Abstract
The northernmost range of beech (Fagus sylvatica L.) is in southern Norway and consists of two distinct and isolated distributions, a single population at Seim in West Norway and several adjacent populations in Vestfold, East Norway. The modest beech pollen deposits beyond these main distributions suggest that the Norwegian beech distribution has never been an extension of the south Scandinavian range. We used genetic markers and historical sources to trace the ancestor populations for the beech at Seim and Vestfold, hypothesising Denmark as the most likely source. Nuclear inter-simple sequence repeat markers, amplified by polymerase chain reaction (PCR), were applied to estimate genetic distances between beech populations in Norway, England and Denmark. The variation in chloroplast DNA polymorphism was estimated using PCR-restriction fragment length polymorphism. The nuclear genetic data indicate Denmark as a source for the beech in Norway, although the data are less certain in the case of Seim than in that of Vestfold. The populations from South England were genetically different from most Scandinavian populations. The genetic variation within Norwegian populations was only slightly lower than that of the English and Danish populations, questioning birds as vectors for dispersal. Thus, the pollen data and our results are in accordance with the intentional introduction and documented human migrations across Skagerrak before and during the Viking Age.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The common beech (Fagus sylvatica L.) is widespread in most of Europe. It is presently distributed from northeastern Spain in the west to the Carpathians and Baltic mountains in the east and from Sicily in the south to southern Sweden and Norway in the north (Hultén and Fries 1986). The Holocene spread of beech to southern Scandinavia occurred about 4,000–3,500 years BP (Björkman 1999; Bradshaw and Lindbladh 2005). The present distribution of beech in its northernmost range is discontinuous and consists of two distinct areas, Vestfold in East Norway, which is the centre of beech in Norway, and Seim, West Norway, which is the world's northernmost beech population (Hultén 1971). The distance between the two occurrences in Norway is 330 km, as the crow flies, and about 500 km along the coast. The seeds of beech are animal-dispersed (Nilsson 1985). In northwest Europe, jays (Garrulus glandarius) are the main vectors for long-distance dispersal (Nilsson 1985), and jays commonly hatch in all the places where beech is presently distributed in Scandinavia (Benz 1988).
Due to its extensive isolation, the geographic origin of the Seim population has been a matter of interest for almost two centuries (Fægri 1954). The population is about 30 ha, and pollen analyses have shown that it was established between 500 and 1,000 AD (Fægri 1954). Based on the appearance after agriculture, intentional planting was considered the most likely mechanism of establishment (Fægri 1954). This is in agreement with the fact that the spread of beech north of the Alps benefited from human forest clearings (Küster 1997; Bradshaw and Lindbladh 2005). The transfer of beech seeds by jays is usually several 100 m, but it has not been reported to exceed 6 km (Nilsson 1985). Accepting human transfer as the mode of spread, Fægri (1954) indicated three possible origins for the Seim beech based on interpretation of historical sources. The preferred assumption is a transfer from the populations in Vestfold, based on information given by the saga writer Snorre Sturlason in the early thirteenth century (Sturluson 1941) about Seim belonging to King Harald Finehair, and that Harald's point of departure was Vestfold. Both King Harald Finehair and Håkon, the son of Harald Finehair, were related to Seim (Sturluson 1941). Second, Fægri (1954) points to King Harald's relation with the English King Æthelstan who fostered Håkon. Æthelstan was the King of Wessex (924–939) who held Winchester as a capital, although the king's place of residence cannot be determined (Keene 1976). Finally, Denmark was suggested as a possible source of the beech. Beech had spread to all the possible origins suggested here by the predicted time of establishment at Seim (Bradshaw 2004). The beech forests in Vestfold are considerably larger than the Seim population and comprise about 2,000 ha distributed over several adjacent populations (Øyen 2010). The dating of the establishment of the beech populations in Vestfold varies from 650 AD (Jerpåsen 1996) to 800 AD (Henningsmoen 1988; Sørensen et al. 2007), depending on the site. Due to the proximity to beech in southern Sweden and Denmark, the origin of the Vestfold beech has not attracted the same interest as the beech at Seim. Anthropogenic introduction has, however, been suggested, based on the scarce pollen records beyond the Vestfold populations (e.g. Henningsmoen 1988; Jerpåsen 1996), which complies with the close contact with Denmark during the period that beech became established in Vestfold (e.g. Ilkjær 2000; Skre 2007).
