Abstract
In this study, different culture-dependent methods were used to examine the cultivable heterotrophic bacteria in the sediments associated with two deep-sea hydrothermal vents (named HV1 and HV2) located at Iheya Ridge and Iheya North in Okinawa Trough. The two vents differed in morphology, with HV1 exhibiting diffuse flows while HV2 being a black smoker with a chimney-like structure. A total of 213 isolates were identified by near full-length 16S rRNA gene sequence analysis. Of these isolates, 128 were from HV1 and 85 were from HV2. The bacterial community structures were, in large parts, similar between HV1 and HV2. Nevertheless, differences between HV1 and HV2 were observed in one phylum, one class, 4 orders, 10 families, and 20 genera. Bioactivity analysis revealed that 25 isolates belonging to 9 different genera exhibited extracellular protease activities, 21 isolates from 11 genera exhibited extracellular lipase activities, and 13 isolates of 8 genera displayed antimicrobial activities. This is the first observation of a large population of bacteria with extracellular bioactivities existing in deep-sea hydrothermal vents. Taken together, the results of this study provide new insights into the characteristics of the cultivable heterotrophic bacteria in deep-sea hydrothermal ecosystems.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Since their first discovery in 1977, hydrothermal vents have drawn a great deal of attention. Nowadays it is known that hydrothermal vents possess unique geological structures, complex physicochemical characteristics, and diverse biotic communities (Anderson et al. 2014; Kilias et al. 2013; Nakamura et al. 2013). The vent biome contains macrofauna and microorganisms, the former including tube worms, mussels, shrimps, crabs, barnacle, and sponges (Buckeridge et al. 2013; Edwards and Nelson 1991; Kim et al. 2013; Nishijima et al. 2010; Plouviez et al. 2013; Watsuji et al. 2010), and the latter including various bacteria and archaea (Polz and Cavanaugh 1995; Takai et al. 2001, 2004; Zhao et al. 2014). Microorganisms play important roles in deep-sea hydrothermal ecosystems. In the absence of photosynthesis, the food chain in the vent system relies on chemoautotrophic prokaryotes, which produce complex organic molecules with energy obtained by oxidizing the inorganic compounds available in the environments. Studies have shown that some chemoautotrophic microorganisms, such as Thioreductor, Sulfurovum, and Sulfurimonas, can use reduced sulfur compounds, Fe(II), Mn(II), H2, and CH4 as electron donors to obtain energy and fix carbon, thus providing a basis for hydrothermal vent ecosystems (Biddle et al. 2012; Inagaki et al. 2004; Kanaparthi et al. 2013; Mino et al. 2014; Nakagawa et al. 2005a). In addition, heterotrophic bacteria were also rich in hydrothermal systems (Rajasabapathy et al. 2014), and they play important roles in carbon and nitrogen cycling by decomposing particulate organic materials (Zhou et al. 2009a, b).
To date, a number of studies on the microbial communities in deep-sea hydrothermal vents have been reported, mainly focusing on habitats inside or outside of the chimney structure (Takai et al. 2001), on the outer surface of deposits around the vents, typically as mats (Moyer et al. 1995), on and within macrofauna as symbionts (Watsuji et al. 2010), and within the plume of hydrothermal fluid in the overlying seawater (Takai et al. 2004). In most documented reports, the microbial diversity is investigated by culture-independent methods, because the majority of microbes in the vent systems are difficult to cultivate (Amann et al. 1995; Inagaki et al. 2006; Zhou et al. 2009a, b). Nevertheless, culture-dependent method is important, as it is essential for the discovery of new species, novel drug, and natural products with industrial application potential (Guan et al. 2011; Thornburg et al. 2010). Moreover, culture-dependent methods help us understand the complex process that occurs at hydrothermal vents by providing us with model organisms to test the hypothesis, often raised by culture-independent techniques and that thus both methods need to be interrelated for a more comprehensive understanding of the system. Recent studies indicate that cultivable bacteria in shallow water hydrothermal vents are abundant and diverse, and that some of these bacteria possess strong enzyme-producing capacities (Mohandass et al. 2012; Raghukumar et al. 2008; Rajasabapathy et al. 2014). However, similar bioactivity studies of microbes from deep-sea hydrothermal vents are scarce.
