Abstract
Global transcriptional regulation during apple fruit maturation and associated texture changes were assessed by transcriptome profiling and systematic characterization of maturation progression of two cultivars, ‘Honeycrisp’ (HC) and ‘Cripps Pink’ (CP). A high-density long-oligo apple microarray consisting of duplex 190,135 cross-hybridization-free 50–70-mer isothermal probes, representing 23,997 unigenes, was designed for and manufactured on a NimbleGen array platform. Cortex tissues from both HC and CP at three maturation stages, i.e., 4, 2, and 0 week(s) before physiological maturity, were utilized for transcriptome profiling. A total of 1,793 and 1,209 differentially expressed unigenes, 7.47 % and 5.04 % of all unigenes deposited on the array, were identified from HC and CP, respectively. Unigenes associated with ethylene biosynthesis and response, auxin homeostasis and transport, gibberellin reception and metabolism as well as degradation of hemicelluloses may contribute to the observed phenotypic variations in apple maturation patterns and texture attributes such as fruit firmness and crispness. Microarray data validation indicated that more than 85 % of randomly selected unigenes showed consistent expression patterns with qRT-PCR results. Physiological characterization demonstrated substantial differences in maturation progression between these two cultivars, and a remarkable transformation in fruit texture occurred from week −4 to week 0.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Apple (Malus × domestica Borkh.) is one of the most popular perennial tree fruits. A wealth of knowledge exists regarding varietal differences, physiological changes, and horticultural management in relation to fruit maturation, ripening, and quality. Due to its long history and widespread cultivation, and more importantly its out-crossing nature, apple exhibits a high level of heterozygosity and great variation in ripening behavior and quality attributes. The ripening season of apple can differ up to 3 months among the elite apple cultivars under the same weather condition. There are considerable variations of apple fruit texture attributes such as fruit firmness and crispness, which significantly influence consumer preference and fruit industry profitability. Accordingly, apple is one of few fruit crops for which consumers recognize and prefer fruit from specific cultivars.
Fruit ripening, characterized by texture modification, aroma production, and color change, is under tight genetic control (Giovannoni 2004; Seymour et al. 2002), and cultivar-specific patterns of maturation and ripening intimately influence fruit quality. The processes of cell division and expansion have largely attenuated in apple fruit at later stages of maturation (Janssen et al. 2008). However, continuing changes inside the cell and cell wall transform the starchy, hard, and astringent flesh into a state more palatable for human consumption and amenable for natural seed dispersion. In regard to fruit texture changes, microscopic and cellular features such as cell size and density, cell wall properties, cell turgor, and cell–cell adhesion may contribute to the observed phenotypes (Johnston et al. 2002). Currently, a molecular model describing apple fruit maturation, ripening, and texture change is lacking, and its molecular characterization is less represented by other model plant systems due to the unique physiology of pome fruit maturation and ripening.
Plant hormones, especially ethylene, are known to regulate apple fruit maturation, ripening, and quality (Bleecker and Kende 2000; Costa et al. 2005; Harada et al. 2000). Apple cultivars with different allelotypes of two ethylene biosynthesis genes, ACS1 and ACO1, have been shown to correlate with apple fruit firmness in a wide selection of germplasm (Oraguzie et al. 2004; Zhu and Barritt 2008). Recent reports suggest that auxin may also play a critical role in regulating fruit ripening and quality of apple and peach (Li and Yuan 2008; Kondo et al. 2009; Trainotti et al. 2007; Ziliotto et al. 2008). Cell wall metabolism has been closely associated with fruit texture changes, and numerous gene families could potentially contribute to cell wall biosynthesis and modification (Bennett and Labavitch 2008; Harker et al. 1997; Johnston et al. 2002). Many other pathways and physiological processes, such as those related to intra-cellular turgor (water and metabolite transport) and epidermal structural deposition (wax and cutin), may also contribute to fruit texture attributes. Yet little is known regarding the individual genes and pathways differentiating cultivar-specific apple fruit ripening patterns and quality attributes. Apple cultivars ‘Honeycrisp’ (HC) and ‘Cripps Pink’ (CP) exhibit distinct texture attributes and ripening behavior. While fruit of late-ripening CP demonstrate outstanding firm flesh, HC is a mid-early ripening cultivar with extraordinarily crisp but less firm flesh. Both are commonly utilized as apple breeding parents for their exceptional texture attributes and other quality traits.
Several recent studies using various apple microarrays were reported including those focusing on early fruit development (Lee et al. 2007); the roles of ethylene on volatile biosynthesis pathways (Schaffer et al. 2007); the comprehensive transcriptome analysis from flowering to fruit ripening on the apple cultivar ‘Gala’ (Janssen et al. 2008); the effect of different apple root stocks in relation to gene expression in scion tissues (Jensen et al. 2010); and using apple EST array and heterologous tomato array to study the ripening dynamics of ‘Mondial Gala’ (Costa et al. 2010). Various formats of microarray have also developed for transcriptome profiling on other rosaceous crops (Falara et al. 2011; Vizoso et al. 2009; Ziliotto et al. 2008). In this study, a high density long-oligo apple array was designed for and manufactured on NimbleGen platform based on Malus unigene assembly version 4 (http://www.bioinfo.wsu.edu/cgi-bin/gdr/gdr_unigeneV4_project_description.cgi?genus=Malus). Parallel transcriptome profiling was performed on two cultivars at equivalent maturation stages. Unigenes associated with biosynthesis, homeostasis and transport, reception and metabolism of multiple plant hormones as well as unigenes functioning in cell wall modification were identified; their relationships with cultivar-specific phenotypic variations of apple maturation patterns and texture attributes were discussed.
Materials and methods
Physiological characterization of apple fruit ripening
Physiological characterization of weekly fruit samples was used to define the maturity level and maturation process of two apple cultivars, ‘Honeycrisp’ (HC) and ‘Cripps Pink’ (CP, also known as Pink Lady™). Fruit were harvested from two separate commercial orchards close to Wenatchee WA in 2007. Fruit with uniform size and appearance were randomly harvested from a number of trees for weekly maturity testing. Based on the ripening data in previous years, fruit sample collection started approximately 6 weeks before and continued to 2 weeks after the projected harvest date for obtaining fruit with the desired range of maturity. Fruit firmness and crispness were measured on pared fruit surfaces using the Mohr Digi-Test, MDT-1 (Mohr and Associates, Richland, WA, USA), which is a computerized penetrometer with 11-mm probe diameter. Crispness (Cn) values from the MDT-1 are the Fourier transformation of the force profile as the probe goes through fruit cortex tissues, and these values therefore are theoretically similar to the energy released during a bite; a recent study showed there was a good correlation between the Cn value and human sensory evaluation regarding fruit crispness (Evans et al. 2010). Fruit internal ethylene concentration (IEC) was measured using a Hewlett Packard 5880A series gas chromatograph equipped with a Porapak Q column (0.3 cm × 30 cm i.d., 80–100 mesh) and flame ionization detector (Blanpied and Silsby 1992; Argenta et al. 2002). Fruit maturity which is primarily determined by starch pattern indices (SPI) were calculated by averaging the values (1–6 in grade) (Brookfield et al. 1997) of 15 apples. The weekly fruit samples with starch pattern indices close to an average of 3.5 were arbitrarily designated as week 0 or physiological maturity in this study. Fruit with various maturity levels were defined in relation to week 0 samples, i.e., fruit collected at 4, 3, 2, or 1 week(s) before week 0 were retrospectively assigned as week −4, −3, −2, or −1 samples. Values of individual fruit diameter were recorded automatically by Digi-Test; the significance of difference between weekly samples was analyzed by t test (Microsoft Excel). Cortex tissues collected at weeks −4, −2, and 0 were used for transcriptome profiling, and each weekly sample was represented by four biological replicates. Each replicate consisted of a pool of cortex tissues from five apples. Once collected, fruit cortex tissues were immediately frozen in liquid nitrogen and stored at −80 °C until RNA isolation.
