Abstract
Interspecific hybrids of Populus species are known for their superior growth. In this study, we examined the effect of the genetic background and contrasting environmental conditions on growth and searched for quantitative trait loci (QTL) affecting growth traits. To this end, two hybrid poplar families resulting from controlled crosses, Populus deltoides ‘S9-2’ × P. nigra ‘Ghoy’ (D × N, 180 F1) and P. deltoides ‘S9-2’ × P. trichocarpa ‘V24’ (D × T, 182 F1), were grown at two contrasting sites, Northern Italy and Central France. At the end of the second growing season, tree dimensions (stem height, circumference, and volume) were assessed. The performances of both families significantly differed within and between sites. Tree volume was significantly larger at the Italian site as compared to the French site. Genotype by environment interactions were significant but low for both families and for all growth traits. Tight correlations among the individual growth traits indicated that there may be a common genetic mechanism with pleiotropic effects on these growth traits. In line with previous studies, linkage groups I, VII, IX, X, XVI, XVII, and XIX appeared to have genomic regions with the largest effects on growth traits. This study revealed that (1) both families have high potential for selection of superior poplar hybrids due to the pronounced heterosis (hybrid vigor) and the large genetic variability in terms of growth and (2) the choice of site is crucial for poplar cultivation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Currently, there is a growing interest in the fast production of biomass as feedstock to displace fossil fuels and to reduce greenhouse gas emissions. Considerable amounts of energy are locked-up in lignocellulose, the main component of plant cell walls (Möller et al. 2007). Lignocellulose can be used as raw material for the manufacture of various bio-based products, for example bioethanol (Schubert 2006). Poplar is a good candidate as biorefinery feedstock (together with Salix, Miscanthus, and Triticum; Möller et al. 2007) due to its rapid juvenile growth (Bergez et al. 1989; Bradshaw et al. 2000), good coppicing ability (Ceulemans and Deraedt 1999), and completely sequenced genome (Boerjan 2005; Tuskan et al. 2006). However, the variability in quantitative and qualitative biomass production is huge in the Populus genus and among the vast number of interspecific hybrids resulting from crosses among the 30 species belonging to the genus (Ceulemans 1990; Cervera et al. 2005). Consequently, hybridization and breeding programs have been developed to screen for the most promising genotypes.
The three main goals of hybridization are: (1) to combine desirable traits from different species into the F1 progeny; (2) to obtain heterosis, or hybrid vigor and; (3) to achieve increased homeostasis, i.e. greater phenotypic stability among different environments (Stettler et al. 1996). The most common species used for hybridization in Europe and North America are Populus deltoides (Bartr. ex Marsh.), Populus trichocarpa (Torr. & Gray) and Populus nigra (L.). Natural hybrids of P. deltoides and P. nigra were the first intercontinental poplars to be used in plantation culture but also hybrids of P. deltoides and P. trichocarpa appeared to be extremely well-suited for cultivation in Europe and North America (Eckenwalder 2001). Although P. deltoides × P. trichocarpa hybrids are known to be much more productive than P. deltoides × P. nigra hybrids, high heterosis values are usually found for both hybrid types (Ceulemans 1990; Stettler and Ceulemans 1993; Marron et al. 2006).
A better understanding of the genetic basis of growth under contrasting environments is required to improve the efficiency of tree breeding. Genotype by environment interactions are generally defined as the differential performance of genotypes among environments (Falconer 1989) and may occur in two ways: (1) a range of genotypes rank differently in different environments, or (2) genotypes do not rank differently, but differences between genotypes vary in magnitude among environments. The latter type of interaction, however, causes no problem for breeding because a well-performing genotype selected in one environment will remain well-performing when grown in a different environment. Besides the desired stability of genotypes among environments, breeders are also searching for stability in time, i.e. genotypes that remain highly productive over successive years.
To gain insight into the genetic basis of growth in poplar, quantitative trait locus (QTL) detection has been accomplished for traits such as height, circumference, and branchiness (Bradshaw and Stettler 1995; Wu et al. 1998; Tsahouras et al. 2002; Rae et al. 2008). High genetic correlations have been found among various growth traits suggesting possible pleiotropic effects of common genomic regions on multiple phenotypic traits (Wu and Stettler 1997; Marron et al. 2006; Rae et al. 2008). In this study, QTL analysis was performed with MultiQTL (http://www.multiqtl.com/, MultiQTL Ltd, Institute of Evolution, Haifa University, 31905 Haifa, Israel). MultiQTL allows the simultaneous analysis of data collected from populations grown in different environments, thereby increasing the statistical power of the QTL detection and the precision of estimates of QTL positions and effects (Jansen et al. 1995; Korol et al. 1998; Rae et al. 2008).
This study presents data from two cloned full-sib families resulting from controlled crosses of the same female parent Populus deltoides ‘S9-2’ with P. nigra ‘Ghoy’ and P. trichocarpa ‘V24’, grown at two contrasting sites, i.e. Northern Italy and Central France. Genetic maps for the three parents have previously been established (Cervera et al. 2001). The objectives of this study were: (1) to examine the extent of genetic and phenotypic variation in tree dimensions for 1- and 2-year-old hybrid poplar families in two contrasting environments; (2) to examine the extent to which the growth performances of the parental species are combined in the interspecific hybrids; (3) to identify QTL for growth traits to better understand the genetic basis for growth in F1 interspecific families.
Material and methods
Plant materials and plantation layout
Two F1 interspecific Populus families resulting from controlled crosses with the same female parent were examined in this experiment. One family is composed of 180 F1 genotypes resulting from an interspecific cross between Populus deltoides (Bartr. ex Marsh.) ‘S9-2’ and P. nigra (L.) ‘Ghoy’ (D × N family) (Cervera et al. 1996, 2001). The second family included 182 F1 genotypes and was generated from an interspecific cross involving P. deltoides ‘S9-2’ and P. trichocarpa (Torr. & Gray) ‘V24’ (D × T family). Both crosses were made by the Research Institute for Nature and Forest (INBO, Geraardsbergen, Belgium) in 1987 and repeated in 1995 to enlarge the progeny.
The field trials were established in April 2003 from 25-cm uniform hardwood cuttings at a density of 6,670 trees ha−1, planted at an initial spacing of 0.75 × 2 m. The two experimental plantations were established according to a randomized block design. For each family, six complete blocks were made; each block contained one randomly planted replicate of each F1 genotype and each of the parents. To reduce border effects (Zavitkovski 1981; Van Hecke et al. 1995), a double border row was planted around the plantations (P. × euramericana ‘I-214’ at the Italian site and P. × euramericana ‘Robusta’ at the French site). Throughout each growing season, the plantation management included irrigation and the use of insecticides and fungicides as needed. However, during the establishment year, some trees did not survive, especially at the Italian site probably due to rooting difficulties (Marron et al. 2006). Irrespective of site and family, the mean number of available replicates per F1 genotype was >3.
Site description
The experimental plantations were located at two sites, i.e. in Northern Italy (Cavallermaggiore, Po valley, 44°42′ N, 7°40′ E) and in Central France (Ardon, Loire valley, 47°46′ N, 1°52′ E). During both growing seasons (April–October 2003 and 2004), the mean temperature was lower at the French site (16.3°C) as compared to the Italian site (18.4°C). Overall, annual rainfall was higher at the French site, but irrigation allowed the control of water availability at both sites. With regard to soil conditions, the Italian soil texture is pure loam whereas the French soil is composed of 75% sand. More details concerning location and climate at the two sites can be found in Dillen et al. (2007).