Molecular markers have been proven useful in tracing refugia and colonisation routes of the European biota following the last glaciation (Hewitt 1999). In angiosperms such as beech, variation in the non-recombining and maternally inherited chloroplast DNA (cpDNA; e.g. Demesure et al. 1996) has allowed the construction of maps for the Holocene migration of numerous European tree species (Taberlet et al. 1998; Hewitt 1999). Several complementary molecular markers should be used in concert to provide a more complete picture (Godinho et al. 2008), and in addition to the cpDNA markers, we used inter-simple sequence repeat (ISSR) markers as nuclear markers to trace possible origins for the beech at Seim and Vestfold. Many ISSR primers are available, and they are easy to design as no prior knowledge of sequence is required (Tsumura et al. 1996; Yin et al. 2002). These features, as well as the high polymorphism and strong discriminative power of ISSR markers (e.g. Tantasawat et al. 2010) suggest their applicability in phylogenetic studies (e.g. Mort et al. 2003). However, tracing the origin of the northern outskirt populations of beech is complicated by the long time since the introductions. Thus, the present work represents a special case in which historical sources have been applied to identify possible origin populations, which have been collected and analysed by chloroplast and nuclear genetic markers. Based on the historical sources and their recent interpretations, we assume that the ancestor populations for the Norwegian beech populations are in Denmark. Alternatively, the Seim beech may originate from South England or Vestfold. We further used the level of genetic diversity to infer mode of dispersal. High genetic diversity would strongly suggest intentional introduction based on many seeds, whereas limited genetic diversity could imply historic founder effects (e.g. Aizawa et al. 2009) associated with random dispersal by birds.
Material and methods
Sampling and DNA extraction
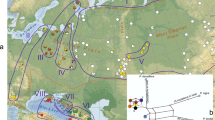
In Norway, beech material was collected from populations in Vestfold County in East Norway and from the isolated population at Seim in West Norway. In Denmark, populations were collected from the main islands, Sjælland (Sor) and Fyn (Fyn), as well as the eastern side of Jutland (Rin, Sil) on the Danish mainland (Table 1, Fig. 1). Rin is located between the Viking cities of Ribe (present Denmark) and Hedeby (present Germany), about 60 km from each of them. Sor is located close to several marked places of the first millennium AD (Skre 2007). The English populations were fairly concentrated to the surroundings of Winchester, South England. To obtain autochthonous beech material, all populations were carefully selected from ancient natural or semi-natural beech woods, in which human disturbance (e.g. silviculture) has been absent or limited (Peterken 1996).
Origins of the beech populations studied (open symbols) and northern distribution limits of beech in Scandinavia and England (Björkman 1996; EUFORGEN 2009). The solid line refers to the range of beech forests, whereas the closed symbols show the distribution of outlying beech stands or scattered trees where beech has always been rare (Björkman 1996). The longest distance between the subpopulations at Seim is 1.7 km, and the longest distance between the individual populations in Vestfold and England are 50 and 36 km, respectively
At each site, one twig (≈20–30 cm) was collected from about 50 trees and then kept in the freezer at −20°C. The trees had an estimated age of 80 to 100 years and were separated by 50 to 100 m. The distance between sample trees was chosen due to the limited pollen flow in beech stands and associated clustering of relatives within short distances (Vornam and Herzog 2004). Sampling of the Seim population was split into four subpopulations, according to the spatial distribution, which also offered a possibility to study the uniformity of the population.
Total DNA from bud tissue was extracted using Qiagen DNEasy plant mini extraction kit (Qiagen, #69106). Five randomly chosen individuals from each population were subjected to further PCR-restriction fragment length polymorphism (RFLP) analysis, and 48 individuals from each population were used for ISSR analysis.
Chloroplast DNA analysis
Chloroplast DNA polymorphism was estimated using PCR-RFLPs. A set of five pairs of universal primers: trnD—trnT (DT), trnC—trnD (SD), psaA—trnS (AS), trnS—trnT (ST) and trnS—trnf M (SfM) were chosen as described by Demesure et al. (1996). Primer sequences and PCR amplification parameters were obtained from Demesure et al. (1995), Dumolin-Lapegue et al. (1997) and notation follows Grivet et al. (2001). PCR products (10 μl) were digested with the restriction enzymes TaqI, MseI and HinfI (New England Biolabs, USA).
ISSR PCR amplification
Seven ISSR primers with dinucleotide motifs were assayed (Table 2). PCR amplification reactions (15 μl total volume) were performed using touchdown PCR in 1× PCR buffer, 1× Q-solution, 200 μM of each dNTPs, 500 μM of primer, 0.5 U of Taq polymerase and 10 ng of genomic DNA using QIAGEN Taq DNA polymerase kit (Qiagen, #201205). The touchdown PCR cycles started with initial denaturation at 95°C for 5 min. The programme proceeded with ten cycles of touchdown PCR: 95°C for 30 s, annealing temperature (Ta) for 30 s, then 2 min at 72°C. The Ta was decreased from 52 to 48°C by 0.4°C steps for each of the ten subsequent touchdown cycles. Then, PCR followed by 30 cycles of 30 s at 95°C, 45 s at 48°C, 2 min at 72°C and a final extension step of 7 min at 72°C. PCRs were carried out in a GeneAmp® PCR System 9700 (Applied Biosystems). The PCR products were separated on 2% (w/v) agarose gels in TBE buffer at 150 V during 1.5 h. Gels were stained with ethidium bromide and photographed using SYNGENE BioImaging System.