Investigations on the microbial community in the hydrothermal vents in the vicinity of Okinawa Trough have been reported (Nakagawa et al. 2005b, c; Takai et al. 2003, 2004). In these reports, systematic survey with culture-dependent method was used only for microbes associated with hydrothermal plume, chimney structures, and vent fluids (Nakagawa et al. 2005b, c). In addition, most of these studies target on autotrophic bacteria, while the heterotrophic bacteria in these vents are largely overlooked. Therefore, the focus of our present study is heterotrophic bacteria. We aimed to investigate the characteristics of the cultivable heterotrophic bacteria in the sediments associated with two deep-sea hydrothermal vents in Iheya North and Iheya Ridge of Okinawa Trough, which exhibited different geochemical features. Moreover, since the current knowledge on the bioactive compounds secreted by deep-sea hydrothermal microorganisms is very limited, we also aimed to investigate the potential of the identified bacteria to produce extracellular protease, lipase, and antimicrobial molecules.
Materials and methods
Sample collection and processing
The samples used in this study were obtained by means of the scientific research vessel KEXUE during the cruise of April 2014. The cruise was part of the research project “Western Pacific Ocean System: Structure, Dynamics, and Consequences—Deep Ocean Environments and Ecosystems” supported by Chinese Academy of Sciences (Tollefson 2014). The sediments approximately 40 meters from Hydrothermal vent 1 (HV1) and Hydrothermal vent 2 (HV2) located in Iheya Ridge and Iheya North respectively, of Okinawa Trough were collected by push-cores on the Remotely Operated Vehicle (ROV) equipped on the KEXUE vessel. The temperature and dissolved oxygen concentration of the sample sites were detected by a conductivity-temperature-depth sampler and a dissolved oxygen sensor, respectively. Undisturbed portions of the samples were immediately processed on board by serial dilutions in sterilized seawater and plating on different solid mediums (see below) in an aseptic environment. The remaining samples were stored at 4 and −80 °C.
Bacterial isolation
Three different media were used for bacterial isolation, namely Marine Agar 2216 (Kobayashi et al. 2008), modified R2A (Yin et al. 2013), and Leibovitz’s L-15 Medium (Thermo Scientific HyClone, Beijing, China) (Futai et al. 2006). Marine 2216 is a common medium for isolation of marine heterotrophic bacteria. However, compared to marine environments, the nutrients of 2216E medium are rich. To obtain more diverse heterotrophic bacteria, modified R2A and Leibovitz’s L-15 were also used. Modified R2A medium prepared with seawater is less nutritious than marine 2216 and widely used for isolation of marine bacteria. Leibovitz’s L-15 Medium, purchased from Thermo Scientific HyClone, Beijing, China, is a kind of synthetic medium. It is the least nutritious medium and contains inorganic salts, a small amount of amino acids, and vitamins. It is used to isolate bacteria that demand little nutrient. The plates were incubated at the temperatures of 4, 16, 28, and 40 °C under aerobic condition for 3 days to 1 month. The bacterial isolates emerged on the plates were first screened on the basis of colony characteristics including size, shape, margin, color, and opacity (Valiente Moro et al. 2013). One colony per type was selected per plate and re-cultured for further purification. The purified isolates were resuspended in medium containing 15 % (v/v) glycerol and stored at −80 °C.
Bacterial identification and phylogenetic analysis
Genomic DNA was prepared from the bacteria with TIANamp bacteria DNA kit (TIANgen, Beijing, China). The nearly complete 16S rRNA genes (~1400 bp) of the bacteria were amplified using primers 27F and 1492R (Kobayashi et al. 2008) with Ex Taq DNA polymerase (Takara, Dalian). The PCR products were purified and sequenced. The 16S rRNA gene sequence similarities were searched using the BLASTn program in the National Center for Biotechnology Information database, the RDP Classifier (Wang et al. 2007), and the EzTaxon server 2.1 (Kim et al. 2012). Evolutionary distances were calculated according to the two-parameter model of Kimura (Kimura 1980). Phylogenetic dendrograms were constructed with maximum likelihood method (Rogers 1997), and tree topologies were evaluated by performing bootstrap analysis of 1000 data sets using MEGA 4.0 software (Tamura et al. 2007).
Geochemical analysis
Chemical composition of the samples was analyzed by the Research Center of Analysis and Measurement (RCAM), Institute of Oceanology, Chinese Academy of Sciences. Major and trace elements were analyzed by an inductively coupled plasma atomic emission spectroscopy (ICP-AES) (Perkin Elmer, USA) using a strong acid digestion method (Lee et al. 2006). The concentrations of total organic carbon (TOC) and total nitrogen (TN) were determined using a PE 2400 Series II CHNS/O analyzer (Perkin Elmer, USA).