Traits measured
Tree growth was assessed at the two sites at the end of the second growing season (November 2004) for all replicates of each F1 and of the parental genotypes (for first-year results, see Marron et al. 2006). Total stem height (height2) was measured to the nearest cm with an extendable height pole. Stem circumference (circum2) was measured at 1 m above ground level to the nearest mm using a tape. Stem volume (vol2) was calculated for each individual stem from height2 and circum2 assuming a conical shape (Pontailler et al. 1997; Marron et al. 2006). The ratio of the stem height to the circumference (htcc2, cm mm−1)—a measure for stem taper—was calculated for data measured at the end of the second growing season. Growth increment during the second growing season was represented by deltaH and deltaC (deltaH = height2 – height1 and deltaC = circum2 – circum1, where height1 = total stem height at the end of the first growing season and circum1 = stem circumference at the end of the first growing season).
Data analyses
Statistical analyses were performed with the R software (Version 2.6.1; http://cran.r-project.org/). Genotypes for which there were less than three surviving replicate trees were removed from all further analyses (i.e. for the D × N family: 62 F1 genotypes in Italy and 13 in France and, for the D × T family: 12 F1 genotypes in Italy and 4 in France). Assumptions on residual distributions of the linear models were checked with the Shapiro–Wilk statistic. When necessary, original values were transformed according to the Box-Cox procedure (Venables and Ripley 2002). The following mixed models were used.
-
1.
For adjustment of individual data to block effect: \(Y_{{\text{ij}}} = {\text{ $ \mu $ }} + B_{\text{i}} + {\text{ $ \varepsilon $ }}_{{\text{ij}}} \) where μ is the general mean and B i is the effect of block i considered as fixed. B i was calculated, for each family and site, as the difference between the mean of block i and the general mean of the whole family.
-
2.
For comparison between families and sites: \(Y_{{\text{ijkl}}}^{\text{'}} = {\text{ $ \mu $ + Fam}}_{\text{l}} + G_{{\text{j}}\left( {\text{l}} \right)} + S_{\text{k}} + {\text{Fam}}_{\text{l}} \times S_{\text{k}} + G_{{\text{j}}\left( {\text{l}} \right)} \times S_{\text{k}} + {\text{ $ \varepsilon $ }}_{{\text{ijkl}}} \) where Y’ijkl are individual values adjusted for the within-site block effects \(\left( {Y_{\text{i}}^{\text{'}} = Y - B_{\text{i}} } \right)\), Faml is the family effect (fixed), G j(l) is the genotype effect (random), S k is the site effect (fixed), Faml × S k is the family by site interaction effect (fixed) and G j(l) × S k is the genotype by site effect (random). The mean of Y’ at the genotypic level is further referred to as the genotypic mean of a particular genotype. These values were used as input values for the QTL analysis.
-
3.
For comparison between parental performances: \(Y_{{\text{ij}}}^{\text{'}} = {\text{ $ \mu $ + }}P_{\text{j}} + \varepsilon _{{\text{ij}}} \) where P j is the effect of parent species j considered as fixed. The Scheffé method was chosen as post-hoc analysis due to different numbers of replicates for the three parental species (Maxwell and Delaney 2004).
Differences between means were considered significant when the P-value of the ANOVA F-test is <0.05. To characterize genetic variation present in each family separately, the following random models were used.
-
1.
Within a site: \(Y_{{\text{ij}}}^{\text{'}} = {\text{ $ \mu $ + }}G_{\text{j}} + \varepsilon _{{\text{ij}}} \) where G j is the effect of genotype j considered as random. Genetic and residual variance components (\(\sigma _G^2 \) and \(\sigma _\varepsilon ^2 \)) were calculated by equating observed mean squares to expected mean squares and solving the resulting equations according to the Henderson III procedure (Henderson 1953; Searle et al. 1992). The coefficient of genetic variation (CVG) was estimated as σG/MeanFamily. Broad-sense heritabilities were estimated at each site and for each family on a genotypic basis, \({\text{H}}_{{\text{Genotype}}}^{\text{2}} = {{\sigma _{\text{G}}^2 } \mathord{\left/ {\vphantom {{\sigma _{\text{G}}^2 } {\left( {\sigma _{\text{G}}^2 + \left( {{{\sigma _\varepsilon ^2 } \mathord{\left/ {\vphantom {{\sigma _\varepsilon ^2 } {n_{\text{j}} }}} \right. \kern-\nulldelimiterspace} {n_{\text{j}} }}} \right)} \right)}}} \right. \kern-\nulldelimiterspace} {\left( {\sigma _{\text{G}}^2 + \left( {{{\sigma _\varepsilon ^2 } \mathord{\left/ {\vphantom {{\sigma _\varepsilon ^2 } {n_{\text{j}} }}} \right. \kern-\nulldelimiterspace} {n_{\text{j}} }}} \right)} \right)}}\), where n j is the average number of replicates per genotype, and on an individual basis, \({\text{H}}_{{\text{Individual}}}^{\text{2}} = {{\sigma _{\text{G}}^2 } \mathord{\left/ {\vphantom {{\sigma _{\text{G}}^2 } {\left( {\sigma _{\text{G}}^2 + \sigma _\varepsilon ^2 } \right)}}} \right. \kern-\nulldelimiterspace} {\left( {\sigma _{\text{G}}^2 + \sigma _\varepsilon ^2 } \right)}}\) (Nyquist 1991). We reported \(H_{Genotype}^2 \) to estimate the efficiency of clonal selection and to give information about the precision of estimated genetic values when genotypic means were used as phenotypic predictors. \(H_{Individual}^2 \) can be considered as a reference value, calculated for one individual, and more easily comparable to literature values. The standard errors of broad-sense heritability were calculated as described by Singh et al. (1993).
-
2.
Between sites: \(Y_{{\text{ijk}}}^{\text{'}} = {\text{ $ \mu $ + }}G_{\text{j}} + S_{\text{k}} + \left( {G \times S} \right)_{{\text{jk}}} + {\text{ $ \varepsilon $ }}_{{\text{ijk}}} \), where G j is the genotype effect (random), S k is the site effect (random) and (G × S)jk is the genotype by site interaction effect (random). In order to quantify the relative importance of each effect, variance components, \(\sigma _G^2 \), \(\sigma _S^2 \), \(\sigma _{G \times S}^2 \), and \(\sigma _\varepsilon ^2 \) were calculated by equating observed mean squares to expected mean squares and solving the resulting equations according to the Henderson III procedure (Henderson 1953; Searle et al. 1992).