Data analysis
The ISSR banding patterns were scored as presence (1)/absence (0) of each amplified band. Genetic diversity data were calculated using PopGene32 (Yeh et al. 1999) and TFPGA software (Miller 1997). Population genetic structure was also analysed using Bayesian approaches (Holsinger and Lewis 2003). Following the notation in Holsinger and Wallace (2004), f refers to F IS, and θ B refers to F ST. Cluster analysis of individuals by their haplotypes was based on similarity matrices using the unweighted pair group method analysis (UPGMA) and the neighbour-joining (NJ) method programme in the WINBOOT software (Yap and Nelson 1996) and the TreeFit software (Kalinowski 2009), respectively, and the degree of fit to a matrix of genetic distances (R 2; Kalinowski 2009) was calculated. Visualisation of dendrograms was done using the TreeView programme (Page 1996). In addition to the cluster analysis, we used STRUCTURE 2.2 (Pritchard et al. 2000; Falush et al. 2003), which, by means of iterative algorithms, identifies clusters of related individuals from multilocus genotypes. We performed many runs of various lengths with different quires for number of genetic clusters (K) represented by the individuals genotyped, testing all values of K from 1 (single populations) to 17 (all populations).
Results
Chloroplast DNA
None of the nine primer/enzyme pairs used as cpDNA markers was polymorphic in the studied regions, which implies that all samples from Norway, Denmark and South England share the same cpDNA haplotype.
Genetic structure of beech populations
Nuclear DNA analyses using ISSRs showed high levels of diversity for the beech populations in all countries (Table 3). A total of 75 fragments were scored for the seven ISSR primers, of which seven bands were monomorphic and 68 were polymorphic. Estimates of total genetic diversity within countries (H T) and populations within countries (H S) showed slightly higher levels in English and Danish populations than in Norwegian populations (Table 3). Among the Norwegian populations, the genetic diversity of the Seim population was similar to the populations in Vestfold (Table 4). There were rather small differences in the observed (n a) and effective (n e) number of alleles between countries (Table 3) and populations (Table 4). ISSRs are dominant markers, and we do not expect the Hardy–Weinberg equilibrium (HWE) in this study where several populations (at Seim) are introduced. Therefore, instead of h and its related parameters, it may be better to consider Shannon's index (I) estimates to compare genetic variation because it does not assume HWE (Table 3). I varies from 0.28 in Seim 1 till 0.41 in the Rin population (average, 0.34). Thus, in total, the studied populations did not differ considerably in the genetic diversity estimates. The Norwegian populations had high interpopulation differentiation (G ST = 0.203) and a low rate of gene flow (N m = 1.961), whereas English populations had a low level of interpopulation differentiation (G ST = 0.103) and a high rate of gene flow (N m = 4.373). The beech populations from Denmark had intermediate estimates (Table 3).
We evaluated the point and interval estimates of f and θ B for the full model considered optimal (supplementary material S1) and calculated posterior means and 95% confidence intervals together with posterior distributions and sample traces for f and θ B estimates (supplementary material S2). The posterior mean of f is 0.3, with the 95% confidence interval varying from 0.022 to 0.754. Thus, the average level of inbreeding in the populations was quite high. The posterior mean of θ B is 0.439 (0.376–0.506). In total, this Bayesian approach gave very robust estimates of genetic diversity and interpopulation differentiation.
Genetic relations among populations
Genetic similarity between the 17 populations was analysed on the basis of genetic distance matrix calculated using Nei's (1978) unbiased distance (D N) and shows that the connection was equally strong between the populations in Vestfold and Denmark as between the adjacent populations in Vestfold (Table 5). In the case of the Seim population, only one subpopulation was significantly related to a Danish population. The dendrogram structures of UPGMA (R 2 = 0,604) and NJ (R 2 = 0,843) were rather similar (supplementary material S3) and in agreement with Nei's genetic distances (Table 5), clearly clustering populations from South England and Seim, respectively, and less distinctly, populations from Denmark and East Norway. This further conforms to the 3-cluster result of STRUCTURE 2.2 (supplementary material S4), which, in agreement with the NJ dendrogram, also suggests genetic proximity between the populations from Denmark and Seim. The last cluster of STRUCTURE 2.2 combines all populations from England and one population from Vestfold (Sto). The 7-cluster result combines adjacent populations within countries and, in some cases, only individuals within populations.