Screening for extracellular protease, lipase, and antibacterial activity
For protease activity assay, the bacteria were plated on Marine 2216 medium supplemented with 1 % (w/v) skim milk. The plates were incubated as described above (section “Bacterial isolation”). The diameter of the colony (C) and the diameter of the hydrolytic zone (Z) around the colony were recorded. The ratio of Z/C was calculated and used as an indicator of enzymatic activity. For lipase activity assay, the bacteria were plated on Marine 2216 medium supplemented with 1 % olive oil, 1 % Tween-80, and 1.6 ‰ Bromocresol purple. Lipid was decomposed to fatty acid, which makes Bromocresol purple changing from purple to yellow. For each isolate, the Z/C ratio was determined by the method above. For antimicrobial activity assay, the bacterial cultures were spotted on a sterile filter-paper disk placed on a plate spread with the Gram-positive bacterium Micrococcus luteus (purchased from CGMCC, Beijing, China) or the Gram-negative bacterium Escherichia coli DH5α (purchased from Tiangen, Beijing, China). The Z/C value was determined as above and used as an indicator of inhibition.
Nucleotide sequence
The 16S rRNA gene sequences of the isolates examined in this study have been deposited in GenBank databases under the accession numbers KM978994-KM979121 (for HV1 isolates) and KM979122-KM979206 (for HV2 isolates).
Results
Sampling sites
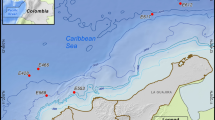
Sediment samples were collected from two deep-sea hydrothermal vent fields in Iheya Ridge and Iheya North in Okinawa Trough (Fig. 1). The two vents exhibited different morphologies. Hydrothermal vent 1 (HV1) had no obvious chimney structure but had diffuse flows, while Hydrothermal vent 2 (HV2) was a typical black smoker with a chimney-like structure by the ROV observation at the time of sampling. Samples of HV1 were brown polymetallic ooze, while samples of HV2 were black sulfides. The characteristics of the samples are indicated in Table 1, which shows that there existed apparent differences in physicochemical features between the fields of HV1 and HV2 in terms of the contents of total organic carbon (TOC), total nitrogen (TN), and metal irons. The TOC and TN in HV2 were 5.0 and 10.0 times respectively, more than those in HV1. Of the metal irons that had been identified, Cu, Pb, and Zn were 725.7, 345.0, and 1058.1 times more abundant in HV2 than in HV1, while Fe and Ba were 2–14 times higher in HV2 than in HV1. In contrast, the contents of Ca, Mg, and Mn in HV1 were 10.6, 2.4, and 3.0 times, respectively, more than those in HV2 (Table 1). The temperature and dissolved oxygen concentrations of the two sites were similar (Table 1).
Structure of the cultivable bacterial community in HV1
Of the isolates obtained from HV1 (Fig. 2), 128 were identified by near full-length 16S rRNA gene sequence analysis. The results showed that the 128 isolates were classified into 4 phyla, 5 classes, 11 orders, 19 families, 26 genera, and 68 species (Fig. 3). The species classification was according to a 97 % identity threshold (Guan et al. 2011). At the phylum level, Proteobacteria (87 isolates) was the dominant group, which was followed in abundance by Firmicutes (37 isolates), Bacteroidetes (2 isolates), and Actinobacteria (2 isolates) (Fig. 4a). Except for the Proteobacteria phylum, which contains two classes, i.e. Gammaproteobacteria (80 isolates) and Alphaproteobacteria (7 isolates), the other three phyla each contains only one class, i.e. Bacilli (37 isolates), Actinobacteria (2 isolates), and Flavobacteria (2 isolates) (Fig. 4b). At the order level, Alteromonadales (46 isolates) was the largest group, which was followed in size by Bacillales (37 isolates), Pseudomonadales (15 isolates), and Oceanospirillales (14 isolates) (Fig. 4c). The other 7 orders each contains less than four isolates. The largest family was Bacillaceae 1 (35 isolates), which was followed by Pseudoalteromonadaceae (17 isolates), Alteromonadaceae (16 isolates), Halomonadaceae (12 isolates), Moraxellaceae (12 isolates), and Alteromonadaceae (6 isolates) (Fig. 4d). For the remaining 13 families, each contains less than five isolates. Bacillus (29 isolates) was the dominant genus and comprised of the largest amount of species (15 in number). Next to Bacillus in species diversity were Pseudoalteromonas, Halomonas, Alteromonas, and Idiomarina, which contain 9, 5, 4, and 4 species respectively.