RNA isolation
Total RNA was isolated as described by Gasic et al. (2004) with modification. Apple cortex tissue (2–3 g) ground (IKA A 11 basic; IKA®-WORK Inc., Wilmington, NC, USA) in liquid nitrogen was transferred to 50-mL polypropylene tubes with 10 mL 2× CTAB extraction buffer, vortexed, then incubated at 60 °C for 15 min with occasional inversions. Then an equal volume of chloroform/isoamyl alcohol (24:1, v/v) was added and the mixture vortexed for 2 min. The mixture was then centrifuged at 10,000 × g for 10 min at 4 °C and the supernatant transferred to a clean 50-mL polypropylene tube. After repeating this process, the supernatant was transferred to a 15-mL polypropylene tube where a one third volume of 7.5 M LiCl was added; the tube was inverted several times to mix and then incubated at 4 °C for 16 h. Following incubation, tubes were centrifuged at 14,000 × g for 30 min at 4 °C, the supernatant was discarded, and the RNA pellet washed with 750 μL 70 % ethanol. The RNA pellet was re-suspended in 500 μL DEPC water and transferred to a 1.5-mL micro-centrifuge tube. Total RNA was precipitated by adding 1/10 vol of 3 M sodium acetate (pH 5.5) and 2 vol of 100 % ethyl alcohol followed by incubation for 1–3 h at −80 °C. After centrifugation at 14,000 × g for 30 min at 4 °C, the supernatant was discarded; the pellet was washed with 70 % ethyl alcohol and then air-dried for 5 min. The pellet was suspended in 150 μL DEPC water then stored at −80 °C. Isolated total RNA was quantified using an ND-1000 NanoDrop spectrophotometer (NanoDrop Technologies, Wilmington, DE, USA) and RNA quality was verified by agarose gel electrophoresis.
Microarray design
An isothermal long-oligo apple microarray was designed based on the Malus unigene V4 sequences which are available at the Genome Database for Rosaceae (GDR) (Jung et al. 2008). A total of 260,581 Malus EST sequences were downloaded from NCBI dbEST (Benson et al. 2007), filtered for contamination and low quality sequence, and assembled into 23,284 contigs and 53,200 singletons using CAP3 (Huang and Madan 1999) with an overlap percentage parameter of 90 (−p 90). The unigenes, comprising of the combined contigs and singlets, were computationally annotated for putative function by pairwise comparison against the Arabidopsis thaliana protein database (http://www.arabidopsis.org), and the Uniprot Swiss-Prot and TrEMBL databases (Wu et al. 2006; Mulder et al. 2007) using the BLASTX algorithm. Only matches with an E value of less than 1.0 e−6 were recorded. Based on the similarity search results, 55,960 (73 %) of the Malus unigene sequences had significant matches with proteins from these databases. Duplicate transcripts present in both the contigs and singletons were eliminated by removing singlets with a common protein match with the contigs. Following detection of Open Reading Frames (ORFs) using the EMBOSS getORF program (http://bioweb2.pasteur.fr/docs/EMBOSS/getorf.html), 23,106 contigs and 17,060 singletons, totaling 40,166 unigenes, were available for oligo design. The isothermal oligonucleotides of 50–70-mer were designed from the coding region of the 40,166 unigenes using custom perl scripts. Oligos designed from contigs had three to 10 oligos per contig with an average of seven; those designed from singlets had seven to 10 oligos with an average of eight oligos per singlet. Including control sequences, a probe set containing 190,135 oligos representing 23,997 unigenes was designed for NimbleGen array format. The unigene, oligo sequences, and protein homology results are available at http://www.bioinfo.wsu.edu/cgi-bin/gdr/gdr_unigeneV4_project_description.cgi?genus=Malus .
Microarray fabrication, hybridization, data acquisition, and analysis
Array fabrication, cDNA conversion and labeling, hybridization, image scanning, and data normalization were performed at NimbleGen (http://www.nimblegen.com/). Using a single color labeling system, a total of 24 microarray slides were utilized for transcriptome profiling, i.e., two cultivars × three maturation stages × four biological replicates. The normalized signal intensity based on the internal control probe set were imported from the NimbleGen expression files and were analyzed using Arraystar (http://www.dnastar.com/). After performing an ANOVA and multiple testing correction (Benjamini and Hochberg 1995), the probes (or oligos) that displayed a 2-fold or greater change in transcript abundance, across any two adjacent maturation stages (i.e., week −4, week −2, and week 0) within each cultivar, were extracted and converted to differentially expressed (p < 0.01) unigenes. Tables S1 and S2, displaying the expression patterns of individual identified unigenes, were prepared using CORELDRAW (http://www.corel.com/servlet/Satellite/us/en/Content/1150905725000).
Validation of microarray data by qRT-PCR
Individual unigenes were selected randomly from the list of identified unigenes for data validation by qRT-PCR. EST sequences were obtained from the GDR (Jung et al. 2008). Forward and reverse primers were designed using web-based software Primer3plus (http://www.bioinformatics.nl/cgi-bin/primer3plus/primer3plus.cgi) and IDT oligo analyzer (http://www.idtdna.com/analyzer/Applications/OligoAnalyzer/). Where possible, an optimum annealing temperature of 59 to 60 °C, GC content 40–60 %, amplicon length 150–180 bp, and primer length 20 bp were applied. As needed, unigenes from multi-member family were analyzed using ClustalW (http://www.ebi.ac.uk/tools/clustalw/) and BLAST (http://www.ncbi.nlm.nih.gov/BLAST/) to choose divergent regions for designing gene-specific primer sets. Primers used in this study were listed in Table S5. Total RNAs isolated from the same tissues used for microarray analysis were treated with DNase I to eliminate co-purified residual genomic DNA and further purified using Qiagen RNeasy columns (Qiagen, Valencia, CA, USA). For a 20-μL reverse transcription (RT) reaction, 2 μg total RNA and 1 μL 100 μM poly dT primer were incubated at 70 °C for 10 min, then put on ice for 2 min. Then 15 U AMV reverse transcriptase and 4 μL of 5× reverse transcription buffer (Promega, Madison, WI, USA), 2 μL of 10 mM dNTP, and 20 U recombinant RNasin® ribonuclease inhibitor (Applied Biosystems, Foster City, CA, USA) were added to a final volume of 20 μL with H2O. The reactions were incubated at 42 °C for 1 h and then at 70 °C for 15 min. For each cortex tissue sample, two independent total RNA isolations were obtained, and thereafter two separate cDNA preparations. PCR reactions were carried out in triplicate for each cDNA preparation and repeated at least once. The volume of quantitative PCR was 15 μL, which contained 1× PCR buffer, 2.5 mM MgCl2, 2.4 μL cDNA (20× dilution from original RT reaction), 200 nM forward and reverse primers, 200 μM dNTP, 0.3 U iTaq DNA polymerase (Bio-Rad Lab, Hercules, CA, USA), and 0.45 μL 2,000× dilution of SYBR I green (Invitrogen, Carlsbad, CA, USA). Reactions started with a denaturation stage for 3 min at 94 °C, then amplified for 40 cycles (94 °C for 15 s and 59 °C for 30 s) using a Bio-Rad iQ5™ real-time PCR detection system (Bio-Rad). Melting curve and amplification efficiency were analyzed for each primer set, and “no template control” and “reverse transcriptase minus control” were routinely included. The sequence of a unigene for actin (Contig13803) was used to design primers as a reference gene for normalization. Data was normalized against an apple actin encoding unigene and expression analysis was performed using the ddCt method (Bio-Rad).
