Abstract
Carob (Ceratonia siliqua L.) is an important evergreen tree of the Mediterranean landscape. Its economic interest is increasing thanks also to the presence, in the seeds, of the locust bean gum (LBG), a galactomannan largely used by the food industry as a stabilizer and thickening agent. Its economic and ecological values make the understanding of carob genetic diversity of great interest both for breeding and conservation purposes. The world’s largest carob germplasm collection was genotyped using both eight carob-specific nuclear short sequence repeat (nSSR) and the sequencing of a chloroplast locus. The collection is composed of 215 accessions introduced from 12 countries of origin spanning from traditional to novel areas of cultivation. To assess the genetic diversity of the collection, several approaches were coupled: structure analysis, principal component analysis (PCA), and graphic clustering either from dissimilarity data and coancestry data. Structure analysis suggested the presence of two distinct genetic pools: one characterizing northeastern Spain and the second spread in other countries and southern Spain. The PCA and discriminant analysis of principal component (DAPC) complemented the structure results allowing a better understanding of the genetic differences between countries while the network-joining analysis provided additional insights on the similarity between individuals. Short sequence repeat (SSR) data coupled with phenotypic data (floral sex and status) were also used to define the first core collection of carobs. Multi-approach analysis of genetic diversity together with the definition of a core collection represent useful tools for the setup of genetic-guided intervention both for conservation and breeding purposes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Carob (Ceratonia siliqua L.) is a diploid species (2n = 24) that belongs to the Fabaceae family, sub-family Caesalpinioideae (Goldblatt 1981; Arista and Talavera 1990). Carob is an evergreen, dioecious tree and it is a constituent of the Mediterranean thermophilus woodlands vegetation (Zohary 2002). Carob is widespread in all Mediterranean countries in different pedoclimatic environments, making this plant one of the most characteristic of the coastal Mediterranean landscape in which it is particularly common in marginal areas due to its lower agronomic requirements compared to other fruit species (Batlle and Tous 1997; Caruso et al. 2008a; Tous et al. 2013).
Ceratonia is one of the most archaic genera within the Fabaceae family (Tucker 1992), its center of origin is still debated varying from Turkey and Syria (Vavilov 1951) to southern Arabia (Schweinfurth 1894) or Indo-Malesia (Zohary 1973). Written records related to carob cultivation can be found in the Mishnah (first century BCE–second century CE) and the Talmud (third–fifth centuries CE) in which a clear distinction is made between carob cultivars used as human food (grafted) or animal feed (non-grafted). In fact, the cultivation of the species took advantages by the diffusion of selected genotypes by grafting methods starting about 3000 years ago (Meyer et al. 2012).
The analysis of paleontological and archeological evidences clearly indicated that carob is native to the east Mediterranean area and its presence can be dated back long before the emergence of horticulture (Zohary 2002). More recently, Viruel et al. (2016) confirmed the hypothesis of natural distribution of carob in the Mediterranean before the emergence of agriculture, hypothesizing a multiple-origins model of domestication from native populations. The diffusion of carob to all Mediterranean countries occurred during Greek and Arabs civilizations; in particular, Greeks contributed for the first spread of this species to Greece and southern Italy, while centuries later, Arabs extended the area of cultivation along the coast of northern Africa into the south and east of Spain (Batlle and Tous 1997). The domestication of this species took place relatively late (only in Roman times), as carob tree is not suitable for simple vegetative propagation (cuttings, suckers, etc.), and cultivation of selected clones had to wait until the introduction of scion grafting into the Mediterranean basin (Zohary 2002).
Carobs have been extensively grown for their fruits. The pod is a sweet bean mainly constituted by pulp (90%) and seed (10%). The pulp has been traditionally used both for human consumption and for animal feeding (Coit 1951; Sahle et al. 1992; Batlle and Tous 1997) and selection effort was consequently made to increase the pulp/seed ratio. Under domestication, heavy-fruiting cultivars have been improved; bearing larger, fleshier, and sweeter pods (Zohary 2002). Currently, the most valuable component of the carob pod is the seed, its endosperm contains the galactomannan locust bean gum (LBG) characterized by an outstanding water-binding capacity (Batlle and Tous 1997). LBG is widely used in the food industry as a stabilizer and thickening agent due to its high viscosity in water over a wide range of temperature and pH (Garcia-Ochoa and Casas 1992). Apart from its increasing economic value, carob tree can be used to reforest arid and marginal areas and for landscaping (Winer 1980).
The presence of C. siliqua varies across geographical areas, while in eastern Mediterranean and North Africa carobs grow mainly wild; in Western Mediterranean countries, carob tree is mainly present in cultivated form (Batlle and Tous 1997; La Malfa et al. 2014; Tous et al. 2013). Traditionally, carob plantations have been established by field budding of scions from selected genotypes (mostly female, and to a lesser extent, hermaphrodite) onto seed rootstocks. Nevertheless, the increase in bud propagation together with the relative lower economic interest compared to other fruit crops have contributed to a severe genetic erosion (La Malfa et al. 2014). To this extent, the collection, characterization, and maintenance of carob germplasm are of pivotal interest. Currently, there are only a few studies investigating the level of genetic diversity existing in carob tree through molecular markers, while to the best of our knowledge, no studies focused on marker-trait association nor on the setup of a core collection. RAPD (Random Amplified Polymorphic DNA) and AFLP (Amplified Fragment Length Polymorphism) markers have first been used to highlight the genetic diversity within carob trees (Talhouk et al. 2005; Konate et al. 2007; Caruso et al. 2008a). The first set of carob tree short sequence repeat (SSR) marker was developed by Caruso et al. (2008b) from floral gene sequences (Expressed Sequence Tags, ESTs). This set of markers was used to elucidate the genetic differences of 71 cultivars collected in Italy, Malta, and Spain (La Malfa et al. 2014).