Phylogenetic tree of the isolates from HV1 based on 16S rRNA gene sequences (1323 bp). The phylogenetic tree was constructed using maximum likelihood method. The numbers on the branches indicate bootstrap values based on 1000 replications. The numbers in parentheses indicate strain numbers. Scale bar represents number of changes per nucleotide position
Structure of the cultivable bacterial community in HV2
A total of 85 isolates from HV2 were identified by near full-length 16S rRNA gene sequence analysis. The 85 isolates were classified into 3 phyla, 4 classes, 9 orders, 15 families, 20 genera, and 50 species (Fig. 5). The three phyla were Proteobacteria (68 isolates), Firmicutes (16 isolates), and Bacteroidetes (1 isolates) (Fig. 4a). The Proteobacteria phylum contains two classes, i.e. Gammaproteobacteria (66 isolates) and Alphaproteobacteria (2 isolates), while the other two phyla each contains only one class, i.e. Bacilli (16 isolates) and Flavobacteria (1 isolates) (Fig. 4b). At the order level, Alteromonadales (53 isolates) was the largest group, which was followed in size by Bacillales (16 isolates) and Pseudomonadales (7 isolates) (Fig. 4c). For the other 6 orders, each has less than four isolates. The dominant families were Pseudoalteromonadaceae (33 isolates), Alteromonadaceae (15 isolates), and Bacillaceae 1 (14 isolates) (Fig. 4d). The other 12 families each contains less than five isolates. The most populous genus was Pseudoalteromonas (33 isolates), which contains 11 species. Next to Pseudoalteromonas in species diversity were Bacillus, Pseudomonas, and Alteromonas, which contain 10, 4, and 4 species respectively.
Phylogenetic tree of the isolates from HV2 based on 16S rRNA gene sequences (1267 bp). The phylogenetic tree was constructed using maximum likelihood method. The numbers on the branches indicate bootstrap values based on 1000 replications. The numbers in parentheses indicate strain numbers. Scale bar represents number of changes per nucleotide position
Comparison of the bacterial communities in HV1 and HV2
Of the four phyla identified in this study, Actinobacteria was found only in HV1, while the other three phyla (Proteobacteria, Firmicutes, and Bacteroidetes) occurred in both HV1 and HV2 (Fig. 6a). At the class level, Gammaproteobacteria, Alphaproteobacteria, Bacilli, and Flavobacteria were shared by HV1 and HV2, whereas Actinobacteria occurred only in HV1 (Fig. 6b). Eight orders, i.e. Flavobacteriales, Bacillales, Rhizobiales, Chromatiales, Vibrionales, Oceanospirillales, Pseudomonadales, and Alteromonadales, occurred in both HV1 and HV2, while Rhodospirillales was isolated only from HV2; the other three orders, i.e. Rhodobacterales, Caulobacterales, and Actinomycetales were isolated only from HV1 (Fig. 6c). At the family level, Rhodobacteraceae, Rhizobiaceae, Caulobacteraceae, Microbacteriaceae, Intrasporangiaceae, Oceanospirillaceae, and Idiomarinaceae occurred only in HV1, and Rhodospirillaceae, Aurantimonadaceae, and Colwelliaceae occurred only in HV2. The other 12 families were common to HV1 and HV2 (Fig. 6d). Of the 26 genera from HV1 and the 20 genera from HV2, 13 were shared by both sites (Fig. 6e). These 13 genera were Pseudoalteromonas, Marinobacter, Alteromonas, Halomona, Bacillus, Olleya, Exiguobacterium, Rheinheimera, Vibrio, Acinetobacter, Pseudomonas, Shewanella, and Fictibacillus. For the other genera, Zunongwangia, Halobacillus, Leucobacter, Domibacillus, Sulfitobacter, Rhizobium, Brevundimonas, Photobacterium, Marinomonas, Cobetia, Paraglaciecola, Idiomarina, and Phycicoccus were only found in HV1, while Virgibacillus, Brevibacterium, Thalassospira, Aureimonas, Psychrobacter, Colwellia, and Marinobacterium were found only in HV2. In summary, the 213 isolates obtained from HV1 and HV2 belonged to 4 phyla, 5 classes, 12 orders, 22 families, 33 genera, and 96 species.