Discussion
Genetic relations between and within the populations
According to pollen records, beech colonised southern Scandinavia 4,000–3,500 BP (Björkman 1999; Bradshaw and Lindbladh 2005), whereas the first establishment in East Norway did not take place until about 650 AD (Henningsmoen 1988; Sørensen et al. 2007). The pollen records also show that the beech in Norway has never been a continuous extension of the south Scandinavian range (Fig. 1; e.g. Hafsten 1956; Danielsen 1970; Björkman 1996), leaving a question of the origin of the beech in Norway. In our data, there was a clear genetic similarity between the populations in Vestfold and Denmark (Table 5). In fact, the individual Vestfold populations were as equally related to the distant Danish populations as the adjoining populations in Vestfold (Table 5), which indicates a Danish genetic legacy of the Vestfold beech. Although the bootstrap support for this node was limited (supplementary material S3), analysis of population structure by means of STRUCTURE 2.2 (Pritchard et al. 2000; Falush et al. 2003) also suggests a genetic proximity between the Danish and East Norwegian populations (supplementary material S4). The subpopulations at Seim were genetically uniform, and the UPGMA dendrogram (supplementary material S3) does not suggest that the Seim population was related to any of the other populations. The STRUCTURE 2.2 approach, however, (Pritchard et al. 2000; Falush et al. 2003) combines populations from Denmark and Seim in a separate cluster (supplementary material S3), as also indicated by the NJ dendrogram (supplementary material S3) and by Nei's genetic distances (Table 5). This result challenges the prevailing assumption that Vestfold was the origin of the beech at Seim (e.g. Fægri 1954). If the Seim population became established in the initial part of the suggested dating (500–1,000 AD; Fægri 1954), chronology also questions descent from the beech at Vestfold, established 650–800 AD (Henningsmoen 1988; Jerpåsen 1996; Sørensen et al. 2007). The beech populations from England were distinct and generally unrelated to the Scandinavian beech populations (Table 5; supplementary material S4), leaving limited genetic evidence that introduction to Norway took place from across the North Sea. Thus, the results are in favour of introduction from Denmark, which is consistent with our prime hypothesis.
An alternative to introduction of beech from Denmark is spread by natural means from southwestern Sweden. During the Holocene, beech populations did not spread as a moving front, but with diffuse spread from scattered nuclei (Magri 2008). Austerlitz et al. (2000) has emphasised the importance of the long juvenile phase of trees for avoiding founder effects during repeated establishments, which may allow a large increase in the number of initial founders and thus, accumulation of genetic variation, before reproduction begins. The requirement for delayed reproduction is fully met in beech, which starts to flower earliest at the age of 40 to 50 years (Giesecke et al. 2007). The caching behaviour of jays is significant for the spread and population dynamics of beech (Johnson et al. 1997), and the seeds are usually transported several 100 m into open habitats (Vander Wall 2001; Kunstler et al. 2004). Thus, the role of human forest clearings for the spread of beech (e.g. Björkman 1996; Bradshaw and Lindbladh 2005) could be related to the targeted disposal of beech seeds at such open sites, with possible importance for maintenance of a robust metapopulation structure (e.g. Hilfiker et al. 2004) during expansion. The possibility that the beech populations in Vestfold are natural outliers is, however, questionable. Magri (2008) used the standard rule of 2% threshold of pollen for presence of beech in local vegetation. The amount in the southern part of the Norwegian–Swedish border area, however, rarely exceeds 1% (Danielsen 1970; Björkman 1996). The northern distribution limit of beech in Sweden (Fig. 1) has possibly been stable over the last 1,000 years (Björkman 1996), which complies with the view that the modern distribution of beech roughly corresponds to its maximum extension (Magri et al. 2006). The critical question is not merely whether the modest pollen finds in the Norwegian–Swedish border area could imply that migration has taken place but also whether the pollen levels may reflect a sufficiently large number of migrants to account for the large genetic variation of the beech populations in Vestfold (cf. Austerlitz et al. 2000; Table 4). Such spatially isolated and marginal tree populations are often characterised by a substantially reduced genetic variability (cf. Tamaki et al. 2008), as in common ash (Fraxinus excelsior) in southern Finland (Höltken et al. 2003). By contrast, the populations in Vestfold and Seim harbour only slightly lower amounts of diversity than the populations in England and Denmark (Table 4). Random spread by jays, the main vector for natural dispersal, would possibly lead to a profound founder effect (Nei et al. 1975; Aizawa et al. 2009). Bird-mediated dispersal is also unlikely once the long distance to the present distribution of beech in southern Sweden or Denmark is taken into account (Nilsson 1985). Thus, it can be stated as feasible, but not likely, that Vestfold represents the natural range margin of beech in northern Europe, but there is no reason to assume the remote beech at Seim was established by natural means (cf. Fægri 1954).