A germplasm core collection consists of a limited set of accessions, within a larger pool, chosen on the basis of either phenotypic or genomic variability to represent the maximum genetic variation of the crop with minimum repetition. In carob, as well as for all other tree species, genetic resources maintained in germplasm collections are sampled to obtain a subset of accessions to preserve as much of the original diversity as possible thus reducing the costs related to the maintenance. Basic goals of a core collection are to introduce, maintain, document, and make accessible representative genetic diversity of the crop of interest. The selection of a core collection can be based on molecular data and/or phenotypic traits (Thachuk et al. 2009). Several algorithms have been proposed to guide the selection of a core subset, some, such as the Modified Rogers (MR, Wright 1969) and Cavalli Sforza and Edwards (CE, Cavalli-Sforza and Edwards 1967) are based on genetic distance while other methods, such as Shannon’s diversity index (SH, Shannon 1948) or the expected proportion of heterozygous loci (HE), are instead based on genetic diversity calculation. Genetic distance estimation methods are based on a pairwise measure of the degree of dissimilarity between 2 samples; the larger the genetic distance, the more genetically different the 2 samples are. On the other side, genetic diversity takes into account several descriptors parameters (such as expected PIC or expected heterozygosity) in order to evaluate the overall genetic variability of a sample as a whole (Thachuk et al. 2009).
In this study, both nuclear SSR and the sequencing of the matK4LF locus of the chloroplast genome were used to genotype a carob tree collection composed by plants sampled in 12 different countries spanning from traditional areas of cultivation (Mediterranean countries) to countries such as Australia and the USA in which the crop was introduced in the last century (Batlle and Tous 1997). The assessment of the genetic diversity within the germplasm collection was also used for the definition of a core collection (the first ever published for carob) within the largest germplasm repository of carob tree worldwide.
Materials and methods
Plant material
The carob germplasm collection used in this study was composed by 215 accessions held “ex situ” as trees grafted on seedling rootstocks in the field at the IRTA Mas Bové, Constantí (Tarragona, northeastern Spain, lat. 41° 10′ 9″ N, long 1° 10′ 28″ E, altitude 110 m). This collection was initiated in 1985 by IRTA and includes accessions covering the most important areas of cultivation of carob tree. Along the years, cultivars and wild types were collected gathering as much as possible the existing carob diversity. Individuals were collected from all Spanish growing regions (Batlle and Tous 1994, 1997; Tous et al. 2008) and other producing countries through surveys like in Tunisia (Tous et al. 2006) and Morocco and/or provided by different international institutions (Tous et al. 2008). As for the productive attitude, female and hermaphrodite accessions were preferred over males. Currently, this carob germplasm collection represents the largest repository of cultivated material in the world (Batlle and Tous 1997; Tous et al. 2013). This plant material includes accessions from 12 different countries: Algeria (1), Australia (7), Croatia (2), Cyprus (3), Israel (1), Italy (8), Morocco (24), Portugal (6), Spain (142), Tunisia (15), Turkey (4), and the USA (2). The name, origin, sex, and status of each accession are provided in Supplementary Table 1.
DNA extraction and genotyping
The DNA extraction was performed on young leaves; 100 mg of fresh tissue was grounded using Qiagen Tissuelyser II (Qiagen, Hilden, Germany) and genomic DNA was extracted using isolate Plant DNA Mini Kit (Bioline, London, UK). Quantity and quality of DNA extracted were assessed with electrophoresis on 1% agarose gel and with a spectrophotometer (Nanodrop 2000, Thermo Scientific, Waltham, MA, USA). Genotyping was performed through the analysis of eight nuclear SSR markers and the sequencing of a chloroplastic locus. The eight SSRs used in this study were selected on the base of their performances from a set of 11 EST-SSRs developed by Caruso et al. (2008a). The primer sequences of the SSRs were retrieved from the same paper. The amplification of the eight SSRs was carried out in 15-μl volumes with 30 ng genomic DNA, 1X NH4 reaction buffer (Bioline), 0.2 mM dNTPs (Bioline), 1 mM MgCl2, 0,167 nM of forward and reverse primer, 0.13 nM of M13F labeled with a fluorescent dye (FAM, PET, NED, or VIC; MWG Operon, Ebersberg, Germany), and 1 U of Taq Polymerase (Bioline). Amplification was performed in two thermal cycler GeneAmp PCR system 9700 and 2700 (Applied Biosystems, Foster City, USA) with an initial denaturation at 94 °C for 5 min, followed by 40 cycles of 94 °C for 30 s, followed by 30 s at 56 °C and 60 s at 72 °C with a final extension at 72 °C for 30 min. An aliquot of 0.4–0.6 μl of PCR product (depending on the performance of amplification of each primer pair) was mixed with 13 μl of formamide and 0.3 μl of LIZ-500 size standard and denatured at 95 °C for 5 min. Up to three PCR products labeled with FAM, PET, NED, or VIC were pooled before separation in the ABI 310 Genetic Analyzer (Applied Biosystems), amplicons were visualized and analyzed using Genemapper® software 4.0. All the samples were analyzed with the same methodology and in the same conditions to make the results comparable. Sequence analysis of chloroplastic locus matK4LF was performed according to the information kindly provided by Dr. Alex Baumel, Aix Marseille University, Marseille, France. After quantification, amplicons were directly sequenced by Eurofin (Eurofins MWG Synthesis GmbH, Anzinger Straße 7a, D-8556 Ebersberg, Germany).
Statistical analysis
Prior to genetic analysis, the germplasm was screened for the presence of putative clones running an identity analysis using Cervus software (Kalinowski et al. 2007). Major allele frequency, number of genotypes detected, number of alleles per locus, observed and expected heterozygosity (Ho and He respectively), and polymorphism information content (PIC) were determined using Cervus software (Kalinowski et al. 2007). Genetic data were also used to compute a genotypic accumulation curve using the “poppr” R package (Kamvar et al. 2014; R Core Team 2016). Pairwise FST values for samples collected in the same country/region were calculated using Genepop software (Raymond and Rousset 1995). The level of genetic stratification within the carob collection was assessed with STRUCTURE v2.3.1 (Pritchard et al. 2000). An admixed model with 500.000 burn-in period followed by 500.000 iterations was computed. Each Markov chain simulation was run ten times for a number of subpopulations (k) ranging from one to ten. The most probable k was identified and visualized with Structure Harvester (Earl and vonHoldt 2012) following the ΔK method proposed by Evanno et al. (2005). Replicates were integrated with CLUMPP software (Jakobsson and Rosenberg 2007) using the “full search” algorithm and default settings while the Q matrix was plotted using an in-house R script (available upon request). Samples were assigned to a subpopulation when the membership coefficient Qi was greater than or equal to 0.8. Principal component analysis (PCA) was performed using the “prcomp” function of the “stat” package in R (R Core Team 2016), while PCA plots were produced using either the “Factoextra” package (Kassambara and Mundt 2016) or the in-house R scripts (also available upon request). “Factoextra” package was also used to compute and visualize the dissimilarity matrix through the generation of a heatmap and a neighbor-joining dendrogram. Discriminant analysis of principal component (DAPC) has been performed using “adegenet” package in R (Jombart 2008). Coancestry between cultivars was assessed using COANCESTRY software (Wang 2011) while the “igraph” R package (Csardi and Nepusz 2006) was used to display the unweighted and undirected network generated on the TrioML values.