Screening for isolates with extracellular activities
Isolates that exhibited extracellular protease activity
Ninety-six isolates representing the 96 different species of HV1 and HV2 were examined for the potential to produce extracellular protease/lipase or antimicrobial activity. Forty-four isolates (45.8 %) were found to exhibit at least one type of activity, but no isolates exhibited all three types of activities (Table 2). Representative isolates with protease, lipase, and antimicrobial activities are shown in Fig. 7. Extracellular protease activity was identified in 25 isolates of 9 different genera (Fig. 8). Of these isolates, 13 are of the Bacillus genus, two, three, and two belong to the genera of Exiguobacterium, Fictibacillus, and Alteromonas respectively, while the remaining five isolates belong to the genera of Brevibacterium, Rheinheimera, Marinomonas, Halomonas, and Pseudoalteromonas respectively (Fig. 8). Relatively high levels of activity (Z/C ≥ 3) were displayed by 10 isolates, i.e. Fictibacillus barbaricus, Fictibacillus solisalsi, Bacillus tequilensis, Bacillus safensis, Bacillus megaterium, Bacillus aerophilus, Bacillus firmus, Bacillus altitudinis, Bacillus subtilis subsp. Inaquosorum, and Rheinheimera aquimaris (Table 2).
Representative isolates with extracellular protease activity (a), lipase activity (b), and antibacterial activity (c). a Cultures of three bacteria were spotted on a plate containing skim milk. b Cultures of three bacteria were spotted on a plate containing olive oil, Tween-80, and Bromocresol purple. c Cultures of three bacteria were spotted onto filter discs placed on a plate spread with Micrococcus luteus. For all panels, arrows indicate strains with extracellular bioactivity. (Color figure online)
Isolates that exhibited extracellular lipase activity
In the screening of extracellular lipase activity, 21 isolates were found to be positive. Seven of the isolates belong to the genus of Pseudoalteromonas, and the other 14 isolates belong to the genera of Exiguobacterium, Domibacillus, Brevibacterium, Brevundimonas, Halomonas, Acinetobacter, Shewanella, Colwellia, Paraglaciecola, and Fictibacillus (Fig. 8). Nine isolates of the genera Exiguobacterium, Domibacillus, Halomonas, Acinetobacter, Pseudoalteromonas, and Paraglaciecola showed high levels of activity (Z/C ≥ 3) (Table 2).
Isolates that exhibited antimicrobial activity
When the Gram-positive bacterium M. luteus was used as a target strain, 12 isolates were found to be able to inhibit M. luteus growth to different extents. Six of the 12 isolates belong to the genus Bacillus, and the other six isolates belong to the genera of Virgibacillus, Rheinheimera, Marinomonas, Pseudoalteromonas, Alteromonas, and Halomonas (Fig. 8). Four of these isolates, i.e. Bacillus tequilensis, Bacillus safensis, Bacillus aerophilus, and Bacillus altitudinis, exhibited relatively strong inhibitory effects (Z/C ≥ 2) (Table 2). When the Gram-negative bacterium E. coli was used as a target strain, only one isolate, Vibrio neocaledonicus, showed apparent inhibitory activity (Table 2).
Isolates that exhibited two types of extracellular activity
There were 15 isolates showing two types of extracellular activity. Specifically, four isolates from the genera of Exiguobacterium, Fictibacillus, and Brevibacterium exhibited both protease and lipase activities; 11 isolates from the genera of Bacillus, Halomonas, Pseudoalteromonas, Alteromonas, Marinomonas, and Rheinheimera exhibited protease and antibacterial activities (Table 2). No isolates showing both lipase and antibacterial activities were found.