Mean heterosis or hybrid vigor was expressed for each family and at each site as the percentage of superiority of hybrids over the mean of the two parents \(\left( {{{\left( {{\text{Mean}}_{{\text{Family}}} - {\text{Mean}}_{{\text{Parents}}} } \right)} \mathord{\left/ {\vphantom {{\left( {{\text{Mean}}_{{\text{Family}}} - {\text{Mean}}_{{\text{Parents}}} } \right)} {{\text{Mean}}_{{\text{Parents}}} \times 100\left[ \% \right]}}} \right. \kern-\nulldelimiterspace} {{\text{Mean}}_{{\text{Parents}}} \times 100\left[ \% \right]}}} \right)\) (Li and Wu 1997). Spearman rank correlation coefficients (based on genotypic means) were calculated to assess the dependency of the genotype performance on the site. The genetic correlation of the performance of a trait between sites a and b was estimated with the following formula: \(r_{{\text{gab}}} = {{r_{{\text{ab}}} } \mathord{\left/ {\vphantom {{r_{{\text{ab}}} } {\left( {{\text{H}}_{{\text{Genotype}}} {\text{a}} \times {\text{H}}_{{\text{Genotype}}} {\text{b}}} \right)}}} \right. \kern-\nulldelimiterspace} {\left( {{\text{H}}_{{\text{Genotype}}} {\text{a}} \times {\text{H}}_{{\text{Genotype}}} {\text{b}}} \right)}}\), with r ab as the Pearson correlation coefficient between group means at sites a and b, and HGenotypea and HGenotypeb as the square root of genotypic heritabilities at sites a and b, respectively (Burdon 1977). Genetic correlations among traits were calculated from the variance–covariance matrices obtained from the MANOVA as \(r_{\text{g}} = {{{\text{Cov}}_{\text{G}} \left( {x{\text{,}}y} \right)} \mathord{\left/ {\vphantom {{{\text{Cov}}_{\text{G}} \left( {x{\text{,}}y} \right)} {\sqrt {\sigma _G^2 \left( x \right) \times \sigma _G^2 \left( y \right)} }}} \right. \kern-\nulldelimiterspace} {\sqrt {\sigma _G^2 \left( x \right) \times \sigma _G^2 \left( y \right)} }}\), where CovG(x,y) is the genetic covariance between traits x and y, estimated by equating the mean co-products with their expected values according to the Henderson III procedure (Henderson 1953; Becker 1984).
Genetic map construction
Amplified fragment length polymorphism (AFLP) analysis was performed as described by Cervera et al. (2001) on an extra 96 offspring of the D × N family leading to a total of 217 individual genotypes and on an extra 134 offspring of the D × T family leading to a total of 235 genotypes. A χ2-test (d.f. = 1, P < 0.01) was used to identify deviations from Mendelian ratios. AFLP markers deviating at the 1% significance level were excluded from the linkage analysis. Linkage analysis was performed with MAPMAKER Unix Version 3.0 (Lander et al. 1987) as described previously in Cervera et al. (2001) with the new marker data and with the marker data obtained before (Cervera et al. 2001). Markers ordered at a LOD score of 2.0 were used as framework markers. Markers that could not be ordered with equal confidence were positioned relative to a framework marker and were called accessory markers. Four genetic maps were obtained. Throughout the paper the notations D1 and D2 refer to the P. deltoides ‘S9-2’ map generated from the D × N and D × T families, respectively. Microsatellites (SSRs) have been used in the past (Cervera et al. 2001) as bridge markers to align the genetic map to other available genetic maps of Populus and to the Populus genome sequence (Morreel et al. 2006). The framework maps were further used for QTL analysis. The percentage of missing marker data in these framework maps was 22, 20, 28, and 22% for P. deltoides (D1), P. nigra (N), P. deltoides (D2), and P. trichocarpa (T), respectively.
QTL analysis
QTL analysis was based on 160 and 92 phenotypes for the D × N family and on 171 and 163 phenotypes for the D × T family, for the French and the Italian site respectively. The Anderson–Darling normality test was performed on the block-adjusted (genotypic) mean values of all growth traits at the end of the first growing season (see Marron et al. 2006 for circum1, height1, and vol1; htcc1 was added in this analysis) and at the end of the second growing season (circum2, height2, vol2, htcc2, deltaC, and deltaH) at each site to verify whether residuals were normally distributed using the proc univariate procedure (SAS Version 9.1.2, SAS Institute, Cary, NC, USA). QTL significance thresholds were determined by permutation tests and thus determined on the actual data, avoiding the necessity to transform the original data (Doerge and Rebaï 1996). QTL analyses were performed on the framework maps with MultiQTL (http://www.multiqtl.com/) for the parental maps separately using a pseudo-testcross analysis (Grattapaglia and Sederoff 1994). This program was chosen for its potential to perform QTL analysis across multiple environments, to test for two-linked QTL models, and to calculate confidence intervals. The simultaneous treatment of data from multiple environments provided a significant increase in power of QTL detection and accuracy of the estimated QTL position and effect (Jansen et al. 1995). The use of clonal replicates also increased the statistical power. The phenotypes were better estimated and microenvironmental noise could be excluded by correcting for block effects. The QTL analysis was performed as described in Rae et al. (2008). We report QTL with a chromosome-wise significance level of 0.05. Experiment-wise (genome-wide and over all traits) significances were based on the false discovery rate (Benjamini and Hochberg, 1995). Note that MultiQTL recalculates the (Kosambi) distances between the markers based on genotype data. However, these distances differ from the distances given by MAPMAKER because the latter has the ‘error detection’ option that prevents the distances to inflate whenever a possible error has been made. Most SSRs were genotyped on only part of the offspring, and were thus not put into the framework map. These SSRs were therefore placed to the MultiQTL framework maps at their approximate positions. All map drawings were done with Mapchart 2.2 (Voorrips 2002).
Results
Within site variability
Within each site, highly significant differences between both hybrid families were obtained for all traits (Table 1). The stem volumes of the D × T family were larger than those of the D × N family irrespective of site (7.2 vs. 5.4 dm3 in Italy and 4.3 vs. 1.9 dm3 in France). At the French site, the P. trichocarpa ‘V24’ parent showed overall a higher growth rate than P. nigra ‘Ghoy’ and P. deltoides ‘S9-2’ (Table 2). Heterosis values for height2, circum2, and vol2 were positive for both families (Table 1). The values ranged from 16.1 to 176.5%, for height2 of the D × N family and for vol2 of the D × T family, respectively. Htcc2 values were higher for the parents than for the F1 hybrids of both families as shown by the negative heterosis values, reflecting that the heterosis for circum2 was larger than for height2 (Table 1). Performance of P. deltoides ‘S9-2’ (Table 2) and heterosis values (Table 1) could not be defined for trees grown at the Italian site.
The coefficients of genetic variation (CVG) ranged from 6.6 to 35.3% and were higher for the D × N than for the D × T family (Table 3). Values of heritability were moderate to high: \(H_{Genotype}^2 \) ranged from 0.61 to 0.89 and \(H_{Individual}^2 \) ranged from 0.25 to 0.62 (Table 3). For heritability values the same trend was observed as for CVG, i.e. slightly higher values for the D × N family than for the D × T family.
Between site variability
All traits significantly differed between the two sites for both families (Table 1). The two families were more productive in Italy than in France and also the male parent P. nigra ‘Ghoy’ showed its highest growth under Italian conditions (Table 2). Conversely, circum2 and vol2 of P. trichocarpa ‘V24’ were significantly larger in France as compared to Italy. No data were available for P. deltoides ‘S9-2’ at the Italian site.
