman, G. J. (2017). Assessment of key plasma metabolites in combat casualties. Journal of Trauma and Acute Care Surgery, 82(2), 309. https://doi.org/10.1097/TA.0000000000001277.
Maddocks, O. D. K., Labuschagne, C. F., Adams, P. D., & Vousden, K. H. (2016). Serine metabolism supports the methionine cycle and DNA/RNA methylation through De Novo ATP synthesis in cancer cells. Molecular Cell, 61(2), 210–221. https://doi.org/10.1016/j.molcel.2015.12.014.
McCurdy, C. E., Schenk, S., Hetrick, B., Houck, J., Drew, B. G., Kaye, S., et al. (2016). Maternal obesity reduces oxidative capacity in fetal skeletal muscle of Japanese macaques. JCI Insight, 1(16), e86612. https://doi.org/10.1172/jci.insight.86612.
Mondanelli, G., Bianchi, R., Pallotta, M. T., Orabona, C., Albini, E., Iacono, A., et al. (2017). A relay pathway between arginine and tryptophan metabolism confers immunosuppressive properties on dendritic cells. Immunity, 46(2), 233–244. https://doi.org/10.1016/j.immuni.2017.01.005.
Nemkov, T., D’Alessandro, A., & Hansen, K. C. (2015). Three-minute method for amino acid analysis by UHPLC and high-resolution quadrupole orbitrap mass spectrometry. Amino Acids, 47(11), 2345–2357. https://doi.org/10.1007/s00726-015-2019-9.
Nemkov, T., Hansen, K. C., & D’Alessandro, A. (2017). A three-minute method for high-throughput quantitative metabolomics and quantitative tracing experiments of central carbon and nitrogen pathways. Rapid Communications in Mass Spectrometry: RCM, 31(8), 663–673. https://doi.org/10.1002/rcm.7834.
Palau-Rodriguez, M., Tulipani, S., Isabel Queipo-Ortuño, M., Urpi-Sarda, M., Tinahones, M., Tinahones, F. J., & Andres-Lacueva, C. (2015). Metabolomic insights into the intricate gut microbial–host interaction in the development of obesity and type 2 diabetes. Frontiers in Microbiology. https://doi.org/10.3389/fmicb.2015.01151.
Reisz, J. A., Slaughter, A. L., Culp-Hill, R., Moore, E. E., Silliman, C. C., Fragoso, M., et al. (2017). Red blood cells in hemorrhagic shock: A critical role for glutaminolysis in fueling alanine transamination in rats. Blood Advances, 1(17), 1296–1305. https://doi.org/10.1182/bloodadvances.2017007187.
Townsend, M. K., Bao, Y., Poole, E. M., Bertrand, K. A., Kraft, P., Wolpin, B. M., et al. (2016). Impact of pre-analytic blood sample collection factors on metabolomics. Cancer Epidemiology, Biomarkers & Prevention, 25(5), 823–829. https://doi.org/10.1158/1055-9965.EPI-15-1206.
Uppal, K., Walker, D. I., Liu, K., Li, S., Go, Y.-M., & Jones, D. P. (2016). Computational metabolomics: A framework for the million metabolome. Chemical Research in Toxicology, 29(12), 1956–1975. https://doi.org/10.1021/acs.chemrestox.6b00179.
Wang, Q., Sun, T., Cao, Y., Gao, P., Dong, J., Fang, Y., et al. (2016). A dried blood spot mass spectrometry metabolomic approach for rapid breast cancer detection. OncoTargets and Therapy, 9, 1389–1398. https://doi.org/10.2147/OTT.S95862.
Wilson, I. (2011). Global metabolic profiling (metabonomics/metabolomics) using dried blood spots: advantages and pitfalls. Bioanalysis, 3(20), 2255–2257. https://doi.org/10.4155/bio.11.221.
Wishart, D. S., Feunang, Y. D., Marcu, A., Guo, A. C., Liang, K., Vázquez-Fresno, R., et al. (2018). HMDB 4.0: The human metabolome database for 2018. Nucleic Acids Research, 46(D1), D608–D617. https://doi.org/10.1093/nar/gkx1089.
Yin, P., Lehmann, R., & Xu, G. (2015). Effects of pre-analytical processes on blood samples used in metabolomics studies. Analytical and Bioanalytical Chemistry, 407(17), 4879–4892. https://doi.org/10.1007/s00216-015-8565-x.
Zukunft, S., Sorgenfrei, M., Prehn, C., Möller, G., & Adamski, J. (2013). Targeted metabolomics of dried blood spot extracts. Chromatographia, 76(19–20), 1295–1305. https://doi.org/10.1007/s10337-013-2429-3.
Acknowledgements
Research reported in this publication was supported in part by funds from the Boettcher Webb-Waring Biomedical Research Award—Early Career grant (ADA) and the Shared Instrument grant by the National Institute of Health (S10OD021641). We would like to thank Seventh Sense for kindly and gratuitously donatng prototype TAP devices for the sake of this pilot study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Though unrelated to the contents of this manuscript, ADA and TN are founders of Omix Technologies Inc and ALTIS Biosciences LLC. ADA is a consultant for New Health Sciences Inc. All the remaining authors have no conflicts of interest to disclose. No economic support was provided by Seventh Sense Biosystems, except for the kind donation of a limited number of prototypes tested in this study.
Electronic supplementary material
Below is the link to the electronic supplementary material.

Rights and permissions
About this article
Cite this article
Catala, A., Culp-Hill, R., Nemkov, T. et al. Quantitative metabolomics comparison of traditional blood draws and TAP capillary blood collection. Metabolomics 14, 100 (2018). https://doi.org/10.1007/s11306-018-1395-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11306-018-1395-z