The monomorphism of the cpDNA markers shows that the Norwegian beech populations correspond with the previously reported cpDNA monomorphism of beech to the north of the Alps studied with PCR-RFLP markers (Demesure et al. 1996; Magri et al. 2006). This may seem obvious, but beech nuts have accompanied man as a natural foodstuff over the history (Fægri 1954), and the similar cpDNA haplotypes serve as a useful exclusion of beech from other glacial refugia in southern Europe (Magri et al. 2006). Despite the common glacial history of the studied populations, estimates of total (H T) and intrapopulation (H S) nuclear genetic diversity showed that a large proportion of the variation (about 20%) was located between populations (supplementary material S2). In F. sylvatica, population differentiation in isozyme markers rarely exceeds 7% of the total variation (Larsen 1996; Comps et al. 2001; Gömöry et al. 2003). The difference could be partly related to the inclusion of only northern populations in this study, which are usually more differentiated than southern populations (Eckert et al. 2008). Another explanation refers to the fact that differentiation generally increases with distance (Petit et al. 2005). This study covers a large geographic area compared to, e.g. a purely Danish approach (Larsen 1996), and higher differentiation can be expected. Finally, and probably most importantly, the high differentiation reported here is a reflection of the marker per se (Karhu et al. 1996), showing that direct comparisons of results across markers should be done with care (cf. Jump and Penuelas 2007). In particular, we suspect that the dominance of the ISSR markers applied to yield high differentiation, as compared to codominant isoenzyme markers (e.g. Larsen 1996; Gömöry et al. 2003). The high inbreeding estimate for all populations (f = 0.300; supplementary material S2) is not surprising, given the large differentiation, but rather, an important explanation for the differentiation itself (cf. Hamrick and Nason 1996).
Historical context
The time of initial beech establishment in Norway, 500 to 1,000 AD, took place during a period when archaeological excavations have shown that there had been regular contact between Norway and Denmark across the Skagerrak for a very long time (Ilkjær 2000), and it has been suggested that the Danes were responsible for the subsequent foundation of the town of Kaupang in Vestfold at about 800 AD (Skre 2007). Between 400 and 700 AD, the agriculture in Vestfold became more intensive, and forests were cleared and converted to arable land on a large scale (Jerpåsen 1996) which may have paved the way for the following success of beech in the area (Björkman 1999; Bradshaw and Lindbladh 2005). Wooden objects made of beech were found in the Viking ship Osebergskipet in Vestfold (c. 830 AD; Bonde and Christensen 1993), and beech nuts were known as a source of swine fodder (Fægri 1954).
Fægri's assumption (1954) linking Vestfold and Seim is not supported by recent research. The description of Harald Finehair as descending from Vestfold is dismissed as a construction made by the saga writer Snorre Sturlason in the early thirteenth century (Krag 2003). There is no genetic evidence for beech transfer from England to Seim, and any suggestions to connect persons or events in sagas to the establishment of beech, either at Seim or in Vestfold, must be ruled out as mere speculation. The oldest written source giving an account of Norway and the Norwegians is given by Ohthere, a magnate from the north of Norway who visited the court of King Alfred in Winchester in the late ninth century. He had sailed all along the coast of Norway to Schirings Heal (Kaupang in Vestfold) and then to Hedeby in the south of Denmark (Bately 2007) and, though it is not explicably stated, probably from Hedeby to Winchester. Ohthere's journey covers the whole region discussed in connection with the beech populations in Vestfold and Seim (Bately 2007) and exemplifies how long-distance trade connected different parts of Scandinavia and England. Such peaceful enterprises must be considered more relevant, as facilitating the intentional planting of beech, than the political incidents reported in the sagas or the Viking raids for booty.
Concluding remarks
Tracing ancestor populations by means of molecular markers is associated with various pitfalls. Even though the populations of this study were critically selected among ancient natural or semi-natural woods, the guarantee is only as far as written sources and memory hold. Irrespective of actual population history, gene flow over more than a thousand years from adjoining managed populations would have influenced gene frequencies and gradually increased the genetic distance between the assumed parental populations in Denmark and filial populations in Norway. It is also unlikely that we would succeed in pinpointing the actual population from where the seeds of the different populations have been collected. Although there was a clear genetic relationship between populations from Denmark and Norway, we cannot exclude the possibility that populations from southern Sweden also might have contributed in the Vestfold area, although there were no towns or ports of trade in southern Sweden in the area where beech was also present at the time (Skre 2007). Contribution from the south of Denmark or along the Baltic Sea is also conceivable since these regions also were connected to trade routes with Scandinavia. The southernmost population in Denmark (Rin; Fig. 1) is, however, located close to Hedeby (presently northern Germany), an important town for trade in Scandinavia during the Viking time. Notwithstanding these considerations, the likely introduction rather than natural spread suggests that beech should be regarded as part of the cultural heritage of Norway rather than a native species.