Genotype by site (G × S) interactions were significantly different from zero (P ≤ 0.001) (results not shown), but their relative contributions to the phenotypic variation (\(\sigma _p^2 \)) were quite low (\({{\sigma _{G \times S}^2 } \mathord{\left/ {\vphantom {{\sigma _{G \times S}^2 } {\sigma _p^2 }}} \right. \kern-\nulldelimiterspace} {\sigma _p^2 }}\) ranging from 3.8 to 11.0%, Table 4). The site, however, contributed much more to the phenotypic variation (\({{\sigma _S^2 } \mathord{\left/ {\vphantom {{\sigma _S^2 } {\sigma _p^2 }}} \right. \kern-\nulldelimiterspace} {\sigma _p^2 }}\) ranging from 16.8 to 81.3%; Table 4). Highly significant (P ≤ 0.001), but moderate values for the Spearman rank coefficients (0.50 to 0.68 for the D × N family, 0.36 to 0.52 for the D × T family) suggested some differences in rank order of genotypes between both sites (Table 5 and Fig. 1). The Spearman rank coefficients were generally lower for the D × T family than for the D × N family. The values of genetic correlation between the two sites were often more than 0.65, particularly for circum2, height2, and deltaC (Table 5).
Relationships for height2 (stem height at the end of the second growing season, cm) between the two sites Cavallermaggiore (Italy) and Ardon (France). Lines of best linear fit are shown (full line for the D × N family and dotted line for the D × T family). See Table 5 for the corresponding Spearman rank coefficients. D = Populus deltoides ‘S9-2’; N = P. nigra ‘Ghoy’; T = P. trichocarpa ‘V24’
Between year variability
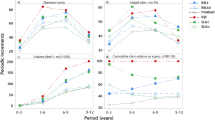
The Spearman rank coefficients between years varied from 0.41 to 0.76 in the D × T family indicating a quite stable rank order of the D × T genotypes over successive years, especially when selected on circumference (Table 6). Stability of genotypic means over the 2 years was lower in the D × N family (0.38 to 0.62) than in the D × T family, except for height increase in France. For the D × T family, there was a shift in growth performance between the two sites: at the end of the first growing season, the D × N family was more productive in Italy as compared to France, while the D × T family performed better in France (Marron et al. 2006). At the end of the second growing season, both families displayed a larger productivity at the Italian site (Table 1 and Fig. 2). Nevertheless, the D × T family was more productive than the D × N family during both growing seasons.
Relationships between circum1 (stem circumference at the end of the first growing season, mm) and deltaC (circumference increase during the second growing season, mm) at the two sites, Cavallermaggiore (Italy) and Ardon (France) for the D × N (full line) and D × T (dotted line) families. See Table 6 for corresponding Spearman rank coefficients. D = Populus deltoides ‘S9-2’; N = P. nigra ‘Ghoy’; T = P. trichocarpa ‘V24’
QTL analysis
Distribution and number of QTL
Summary statistics of the genetic maps are outlined in Table 7. The QTL results are listed in Table 8 and displayed in Fig. 3 where only framework maps are shown as calculated by MultiQTL. For the D × N family, residuals exhibited a normal distribution for most traits (data not shown). However, residual distributions of the D × T family at the French site exhibited a strong left tail, except for htcc1 and htcc2 that exhibited a pronounced right tail. This skewness was probably caused by a difficult establishment of the least performing genotypes. For the D × N family, 21 QTL were mapped on the maternal P. deltoides ‘S9-2’ map and 29 on the paternal P. nigra ‘Ghoy’ map. For the D × T family, 48 QTL were mapped on the maternal P. deltoides ‘S9-2’ map and 20 on the paternal P. trichocarpa ‘V24’ map, at the 5% linkage group level. The average number of QTL per trait was 2.1 in D1, 4.8 in D2, 2.9 in P. nigra ‘Ghoy’ and 2.0 in P. trichocarpa ‘V24’. At the 5% experiment-wise level, 3, 3, 39 and 11 QTL were detected for P. deltoides (D1), P. nigra, P. deltoides (D2), and P. trichocarpa, respectively.
QTL for stem traits related to plant growth for the first and second growing seasons. Abbreviations: D1 linkage groups of P. deltoides ‘S9-2’ (D × N family), D2 linkage groups of P. deltoides ‘S9-2’ (D × T family), N linkage groups of P. nigra ‘Ghoy’, T linkage groups of P. trichocarpa ‘V24’. AFLP markers are in black, microsatellite markers in green, and gene markers are in pink. Microsatellite markers that do not belong to the framework are in italic. Solid boxes denote QTL that showed a positive effect in Italy and France; empty boxes denote QTL that showed a negative effect at both sites; and hatched boxes denote QTL that showed an opposite effect at both sites. The middle of the box indicates the peak of the QTL, and the total length (box + lines) represents the 95% confidence interval. Only linkage groups (LG) for which QTL have been identified are presented. The traits have been explained in Table 1 and in the text
Percentage of variance explained
The percentage of variance explained (PEV) by individual QTL (single QTL model; note that for 2-linked QTL models, the individual contribution of each QTL is not known) reached a maximum of 18.1% in D1 (for circum1), 17.2% in P. nigra ‘Ghoy’ (for vol1), 16.3% in D2 (for vol2) and 23.7% in P. trichocarpa ‘V24’ (for height2; Table 8, values underlined in right-hand column). Maximum PEV for individual QTL was found mostly at the Italian site with the exception for QTL on LG T-VII. Total PEV per trait and per site was maximally 44.0% (Table 9). QTL with PEV >20% were found on LG T-VII only (Table 8).
Co-locating QTL
Assuming that co-locating QTL can be considered as a single pleiotropic QTL, the number of QTL reduced in D1 from 21 to 9, in P. nigra ‘Ghoy’ from 29 to 9, in D2 from 48 to 11 and in P. trichocarpa ‘V24’ from 20 to eight. Co-locating QTL were found on LG VII (D1, D2, and T), LG IX (N and T), LG X (D2), LG XIII (N and D2), LG XIV (N, D1 and D2), LG XVI (N), LG XVII (D2), and LG XIX (D2). Some LG were found with coinciding QTL regions for the same traits on different parental maps (LG II: vol1, circum1; LG XIII: circum2, vol2, htcc2, deltaC; XVI: deltaH, htcc2).
Directionality and site differences
At each site, a high genetic correlation was observed between the first and the second growing seasons for the traits height, circum, vol, and htcc (from 0.74 to 0.92; Table 10) with the only exception of htcc in France (0.49). The traits height, circum, and vol were highly genetically (and positively) correlated with each other (correlation coefficients from 0.68 to 0.99; Table 10). There was also a genetic correlation, albeit to a lower extent and negative, between htcc on the one hand, and height, circum, and vol on the other hand (Table 10). This negative correlation is reflected in the opposite direction of the QTL effect (e.g. LG D1-VII, T-VII, T-IX, D2-X, N-XIII, D2-XIII, D1-XIV, N-XVI, D1-XVII, D2-XVII, and D2-XIX). The QTL on LG II, IV, and XIII had a different directionality and the QTL on LG XVI resulted in a lower performance (Fig. 3).