Definition of the core collection
The construction and composition of the core collection were calculated on the obtained genetic data using the R package “corehunter” (De Beukelaer et al. 2018). The number of individuals selected for the core collection (sampling intensity) was 20%, in agreement with previous works (van van Hintum 1999; van Hintum et al. 2000; Thachuk et al. 2009). The analysis was carried out using the average entry-to-nearest-entry distance method (EN). Matrix of distances was calculated for genotype and phenotypes using the Modified Rogers (MR) and Gower distances (GD) algorithms respectively as implemented in the corehunter package. EN maximizes the average distances between a given individual and another selected item of the core collection. The rationale is to include in the core the individuals that are sufficiently different from the most similar individual already included in the core collection.
Results
Carob collection
The germplasm collection was selected to encompass the most important areas of cultivation of carob. All individuals were phenotyped for the sex of the flower and the status. To this extent, 159 individuals showed monosexual flower (156 females, 3 male), while the remaining 44 showed hermaphrodite flower. On the other side, 75 and 128 plants were considered cultivar or wild type respectively, based on their status (Supplementary Table 1). The relative frequency of the two status within individuals showing a female flower was similar (44% cultivar and 56% wild) while a more skewed proportion was registered for carobs showing hermaphrodite (16% cultivar and 84% wild) and male flowers (100% wild) (Supplementary Table 1).
Genotyping of the carob collection
The 215 accessions composing the carob germplasm collection were genotyped both with eight nuclear SSRs and through the sequencing of the matK4LF locus of the chloroplast genome (Supplementary Table 2). The genotyping analysis allowed the detection of 12 putative clones that were subsequently excluded from the analysis (Supplementary Table 1) leading to a final number of 203 unique individuals. The SSRs detected a total of 43 alleles. The allelic richness estimation across the whole dataset allowed efficient discrimination of the 94% of the samples as showed by the genotypic accumulation curve displayed in Supplementary Fig. 1. The mean number of alleles per loci and genotypes were 5.38 and 10.25 respectively (Table 1). Marker AT15 showed the highest allelic and genotypic variability (9 alleles and 29 unique genotypes) while markers TA5TG6 and TA7 showed the lowest number of alleles (3) forming 5 and 4 genotypes respectively. The mean He and Ho across the eight SSRs were 0.47 and 0.49; TTA7 showed the highest Ho (0.83) while the lowest value (0.13) was registered for CTTT7 (Table 1). Thus, it is not surprising that the latter marker showed the lowest values for He (0.14) polymorphic index content (PIC, 0.13) across the whole set of SSR used. In contrast, the highest PIC (0.77) and Ho were detected for marker AT15 (Table 1). When only alleles having a minor allelic frequency (MAF) higher than 5% were considered, the mean number of alleles and genotypes was 3 and 6.62 respectively. The highest changes in the absolute value of Ho, He, and PIC, compared to the complete dataset, were detected for marker GA12 that showed a decrease of 0.5 for the three parameters (Table 1).
To better assess the genetic diversity and to investigate the role of maternal contribution in the carob collection, the matK4LF locus of the chloroplast DNA was sequenced resulting in a 668 bp amplicon. The sequence alignment allowed the detection of two haplotypes originated by the presence of a T/G polymorphism located approximately 622 bp downstream (Supplementary Fig. 2, Supplementary Table 2). The 80% of the samples presented the first haplotype (G allele, hereafter called hap-1) while the remaining showed the presence of the alternative haplotype (T allele, hap-2). The relative frequency of the two haplotypes did not change significantly across different countries (data not shown).
To assess the genetic diversity of the carob germplasm three approaches have been used: Bayesian clustering, multivariate analysis, and graphic clustering either from dissimilarity and coancestry data.
Structure
The eight SSRs were used to highlight the presence of genetic stratification within the carob collection. To this extent, the most probable value of k detected by STRUCTURE software was two (ΔK = 354, Supplementary Fig. 3a). Figure 1a showed the genetic configuration of the 203 individuals composing the germplasm collection, all samples were grouped according to the country of origin, Spanish samples (65% of the whole germplasm) were instead grouped according to their region of provenience. Averaging the individuals’ membership coefficients by countries, Subpop 1 resulted predominant (average value higher than 0.8) in accessions collected in Croatia, Cyprus, Morocco, Tunisia, and the USA, while no countries showed a decisive prevalence of Subpop-2 (Supplementary Table 3, Fig. 2a). Spanish germplasm showed an average incidence of the two subpopulations equal to 39% and 61% respectively (Supplementary Table 3), while when samples were grouped according to their region of origin a much clear genetic differentiation was evident (Figs. 1a and 2a, Supplementary Table 4). Accessions originated from Andalusia showed a predominance of Subpop1 (mean value of 0.76) while samples collected in Catalunya, Valencia, Murcia, and the Balearic Islands showed a prevalence of Subpop 2 with mean values ranging from 0.68 to 0.93 (Fig. 2a, Supplementary Table 4). Subpop 2 is therefore located predominantly in the northeastern part of Spain while all other countries and Andalusia showed a clear predominance of Subpop 1 (Fig. 2a). This finding was confirmed by the examination of the pairwise FST value: the highest FST among Spanish regions was calculated for Catalunya VS Andalusia (FST = 0.156), followed by Valencia VS Andalusia (FST = 0.117). Comparing the results obtained with STRUCTURE with phenotypic data, we found that carobs with hermaphrodite flowers were mostly classified as Subpop 2 or admixed (70% and 23% respectively) while only three samples with hermaphrodite flowers belong to Subpop 1 (Supplementary Table 1). On the contrary, the 94% of samples that showed a clear predominance of Subpop 1 showed female flowers (Supplementary Table 1 and Fig. 3) (Pearson’s chi-square test = 41.263, df = 4, p value = 2.371e-08). When the status is taken into account, the differences between the subpopulations defined by structure are less evident: the 72% of samples having a prevalence of Subpop1 had a wild status while the proportion changed to 53% for samples assigned to Subpop 2 (Pearson’s chi-square test = 5.271, df = 2, p value = 0.07). When the country/region of origin are used as a factor to differentiate plants according to sex, we found that 87% of the hermaphrodite accessions were collected in Catalunya, Valencia, Murcia, or the Balearic Islands, while the remaining 13% were sampled in Andalusia, Italy, Tunisia, and the USA (Supplementary Table 1, Supplementary Fig. 4) (Pearson’s chi-square test = 35.728, df = 1, p value = 2.268e-09). When the habitus is considered the proportion of cultivated carobs in northeastern Spain is significantly higher than the other countries/region (Pearson’s chi-square test = 18.962, df = 1, p value = 1.333e-05) The comparison between the two haplotypes detected through the sequencing of the matK4LF locus of the chloroplast did not highlight significant relations with the structure results (data not shown).