References
Aizawa M, Yoshimaru H, Saito H, Katsuki T, Kawahara T, Kitamura K, Shi F, Sabirov R, Kaji M (2009) Range-wide genetic structure in a north-east Asian spruce (Picea jezoensis) determined using nuclear microsatellite markers. J Biogeogr 36:996–1007
Austerlitz F, Mariette S, Machon N, Gouyon P-H, Godelle B (2000) Effects of colonisation processes on genetic diversity: differences between annual plants and tree species. Genetics 154:1309–1321
Arcade A, Anselin F, Rampant PF, Lesage MC, Paques LE, Prat D (2000) Application of AFLP, RAPD and ISSR markers to genetic mapping of European larch and Japanese larch. Theor Appl Genet 100:299–307
Bately J (2007) Text and translation: the three parts of the known world and the geography of Europe north of the Danube according to Orosius' Historiae and its Old English version. In: Bately J, Englert A (eds) Othere's voyages, a late 9th-century account of voyages along the coasts of Norway and Denmark and its cultural context. Viking ship Museum, Roskilde, pp 46–47
Benz P-G (1988) Norsk fuglehåndbok. Aschehoug, Oslo
Björkman L (1996) Long-term population dynamics of Fagus sylvatica at the northern limits of its distribution in southern Sweden: a paleoecological study. Holocene 6:225–234
Björkman L (1999) The establishment of Fagus sylvatica at the stand scale in southern Sweden. Holocene 9:237–245
Bonde N, Christensen AE (1993) Dendrochronological dating of the Viking Age ship burial at Oseberg, Gokstad and Tune, Norway. Antiquity 67:575–583
Bradshaw RHW (2004) Past anthropogenic influence on European forests and some possible genetic consequences. For Ecol Manage 197:203–212
Bradshaw RHW, Lindbladh M (2005) Regional spread and stand-scale establishment of Fagus sylvatica and Picea abies in Scandinavia. Ecology 86:1679–1686
Comps B, Gömöry D, Letouzey J, Thiebaut B, Petit RJ (2001) Diverging trends between heterozygosity and allelic richness during postglacial colonization in the European beech. Genetics 157:389–397
Danielsen A (1970) Pollen-analytical late Quaternary studies in the Ra district of Østfold, southeastern Norway. Årbok for Universitetet i Bergen (1969, no. 14), Norwegian University Press
Demesure B, Sodzi N, Petit RJ (1995) A set of universal primers for amplification of polymorphic non-coding regions of mitochondrial and chloroplast DNA in plants. Mol Ecol 1:129–131
Demesure B, Comps B, Petit RJ (1996) Chloroplast DNA phylogeography of the common beech (Fagus sylvatica L) in Europe. Evolution 50:2515–2520
Dumolin-Lapegue S, Pemonge MH, Petit RJ (1997) An enlarged set of consensus primers for the study of organelle DNA in plants. Mol Ecol 4:393–397
Eckert CG, Samis KE, Lougheed SC (2008) Genetic variation across species' geographic ranges: the central-marginal hypothesis and beyond. Mol Ecol 17:1170–1188
EUFORGEN (2009) Distribution map of beech (Fagus sylvatica). http://www.euforgen.org/distribution_maps.html, accessed 18 November 2010
Fægri K (1954) On age and origin of the beech forest (Fagus sylvatica L) at Lygrefjorden, near Bergen (Norway). Danm Geol Unders II rk 80:230–249
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure: extensions to linked loci and correlated allele frequencies. Genetics 164:1567–1587
Giesecke T, Hickler T, Kunkel T, Sykes MT, Bradshaw RHW (2007) Towards an understanding of the Holocene distribution of Fagus sylvatica L. J Biogeogr 34:118–131
Godinho R, Crespo EG, Ferrand N (2008) The limits of the mtDNA phylogeography: complex patterns of population history in highly structured Iberian lizard only revealed by the use of nuclear markers. Mol Ecol 17:4670–4683
Gömöry D, Paule L, Shvadchak IM, Popescu F, Suskowska M, Hynek V, Longauer R (2003) Spatial patterns of the genetic differentiation in European beech (Fagus sylvatica L) at allozyme loci in the Carpathians and the adjacent regions. Silvae Genet 52:78–83
Grivet D, Heinze B, Vendramin GG, Petit RJ (2001) Genome walking with consensus primers: application to the large single copy region of chloroplast DNA. Mol Ecol Notes 1:345–349
Hafsten U (1956) Pollen-analytical investigations on the late Quaternary development in the inner Oslofjord area. Årbok for Universitetet i Bergen, AS John Griegs trykkeri
Hamrick JL, Nason JD (1996) Consequences of dispersal in plants. In: Rhodes E, Chesser RK, Smith MH (eds) Population dynamics in ecological space and time. University of Chicago Press, Chicago
Henningsmoen KE (1988) The beech forest in Larvik—a pollen analytic investigation. Blyttia 46:203–207