In 16 cases, there was a significantly different effect of the QTL between the two sites (Table 8), supporting the significant G × S interactions. The P. trichocarpa ‘V24’ map showed most QTL with significant differences in effect between both sites. Five were located on LG VII, and three on LG III. P. deltoides ‘S9-2’ (D1) showed three QTL with significant differences in effect between both sites, P. deltoides ‘S9-2’ (D2) four and P. nigra ‘Ghoy’ two. Most QTL showed a negative effect on the trait (Table 8). From the 21 QTL found on D1, only four were found as well on D2. They were situated on LG XIV and XVII (Fig. 3). However, the effect on LG XVII was opposite, leaving only one QTL found in common between D1 and D2.
Discussion
Genotypic variability
In line with previous studies, average values of stem height were around 6 m for the single-stem F1 poplar hybrids at the end of the second growing season (Heilman and Stettler 1985; Ceulemans et al. 1992, 1996; Wu et al. 1992). Irrespective of site, the D × T family showed a significantly higher growth performance than the D × N family in terms of stem height, circumference, and volume. Marron et al. (2007) indicated that the differences in productivity between these families could be related to a large extent to differences in the length of the growing season than to inherent differences in growth rates. The high productivity of the D × T hybrids has promoted the inter-American hybrids to a superior position in poplar plantation culture in Pacific Northwest America as well as in Central and Western Europe (Eckenwalder 2001). However, D × T hybrids are generally highly susceptible to leaf rust (Melampsora larici-populina) and, therefore, proper management including treatment with fungicides is needed to prevent severe reductions in growth (Pinon 1992; Laureysens et al. 2005).
During the second growing season, both families displayed pronounced heterosis or hybrid vigor. Possibly the heterosis values were slightly overestimated because the parents growing in the same plantation as the F1 hybrids might have experienced effects of competition. Heterosis for Populus hybrids is a well-known phenomenon (Muhle Larsen 1970; Stettler et al. 1996; Li and Wu 1997; Marron et al. 2006). Heterosis is determined by non-mutually exclusive mechanisms, including genome-wide dominance complementation, locus-specific overdominance effects and epistasis, although the relative contribution of each of these mechanisms is still unclear (Lippman and Zamir 2007). Higher values of heterosis were observed for the D × T family as compared to the D × N family. The superiority of the D × T hybrids could be partly explained by the combination of the rapid height growth of the P. trichocarpa parent with the high diameter growth of P. deltoides in the F1 hybrids (Eckenwalder 2001). As expected, heterosis values were highest for stem volume because volume is a multiplicative function of stem height and circumference (Li and Wu 1997).
Both families showed moderate to high values of broad-sense heritability (\(H_{Individual}^2 \) ranging from 0.25 to 0.62) at the two sites indicating that the amount of phenotypic variation attributable to genetic variation is quite high for these growth characteristics. Comparable values were reported for F1 hybrids of P. deltoides and P. trichocarpa (Marron et al. 2006, 2007). Overall, F2 hybrids of the same species displayed higher values of heritability (Wu and Stettler 1997; Rae et al. 2004, 2008). In contrast to F1 breeding where species-specific linkages are left intact, F2 breeding highlights recombination and maximizes genetic variance (Stettler et al. 1996).
Higher values for \(H_{Individual}^2 \) and CVG were observed for the D × N family than for the D × T family. Differences in length of the growing season appeared to be related to differences in productivity between both families (Marron et al. 2007). More variability in cessation of growth was found for the D × N family than for the D × T family (data not shown), which could induce more genetic variation in growth traits. However, both families showed high potential for breeding, the D × N family based on its relatively wide genetic variation, and the D × T family because of its high heterosis and superior productivity.
Genotype by site interactions
Genotype by site interactions (G × S) were highly significant but low for all growth traits. Both families were more productive in Italy as compared to France. This could be partly explained by the soil as loamy soils (as in the Italian site) generally present a better nutrient- and water-holding capacity in comparison with more sandy soil types (Yu and Pulkkinen 2003). Due to its southern location, the Italian site was furthermore characterized by high-radiation conditions while sufficient water supply was guaranteed by irrigation. Therefore, the Italian conditions could be considered as more favorable than the French ones. Moreover, higher stem growth for D × T hybrids has been shown in warm, high-radiation and well-watered conditions as compared to cooler coastal conditions (Wu and Stettler 1997). These authors compared growth of poplar hybrids in Boardman (OR, USA), which is known for its warm continental climate, with hybrids grown in Clatskanie (OR, USA), which has a wetter maritime climate.
The French site was characterized by higher values of heritability in comparison with the Italian site for all studied traits. As many contradictory results have been reported, no generalizations have emerged from previous studies on how heritability estimates change under different environmental conditions (Hoffmann and Parsons 1991). Thus, care should be taken in extrapolating results beyond the environment in which they were obtained (Lynch and Walsh 1998).
In line with previous studies, changes in ranking of genotypes between the two distinct sites were suggested by the moderate values of Spearman rank coefficients between sites (Namkoong et al. 1992; Marron et al. 2006; Rae et al. 2008). However, no complete trade-off in performance was observed because G × S interactions were low; their relative contribution to the phenotypic variation was often less than 10%. This did not imply that ranking for stem growth was consistent between the Italian and French sites. In fact, superior genotypes did not automatically perform well in both environments, although poor genotypes were generally poor in both environments, in line with observations on F2 P. trichocarpa × P. deltoides hybrids by Wu and Stettler (1997), who hypothesized that the F2 genotypes performing poorly in two different environments represented unfavorable recombinants expressing developmental disharmony or were affected by deleterious recessive alleles due to inbreeding. Interestingly, we made similar observations in our two outbred families (F1), indicating that this phenomenon is not typical to inbred populations (F2).
Stability over time
The D × T family was more productive than the D × N family during both growing seasons (Marron et al. 2006, 2007; Dillen et al. 2007). During the second growing season, both families at the Italian site experienced a spectacular boost in growth resulting in an outperformance compared to the production at the French site. Consequently, the site ranking of the D × T family changed for growth traits (height, circum, and vol) between the first and the second growing seasons: France > Italy, at the end of the first growing season versus Italy > France at the end of the second growing season (Marron et al. 2006). The relative performances of both families at the two sites at the end of the first growing season appeared to correlate with the different preferences of the male parent species for the climate (Marron et al. 2006). P. nigra often performs better in warmer and drier conditions (Wu and Stettler 1997; Rae et al. 2004, 2008; Marron et al. 2006, 2007). In contrast, P. trichocarpa has a preference for cooler and wetter conditions (Farmer 1996; Wu and Stettler 1997; Marron et al. 2006). During the first growing season the D × N family was indeed more productive under the warmer and drier Italian conditions, while the D × T family grew better at the wetter site in France, in line with the respective preferences of their male parents. For the second growing season, however, this hypothesis did no longer explain the observations.
Four possible explanations for the outperformance of the two hybrid families at the Italian site during the second growing season can be formulated as follows:
-
1.
The Italian site was characterized by more favorable growth conditions, namely soil characteristics and irradiance.
-
2.
A C-effect, i.e. a physiological preconditioning of the woody cuttings used to establish the field trial due to differences in quality among cuttings of the same genotype, might have affected growth during the first growing season (Lerner 1958). The C-effect is generally expressed during the first growing season when variance in shoot growth attributable to primary ramet effects is often as large as the variance due to differences among genotypes (Wilcox and Farmer 1968; Farmer et al. 1989; Dunlap et al. 1992).