Results
Cultivar-specific fruit maturity progression
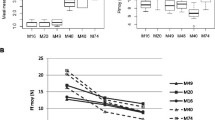
Based on weekly maturity data, ‘Honeycrisp’ (HC) fruit harvested on Sep. 2 and ‘Cripps Pink’ (CP) fruit harvested on Nov. 4 were defined as week 0 samples, i.e., HC0 and CP0, when their starch pattern indices (SPI) were close to average values of 3.5 (Table 1). Fruit firmness decreased as maturation progressed; however, the rate of firmness loss varied substantially between these two cultivars (Table 1). At week −4, a difference of fruit firmness between these two cultivars was 11.2 N (Newton), yet a more substantial 33.3-N difference was observed at week 0. Apple fruit crispness measured by the Digi-Test expressed as Cn value also showed remarkable differences between these two cultivars. Most of the Cn values (four out of five) from the fruit of HC were close to or higher than 250, while most of the values (four out of five) for those of CP were close to or lower than 200. In both cultivars, the average values of SPI steadily increased as maturation progressed, and the comparable extent starting from an average value of about 1 to a value close to 3.5 among the five weekly samples were observed. No substantial and consistent ethylene production was observed, indicating that climacteric ripening had not initiated in these fruit. No significant changes of fruit diameter were observed among weekly fruit samples except for week −4 and −3 of HC, indicating fruit have attained their mature size at late maturation.
Transcriptome profiling of apple fruit cortex tissues
The identification of differentially expressed unigenes was performed within each cultivar and based on ANOVA analyses using a cutoff value of 2-fold of normalized signal intensity and a non-adaptive false discovery rate (FDR) of 0.01. A total of 1,793 differentially expressed unigenes from HC and 1,209 from CP were identified, which represent 7.47 % and 5.04 % of all unigenes deposited on the array, respectively (Table 2). The volcano plots showed an asymmetric shape between two adjacent stages (−4 to −2 and −2 to 0) for HC which reflected more unigenes with “transitional expression patterns”, i.e., those with down- then up-regulated patterns or vice versa; this trend was not observed in CP (Fig. 1 and Fig. S1a). In both cultivars, unigenes showing down-regulated expression patterns substantially outnumbered the ones with up-regulated expression patterns (Table 2 and Fig. S1a). The raw dataset was deposited in GEO with Series accession GSE24523 (http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE24523).
Volcano plots showing the distribution patterns of identified unigenes from two cultivars. a Identified unigenes from week −4 to week −2 in the cortex tissue of HC. b Identified unigenes from week −2 to week 0 in the cortex tissue of HC. c Identified unigenes from week −4 to week −2 in the cortex tissue of CP. d Identified unigenes from week −2 to week 0 in the cortex tissue of CP
Functional categories of differentially expressed unigenes
The differentially expressed unigenes were classified via their functional annotations and grouped into finer functional categories in order to derive biological meaning. Unigenes with genotype-specific expression patterns may contribute to the observed phenotypes of maturation patterns and texture attributes.
Hormonal metabolism and response
Over half of the unigenes in this group (41 out of total 74) were those related to the biosynthesis, metabolism, transport, and response of ethylene and auxin, indicating the central roles of these two hormones during apple maturation. Unigenes implicated in metabolism and response of other hormones including gibberellins (GA), brassinosteroid (BR), jasmonic acid (JA), abscisic acid (ABA), and cytokinins (CK) were also identified (Table 3). About 50 % of the unigenes were shared between these two cultivars, and the majority of them exhibited similar expression patterns (Table 3, Table S1, Fig. 2a, and Fig. S2).
Venn diagram demonstrating overlapping differentially expressed unigenes from each cultivar for selected functional groups, and those between cultivars. a Unigenes with annotated functions of hormone metabolism and responses. b Unigenes with annotated functions of cell wall metabolism and modification. c Transcription factor encoding unigenes. d Unigenes with other cellular functions
Ethylene biosynthesis and responses
Consistent with the critical roles of ethylene during pome fruit maturation and ripening, the majority of unigenes related to ethylene functions showed up-regulated expression patterns as fruit maturation progressed (Table 3, Table S1, and Fig. S2a). In fact, this was one of a few sub-groups which consisted of more up-regulated unigenes than down-regulated ones. Two unigenes, contig21202 and 21169, annotated as “1-aminocyclopropane-1-carboxylate synthase 7”, and sequence alignment indicated that they encode apple MdACS3 for pre-climacteric ethylene biosynthesis (Rosenfield et al. 1996; Varanasi et al. 2011), showed consistently up-regulated expression patterns in both cultivars, although a higher fold increase was observed in CP. Two identified unigenes (contig13926 and 21937) encoding 1-aminocyclopropane-1-carboxylate oxidase (ACO) were similarly up-regulated, though a much higher fold increase from week −4 to week −2 was observed in HC. A unigene (contig18342) encoding MdETR2 (Wiersma et al. 2007), an ethylene receptor which negatively regulates ethylene responses, exhibited up-regulation but only in CP. Four unigenes encoding the EIN3-binding F-box protein (Contig17900, 18538, 21897, and 21173), a negative regulator of ethylene action (Gagne et al. 2004), were also identified exclusively from CP with up-regulated expression patterns. A unigene (Contig12329) encoding an “Ethylene-overproduction protein 1”, a negative regulator of ethylene evolution (Yoshida et al. 2005), also showed a down-regulated pattern and only in CP.
Auxin biosynthesis, transport, and responses
In contrast to up-regulated expression patterns for most ethylene function-related unigenes, the majority of the identified unigenes associated with auxin function exhibited down-regulated expression patterns, particularly those from CP (Table 3; Table S1 and Fig. S2b). Four unigenes annotated as “auxin efflux carrier component” (contig7586, 18662, Malus_CV794150, and Malus_CV880396), which encode auxin transporter PIN1 and PIN8 homologues (Petrášek et al. 2006), were identified only from CP with down-regulated expression patterns. Additionally, one unigene from each cultivar (Malus_CN945995 from CP and contig17221 from HC) with the similar annotated function showed the comparable up- and then down-regulated expression patterns. Three unigenes with annotated functions of auxin homeostasis regulation were identified from both cultivars. Contig3460 encoding a protein homologous to rice “indole-3-acetic acid-amido synthetase GH3.5”, which catalyzes the synthesis of IAA–amino acid conjugates (Jain et al. 2006), showed down-regulated expression patterns in both cultivars, though with a double-digit fold decrease in HC at later stages (from −2 to 0 week), while another unigene with similar function (Malus_CN909148) was up-regulated in CP. Contig196 encoding a protein homologue to “indoleacetamide hydrolase” (Mazzola and White 1994), which releases bioactive IAA from amide conjugation, was down-regulated only in CP. In addition, more than a dozen unigenes encoding “auxin induced proteins” or “auxin responsive proteins” were identified from both cultivars, but more unigenes showed down-regulated patterns in CP.
Metabolism and responses of other plant hormones
Thirteen unigenes with the annotated functions in gibberellin biosyntheses and responses were identified (Table 3, Table S1, and Fig. S2c). Two unigenes from each cultivar (contig21434 and 21375 from CP, contig21386 and 22883 from HC) which encode “DELLA proteins”, the repressor of GA responses (Chandler et al. 2002), were identified and all showed down-regulated expression patterns except the up-regulated contig22883 in HC at the later stages. Four unigenes (contig16116, 9239, 16825, and 9114) encoding proteins homologous to “gibberellin 3 (or 2)-beta-dioxygenase”, which convert the inactive precursors to the bioactive GA (Thomas et al. 1999), showed mostly up-regulated expression patterns in both cultivars. Contig16116 showed slight down-regulation first and then a double-digit fold change of up-regulated expression in HC, compared to a moderately increased expression in CP. A unigene (contig5705) encoding protein homologous to “rice gibberellin receptor GID1”, a soluble gibberellin receptor (Ueguchi-Tanaka et al. 2005), showed up- and then down-regulated expression patterns in both cultivars, though a greater fold change was observed in HC. Contig8101 encoding another “gibberellin receptor” was identified only from CP with moderate but progressive up-regulation. A total of nine unigenes annotated as “brassinosteroid insensitive 1-receptor kinase 1” were identified from both cultivars. All seven of those identified from CP showed down-regulated expression patterns; similarly, all five from HC were down-regulated except two unigenes which showed slight up-regulation during the early maturation stage. Among four “Jasmonate O-methyltransferase” (contig8892, 17950, 23142, and 17848) encoding unigenes identified from both cultivars, all three from HC (except one value) were up-regulated, but in CP one was down-regulated and another up-regulated. Two unigenes encoding proteins homologous to “cytokinin dehydrogenase” (contig15546 and Malus_CN910875) and three unigenes for “cytokinin-N- (or O-) glucosyltransferase” (contig21874, 21694, and Malus_CN994862), which regulate homeostasis by deactivating CK (Hou et al. 2004; Schmuelling et al. 2003), were identified from both cultivars; more than 50 % of the values indicated up-regulated patterns of three unigenes in HC, but both unigenes from CP showed down-regulation. A unigene for “abscisic acid 8′-hydroxylase (contig1068)” was down-regulated in CP, but displayed an up- and then down-regulated pattern in HC. An “abscisic acid response protein” (contig5535) encoding unigene was down- and then up-regulated only in CP.