Discussion
In this study, we examined the cultivable heterotrophic bacterial communities in sediments associated with two deep-sea hydrothermal vents, HV1 and HV2, in Iheya Ridge and Iheya North respectively, via culture-dependent methods. A total of 213 isolates were obtained from the two vent fields, which were classified into 96 species. This species ratio (96/213) or operational taxonomic units (OTU) in all isolates was similar to that (95/221) observed in a shallow water hydrothermal vent in Espalamaca (Rajasabapathy et al. 2014). For both HV1 and HV2, Proteobacteria and Gammaproteobacteria were most abundant at the phylum and class level respectively. Gammaproteobacteria appears to be a common inhabitant of hydrothermal vents and has been reported to exist in different vent systems. For example, Perez-Rodriguez et al. (2013) found a large amount of Gammaproteobacteria in cooling and less reducing vent fields, and Rajasabapathy et al. identified abundant members of Gammaproteobacteria in a shallow hydrothermal field (Rajasabapathy et al. 2014). Previous reports showed that some Pseudoalteromonas and Marinobacter, both belonging to Gammaproteobacteria, may take part in the Mn-oxidizing process in hydrothermal vents (Mason et al. 2009; Orcutt et al. 2011), and that Marinobacter is thought to participate in Fe oxidation (Edwards et al. 2003; Kaye et al. 2011). In our study, Pseudoalteromonas was found to be the dominant genus in both HV1 and HV2, and Marinobacter also occurred in HV1 and HV2. Considering the possible role of Pseudoalteromonas and Marinobacter in metal processing, these observations are consistent with the fact that the contents of Mn and Fe were relatively high in the sampling sites of HV1 and HV2. In addition to Pseudoalteromonas and Marinobacter, seven other genera, i.e. Alteromonas, Halomona, Vibrio, Acinetobacter, Shewanella, Pseudomonas, and Rheinheimera, of the Gammaproteobacteria phylum were also isolated from both HV1 and HV2. Of these 7 genera, the first 6 have been detected in hydrothermal fields around the world (Kaye et al. 2011; Martins et al. 2013; Pena et al. 2012; Raguenes et al. 2003; Rajasabapathy et al. 2014), while the last one was identified in vent environments for the first time. The roles of these bacteria in hydrothermal vent ecosystems remain to be elucidated. It should be noted that V. alginolyticus, which is an aquaculture pathogen to fish and invertebrate marine animals, was isolated from HV1. It will be interesting for future studies to investigate whether it is pathogenic to vent fauna.
Firmicutes and Bacilli were the second largest phylum and class respectively, in both HV1 and HV2. Within the Firmicutes phylum, three genera, i.e. Bacillus, Exiguobacterium, and Fictibacillus, were shared by HV1 and HV2. In the past, reports on Firmicutes found in hydrothermal vents were rare. However, recent studies indicate that this bacterial group, which can tolerate a wide range of temperatures, exists in varying phenotypes in different hydrothermal environments (Cao et al. 2014; Price et al. 2013). In a shallow water hydrothermal vent at D. João de Castro Seamount (DJCS), researchers isolated 22 strains, 15 of which were within the genus Bacillus, and the growth of these bacteria was promoted by Fe and Mn (Mohandass et al. 2012). Another report showed that diverse spore-forming Bacillus spp. from Guaymas Basin hydrothermal sediments are involved in copper-mediated Mn(II) oxidation and precipitation (Dick et al. 2006). Given these findings and the observation that the sediments of HV1 and HV2 were rich in metal ions, it is possible that the abundant Bacillus found in our study may play a role in metal oxidation.
Of the phyla shared by HV1 and HV2, Bacteroidetes was the smallest one. The percentages of Bacteroidetes isolates in the two fields were much less than that reported for a shallow hydrothermal vent field in Espalamaca (Rajasabapathy et al. 2014). This is probably due to the difference in the geochemical features of these vent systems. The shallow water hydrothermal vent in Espalamaca is exposed to the sunlight, and consequently both photosynthesis and chemosynthesis exist, which leads to production of more abundant organic matters relative to deep-sea hydrothermal vents. Since members of Bacteroidetes are known to play an important role in degradation of organic compounds (Gomez-Pereira et al. 2012), its scarcity in HV1 and HV2 may reflect a harshness of the environments in these two vent niches.
In addition to similarities, we also observed dissimilarities between HV1 and HV2 in terms of bacterial diversity. HV1 and HV2 differed in one phyla, one class, four orders, ten families, and 20 genera. It is likely that the differences in the geological, physical, and chemical characteristics of the vent systems, which are known to select and shape the population structures of the ecosystem (Klochko et al. 2012; Lai et al. 2014), may account for the difference in the bacterial communities observed in HV1 and HV2. Huston and Deming (2002) reported that the amounts of organic carbon and organic nitrogen in water would affect heterotrophic bacterial communities. In our study, we found that the contents of TOC and TN were significantly different between HV1 and HV2, which may contribute at least in part to the difference in the heterotrophic bacterial communities between the two sites.