-
3.
In Italy, a higher mortality, possibly due to rooting difficulties, was recorded as compared to the French site. Given the high bulk density of the soil, the poor rooting capacity of the female parent P. deltoides (Dickmann and Stuart 1983; Laureysens et al. 2003) could have contributed to the high mortality and lower growth performance during the establishment year. Likely, the cuttings of 25 cm were too small for a good establishment in the heavier Italian soil (loam) as compared to the French sandy soil.
-
4.
Due to the higher mortality at the Italian site, planting density decreased, resulting in a lower competition and better light conditions. These improved light conditions could be another reason for the superior growth during the second year of both families in Italy as compared to France.
In conclusion, the preferences of the parents in terms of climate and soil, the C-effect and the rooting difficulties could have influenced the response of the hybrids to the different sites during the year of planting only. Afterwards, when the plants were established and acclimated to the different environments, their growth was predominantly affected by the limiting growth conditions. This was reflected by the changes in relative contributions of the different components of phenotypic variation, i.e. genetic (\(\sigma _G^2 \)), site (\(\sigma _S^2 \)) and G × S (\(\sigma _{G \times S}^2 \)) components, for both families. During the first growing season, the genetic component contributed mainly to the phenotypic variation, while at the end of the second growing season the relative contribution of the site component gained in importance (ranging from 20 to 80%). Hence, the choice of the site appears to be crucial for poplar cultivation (Marron et al. 2006).
QTL
As already observed, the tree dimension traits (height, circum, and vol) highly correlated with each other (Brown et al. 1997; Ketterings et al. 2001; Dillen et al. 2007), indicating that there may be a common genetic mechanism that has a pleiotropic effect on multiple growth traits. The correlations between the values measured at the end of the first and at the end of the second growing season were also high. In P. nigra ‘Ghoy’, two QTL for traits measured at the end of the first growing season, co-located with QTL for traits measured at the end of the second growing season. In D2, 13 QTL for traits measured at the end of the first growing season, co-located with QTL for traits measured at the end of the second growing season and in P. trichocarpa ‘V24’ eight. Co-location of QTL over successive years was also observed by Rae et al. (2008) suggesting that the same genomic regions influence the traits during this first 2 years. Remarkably, no QTL collocated over the 2 years in D1.
Comparing the QTL data presented by Bradshaw and Stettler (1995), Wu et al. (1998), Rae et al. (2008) and the data presented in this study, LG I, VII, IX, X, XVI, XVII, and XIX appeared to contain genomic regions with the largest effects on growth traits. The precision, however, was limited to the linkage group level as a result of a lack of homologous markers between the maps. The number of QTL found per trait was in the range of what is normally expected in a F1 (pseudo-test)cross or backcross with 100–200 progeny (Grattapaglia et al. 1996; Beavis 1998; Shepherd et al. 2002; Wullschleger et al. 2005; Zhang et al. 2006). Surprisingly, many more QTL were detected in P. deltoides (D2) than in P. deltoides (D1). A possible explanation is that P. deltoides and P. trichocarpa belong to two different sections. This could lead to hemizygous loci in the hybrid offspring, potentially revealing more QTL. On the other hand, the percentage of missing data was quite large in P. deltoides (D2). To reduce the effect of missing information, MultiQTL calculates probabilities of the missing marker status based on scores of the neighbor markers. Hence, QTL estimates might be less reliable. This study did also reveal only few loci with a relatively large effect (> 20%) as was the case in Bradshaw and Stettler (1995).
Wu et al. (1998) identified more QTL involved in basal area than in stem height, which they explained by the fact that secondary growth is a more complex trait than height growth. Our study also identified slightly more QTL for circumference than for height, as expected since circumference had a higher CVG and H 2. However, on average the PEV per locus was the same for both traits. QTL associated with growth were also investigated by Li et al. (1999) on a F2 population of P. deltoides × P. cathayana and by Zhang et al. (2006) on an interspecific backcross of P. tomentosa × P. bolleana, but unfortunately, the maps were based on RAPD (Li et al. 1999) or AFLP (Zhang et al. 2006), making it impossible to compare the QTL regions with our data.
In conclusion, we showed that there were significant but low G × S interactions in terms of the growth performance of both related families. Close correlations between growth traits suggested that common genetic mechanisms are at the basis of these growth traits. Since the maps are still poorly anchored, only very general comparisons on QTL positions between the maps were possible. Future efforts need to focus on integrating the various maps. Given the large size of the confidence intervals, candidate gene selection based on map position is impossible. Yet, combining results from other approaches, such as genetical genomics (Street et al. 2006; Morreel et al. 2006) may help narrowing down the list of candidate genes, which can then be further refined by association genetics (Neale and Savolainen 2004; Ingvarsson et al. 2008).
References
Beavis WD (1998) QTL analyses: power, precision and accuracy. In: Paterson AH (ed) Molecular dissection of complex traits. CRC, New York, pp 145–162
Becker WA (1984) Manual of quantitative genetics. Pullman Academic Enterprises, Washington, pp 191
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B-Stat Methodol 57:289–300
Bergez JE, Auclair D, Bouvarel L (1989) First-year growth of hybrid poplar shoots from cutting or coppice origin. Forest Sci 35:1105–1113
Boerjan W (2005) Biotechnology and the domestication of forest trees. Curr Opin Biotechnol 16:159–166
Bradshaw HD Jr, Stettler RF (1995) Molecular genetics of growth and development in Populus. IV. Mapping QTLs with large effects on growth, form and phenology traits in a forest tree. Genetics 139:963–973
Bradshaw HD Jr, Ceulemans R, Davis J, Stettler R (2000) Emerging model systems in plant biology: poplar (Populus) as a model forest tree. J Plant Growth Regul 19:306–313
Brown S, Schroeder P, Birdsey R (1997) Aboveground biomass distribution of US eastern hardwood forests and the use of large trees as an indicator of forest development. Forest Ecol Manage 96:37–47