Carbohydrate metabolism and cell wall modification
Identified unigenes with annotated functions in polysaccharide metabolism and cell wall modification represent another major transcriptomic change during apple maturation. Overall, comparable numbers of unigenes belonging to this category (57 from CP and 59 from HC) were identified from each cultivar and over 40 % of these unigenes were shared by both cultivars (Fig. S3). Nevertheless, a higher percentage of unigenes (42.9 %) from HC showed up-regulated patterns compared with those from CP (29.8 %) (Table 4, Table S2, and Fig. S3). Across different sub-groups of cell-wall-related genes (Cantarel et al. 2009), fewer unigenes with annotated functions in polysaccharide biosynthesis (glycosyl transferases, GTs) were identified than those implicated in polysaccharide breakdown and cell wall disassembly or remodeling such as glycosyl hydrolases: GHs and polysaccharides lyases and esterases (Table 4 and Fig. S3). Within the GT sub-group, the majority of the unigenes from HC were up-regulated during the later maturation stage from week −2 to week 0, compared to only two unigenes (out of 10) that showed up-regulated patterns in CP. Among GHs and other cell wall modifying proteins, comparable numbers of unigenes were identified between these two cultivars, and in general these unigenes exhibited similar expression patterns. For example, a unigene annotated as polygalacturanase (PG) (contig21179) showed similar up-regulated expression patterns in both cultivars, even with similar fold increases (Table 4 and Table S2). For a few sub-groups including “glucan endo-1, 3-beta-glucosidase”, “carbohydrate esterase”, “polysaccharide lyase”, “expansin”, and “cell wall structural proteins”, equal or more unigenes were with up-regulated expression pattern in cortex tissues of HC, though the extra down-regulated unigenes were commonly identified from CP. For a few gene families, noticeable cultivar-specific regulation patterns were observed (Table 4 and Table S2). Six unigenes encoding “xyloglucan endotransglycosylases/hydrolase (XTH)” (contig17875, 22333, 21660, 21193, 8509, and 21334) were identified from these two cultivars. All four of those from HC showed progressively up-regulated expression patterns during maturation; among five XTHs identified from CP, two unigenes exhibited up-regulated expression patterns, but other three showed up- and then down-regulated patterns. Three unigenes (contig15993, 11693, and Malus_CV793668) annotated as wall-associated-kinase (Verica et al. 2003) were exclusively identified from HC, and two of them showed up- then down-regulated patterns. A unigene (contig13231) encoding a protein homologous to COBRA7 protein (Roudier et al. 2005) and with putative function in cellulose microfibril deposition exhibited down- and up-regulation patterns and only from HC.
Transcription factors (TFs)
A substantial number of TF-encoding unigenes were identified in cortex tissues during apple fruit maturation, i.e., 139 from HC and 81 from CP. Consistent with the critical roles of ethylene and auxin on apple fruit ripening (as shown in Table 3), more than 20 % (39/182) of TFs encoding unigenes belonged to those specifically responding to these two plant hormones (Table 5 and Table S3). From HC, the relatively equal numbers for both auxin- and ethylene-specific TFs showed either up- or down-regulated patterns. However, from CP, an obvious disparity was observed between these two sub-groups of TF-encoding unigenes: the overwhelming majority of auxin-specific TF-encoding unigenes showed down-regulated expression patterns, compared to more up-regulated values for ethylene-specific TFs (Table 5 and Fig. S4a and S4b). This observation suggests that a deficiency in auxin metabolism and/or transport occurs in CP (see previous section). For several TF families, unigenes were commonly identified from both cultivars, such as “bHLH”, “homeobox-leucine zipper protein”, “MYB”, and “squamosa promoter-binding-like proteins” (Table 5 and Table S3). Nevertheless, unigenes belonging to a few other TF families showed strong cultivar-specific expression patterns, and in most cases they were identified almost exclusively from HC, such as “zinc finger protein”, “WRKY”, and “NAC” (Table 5 and Fig. S4c).
Unigenes with other annotated molecular functions
Unigenes with various annotated molecular functions were identified (Table S4). Among them, the largest group consisted of more than 100 unigenes which were annotated as “protein kinase”, “receptor-like protein kinase”, or “phosphatase”. Other groups included those with putative functions in modulating oxidative balances such as “cytochrome p450”, “lipoxygenase”, and “oxidase”; stress-response and other cellular processes such as “heat shock proteins”, “dehydration-responsive and element-binding protein”, “aquaporin” as well as those annotated as “major allergen Mal d1” and “calcium responsive proteins”. For the vast majority of these unigenes, their biological functions and their potential association with apple fruit maturation are largely elusive.
Microarray data validation
Twelve identified unigenes based on transcriptome profiling were randomly selected and their expression profiles were characterized by quantitative reverse transcription PCR (qRT-PCR). As summarized in Table 6 and Fig. S5, among 48 values for 12 unigenes from two cultivars, 87.5 % of the values showed consistency between the microarray data and qRT-PCR result in terms of either up- or down-regulation expression patterns. Due to different chemistries employed by these two approaches, the actual values of fold changes between the microarray analysis and relative expression levels by qRT-PCR may not be comparable in some cases.
Discussion
To gain the insight of global transcriptional networks and identify genotype-specific transcriptome changes over apple fruit maturation patterns and fruit texture attributes, parallel transcriptome profiling was performed on two apple cultivars with contrast phenotypes (Table 1). The high-density microarray developed for this study, to our knowledge, contains the largest unigene collection among reported transcriptomic studies on apple or other Rosaceae crops (Costa et al. 2010; Falara et al. 2011; Schaffer et al. 2007; Vizoso et al. 2009; Ziliotto et al. 2008), which offered better coverage and resolution for analyzing transcriptome activities. A reliable dataset was generated using single color labeling system in array hybridization and proper biological repeats, which was mostly verified by qRT-PCR on randomly selected unigenes (Table 6 and Fig. S5). The majority of gene families or functional groups showed comparable regulation patterns between these two cultivars; those unigenes exhibiting cultivar-specific expression patterns could represent the candidates contributing to the observed phenotypes of fruit maturation patterns, ripening season, cell wall features, and texture attributes. However, it is likely that some of the identified unigenes may result from the variations of external conditions at or before sampling time such as temperature or horticultural practices. The presence of more unigenes with “transitional expression patterns” from HC is not clear, which could derive from either external conditions or the cultivar itself as HC was originally bred for cold hardiness but hot and dry summer is common in central Washington State.