The second most probable number of subpopulations (ΔK = 20.18) was k = 4 (Supplementary Fig. 3a). When four subpopulations were postulated, the difference between the genetic stratification of samples collected from Balearic Islands, Catalunya, and Valencia become more evident (Figs. 1b and 2b): the relative incidence of the subpopulation depicted in orange decreased progressively from Catalunya to Valencia and Balearic Islands, while at the same time, the contribution of the green and blue subpopulations increased. Differences between Morocco, Andalusia, and Tunisia (all showing similar incidences of Subpop1 and Subpop2 when the k = 2 model was considered, Supplementary Tables 3 and 4) were more easily detected. Among the three countries/regions, Tunisia showed the highest and lowest contribution of the “yellow” and “green” subpopulations respectively, Andalusia showed clear predominance of the green subpopulation over the yellow one, while Morocco showed a more balanced contribution of the two subpopulations. In all countries showing a predominance of Subpop1 on a k = 2 model (Supplementary Table 3), the incidence of the orange subpopulation was neglectable (Figs. 1b and 2b).
To further highlight the genetic structure within the subpopulations detected with a 2K model, a second round of STRUCTURE was carried out on two smaller datasets including only those individuals showing a membership coefficient higher than 0.8 for one of the two subpopulations. This resulted in the definition of two subsets composed by 73 (subset 1, individuals showing a predominance of Subpop 1) and 58 (subset 2, individuals showing a predominance of Subpop 2) samples. Subset 1 was composed of individuals coming from all countries except Algeria and Israel, while all Spanish samples but 1 were collected in Andalusia (Fig. 4, Supplementary Table 1). The k showing the highest probability for subset 1 was three (ΔK = 16, Supplementary Fig. 3b). Subset 2 was composed only by individuals coming from Spain, Italy, and Turkey and conversely to what was observed for subset 1; subset 2 did not show clear genetic stratification as highlighted in Supplementary Fig. 3c.
Coancestry between carobs cultivars
For the vast majority of the samples composing the carob collection, there were no pedigree records available. This can be reconducted to the long intergenerational time proper of carob and to the low economical interest for this species compared to other crops such as apple or peach for which extensive pedigree records were reconstructed (Aranzana et al. 2003; Howard et al. 2017) and often used for research and breeding purposes (Bink et al. 2014; Fresnedo-Ramírez et al. 2016; Di Guardo et al. 2017). The lack of pedigree records is a common issue for most Mediterranean tree species; recently, the triadic likelihood estimator (TrioML) was successfully used to get insight on the level of coancestry (r) within a dataset of olive varieties (Diez et al. 2015). Figure 5a shows the relative frequency of the r values calculated for all pairwise comparisons within the carob germplasm collection. The majority of the pairwise comparisons showed an r value lower than 0.05 (55.5%). The r values higher than 0.05 were gradually less frequently moving toward r = 1 with the exception for values comprised between 0.45 and 0.5 showing a much higher frequency than the flanking bins (Fig. 5a). An r value close to 0.5 ideally stands for first-degree relatives, this finding is confirmed in our case by the examination of the r values of the few samples for which a priori knowledge of parent-child relationship was available, for which the lowest r value was 0.5 (data not shown). This value was then used as cut-off to identify first-degree relationship among the carob collection. Interconnection between individuals showing coancestry values higher than 0.5 was depicted in a network analysis Fig. 5b.
Multivariate analysis
The genetic structure of the carob germplasm collection was also investigated using a multivariate approach coupling SSR with chloroplast sequencing data. Together, the first two principal components (PC1 and PC2) explained the 24.8% of the total genetic variability as shown in Fig. 6. The individual values of PC1 corroborate the results obtained by STRUCTURE: samples with positive value of PC1 showed a decisive contribution (membership coefficient > 0.8) of Subpop 1 of STRUCTURE, while individuals with negative values of PC1 belong to Subpop 2. Individuals with PC1 values close to 0 were deemed as admixed, slightly positive, or negative values of PC1 indicates a prevalence of Subpop 1 or Subpop 2 respectively. Genetic data were also used to compute a DAPC analysis using the information on the country/region of origin as group membership factor. Samples showing a clear prevalence of Subpop 2 on the STRUCTURE analysis clustered together on the bottom-right quadrant of the DAPC (Supplementary Fig. 5) while carobs collected in Andalusia were grouped with the samples collected in Turkey and Morocco on the upper-right quadrant. Samples collected in Italy and Algeria were clustered in the bottom-left quadrant forming a separate cluster (Supplementary fig. 5).