Burdon RD (1977) Genetic correlation as a concept for studying genotype-environment interaction in forest tree breeding. Agronomie 17:219–246
Cervera MT, Gusmão J, Steenackers M, Peleman J, Storme V, Vanden Broeck A, Van Montagu M, Boerjan W (1996) Identification of AFLP molecular markers for resistance against Melampsora larici-populina in Populus. Theor Appl Genet 93:733–737
Cervera MT, Storme V, Ivens B, Gusmão J, Liu BH, Hostyn V, Van Slycken J, Van Montagu M, Boerjan W (2001) Dense genetic linkage maps of three Populus species (Populus deltoides, P. nigra and P. trichocarpa) based on AFLP and microsatellite markers. Genetics 158:787–809
Cervera MT, Storme V, Soto A, Ivens B, Van Montagu M, Rajora OP, Boerjan W (2005) Intraspecific and interspecific genetic and phylogenetic relationships in the genus Populus based on AFLP markers. Theor Appl Genet 111:1440–1456
Ceulemans R (1990) Genetic variation in functional and structural productivity determinants in poplar. Thesis Publishers, Amsterdam, The Netherlands, pp 100
Ceulemans R, Deraedt W (1999) Production physiology and growth potential of poplars under short-rotation forestry culture. Forest Ecol Manage 121:9–23
Ceulemans R, Scarascia-Mugnozza G, Wiard BM, Braatne JH, Hinckley TM, Stettler RF, Isebrands JG, Heilman PE (1992) Production physiology and morphology of Populus species and their hybrids grown under short rotation. I. Clonal comparisons of 4-year growth and phenology. Can J Forest Res 22:1937–1948
Ceulemans C, Shao BY, Jiang XN, Kalina J (1996) First- and second-year aboveground growth and productivity of two Populus hybrids grown at ambient and elevated CO2. Tree Physiol 16:61–68
Dickmann DI, Stuart KW (1983) The culture of poplars in eastern North America. Michigan State University Publications, East Lansing, Michigan, USA, pp 168
Dillen SY, Marron N, Bastien C, Ricciotti L, Salani F, Sabatti M, Pinel MPC, Rae AM, Taylor G, Ceulemans R (2007) Effects of environment and progeny on biomass estimations of five hybrid poplar families grown at three contrasting sites across Europe. Forest Ecol Manage 252:12–23
Doerge RW, Rebaï A (1996) Significance thresholds for QTL interval mapping tests. Heredity 76:459–464
Dunlap JM, Heilman PE, Stettler RF (1992) Genetic variation and productivity of Populus trichocarpa and its hybrids. V. The influence of ramet position on 3-year growth variables. Can J Forest Res 22:849–857
Eckenwalder JE (2001) Descriptions of clonal characteristics. In: Dickmann DI, Isebrands JG, Eckenwalder JE, Richardson J (eds) Poplar culture in North America. Part B, Chapter 13. NRC Research, National Research Council of Canada, Ottawa, Canada, pp 331–382
Falconer DS (1989) Introduction to quantitative genetics, 3rd edn. Longman Scientific and Technical, London, UK, pp 438
Farmer RE Jr (1996) The genecology of Populus. In: Stettler RF, Bradshaw HD Jr, Heilman PE, Hinckley TM (eds) Biology of Populus and its implications for management and conservation. Part I, Chapter 2. NRC Research, National Research Council of Canada, Ottawa, Canada, pp 33–55
Farmer RE Jr, Freitag M, Garlick K (1989) Genetic variance and “C” effects in balsam poplar rooting. Silvae Genet 38:62–65
Grattapaglia D, Sederoff R (1994) Genetic linkage maps of Eucalyptus grandis and Eucalyptus urophylla using a pseudo-test-cross: mapping strategy and RAPD markers. Genetics 137:1121–1137
Grattapaglia D, Bertolucci FLG, Penchel R, Sederoff RR (1996) Genetic mapping of quantitative trait loci controlling growth and wood quality traits in Eucalyptus grandis using a maternal half-sib family and RAPD markers. Genetics 144:1205–1214
Heilman PE, Stettler RF (1985) Genetic variation and productivity of Populus trichocarpa and its hybrids. II. Biomass production in a 4-year plantation. Can J Forest Res 15:384–388
Henderson CR (1953) Estimation of variance and co-variance components. Biometrics 9:226–252
Hoffmann AA, Parsons PA (1991) Evolutionary genetics and environmental stress. Oxford University Press, Oxford, UK, pp 284
Ingvarsson PK, Garcia MV, Luquez V, Hall D, Jansson S (2008) Nucleotide polymorphism and phenotypic associations within and around the phytochrome B2 locus in European aspen (Populus tremula, Salicaceae). Genetics 178:2217–2226
Jansen RC, Van Ooijen JW, Stam P, Lister C, Dean C (1995) Genotype-by-environment interaction in genetic mapping of multiple quantitative trait loci. Theor Appl Genet 91:33–37
Ketterings QM, Coe R, van Noordwijk M, Ambagau Y, Palm CA (2001) Reducing uncertainty in the use of allometric biomass equations for predicting above-ground tree biomass in mixed secondary forests. Forest Ecol Manage 146:199–209
Korol AB, Ronin YI, Nevo E (1998) Approximate analysis of QTL-environment interaction with no limits on the number of environments. Genetics 148:2015–2028
Lander ES, Green P, Abrahamson J, Barlow A, Daly MJ, Lincoln SE, Newburg L (1987) MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1:174–181
Laureysens I, Deraedt W, Indeherberge T, Ceulemans R (2003) Population dynamics in a six-year old coppice culture of poplar. I. Clonal differences in stool mortality, shoot dynamics and shoot diameter distribution in relation to biomass production. Biomass Bioenerg 24:81–95
Laureysens I, Pellis A, Willems J, Ceulemans R (2005) Growth and production of a short rotation coppice culture of poplar. III. Second rotation results. Biomass Bioenerg 29:10–21
Lerner IM (1958) The genetic basis of selection. Wiley, New York, pp 298
Li B, Wu R (1997) Heterosis and genotype × environment interactions of juvenile aspens in two contrasting sites. Can J Forest Res 27:1525–1537
Li JH, Su XH, Zhang QW, Zsuffa L (1999) Detection of QTLs for growth and phenology traits of poplar using RAPD markers. Forest Res 12:111–117 (in Chinese)
Lippman ZB, Zamir D (2007) Heterosis: revisiting the magic. Trends Genet 23:60–66
Lynch M, Walsh B (1998) Genetics and analysis of quantitative traits. Sinauer, Sunderland, Massachusetts, pp 980
Marron N, Bastien C, Sabatti M, Taylor G, Ceulemans R (2006) Plasticity of growth and sylleptic branchiness in two poplar families grown at three contrasting sites across Europe. Tree Physiol 26:935–946
Marron N, Dillen SY, Ceulemans R (2007) Evaluation of leaf traits for indirect selection of high yielding poplar hybrids. Env Exp Bot 61:103–116
Maxwell SE, Delaney HD (2004) Designing experiments and analyzing data: a model comparison perspective, 2nd edn. Lawrence Erlbaum Associates, Mahwah, New Jersey, pp 217–218
Möller R, Toonen M, van Beilen J, Salentijn E, Clayton D (2007) Crop platforms for cell wall biorefining: lignocellulose feedstocks. EPOBIO project. CNAP, University of York, UK, pp 162
Morreel K, Goeminne G, Storme V, Sterck L, Ralph J, Coppieters W, Breyne P, Steenackers M, Georges M, Messens E, Boerjan W (2006) Genetical metabolomics of flavonoid biosynthesis in Populus: a case study. Plant J 47:224–237
Muhle Larsen C (1970) Recent advances in poplar breeding. Int Rev Forest Res 3:1–67