Maturation progression and tissue comparability between cultivars
For a meaningful comparison between two cultivars, it is critical that the samples included are with equivalent maturity or at similar developmental stages. Several physiological indicators can practically be used to define apple fruit maturity, such as days after full bloom (DAFB), fruit firmness, background color, internal ethylene concentration (IEC), and fruit starch levels. Fruit texture and maturation pattern are two targeted phenotypes in this study. IEC remains low and fluctuates during apple fruit maturation prior to climacteric ripening. Therefore, fruit starch pattern indices (SPI) (Brookfield et al. 1997), which provide a steady change through the maturation process, were primarily utilized to align the maturation stages between these two cultivars. Several studies indicated that climacteric ethylene production from system II ethylene biosynthesis by the function of MdACS1 had not activated in apple fruit before physiological maturity (Rosenfield et al. 1996; Varanasi et al. 2011; Wiersma et al. 2007; Zhu et al. 2008). Consistent with these results, no up-regulation of MdACS1 was detected in either cultivar; instead, two unigenes encoding MdACS3 and functioning in pre-climacteric ethylene biosynthesis showed similar up-regulation in both cultivars (Table 3 and Table S1). Based on our maturity data, fruit cortex tissues with comparable maturity were selected for transcriptome profiling. For example, similar extent of SPI values from 1 to about 3.5 was observed for both cultivars.
Critical roles of cultivar-specific hormonal regulation
Consistent with the fundamental roles of plant hormones and their crosstalk in plant physiology (Spartz and Gray 2008; Vandenbussche and Van Der Straeten 2007), the identified unigenes associated with plant hormone metabolism and responses were one of the major focal points of transcriptomic changes in apple cortex tissues (Tables 3, 5, S1, and S3; Figs. S2 and S4). Except ethylene biosynthesis, little is known regarding the genotype-specific roles of plant hormones metabolism and response during fruit ripening. Several unigenes encoding negative regulators of ethylene response, such as ethylene receptor and EIN3-binding F-box (EBF) protein (Rosenfield et al. 1996; Gagne et al. 2004), were identified with up-regulated patterns and exclusively from CP, a late-ripening cultivar with extremely firm flesh (Tables 1, 3, and S1). Most unigenes related to auxin metabolism and function showed down-regulated patterns in contrast to up-regulated trends of most ethylene-related genes. The data suggested that the deficiency in auxin transport and availability of its biologically active form may be associated with the prolonged maturation and late-ripening phenotypes of CP (Table 3 and Table S1). Given the synergistic nature between auxin and ethylene interactions (Rahman et al. 2002; Stepanova et al. 2005; Swarup et al. 2002), the variations of auxin metabolism and action may determine the timing and strength of ethylene metabolism. Jasmonate and its methyl ester have been shown to stimulate apple ACS and ACO enzyme activities and gene expression in apple fruit (Fan et al. 1998; Kondo et al. 2009), as similarly demonstrated in vegetative tissues of Arabidopsis (Kazan and Manners 2008; Shinshi 2008). The interplays between GA and other plant hormones in apple fruit tissues are virtually unknown and has been rarely reported, but the role of GA in apple fruit ripening cannot be dismissed based on this dataset. Although ethylene biosynthesis and signaling in apple fruit maturation, ripening, and quality have been extensively studied, ethylene itself is most likely just one node in the plant hormone regulation network (Kondo et al. 2009; Kuppusamy et al. 2009; Shinshi 2008) rather than an independent or isolated pathway. With the progress in elucidating the plant hormone regulation network and their crosstalk in model plant systems (Ding et al. 2008; Murphy et al. 2002; Noh et al. 2001; Swarup et al. 2007; Teale et al. 2006; Yoo et al. 2009), significant advances of understanding the roles of their counterparts in apple fruit are expected.
TFs in transcriptional regulation network of gene expression
Transcriptional regulation is a major control point of gene function, primarily by the actions of transcription factors (TFs) to control the unique sets of genes in response to endogenous and exogenous stimuli (Riechmann et al. 2000). Consistent with the predominant roles of ethylene and auxin in apple fruit ripening regulation, a substantial portion of identified TF-encoding unigenes (39/182) were those specifically responding to these two hormones (Table 5, Table S1, and Fig. S4). Interestingly, more ethylene-specific TF-encoding unigenes identified from CP showed up-regulated expression patterns, suggesting a delayed but essential role of ethylene even for an extremely late-ripening cultivar. In contrast, the majority of auxin-specific TFs showed down-regulated patterns from CP. Whether or not the deficiency of auxin metabolism and transport (Table 3) in late-ripening CP resulted in a less synergistic interaction with ethylene will remain an interesting question. Several TF families, such as NAC, MYB, “zinc finger”, and WRKY, showed larger numbers of up-regulated unigenes in HC (Table 5, Fig. S4, and Table S3). It is likely that some other TFs such as NAC and MYB may be involved in cell wall modification and/or hormone metabolism, but further investigation is needed. It should not be surprising that some of the identified TFs were the result of different environmental factors or horticultural practice in separate commercial orchards. Nevertheless, due to the nature of tree fruit crops, it will be very difficult, if not impossible, to exclude environmental and/or horticultural conditions from sampling strategy. One example is CBF/DREB1 (contig21396), which encodes a stress-regulated transcription factor and was moderately and consistently up-regulated in CP, but showed a double-digit fold change of up-regulation from the week −4 to week −2 stage, and then a double-digit fold decrease from week −2 to week 0 in HC.
Cell wall metabolism and cultivar-specific fruit texture attributes
Plant cell wall metabolism, particularly cell wall degradation and modification, has long been associated with fruit ripening and softening (Brummell and Harpster 2001; Harker et al. 1997; Johnston et al. 2002). Individual genes encoding cell wall enzymes have been studied for their potential roles in fruit ripening and postharvest fruit softening (Brummell 2006; Dotto et al. 2006; Mann et al. 2008; Marín-Rodríguez et al. 2002; Wakasa et al. 2006). Cell wall metabolisms include multiple steps such as substrate generation, synthases and glycosyl transferases activities, secretory pathways, wall assembly, wall dynamics, and wall disassembly (Cantarel et al. 2009). It was estimated that plants devote 10 % of their genome to cell wall biogenesis alone, and among those genes currently annotated as “unknown function”, up to 1,000 of them may encode cell-wall-related proteins (Yong et al. 2005; Penning et al. 2009). From the current dataset, identified unigenes related to cell wall metabolisms represented a major transcriptomic change (Table 4 and Figs. S1c and S3). Coincident with the decreasing fruit firmness as maturation progressed, larger number of unigenes with putative functions in cell wall disassembly were identified, compared to fewer unigenes with putative roles in cell wall biosynthesis (GT) (Table 1, Table 4, and Fig. S3). As GTs are defined as those catalyzing the formation of glycosidic bonds, their functions may encompass the biosynthesis of disaccharides, oligosaccharides and complex carbohydrates, as well as glycosylation of many other molecules beyond cell wall metabolisms including protein, lipid, and plant hormones (Coutinho et al. 2003). A few gene families including XTH, WAK, and a “COBRA-like cell wall protein” showed strong cultivar-specific expression patterns. The apple genome contains at least 11 XTHs, and their activities potentially regulate several aspects of growth and development (Atkinson et al. 2009). An earlier analysis on EST frequency between the apple cultivars ‘Gala’ and ‘GoldRush’ also suggested the roles of XTHs in regulating fruit texture (Park et al. 2006). The roles of expansin, polysaccharide lyases, beta-galactosidases, and polysaccharide esterase encoding genes contributing to genotype-specific textural attributes deserve more investigation. The degradation of hemicelluloses by XTHs may also associate with cultivar-specific fruit firmness and crispness.
Unigenes annotation and their alignment with whole genome sequences
Multiple unigenes belonging to a few gene families (or functional groups), such as “auxin-induced protein 5NG4” and “Jasmonate O-methyltransferease” (Table S1), were observed to share the same annotation and protein ID but with different expression patterns. Blast search of these unigene against recently available apple whole genome sequences (Velasco et al. 2010; http://www.rosaceae.org/projects/apple_genome#publication) suggested that they belong to different individual genes in the same gene family (data not shown). It is likely that the short and incomplete coding sequence and the huge gene family size they belong to are the cause of the ambiguity in annotation. It was also observed that different unigenes were aligned at various coding regions of the same gene model. It is also possible that splicing variants of the same genes or different alleles of the same loci cause multiple ESTs sharing the same protein IDs. Or in some cases, they may simply result from errors in sequencing, assembling, detection, or data analysis.