To our knowledge, no systematic studies on the production of extracellular bioactive compounds by deep-sea hydrothermal microorganisms have been documented. One report by Mohandass et al. showed that bacteria from a shallow water hydrothermal vent in D. João de Castro Seamount (DJCS), Azores, Portugal were able to produce degradative enzymes like amylase (Mohandass et al. 2012). In two other reports, Pseudoalteromonas from the sediments of South China Sea and Bacillus spp. from the coastal sediments of King George Island produced extracellular proteases (Zhou et al. 2009a, b; Zhou et al. 2013). In our study, we identified diverse protease-producing isolates from HV1 and HV2, most of which belonging to the Bacillus genus. We also detected a large number of lipase-producing bacteria, two of which, i.e. Domibacillus and Paraglaciecola, were found for the first time to secret lipase. Hydrothermal vents are considered the next hot spots for novel drug discovery, especially antibacterial drugs (Andrianasolo et al. 2009, 2012; Tasiemski et al. 2014; Taylor 2013; Thornburg et al. 2010). In our study, antibacterial properties were detected in 13 isolates, and that relatively strong inhibitory effects on Gram-positive bacteria were observed with isolates of the Bacillus genus. These results are comparable to the previous reports that antimicrobial activities were observed with Bacillus from widely different sources such as rhizosphere plants and gastrointestinal tract (Mouloud et al. 2013; Thankappan et al. 2014).
In conclusion, the results of this study demonstrate that diverse cultivable bacteria exist in sediments associated with two deep-sea hydrothermal vents, HV1 and HV2, in Okinawa Trough, and that although the bacterial community structures in HV1 and HV2 are largely similar, differences between these two sites do exist, possibly as a result of specific adaptation to the local environmental selection. In addition, we also provide the first evidence that a large population of bacteria with extracellular bioactivities flourishing in deep-sea hydrothermal vents, which suggests a potential for industrial/medical application and provides a basis for future studies of the biological roles of microorganisms in hydrothermal ecosystems.
References
Amann RI, Ludwig W, Schleifer KH (1995) Phylogenetic identification and in situ detection of individual microbial-cells without cultivation. Microbiol Rev 59:143–169
Anderson RE, Sogin ML, Baross JA (2014) Evolutionary strategies of viruses, bacteria and archaea in hydrothermal vent ecosystems revealed through metagenomics. PLoS One 9:e109696
Andrianasolo EH, Haramaty L, Rosario-Passapera R, Bidle K, White E, Vetriani C, Falkowski P, Lutz R (2009) Ammonificins A and B, hydroxyethylamine chroman derivatives from a cultured marine hydrothermal vent bacterium, Thermovibrio ammonificans. J Nat Prod 72:1216–1219
Andrianasolo EH, Haramaty L, Rosario-Passapera R, Vetriani C, Falkowski P, White E, Lutz R (2012) Ammonificins C and D, hydroxyethylamine chromene derivatives from a cultured marine hydrothermal vent bacterium, Thermovibrio ammonificans. Mar Drugs 10:2300–2311
Biddle JF, Cardman Z, Mendlovitz H, Albert DB, Lloyd KG, Boetius A, Teske A (2012) Anaerobic oxidation of methane at different temperature regimes in Guaymas Basin hydrothermal sediments. ISME J 6:1018–1031
Buckeridge JS, Linse K, Jackson JA (2013) Vulcanolepas scotiaensis sp nov., a new deep-sea scalpelliform barnacle (Eolepadidae: Neolepadinae) from hydrothermal vents in the Scotia Sea, Antarctica. Zootaxa 3745:551–568
Cao H, Wang Y, Lee OO, Zeng X, Shao Z, Qian PY (2014) Microbial sulfur cycle in two hydrothermal chimneys on the southwest Indian Ridge. MBIO 5:e00980-13
Dick GJ, Lee YE, Tebo BM (2006) Manganese(II)-oxidizing Bacillus spores in Guaymas Basin hydrothermal sediments and plumes. Appl Environ Microbiol 72:3184–3190