Namkoong G, Jonsson A, Eriksson G (1992) Genetic variation in nutrient response functions. Theor Appl Genet 85:165–172
Neale DB, Savolainen O (2004) Association genetics of complex traits in conifers. Trends Plant Sci 9:325–330
Nyquist WE (1991) Estimation of heritability and prediction of selection response in plant populations. Crit Rev Plant Sci 10:235–322
Pinon J (1992) Variability in the genus Populus in sensitivity to Melampsora rusts. Silvae Genet 41:25–34
Pontailler JY, Ceulemans R, Guittet J, Mau F (1997) Linear and non-linear functions of volume index to estimate woody biomass in high density young poplar stands. Ann Forest Sci 54:335–345
Rae AM, Robinson KM, Street NR, Taylor G (2004) Morphological and physiological traits influencing biomass productivity in short-rotation coppice poplar. Can J Forest Res 34:1488–1498
Rae AM, Pinel MPC, Bastien C, Sabatti M, Street NR, Tucker J, Dixon C, Marron N, Dillen SY, Taylor G (2008) QTL for yield in bioenergy Populus: identifying G×E interactions from growth at three contrasting sites. Tree Genet Genom 4:97–112
Schubert C (2006) Can biofuels finally take center stage? Nature Biotechnol 24:777–784
Searle SR, Casella G, McCulloch CE (1992) Variance components. Wiley, New York, pp 501
Shepherd M, Cross M, Dieters MJ, Henry R (2002) Branch architecture QTL for Pinus elliottii var. elliottii × Pinus caribaea var. hondurensis hybrids. Ann Forest Sci 59:617–625
Singh M, Ceccarelli S, Hamblin J (1993) Estimation of heritability from varietal trials data. Theor Appl Genet 86:437–441
Stettler RF, Ceulemans R (1993) Clonal material as a focus for genetic and physiological research in forest trees. In: Ahuja MR, Libby WJ (eds) Clonal forestry. 1: Genetics and biotechnology. Springer, Berlin, Germany, pp 68–86
Stettler RF, Zsuffa L, Wu R (1996) The role of hybridisation in the genetic manipulation of Populus. In: Stettler RF, Bradshaw HD Jr, Heilman PE, Hinckley TM (eds) Biology of Populus and its implications for management and conservation. Part I, Chapter 4. NRC Research, National Research Council of Canada, Ottawa, Canada, pp 87–112
Street NR, Skogström O, Sjödin A, Tucker J, Rodriguez-Acosta M, Nilsson P, Jansson S, Taylor G (2006) The genetics and genomics of the drought response in Populus. Plant J 48:321–341
Tsahouras V, Gullberg U, Lagercrantz U (2002) An AFLP and RFLP linkage map and quantitative trait locus (QTL) analysis of growth traits in Salix. Theor Appl Genet 105:277–288
Tuskan GA, DiFazio S, Jansson S, Bohlmann J, Grigoriev I, Hellsten U, Putnam N, Ralph S, Rombauts S, Salamov A, Schein J, Sterck L, Aerts A, Bhalerao RR, Bhalerao RP, Blaudez D, Boerjan W, Brun A, Brunner A, Busov V, Campbell M, Carlson J, Chalot M, Chapman J, Chen G-L, Cooper D, Coutinho PM, Couturier J, Covert S, Cronk Q, Cunningham R, Davis J, Degroeve S, Déjardin A, dePamphilis C, Detter J, Dirks B, Dubchak I, Duplessis S, Ehlting J, Ellis B, Gendler K, Goodstein D, Gribskov M, Grimwood J, Groover A, Gunter L, Hamberger B, Heinze B, Helariutta Y, Henrissat B, Holligan D, Holt R, Huang W, Islam-Faridi N, Jones S, Jones-Rhoades M, Jorgensen R, Joshi C, Kangasjärvi J, Karlsson J, Kelleher C, Kirkpatrick R, Kirst M, Kohler A, Kalluri U, Larimer F, Leebens-Mack J, Leplé J-C, Locascio P, Lou Y, Lucas S, Martin F, Montanini B, Napoli C, Nelson DR, Nelson C, Nieminen K, Nilsson O, Pereda V, Peter G, Philippe R, Pilate G, Poliakov A, Razumovskaya J, Richardson P, Rinaldi C, Ritland K, Rouzé P, Ryaboy D, Schmutz J, Schrader J, Segerman B, Shin H, Siddiqui A, Sterky F, Terry A, Tsai C-J, Uberbacher E, Unneberg P, Vahala J, Wall K, Wessler S, Yang G, Yin T, Douglas C, Marra M, Sandberg G, Van de Peer Y, Rokhsar D (2006) The genome of black cottonwood, Populus trichocarpa (Torr. & Gray). Science 313:1596–1604
Van Hecke P, Moermans R, Mau F, Guittet J (1995) Border effects and size inequality in experimental even-aged stands of poplar clones (Populus). Ann Forest Sci 52:193–200
Venables WN, Ripley BD (2002) Modern applied statistics with S. 4th edn. Springer, New York, USA, pp 495
Voorrips RE (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered 93:77–78
Wilcox JR, Farmer RE Jr (1968) Heritability and C effects in early root growth of eastern cottonwood cuttings. Heredity 23:239–245
Wu R, Stettler RF (1997) Quantitative genetics of growth and development in Populus. II. The partitioning in genotype × environment interaction in stem growth. Heredity 78:124–134
Wu R, Wang M-X, Huang M-R (1992) Quantitative genetics of yield breeding for Populus short rotation culture. I. Dynamics of genetic control and selection model of yield traits. Can J Forest Res 22:175–182
Wu R, Bradshaw HD Jr, Stettler RF (1998) Developmental quantitative genetics of growth in Populus. Theor Appl Genet 97:1110–1119
Wullschleger SD, Yin TM, DiFazio SP, Tschaplinski TJ, Gunter LE, Davis MF, Tuskan GA (2005) Phenotypic variation in growth and biomass distribution for two advanced-generation pedigrees of hybrid poplars. Can J Forest Res 35:1779–1789
Yu Q, Pulkkinen P (2003) Genotype-environment interaction and stability in growth of aspen hybrid clones. Forest Ecol Manage 173:25–35
Zavitkovski J (1981) Small plots with unplanted plot border can distort data in biomass production studies. Can J Forest Res 11:9–12
Zhang D, Zhang Z, Yang K (2006) QTL analysis of growth and wood chemical content traits in an interspecific backcross family of white poplar (Populus tomentosa × P. bolleana) × P. tomentosa. Can J Forest Res 36:2015–2023
Acknowledgements
This research was supported by the European Commission through the Directorate General Research within the Fifth Framework Programme for Research—Quality of Life and Management of the Living Resources Programme, contract N° QLK5-CT-2002-00953 (POPYOMICS; http://www.soton.ac.uk/~popyomic). The authors thank Jean Gauvin and Patrick Poursat (INRA Orléans, France), Dr. Fabrizio Nardin (Alasia Vivai company, Cavallermaggiore, Italy), Dr. Maurizio Sabatti, Luca Ricciotti, and Francesco Salani (University of Viterbo, Italy) for their help with the management of field experiments and with the data collection, and Abraham Korol for useful comments on the MultiQTL software. Sophie Y. Dillen is a Research Fellow of the Research Foundation-Flanders (F.W.O.–Vlaanderen).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by: O. Savolainen
Dillen and Storme contributed equally to the work.
An erratum to this article can be found at http://dx.doi.org/10.1007/s11295-009-0199-8
Rights and permissions
About this article
Cite this article
Dillen, S.Y., Storme, V., Marron, N. et al. Genomic regions involved in productivity of two interspecific poplar families in Europe. 1. Stem height, circumference and volume. Tree Genetics & Genomes 5, 147–164 (2009). https://doi.org/10.1007/s11295-008-0175-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11295-008-0175-8