The full bloom dates for these two cultivars are known to be only a few days apart. However, maturation and ripening of HC advances with an accelerated and condensed process, while CP progresses in a much slow-down and expanded fashion resulting in a separation of ripening time for more than 2 months. From current transcriptomics data focusing on the late maturation stages, it can be hypothesized that the inherent genetic variation at both hormone regulations and cell wall metabolisms primarily regulate apple fruit maturation patterns and fruit texture such as fruit firmness and crispness. For example, the deficiency in auxin transport and less available in biologically active auxin (from homeostasis regulation) may lead to the delayed activation of climacteric ethylene biosynthesis. Conceivably, the expanded maturation period as well as less active cell wall metabolism-related genes (such as XTHs) eventually lead to a late-ripening CP with firm fruit, in contrast to early-ripening HC with less firm but crispy fruit texture. A simplified working hypothesis can be outlined as follows: (1) The genetic variations in plant hormone metabolism and responses, such as ethylene responses, auxin homeostasis, and transport, could fundamentally impact the apple fruit ripening process. (2) Cultivar-specific plant hormone metabolism and crosstalk will activate or suppress a unique set of TFs and/or signal transduction components. (3) Consequently, distinct sets of genes in various metabolic pathways, such as those encoding cell wall modifying enzymes of XTHs, are regulated in a cultivar-specific fashion. The overall outcomes of such differential gene expression ultimately determine the unique features of fruit ripening behaviors and quality attributes. Selected unigenes from current analysis are being aligned to apple genome sequences, and their expression profiles are being investigated in an HC × CP cross population consisting of 170 fruiting trees and other more diverse apple germplasm. The expression patterns of these genes will be analyzed against apple ripening and texture phenotypes to further define the specific function of these candidate genes.
References
Argenta L, Fan X, Mattheis J (2002) Responses of ‘Fuji’ apples to short and long duration exposure to elevated CO2 concentration. Postharv Biol Tech 24:13–24
Atkinson RG, Johnston SL, Yauk Y-K, Sharma NN, Schröder R (2009) Analysis of xyloglucan endotransglucosylase/hydrolase (XTH) gene families in kiwifruit and apple. Postharv Biol Technol 51:149–157
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc B (Methodological) 57:289–300
Bennett AB, Labavitch JM (2008) Ethylene and ripening-regulated expression and function of fruit cell wall modifying proteins. Plant Sci 175:130–136
Benson DA, Karsch-Mizrachi I, Lipman DJ, Ostell J, Wheeler DL (2007) GenBank. Nucleic Acids Res 35:D21–D25
Blanpied GD, Silsby KJ (1992) Predicting harvest date windows for apples. Cornell Coop Ext Publ Inform Bul 221:12
Bleecker A, Kende H (2000) Ethylene: a gaseous signal molecule in plants. Annu Rev Cell Dev Biol 16:1–18
Brookfield P, Murphy P, Harker R, MacRae E (1997) Starch degradation and starch pattern indices; interpretation and relationship to maturity. Postharv Biol Technol 11:23–30
Brummell DA (2006) Cell wall disassembly in ripening fruit. Funct Plant Biol 33:103–119
Brummell D, Harpster M (2001) Cell wall metabolism in fruit softening and quality and its manipulation in transgenic plants. Plant Mol Biol 47:311–340
Cantarel BL, Coutinho PM, Rancurel C, Bernard T, Lombard V, Henrissat B (2009) The Carbohydrate-Active EnZymes database (CAZy): an expert resource for glycogenomics. Nucleic Acids Res 37:D233–D238
Chandler PM, Marion-Poll A, Ellis M, Gubler G (2002) Mutants at Slender1 locus of barley cv Himalaya. Molecular and physiological characterization. Plant Physiol 129:181–190
Costa FS, Sara W, Van de Weg E, Guerra W, Cecchinel M, Dallivina J, Koller B, Sansivini S (2005) Role of the genes Md-ACO1 and Md-ACS1 in ethylene production and shelf life of apple (Malus domestica Borkh). Euphytica 141:181–190
Costa F, Alba R, Schouten H, Soglio V, Gianfranceschi L, Serra S, Musacchi S, Sansavini S, Costa G, Fei Z, Giovannoni J (2010) Use of homologous and heterologous gene expression profiling tools to characterize transcription dynamics during apple fruit maturation and ripening. BMC Plant Biol 10:229
Coutinho PM, Deleury E, Davies GJ, Henrissat B (2003) An evolving hierarchical family classification for glycosyltransferases. J Mol Biol 328:307–317
Ding X, Cao Y, Huang L, Zhao J, Xu C, Li X, Wang S (2008) Activation of the indole-3-acetic acid–amido synthetase GH3-8 suppresses expansin expression and promotes salicylate- and jasmonate-independent basal immunity in rice. Plant Cell 20:228–240
Dotto MC, Martínez GA, Civello PM (2006) Expression of expansin genes in strawberry varieties with contrasting fruit firmness. Plant Physiol Biochem 44:301–307
Evans K, Brutcher L, Konishi B, Barritt B (2010) Correlation of sensory analysis with physical textural data from a computerized penetrometer in the Washington State University apple breeding program. HortScience 20:1026–1029
Falara V, Manganaris GA, Ziliotto F, Manganaris A, Bonghi C, Ramina A, Kanellis AK (2011) A β-D: -xylosidase and a PR-4B precursor identified as genes accounting for differences in peach cold storage tolerance. Funct Integr Genomics 11:357–368
Fan X, Mattheis JP, Fellman JK (1998) A role for jasmonates in climacteric fruit ripening. Planta 204:444–449
Gagne JM, Smalle J, Gingerich DJ, Walker JM, Yoo S-D, Yanagisawa S, Vierstra RD (2004) Arabidopsis EIN3-binding F-box 1 and 2 form ubiquitin–protein ligases that repress ethylene action and promote growth by directing EIN3 degradation. Proc Natl Acad Sci USA 101:6803–6808
Gasic K, Hernandez A, Korban S (2004) RNA extraction from different apple tissues rich in polyphenols and polysaccharides for cDNA library construction. Plant Mol Biol Report 22:437a–437g
Giovannoni J (2004) Genetic regulation of fruit development and ripening. Plant Cell 16:170–180
Harada T, Sunako T, Wakasa Y, Soejima J, Satoh T, Niizeki M (2000) An allele of 1-aminocyclopropane-1-carboxylate synthase gene (Md-ACS1) accounts for the low ethylene production in climacteric fruits of some apple cultivars. Theor Appl Genet 101:742–746
Harker RR, Redgwell RJ, Hallett IC, Murray SH (1997) Texture of fresh fruit. Hortic Rev 20:121–224
Hou B, Lim E-K, Higgins GS, Bowles DJ (2004) N-glucosylation of cytokinins by glycosyltransferases of Arabidopsis thaliana. J Biol Chem 279:47822–47832
Huang X, Madan A (1999) CAP3: a DNA sequence assembly program. Genome Res 9:868–877
Jain M, Kaur N, Tyagi AK, Khurana JP (2006) The auxin-responsive GH3 gene family in rice (Oryza sativa). Funct Integr Genomics 6:36–46
Janssen BJ, Thodey K, Schaffer RJ, Alba R, Balakrishnan L, Bishop R, Bowen JH, Crowhurst RN, Gleave AP, Ledger S, McArtney S, Pichler FB, Snowden KC, Ward S (2008) Global gene expression analysis of apple fruit development from the floral bud to ripe fruit. BMC Plant Biol 8:16
Jensen PJ, Makalowska I, Altman N, Fazio G, Praul C, Maximova SN, Crassweller RM, Travis JW, McNellis TW (2010) Rootstock-regulated gene expression patterns in apple tree scions. Tree Genet Genome 6:57–72
Johnston JW, Hewett EW, Hertog MLATM (2002) Postharvest softening of apple (Malus domestica) fruit: a review. N Z J Crop Hortic Sci 30:145–160