Hewitt GM (1999) Post-glacial re-colonisation of European biota. Biol J Linn Soc 68:87–112
Hilfiker K, Gugerli F, Schütz J-P, Rotach P, Holdegger R (2004) Low RAPD variation and female-based sex ratio indicate genetic drift in small populations of the dioecious conifer Taxus baccata in Switzerland. Conservat Genet 5:357–365
Holsinger KE, Lewis PO (2003) Software for analysis of geographic structure in genetic data. Hickory v 0.8. http://darwin.eeb.uconn.edu/hickory/software.html, accessed 18 November 2010
Holsinger KE, Wallace LE (2004) Bayesian approaches for the analysis of population genetic structure: an example from Platanthera leucophaea (Orchidaceae). Mol Ecol 13:887–894
Höltken AM, Tähtinen J, Pappinen A (2003) Effects of discontinuous marginal habitats on the genetic structure of common ash (Fraxinus excelsior L.). Silvae Genet 52:206–212
Hultén E (1971) Atlas of the distribution of vascular plants in northwestern Europe. Generalstabens Litagrafiska Anstalts Förlag, Stockholm
Hultén E, Fries M (1986) Atlas of North European vascular plants north of the Tropic of Cancer. Koeltz Scientific Books, Königstein
Ilkjær J (2000) Den første Norgeshistorien. Illerupfunnet: Ny innsikt i skandinavisk romertid. Kulturhistorisk Forlag AS
Jerpåsen GB (1996) Gunnerød En arkeologisk landskapsanalyse Varia 35. Universitetets Oldsaksamling, Oslo
Johnson WC, Adkisson CS, Crow TR, Dixon MD (1997) Nut caching by blue jays (Cyanocitta cristata L.): implications for tree demography. Am Midl Nat 138:357–370
Jump AS, Penuelas J (2007) Extensive spatial genetic structure revealed by AFLP but not SSR molecular markers in the wind-pollinated tree, Fagus sylvatica. Mol Ecol 16:925–936
Kalinowski ST (2009) How well do evolutionary trees describe genetic relationships between populations? Heredity 102:506–513
Karhu A, Hurme P, Karjalainen M, Karvonen P, Kärkkäinen K, Neale D, Savolainen O (1996) Do molecular markers reflect patterns of differentiation in adaptive traits of conifers? Theor Appl Genet 93:215–221
Keene D (1976) The late Saxon ‘Burh’. In: Biddle M (ed) Winchester Studies I. Winchester in the early Middle Ages. Claredon Press, Oxford
Krag C (2003) The early unification of Norway. In: Helle K (ed) The Cambridge history of Scandinavia, volume I. Cambridge University Press, pp 186–188
Kunstler T, Curt T, Lepart J (2004) Spatial pattern of beech (Fagus sylvatica L.) and oak (Quercus pubescens) seedlings in natural pine (Pinus sylvestris L.) woodlands. Eur J For Res 123:331–337
Küster H (1997) The role of farming in the postglacial expansion of beech and hornbeam in the oak woodlands of central Europe. Holocene 7:239–242
Larsen AB (1996) Genetic structure of populations of beech (Fagus sylvatica L) in Denmark. Scand J For Res 11:220–232
Magri D (2008) Patterns of post-glacial spread and the extent of glacial refugia of European beech (Fagus sylvatica). J Biogeogr 35:450–463
Magri D, Vendramin GG, Comps B, Dupanloup I, Geburek T, Gömöry D, Latalowa M, Litt T, Paule L, Roure JM, Tantau I, van der Knaap WO, Petit RJ, de Beaulieu JL (2006) A new scenario for the quaternary history of European beech populations: palaeobotanical evidence and genetic consequences. New Phytol 171:199–221
Miller MP (1997) Tools for population genetic analysis (TFPGA) 1.3: a Windows program for the analysis of allozyme and molecular population genetic data. http://www.marksgeneticsoftware.net, accessed 18 November 2010
Mort ME, Crawford DJ, Santos-Guerra A, Francisco-Ortega J, Esselman EJ, Wolfe AD (2003) Relationships among the Macaronesian members of Tolpis (Asteraceae: Lactuceae) based upon analyses of inter simple sequence repeat (ISSR) markers. Taxon 52:511–518
Nei M (1973) Analysis of genetic diversity in subdivided populations. Proc Natl Acad Sci USA 70:3321–3323
Nei M (1978) Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89:583–590
Nei M, Maruyama T, Chakraborty R (1975) The bottleneck effect and genetic variability in populations. Evolution 29:1–10
Nilsson SG (1985) Ecological and evolutionary interactions between reproduction of beech Fagus sylvatica and seed eating animals. Oikos 44:157–164
Øyen BH (2010) Høydebonitet og produksjonsevne ved konvertering mellom gran, ask, bøk, eik, platanlønn og svartor i Sør-Norge. Forskning fra Skog og landskap (in press)
Page RDM (1996) TREEVIEW: an application to display phylogenetic trees on personal computers. Comput Appl Biosci 12:357–358