Neighbor-joining analysis
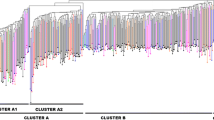
The genetic relationship between individuals was investigated through the construction of a dissimilarity matrix used to compute a neighbor-joining dendrogram (Fig. 7). The 94% of the samples were efficiently discriminated while the remaining 6% includes samples that were not efficiently discriminated by our set of markers due to the presence of missing data. The cluster analysis identified three main groups (Fig. 7) each composed by 122, 80, and 1 samples respectively. Cluster 1 is composed of 56% by individuals belonging to Subpop 1, 40% of the accessions showed instead an admixed genetic configuration while the remaining 4% were samples characterized by a predominance of Subpop 2. On the contrary, cluster 2 grouped mainly individuals with a predominance of Subpop 2 (69% of the total) while the27% and 4% were composed by samples with admixed or Subpop 1 genetic configuration respectively. Cluster 3 is composed only by the carob “T-1,” this sample is characterized by 2 unique alleles at SSR GA12 and TTA7.
Core collection
A total of 41 accessions were selected to compose the core collection (Supplementary Table 1). The selection of the samples has been made using the EN method and both genotypic and phenotypic data were considered. The Ho was slightly lower than in the whole germplasm (0.47), while the PIC was higher in the core collection compared to the whole dataset (0.48 and 0.42 respectively). All three subpopulations detected in structure analysis were represented in the core collection. In particular, 17 accessions from Subpop 1, 6 from Subpop 2, and 18 from admixed were included representing41, 15, and 44% of the total of each group, respectively. All floral sex and status were included in the core collection: the vast majority of the carobs were female (93%) while hermaphrodite and male plants were 5% and 2% respectively while the two status were represented almost in equal measure (54% wild type and 46% cultivated plants).
Discussion
The vast majority of the individuals composing the collection was composed of female or hermaphrodite plants (98.6%). This choice can be reconducted to two considerations: (i) usually carobs showing male flowers do not show significant differences in terms of phenotypes, (ii) since the collection is aimed to assess carob productivity preference has been given to female and to hermaphrodite accessions being the latter used both as pollinators and for pod production.
To better highlight the genetic differences between accessions and to guide the selection of a core collection, two types of markers were used: SSR and the sequencing of the matK4LF locus of the chloroplast genome. The eight SSRs used were ad hoc chosen based on the results presented by La Malfa et al. (2014). The number of alleles (and genotypes) detected in the analyzed carob collection was higher than what was reported by La Malfa et al. (2014) for seven out of eight SSRs; only marker TA5TG6 (previously named Cesi_976_ta5tg6) showed an equal number of alleles and genotypes (3 alleles forming 5 genotypes). These results can be explained by the intrinsic differences in size and geographical areas covered between the two carob collections, while the present collection is composed by 215 genotypes collected in 12 countries, the previous work was composed by 71 cultivars sampled in three countries.
On the whole, the set of markers used in this study allowed to univocally discriminate 203 out of 215 (94.4%) individuals of the analyzed collections. The occurrence of the 12 putative clones could be due either to their clonal origin or to the subtle genetic differences that were undetected by our set of markers (Table 1). The availability of high throughput genotyping platforms could unravel further genetic variability within the current putative clones. The picture about this characterization is also confirmed by the results of genotypic probability curve analysis and the neighbor-joining analysis (Supplemental Fig. 1, Fig. 7) in which the 94% of samples were efficiently discriminated, while the identification of the remaining individuals was hampered by the occasional occurrence of missing data.
The two haplotypes detected within the chloroplast locus matK4LF were evenly distributed not only across countries but also between the two subpopulations detected by STRUCTURE. This result suggests that the T/G mutation might occur before the 1000 A.C. prior to the first diffusion of carobs across the Mediterranean countries made by Greek. Structure analysis detected a genetic model with k equal to 2 or 4 as the most probable number of subpopulations (Supplemental Fig. 3a), the same number of subpopulations was also detected by Viruel and colleagues (Viruel et al. 2018). In both cases, it was possible to detect peculiar genetic stratification according to the area of origin. This is of particular importance since previous works using isoenzymes markers did not find pattern of geographical variation (Tous et al. 1992). The high number of samples collected in Spain allowed a finer differentiation of the carobs according to the region of collection without losing significant statistical power. When a structure model with k = 2 was tested, while all countries showed a decisive prevalence of Subpop 1, Spain had the highest frequency of Subpop 2 both in relative terms (61%) and as number of individuals (96% of the samples within subset 2, Supplementary Table 1). The decisive incidence of Subpop 2 in Catalunya, Valencia, Murcia, and the Balearic Islands, and the subsequent low incidence in Andalusia (Supplementary Table 4) suggested the occurrence of two different genetic pools: one, Subpop 1, characterizing the carob cultivars collected in southern Spain while a second, Subpop 2, is common between the northeastern regions of Spain. A prevalence of Subpop 1 was also detected in all the other Mediterranean countries analyzed in this study (Fig. 2a) suggesting a common past of diffusion between those countries and southern Spain. We also tested a second model based on the identification of four subpopulations (Fig. 2b). Overall, the results still pointed to two subpopulations model with samples collected in Balearic Islands, Catalunya, Murcia, and Valencia forming a separate genetic cluster compared to the other countries/region. Nevertheless, distinctions within the northeastern regions were more evident and Catalunya exhibited a much peculiar genetic configuration showing a prevalence of the orange subpopulation. The nested structure analysis performed on the outcome of the structure k = 2 model highlighted even a finer definition of the genetic structure of individuals with decisive prevalence of Subpop 1. To this extent, carobs collected in Andalusia showed a significant presence of subpopulation 1c that was almost absent in all other countries except the neighboring Morocco (Supplementary Table 1, Fig. 4). This is not surprising since Andalusia and Morocco shared many similarities in terms of phenotypes, origin, and agricultural and forestry practices. The size and quality of pods of carobs originated from those two countries are similar to wild type exhibiting a low pulp content and high seed/pulp ratio, while carobs originated from northeastern Spain showed a higher relatedness to cultivated carob pods. This can be explained by the fact that lower breeding efforts have been made on carobs from Andalusia and Morocco compared to northeastern Spain. Moreover, in Andalusia and Morocco, carob crop intensity was lower compared to other Spanish growing regions and plants are often propagated by seeds rather than by grafting (Sidina et al. 2009; Tous et al. 2013). Interestingly, the group of “Benajarafe” accessions selected in Andalusia (18, representing 42% of the whole Andalusian genotypes) were again trees grown from seeds (mixture of cultigens “Cajiz,” “Ojen-Cajiz,” and “Sayalonga”) or ungrafted seedlings. These entries were named “Benajarafe” (from the name of Malaga’s coastal village where budwoods were collected from). Five of them (28%) were part of the core collection confirming the genetic diversity within this group.