Jung S, Staton M, Lee T, Blenda A, Svancara R, Abbott A, Main D (2008) GDR (Genome Database for Rosaceae): integrated web-database for Rosaceae genomics and genetics data. Nucleic Acids Res 36:D1034–D1040
Kazan K, Manners JM (2008) Jasmonate signaling: toward an integrated view. Plant Physiol 146:1459–1468
Kondo S, Meemak S, Ban Y, Moriguchi T, Harada T (2009) Effects of auxin and jasmonates on 1-aminocyclopropane-1-carboxylate (ACC) synthase and ACC oxidase gene expression during ripening of apple fruit. Postharv Biol Tech 51:281–284
Kuppusamy KT, Walcher CL, Nemhauser JL (2009) Cross-regulatory mechanisms in hormone signaling. Plant Mol Biol 69:375–381
Lee YP, Yu GH, Seo YS, Han SE, Choi YO, Kim D, Mok IG, Kim WT, Sung SK (2007) Microarray analysis of apple gene expression engaged in early fruit development. Plant Cell Rep 26:917–926
Li J, Yuan R (2008) NAA and ethylene regulate expression of genes related to ethylene biosynthesis, perception, and cell wall degradation during fruit abscission and ripening in ‘Delicious’ apples. J Plant Growth Regul 27:283–295
Mann HS, Alton JJ, Kim S, Tong CBS (2008) Differential expression of cell-wall-modifying genes and novel cDNA in apple fruit during storage. J Am Soc Hortic Sci 133:153–164
Marín-Rodríguez MC, Orchard J, Seymour GB (2002) Pectate lyases, cell wall degradation and fruit softening. J Exp Bot 53:2115–2119
Mazzola M, White FF (1994) A mutation in the indole-3-acetic acid biosynthesis pathway of Pseudomonas syringae pv. syringae affects growth in Phaseolus vulgaris and syringomycin production. J Bacteriol 176:1374–1382
Mulder NJ, Apweiler R, Attwood TK, Bairoch A et al (2007) New developments in the InterPro database. Nucleic Acids Res 35:D224–D228
Murphy AS, Hoogner KR, Peer WA, Taiz L (2002) Identification, purification, and molecular cloning of N-1-naphthylphthalmic acid-binding plasma membrane-associated aminopeptidases from Arabidopsis. Plant Physiol 128:935–950
Noh B, Murphy AS, Spalding EP (2001) Multidrug resistance-like genes of Arabidopsis required for auxin transport and auxin-mediated development. Plant Cell 13:2441–2454
Oraguzie NC, Iwanami H, Soejima J, Harada T, Hall A (2004) Inheritance of Md-ACS1 gene and its relationship to fruit softening in apple (Malus × domestica Borkh.). Theor Appl Genet 108:1526–1533
Park S, Sugimoto N, Larson MD, Beaudry R, van Nocker S (2006) Identification of genes with potential roles in apple fruit development and biochemistry through large-scale statistical analysis of expressed sequence tags. Plant Physiol 141:811–824
Penning BW, Hunter CT III, Tayengwa R, Eveland AL, Dugard CK, Olek AT, Vermerris W, Koch KE, McCarty DR, Davis MF, Thomas SR, McCann MC, Carpita NC (2009) Genetic resources for maize cell wall biology. Plant Physiol 151:1703–1728
Petrášek J, Mravec J, Bouchard R, Blakeslee JJ, Abas M, Seifertová D, Wiśniewska J, Tadele Z, Kubeš M, Čovanová M, Dhonukshe P, Skůpa P, Benková E, Perry L, Křeček P, Lee OR, Fink GR, Geisler M, Murphy AS, Luschnig C, Zažímalová E, Friml J (2006) PIN protein perform a rate-limiting function in cellular auxin efflux. Science 312:914–918
Rahman A, Hosokawa S, Oono Y, Amakawa T, Goto N, Tsurumi S (2002) Auxin and ethylene response interactions during Arabidopsis root hair development dissected by auxin influx modulations. Plant Physiol 130:1908–1917
Riechmann JL, Heard J, Martin G, Reuber L, Jiang CZ, Keddie J, Adam L, Pineda O, Ratcliffe OJ, Samaha RR, Creelman R, Pilgrim M, Broun P, Zhang JZ, Ghandehari D, Sherman BK, Yu G-L (2000) Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes. Science 290:2105–2110
Rosenfield CL, Kiss E, Hrazdina G (1996) Md-ACS-2 (accession no. U73815) and Md-ACS-3 (accession no. U73816): two new 1-aminocyclopropane-1-carboxylate synthases in ripening apple fruit (PGR96–122). Plant Physiol 112:1735
Roudier F, Fernandez AG, Fujita M, Himmelspach R, Borner GH, Schindelman G, Song S, Baskin TI, Dupree P, Wasteneys GO, Benfey PN (2005) COBRA, an Arabidopsis extracellular glycosylphosphatidylinositol-anchored protein specifically controls highly anisotropic expansion through its involvement in cellulose microfibril orientation. Plant Cell 17:1749–1763
Schaffer RJ, Friel EN, Souleyre EJF, Bolitho K, Thodey K, Ledger S, Bowen JH, Ma JH, Nain B, Cohen D, Gleave AP, Crowhurst RN, Janssen BJ, Yao JL, Newcomb RD (2007) A genomics approach reveals that aroma production in apple is controlled by ethylene predominantly at the final step in each biosynthetic pathway. Plant Physiol 144:1899–1912
Schmuelling T, Werner T, Riefler M, Krupkova E, Bartrina Y, Manns I (2003) Structure and function of cytokinin oxidase/dehydrogenase genes of maize, rice, Arabidopsis and other species. J Plant Res 116:241–252
Seymour G, Manning K, Eriksson E, Popovich A, King G (2002) Genetic identification and genomic organization of factors affecting fruit texture. J Exp Bot 53:2065–2071
Shinshi H (2008) Ethylene-regulated transcription and crosstalk with jasmonic acid. Plant Sci 175:18–23
Spartz AK, Gray WM (2008) Plant hormone receptors: new perceptions. Gene Dev 22:2139–2148
Stepanova AN, Hoyt JM, Hamilton AA, Alonso JM (2005) A link between ethylene and auxin uncovered by the characterization of two root-specific ethylene-insensitive mutants in Arabidopsis. Plant Cell 17:2230–2242
Swarup R, Parry G, Graham N, Allen T, Bennett M (2002) Auxin cross-talk: integration of signalling pathways to control plant development. Plant Mol Biol 49:411–426
Swarup R, Perry P, Hagenbeek D, Van Der Straeten D, Beemster GT, Sandberg G, Bhalerao R, Ljung K, Bennett MJ (2007) Ethylene upregulates auxin biosynthesis in Arabidopsis seedlings to enhance inhibition of root cell elongation. Plant Cell 19:2186–2196
Teale WD, Paponov IA, Palme K (2006) Auxin in action: signalling, transport and the control of plant growth and development. Nat Rev Mol Cell Biol 7:847–859
Thomas SG, Phillips AL, Hedden P (1999) Molecular cloning and functional expression of gibberellin 2-oxidases, multifunctional enzymes involved in gibberellin deactivation. Proc Natl Acad Sci USA 96:4698–4703
Trainotti L, Tadiello A, Casadoro G (2007) The involvement of auxin in the ripening of climacteric fruits comes of age: the hormone plays a role of its own and has an intense interplay with ethylene in ripening peaches. J Exp Bot 58:3299–3308
Ueguchi-Tanaka M, Nakajima M, Itoh H, Katoh E, Kobayashi M, Chow T-Y, Hsing Y-I, Kitano H, Yamaguchi I, Matsuoka M (2005) GIBBERELLIN INSENSITIVE WARF1 encodes a soluble receptor for gibberellin. Nature 437:693–698
Vandenbussche F, Van Der Straeten D (2007) One for all and all for one: cross-talk of multiple signals controlling the plant phenotype. J Plant Growth Regul 26:178–187
Varanasi V, Mattheis JP, Rudell D, Zhu Y (2011) Expression profiles of MdACS3 gene suggest function as an accelerator of apple (Malus × domestica) fruit ripening. Postharvest Biol Technol 62:141–148
Velasco R, Zharkikh A, Affourtit J, Dhingra A et al (2010) The genome of the domesticated apple (Malus × domestica Borkh.). Nat Genet 42:833–839
Verica JA, Chae L, Tong H, Ingmire P, He Z-H (2003) Tissue-specific and developmentally regulated expression of a cluster of tandemly arrayed cell wall-associated kinase-like kinase genes in Arabidopsis. Plant Physiol 133:1732–1746
Vizoso P, Meisel LA, Tittarelli A, Latorre M, Saba J, Caroca R, Maldonado J, Cambiazo V, Campos-Vargas R, Gonzalez M, Orellana A, Silva H (2009) Comparative EST transcript profiling of peach fruits under different post-harvest conditions reveals candidate genes associated with peach fruit quality. BMC Genomics 10:423