Peterken GF (1996) Natural woodland: ecology and conservation in northern temperate regions. Cambridge University Press, Cambridge
Petit RJ, Duminil J, Fineschi S, Hampe A, Salvini D, Vendramin GG (2005) Comparative organisation of chloroplast, mitochondrial and nuclear diversity in plant populations. Mol Ecol 14:689–701
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Skre D (2007) Towns and markets, kings and central places in south-western Scandinavia c. AD 800–950. In: Skre D (ed) Kaupang in Skiringssal. Kaupang excavation project publication series, volume I; Norske Oldfunn XXII. Aarhus University Press, Aarhus, pp 445–470
Sørensen R, Henningsmoen KE, Høeg HI, Stabell B, Bukholm KM (2007) Geology, soils, vegetation and sea levels in the Kaupang area. In: Skre D (ed) Kaupang in Skiringssal. Kaupang excavation project publication series, volume I; Norske Oldfunn XXII. Aarhus University Press, Aarhus, pp 251–272
Sturluson S (1941) Heimskringla (Íslenzk fornrit) XXVI bindi Heimskringla I. Aðalbjarnarson, Reykjavik
Taberlet PT, Fumagalli L, Wust-Saucy AG, Cosson JF (1998) Comparative phylogeography and postglacial colonisation routes in Europe. Mol Ecol 7:453–464
Tamaki I, Setsuko S, Tomaru N (2008) Genetic variation and differentiation in populations of a threatened tree, Magnolia stellata: factors influencing the level of within-population genetic variation. Heredity 100:415–423
Tantasawat P, Trongchuen J, Prajongjai T, Seehalak W, Jittayasothorn Y (2010) Variety identification and comparative analysis of genetic diversity in yardlong bean (Vigna unguiculata spp. sesquipedalis) using morphological characters, SSR and ISSR analysis. Sci Hortic 124:204–216
Tsumura Y, Ohba K, Strauss SH (1996) Diversity and inheritance of inter-simple sequence repeat polymorphisms in Douglas-fir (Pseudotsuga menziesii) and sugi (Cryptomeria japonica). Theor Appl Genet 92:40–45
Vander Wall SB (2001) The evolutionary ecology of nut dispersal. Bot Rev 67:74–116
Vornam B, Herzog S (2004) Spatial distribution of genetic variation in a natural beech stand (Fagus sylvatica L) based on microsatellite markers. Conservat Genet 5:561–570
Yap IV, Nelson RJ (1996) WINBOOT: a program for performing bootstrap analysis of binary data to determine the confidence limits of UPGMA-based dendrograms. IRRI discussion paper series no. 14, International Rice Research Institute, Manila. http://www.riceworld.org/science/software/winboot.asp, accessed 18 November 2010
Yeh FC, Yang RC, Boyle T (1999) POPGENE version 1.31. Microsoft Windows-based freeware for population genetic analysis. Quick user guide. University of Alberta and Centre for International Forestry Research. http://www.ualberta.ca/∼fyeh/popgene.pdf, accessed 18 November 2010
Yin T, Zhang X, Huang M, Wang M, Zhuge Q, Tu S, Zhu LH, Wu R (2002) Molecular linkage maps of the Populus genome. Genome 45:541–555
Acknowledgements
We wish to thank Hugh Milner and co-workers (UK), J.S. Jensen, B. Ditlevsen (Denmark), E. Finne, C. Kierulf, Å. Østgård and S. Øen (Norway) for organising the collection and shipment of beech buds. We also wish to thank K.E. Henningsmoen, H.I. Høeg, D. Skre and B. Myhre for valuable literature and discussions. J. Hubert, M. Sætersdal, T. Skrøppa and R. Sørensen have critically reviewed the paper, for which we are very grateful. J. Hubert also checked the English. We thank anonymous reviewers for constructive comments that substantially improved this manuscript. This work was supported by grants from the Norwegian Committee on Forest Genetic Resources and the Norwegian Forest and Landscape Institute.
Ethical standards
This work complies with the current laws in the countries in which it has been done.
Conflict of interest
The authors declare that they have no conflicts of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by G. G. Vendramin
Tor Myking and Igor Yakovlev contributed equally to this work.
Electronic supplementary materials
Below is the link to the electronic supplementary material.
Supplementary material, S1
(DOC 35 kb)
Supplementary material, S2
(DOC 84 kb)
Supplementary material, S3
(DOC 71 kb)
Supplementary material, S4
(DOC 60 kb)
Rights and permissions
About this article
Cite this article
Myking, T., Yakovlev, I. & Ersland, G.A. Nuclear genetic markers indicate Danish origin of the Norwegian beech (Fagus sylvatica L.) populations established in 500–1,000 AD. Tree Genetics & Genomes 7, 587–596 (2011). https://doi.org/10.1007/s11295-010-0358-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11295-010-0358-y