Within the Spanish germplasm, the occurrence of hermaphrodite specimens was significantly higher in Subpop 2 (including plants collected mainly in Catalunya, Valencia, and the Balearic Islands) in relation to the others (Subpop 1 and admix). The higher frequency of hermaphrodite forms in those regions is likely correlated with a high presence of cultivated fields in which the presence of hermaphrodites’ varieties or selections has been favored by growers and also due to the deeper exploitation of their productive potential carried out by IRTA’s researchers especially in the three above-cited regions (Zohary 2002).
The PCA, DAPC, coancestry analysis, and the neighbor-joining analysis were in agreement with these findings highlighting a strong genetic differentiation between Catalunya, Valencia, Murcia, and the Balearic Islands on one side and all other countries/region on the other with a subtle differentiation, within the Subpop 1, between Andalusia and all other countries (Figs. 5 and 6, Supplementary Table 1, Supplementary Fig. 5).
Taken together, these results straighten the theory that carob tree originated from the near east (Schweinfurth 1894; Vavilov 1951; Zohary 1973; Zohary 2002) and then it has been spread westward along the coasts of northern Africa till the southern and eastern of Spain (Batlle and Tous 1997). The presence of a different genetic pool in northeastern Spain might be due to a significative diffusion of the cultivation coupled with a reduction in the commercial exchanges between northeastern Spain and all other countries except Portugal. The definition of such a germplasm collection is a first step for the set-up of marker-trait association analysis (genome-wide association studies or genetic selection) to link the observed genotypic variability to a specific trait of interest. The identification of an association between a trait and a polymorphic locus will allow the set-up of the first marker-assisted selection (MAS) approach in carob.
In addition, the definition of a core collection allowed the selection of a subset of individuals accounting for the highest diversity both in terms of phenotype and genotype. Among the algorithms currently employed to define a core collection, the EN method has been widely used when both genetic and phenotypic diversity is taken into account (Thachuk et al. 2009). This is relevant for our conditions in which a couple of basilar phenotypic traits of agronomic value have been taken into account to guide, together with genotypic data, the definition of core collection. Within the 41 entries of the core collection, 18 belong to Subpop 1, 11 to Subpop 2, and 12 were considered admixed. The relatively lower number of individual belonging to Subpop2 supports their closer genetic relationships compared to Subpop1. This is confirmed by the second round of structure that did not highlight the presence of significant genetic stratification within Subpop 2, while three subpopulations were detected for Subpop 1 (Supplementary Fig. 2 B-C). Only 19 entries of the total core collection (46%) can be considered cultivars stricto sensu while the remaining were unnamed accessions (mostly seedlings from wild or feral origin) which provide an insight on the carob genetic diversity and composition. As in other evergreen species like olive (Díez et al. 204), we found admixture to be an important component of carob’s genetic history.
Similarly to other fruit tree species like olive, grapevine, cherimoya, and pear (Escribano et al. 2008; Cipriani et al. 2010; Miranda et al. 2010; Belaj et al. 2012; Diez et al. 2015), a low sampling intensity (20% of the whole collection) was sufficient to efficiently represent the genetic diversity found in the whole collection. Although accessions from Spain represented 64% of the total germplasm collection, only 53% were part of the core collection. A similar relative frequency was found for accessions collected in Tunisia in which, among the 15 accessions, 8 were chosen to form the core collection. Regarding sex representation within the core collection, only 2 accessions out of 41 were hermaphrodite (5%) and 1 male.
This paper is aimed to document the genetic diversity and structure of Ceratonia siliqua L. by combining two different types of molecular data and some phenotypic traits to guide the definition of a core collection. Multi-approach analysis of genetic diversity together with the definition of a core collection represents a useful tool for genetic-assisted conservation and breeding protocols. Future studies may also give interesting insights about the representation of phenotypic carob diversity (morphological traits, seed yield, resistance to disease and pest, seed and gum quality components, etc.) into the selected core subset in comparison to the whole collection.