Wakasa Y, Kudo H, Ishikawa R, Akada S, Senda M, Niizeki M, Harada T (2006) Low expression of an endopolygalacturonase gene in apple fruit with long-term storage potential. Postharv Biol Tech 39:193–198
Wiersma PA, Zhang H, Lu C, Quail A, Toivonen PMA (2007) Survey of the expression of genes for ethylene synthesis and perception during maturation and ripening of ‘Sunrise’ and ‘Golden Delicious’ apple fruit. Postharv Biol Technol 44:204–211
Wu CH, Apweiler R, Bairoch A, Natale DA, Barker WC, Boeckmann B, Ferro S, Gasteiger E, Huang H, Lopez R, Magrane M, Martin MJ, Mazumder R, O'Donovan C, Redaschi N, Suzek B (2006) The universal protein resource (UniProt): an expanding universe of protein information. Nucleic Acids Res 34:D187–D191
Yong W, O’Malley R, Link B, Binder B, Bleecker A, Koch KE, McCann MC, McCarty DR, Patterson S, Reiter WD et al (2005) Genomics of plant cell wall biosynthesis. Planta 221:747–751
Yoo SD, Cho Y, Sheen J (2009) Emerging connections in the ethylene signaling network. Trends Plant Sci 14:270–279
Yoshida H, Nagata M, Saito K, Wang KLC, Ecker JR (2005) Arabidopsis ETO1 specifically interacts with and negatively regulates type 2 1-aminocyclopropane-1-carboxylate synthases. BMC Plant Biol 5:14
Zhu Y, Barritt BH (2008) Apple cultivar genotypes for Md-ACS1 and Md-ACO1 ethylene production genes and implications for breeding. Tree Genet Genome 4:555–562
Zhu Y, Rudell D, Mattheis JP (2008) Characterization of cultivar differences in alcohol acyltransferase and 1-aminocyclopropane-1-carboxylate synthase gene expression and volatile ester emission during apple fruit maturation and ripening. Postharv Biol Technol 49:330–339
Ziliotto F, Begheldo M, Rasori A, Bonghi C, Tonutti P (2008) Transcriptome profiling of ripening nectarine (Prunus persica L. Batsch) fruit treated with 1-MCP. J Exp Bot 59:2781–2791
Acknowledgments
The authors are grateful to Ann Callahan, Chris Dardick, Cindy Tong, and Tianbao Yang for their helpful comments. The authors wish to thank Xia Rui for preparation of Table S1 and Table S2. We wish to thank Mallela Magana, Janie Countryman, Edward Valdez, and Bruce Mackey for their contribution in fruit harvest, maturity test, tissue collection, and other excellent technical assistance. This study was supported by Washington Tree Fruit Research Commission and USDA base fund.
The authors declare that the experiments comply with the current laws of USA. All authors read and approved the final manuscript, and declared no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by A. Dandekar
Yanmin Zhu and Ping Zheng contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S1
Overall distribution of identified unigenes in major functional categories. a Distribution of different expression patterns among identified unigenes. Column with green color represented those identified from HC, and column with pink color was from CP. Values of Y-axis indicated the numbers of unigenes. X-axis indicates four different expression patterns for both cultivars. Refer to Tables 3, 4, and S1, S2, S3 and S4 for detailed information of individual unigenes. b Cultivar specificity of identified unigenes implicated in plant hormone metabolism and responses. c Cultivar specificity of unigenes implicated in cell wall metabolisms. d Cultivar specificity of TF-encoding unigenes. In (b), (c), and (d), Y-axis showed the numbers of up- or down-regulated values (all unigenes have two values of fold change). Column with blue color represented those identified with up-regulated expression pattern, and column with red color represented those identified with down-regulated expression pattern (JPEG 105 kb)
Fig. S2
Cultivar-specific expression patterns for unigenes in plant hormone functions. a Unigenes implicated in ethylene metabolism and response. b Unigenes implicated in auxin metabolism, transport, and response. c Unigenes implicated in GA metabolism and response. d Unigenes implicated in the metabolism and response of other plant hormones. Y-axis showed the up- or down-regulated values of identified unigenes (all unigenes have two values of fold changes). Column with blue color represented those identified with up-regulated expression pattern, and column with red color represented those identified with up-regulated expression pattern (JPEG 90 kb)
Fig. S3
Cultivar-specific expression patterns for unigenes in carbohydrate metabolism and cell wall modification. a Unigenes encoding glycosyl transferase (GT). b Unigenes encoding glycosyl hydrolase (GH). c Unigenes encoding polysaccharide lyase and carbohydrate esterase. d Unigenes encoding expansin and other cell wall proteins. Y-axis showed the number of either up- or down-regulated values (all unigenes have two values of fold changes). Column with blue color represented those identified with up-regulated expression pattern, and column with red color represented those identified with up-regulated expression pattern (JPEG 93 kb)
Fig. S4
Cultivar-specific expression patterns of unigenes encoding TFs. a Unigenes encoding transcription factors which specifically respond to ethylene. b Unigenes encoding transcription factors which specifically respond to auxin. c Unigenes encoding transcription factors belong to other TF families. Y-axis showed the number of either up- or down-regulated value (all unigenes have two values of fold changes). In (b) and (c), column with blue color represented those identified with up-regulated expression pattern, and column with red color represented those identified with up-regulated expression pattern (JPEG 90 kb)
Fig. S5
Relative expression level of randomly selected unigenes by qRT-PCR. Bars represent normalized values of qRT-PCR assay; two independent RNA samples were used in separated cDNA synthesis and PCR reactions were repeated twice in triplicates for each cDNA. Y-axis indicated the relative gene expression, and X-axis showed samples at different maturation stages. Names of the selected unigenes were placed at the top of each square. Values at the bottom of each square were the fold changes based from microarray dataset one for between week −4 and week −2 and another for week −2 to week 0. Numbers highlighted in red color represented the consistent gene expression patterns between microarray analysis and qRT-PCR assay, while those highlighted in blue color represented inconsistent patterns between two methods. HC ‘Honeycrisp’, CP ‘Cripps Pink’ (JPEG 896 kb)
Table S1
Visual representation of differentially expressed unigenes with the annotated function in plant hormone metabolism and responses (DOC 503 kb)
Table S2
Visual representation of differentially expressed unigenes with the annotated function in cell wall metabolism and modification (DOC 604 kb)
Table S3
Differentially expressed transcription factor encoding unigenes (DOC 280 kb)
Table S4
Differentially expressed unigenes with other annotated molecular functions (XLS 70 kb)
Table S5
Primer sequences used for data validation by qRT-PCR (DOC 33 kb)
Rights and permissions
About this article
Cite this article
Zhu, Y., Zheng, P., Varanasi, V. et al. Multiple plant hormones and cell wall metabolism regulate apple fruit maturation patterns and texture attributes. Tree Genetics & Genomes 8, 1389–1406 (2012). https://doi.org/10.1007/s11295-012-0526-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11295-012-0526-3