References
Aranzana MJ, Carb J, Ar P (2003) Microsatellite variability in peach [ Prunus persica ( L .) Batsch ]: cultivar identification , marker mutation , pedigree inferences and population structure. 1341–1352. doi: https://doi.org/10.1007/s00122-002-1128-5
Arista M, Talavera S (1990) Números cromosómicos para la flora española. Lagascalia 16:323–328
Batlle I, Tous J (1994) Carob tree germplasm in Andalusia (Spain). NUCIS Newsl
Batlle I, Tous J (1997) Carob tree. Ceratonia siliqua L. promoting the conservation and use of underutilized and neglected crops
Belaj A, Dominguez-García M d C, Atienza SG et al (2012) Developing a core collection of olive (Olea europaea L.) based on molecular markers (DArTs, SSRs, SNPs) and agronomic traits. Tree Genet Genomes 8:365–378. https://doi.org/10.1007/s11295-011-0447-6
Bink MCAM, Jansen J, Madduri M, Voorrips RE, Durel CE, Kouassi AB, Laurens F, Mathis F, Gessler C, Gobbin D, Rezzonico F, Patocchi A, Kellerhals M, Boudichevskaia A, Dunemann F, Peil A, Nowicka A, Lata B, Stankiewicz-Kosyl M, Jeziorek K, Pitera E, Soska A, Tomala K, Evans KM, Fernández-Fernández F, Guerra W, Korbin M, Keller S, Lewandowski M, Plocharski W, Rutkowski K, Zurawicz E, Costa F, Sansavini S, Tartarini S, Komjanc M, Mott D, Antofie A, Lateur M, Rondia A, Gianfranceschi L, van de Weg WE (2014) Bayesian QTL analyses using pedigreed families of an outcrossing species, with application to fruit firmness in apple. Theor Appl Genet 127:1073–1090. https://doi.org/10.1007/s00122-014-2281-3
Caruso M, Distefano G, Ye X, la Malfa S, Gentile A, Tribulato E, Roose ML (2008a) Generation of expressed sequence tags from carob (Ceratonia siliqua L.) flowers for gene identification and marker development. Tree Genet Genomes 4:869–879. https://doi.org/10.1007/s11295-008-0159-8
Caruso M, La Malfa S, Pavlíček T et al (2008b) Characterisation and assessment of genetic diversity in cultivated and wild carob (Ceratonia siliqua L.) genotypes using AFLP markers. J Hortic Sci Biotechnol 83:177–182. https://doi.org/10.1080/14620316.2008.11512367
Cavalli-Sforza LL, Edwards AWF (1967) Phylogenetic analysis models and estimation procedures. Am J Hum Genet 19:233–257. https://doi.org/10.2307/2406616
Cipriani G, Spadotto A, Jurman I, di Gaspero G, Crespan M, Meneghetti S, Frare E, Vignani R, Cresti M, Morgante M, Pezzotti M, Pe E, Policriti A, Testolin R (2010) The SSR-based molecular profile of 1005 grapevine (Vitis vinifera L.) accessions uncovers new synonymy and parentages, and reveals a large admixture amongst varieties of different geographic origin. Theor Appl Genet 121:1569–1585. https://doi.org/10.1007/s00122-010-1411-9
Coit JE (1951) Carob or St. John’s bread. Econ Bot 5:82–96. https://doi.org/10.1007/BF02984784
Csardi G, Nepusz T (2006) The igraph software package for complex network research. InterJournal, Complex Systems 1695
De Beukelaer H, Davenport GF, Fack V (2018) Core Hunter 3: flexible core subset selection. BMC Bioinformatics 19:203. https://doi.org/10.1186/s12859-018-2209-z
Di Guardo M, Bink MCAM, Guerra W et al (2017) Deciphering the genetic control of fruit texture in apple by multiple family-based analysis and genome-wide association. J Exp Bot 68:1451–1466. https://doi.org/10.1093/jxb/erx017
Diez CM, Trujillo I, Martinez-Urdiroz N, Barranco D, Rallo L, Marfil P, Gaut BS (2015) Olive domestication and diversification in the Mediterranean Basin. New Phytol 206:436–447. https://doi.org/10.1111/nph.13181
Earl DA, vonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361. https://doi.org/10.1007/s12686-011-9548-7
Escribano P, Viruel MA, Hormaza JI (2008) Comparison of different methods to construct a core germplasm collection in woody perennial species with simple sequence repeat markers. A case study in cherimoya (Annona cherimola, Annonaceae), an underutilised subtropical fruit tree species. Ann Appl Biol 153:25–32. https://doi.org/10.1111/j.1744-7348.2008.00232.x
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Fresnedo-Ramírez J, Frett TJ, Sandefur PJ, Salgado-Rojas A, Clark JR, Gasic K, Peace CP, Anderson N, Hartmann TP, Byrne DH, Bink MCAM, van de Weg E, Crisosto CH, Gradziel TM (2016) QTL mapping and breeding value estimation through pedigree-based analysis of fruit size and weight in four diverse peach breeding programs. Tree Genet Genomes 12:25. https://doi.org/10.1007/s11295-016-0985-z
Garcia-Ochoa F, Casas JA (1992) Viscosity of locust bean (Ceratonia siliqua) gum solutions. J Sci Food Agric 59:97–100. https://doi.org/10.1002/jsfa.2740590114
Goldblatt P (1981) Chromosome numbers in legumes II. Ann Mo Bot Gard 68:551–557
Howard NP, Van De Weg E, Bedford DS et al (2017) Elucidation of the ‘Honeycrisp’ pedigree through haplotype analysis with a multi-family integrated SNP linkage map and a large apple (Malus × domestica) pedigree-connected SNP data set. Horticolture Research 4:1–7. https://doi.org/10.1038/hortres.2017.3
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23:1801–1806. https://doi.org/10.1093/bioinformatics/btm233
Jombart T (2008) Adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24:1403–1405. https://doi.org/10.1093/bioinformatics/btn129
Kalinowski ST, Taper ML, Marshall TC (2007) Revising how the computer program cervus accommodates genotyping error increases success in paternity assignment. Mol Ecol 16:1099–1106. https://doi.org/10.1111/j.1365-294X.2007.03089.x
Kamvar Z, Tabima J, Grünwald N (2014) Poppr: an R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2:e281. https://doi.org/10.7717/peerj.281
Kassambara A, Mundt F (2016) Factoextra: extract and visualize the results of multivariate data analyses. R Packag version 1:
Konate I, Filali-Maltouf A, Berraho EB (2007) Diversity analysis of Moroccan carob (Ceratonia siliqua L.) accessions using phenotypic traits and RAPD markers. Acta botanica malacitana 32:79–90
La Malfa S, Currò S, Bugeja Douglas A et al (2014) Genetic diversity revealed by EST-SSR markers in carob tree (Ceratonia siliqua L.). Biochem Syst Ecol 55:205–211. https://doi.org/10.1016/j.bse.2014.03.022
Meyer RS, DuVal AE, Jensen HR (2012) Patterns and processes in crop domestication: an historical review and quantitative analysis of 203 global food crops. New Phytologist 196(1):29–48
Miranda C, Urrestarazu J, Santesteban LG, Royo JB, Urbina V (2010) Genetic diversity and structure in a collection of ancient Spanish pear cultivars assessed by microsatellite markers. Journal of the American Society of Horticultural Science 135:428–437
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959. https://doi.org/10.1111/j.1471-8286.2007.01758.x
R Core Team (2016) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria
Raymond M, Rousset F (1995) GENEPOP (version 1.2): population genetics software for exact tests and ecumenicism. Journal of Heredity 86:248–249 https://ci.nii.ac.jp/naid/10020473626/en/
Sahle M, Coleou J, Haas C (1992) Carob pod (Ceratonia Siliqua) meal in geese diets. Br Poult Sci 33:531–541. https://doi.org/10.1080/00071669208417492
Schweinfurth G (1894) Sammlung arabisch-aethiopischer Pflanzen, Ergebnisse von Reisen in den Jahren 1881, 1888–89, 1891–92. Bullettin de l'herbier Boissier 2:1–114
Shannon CE (1948) A mathematical theory of communication. Bell System Technical Journal 27:379–423. https://doi.org/10.1145/584091.584093
Sidina MM, El Hansali M, Wahid N et al (2009) Fruit and seed diversity of domesticated carob (Ceratonia siliqua L.) in Morocco. Sci Hortic (Amsterdam) 123:110–116. https://doi.org/10.1016/j.scienta.2009.07.009
Talhouk SN, Van BP, Zurayk R et al (2005) Status and prospects for the conservation of remnant semi-natural carob Ceratonia siliqua L. populations in Lebanon. For Ecol Manag 206:49–59. https://doi.org/10.1016/j.foreco.2004.10.053
Thachuk C, Crossa J, Franco J, Dreisigacker S, Warburton M, Davenport GF (2009) Core hunter: an algorithm for sampling genetic resources based on multiple genetic measures. BMC Bioinformatics 10:1–13. https://doi.org/10.1186/1471-2105-10-243
Tous J, Olarte C, Truco M, Arus P (1992) Isozyme polymorphisms in carob cultivars. HortScience 27:257–258
Tous J, Rovira M, Romero A et al (2006) Carob tree germplasm in Tunisia. NUCIS Newsl 13:55–59
Tous J, Romero A, Hermoso JF et al (2008) Fruiting and kernel production characteristics of ten Mediterranean carob cultivars grown in northeastern Spain. J Am Pomol Soc 62(4):144
Tous J, Romero A, Batlle I (2013) The carob tree: botany, horticulture, and genetic resources. In: Horticultural reviews, vol 41. Wiley-Blackwell, New York, pp 385–456
Tucker SC (1992) The developmental basis for sexual expression in Ceratonia siliqua (Leguminosae: Caesalpinioideae: Cassieae). Am J Bot 79:318–327
van Hintum T (1999) The general methodology for creating a core collection. In: John- son RC HT (ed) Core collections for today and tomorrow. International Plant Genetic Resources Instit., (IPGRI), Rome (Italy)
van Hintum TJL, Brown AHD, Spillane C, Hodgkin T (2000) Core collections of plant genetic resources. In: IPGRI Technical Bulletin N. 3
Vavilov NI (1951) The origin, variation, immunity, and breeding of cultivated plants. Ronald Press Co, New York
Viruel J, Médail F, Marianick J, et al. (2016) Mediterranean carob populations, native or naturalized? A continuing riddle. OPTIMA XV Montpellier, Fr. doi: https://doi.org/10.13140/RG.2.2.33681.84328
Viruel J, Haguenauer A, Juin M, Mirleau F, Bouteiller D, Boudagher-Kharrat M, Ouahmane L, la Malfa S, Médail F, Sanguin H, Nieto Feliner G, Baumel A (2018) Advances in genotyping microsatellite markers through sequencing and consequences of scoring methods for Ceratonia siliqua (Leguminosae). Applications in Plant Sciences 6:e01201. https://doi.org/10.1002/aps3.1201
Wang J (2011) COANCESTRY: a program for simulating, estimating and analysing relatedness and inbreeding coefficients. Mol Ecol Resour 11:141–145
Winer N (1980) The potential of the carob (Ceratonia siliqua). Int Tree Crop Journal 1:15–26. https://doi.org/10.1080/01435698.1980.9752711
Wright S (1969) Evolution and the genetics of populations. theory gene Freq Volume 2
Zohary M (1973) Geobotanical foundations of the Middle East. Gustav Fisher Verlag, Stuttgart
Zohary D (2002) Domestication of the carob (Ceratonia siliqua L.). Israel Journal of Plant Sciences 50:141–145. https://doi.org/10.1560/BW6B-4M9P-U2UA-C6NN
Acknowledgments
The authors gratefully acknowledge Alex Baumel, Aix Marseille Universite, Marseille, France, for providing matK4LF sequence. A.N., M.R., J.F.H., and I.B. are grateful to CERCA Institution from the Generalitat of Catalonia for its support.
This study has been partially supported by the international project DYNAMIC “Deciphering sYmbiotic Networks in cArob-based MedIterranean agro-eCosystems” (https://dynamic.cirad.fr/en) supported by the French national research agency (ANR-14-CE02-0016).
Data archiving statement
Raw genotypic data were presented in Supplementary Table 2.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Troggio
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Supplementary Table 1
(DOCX 82 kb)
Supplementary Table 2
(DOCX 48 kb)
Supplementary Table 3
(DOCX 13 kb)
Supplementary Table 4
(DOCX 12 kb)
Supplementary Figure 1
Electropherograms of the sequencing of the matK4LF chloroplast locus. The alignment of the sequencing data allowed the detection of a T/G polymorphism at approximately 622 bp position (PNG 144 kb)
Supplementary Figure 2
Log likelihood curve (ΔK) calculated according to Evanno et al. (2005) plotted against increasing K value. A: plot of the whole carob collection, B: plot of the subset of individuals deemed as Subpop 1, C: plot of the subset of individuals deemed as Subpop 2 (PNG 741 kb)
Supplementary Figure 3
Absolute frequency of the sex of the accessions according to the country/region of origin (PNG 164 kb)
Supplementary Figure 4
Relative frequency of the distribution of the two haplotypes detected through the sequencing of the matK4LF chloroplast locus (PNG 76 kb)
Supplementary Figure 5
Discriminant Analysis of Principal Components (DAPC) based on the genetic data. Countries or regions of collection were used as grouping factor (PNG 753 kb)
Rights and permissions
About this article
Cite this article
Di Guardo, M., Scollo, F., Ninot, A. et al. Genetic structure analysis and selection of a core collection for carob tree germplasm conservation and management. Tree Genetics & Genomes 15, 41 (2019). https://doi.org/10.1007/s11295-019-1345-6
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11295-019-1345-6