Edwards DB, Nelson DC (1991) DNA-DNA solution hybridization studies of the bacterial symbionts of hydrothermal vent tube worms (Riftia pachyptila and Tevnia jerichonana). Appl Environ Microbiol 57:1082–1088
Edwards KJ, Rogers DR, Wirsen CO, McCollom TM (2003) Isolation and characterization of novel psychrophilic, neutrophilic, Fe-oxidizing, chemolithoautotrophic alpha- and gamma-Proteobacteria from the deep sea. Appl Environ Microbiol 69:2906–2913
Futai N, Gu W, Song JW, Takayama S (2006) Handheld recirculation system and customized media for microfluidic cell culture. Lab Chip 6:149–154
Gomez-Pereira PR, Schuler M, Fuchs BM, Bennke C, Teeling H, Waldmann J, Richter M, Barbe V, Bataille E, Glöckner FO, Amann R (2012) Genomic content of uncultured Bacteroidetes from contrasting oceanic provinces in the North Atlantic Ocean. Environ Microbiol 14:52–66
Guan L, Cho KH, Lee JH (2011) Analysis of the cultivable bacterial community in jeotgal, a Korean salted and fermented seafood, and identification of its dominant bacteria. Food Microbiol 28:101–113
Huston AL, Deming JW (2002) Relationships between microbial extracellular enzymatic activity and suspended and sinking particulate organic matter: seasonal transformations in the North Water. Deep Sea Res Part II Top Stud Oceanogr 49:5211–5225
Inagaki F, Takai K, Nealson KH, Horikoshi K (2004) Sulfurovum lithotrophicum gen. nov., sp nov., a novel sulfur-oxidizing chemolithoautotroph within the epsilon-Proteobacteria isolated from Okinawa Trough hydrothermal sediments. Int J Syst Evol Microbiol 54:1477–1482
Inagaki F, Kuypers MMM, Tsunogai U, Ishibashi J, Nakamura K, Treude T, Ohkubo S, Nakaseama M, Gena K, Chiba H, Hirayama H, Nunoura T, Takai K, Jørgensen BB, Horikoshi K, Boetius A (2006) Microbial community in a sediment-hosted CO2 lake of the southern Okinawa Trough hydrothermal system. Proc Natl Acad Sci USA 103:14164–14169
Kanaparthi D, Pommerenke B, Casper P, Dumont MG (2013) Chemolithotrophic nitrate-dependent Fe(II)-oxidizing nature of actinobacterial subdivision lineage TM3. ISME J 7:1582–1594
Kaye JZ, Baross JA (2000) High incidence of halotolerant bacteria in Pacific hydrothermal-vent and pelagic environments. FEMS Microbiol Ecol 32:249–260
Kaye JZ, Sylvan JB, Edwards KJ, Baross JA (2011) Halomonas and Marinobacter ecotypes from hydrothermal vent, subseafloor and deep-sea environments. FEMS Microbiol Ecol 75:123–133
Kilias SP, Nomikou P, Papanikolaou D, Polymenakou PN, Godelitsas A, Argyraki A, Carey S, Gamaletsos P, Mertzimekis TJ, Stathopoulou E, Goettlicher J, Steininger R, Betzelou K, Livanos I, Christakis C, Bell KC, Scoullos M (2013) New insights into hydrothermal vent processes in the unique shallow-submarine arc-volcano, Kolumbo (Santorini), Greece. Sci Rep 3:2421–2434
Kim OS, Cho YJ, Lee K, Yoon SH, Kim M, Na H, Park SC, Jeon YS, Lee JH, Yi H, Won S, Chun J (2012) Introducing EzTaxon-e: a prokaryotic 16S rRNA gene sequence database with phylotypes that represent uncultured species. Int J Syst Evol Microbiol 62:716–721
Kim SJ, Pak SJ, Ju SJ (2013) Mitochondrial genome of the hydrothermal vent shrimp Nautilocaris saintlaurentae (Crustacea: Caridea: Alvinocarididae). Mitochondrial DNA. doi:10.3109/19401736.2013.815169
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide-sequences. J Mol Evol 16:111–120
Klochko VV, Zelena LB, Voychuk SI, Ostapchuk AN (2012) Peculiarities of Alteromonas macleodii strains reflects their deep/surface habitation rather than geographical distribution. J Gen Appl Microbiol 58:129–135
Kobayashi T, Koide O, Mori K, Shimamura S, Matsuura T, Miura T, Takaki Y, Morono Y, Nunoura T, Imachi H, Inagaki F, Takai K, Horikoshi K (2008) Phylogenetic and enzymatic diversity of deep subseafloor aerobic microorganisms in organics- and methane-rich sediments off Shimokita Peninsula. Extremophiles 12:519–527
Lai QL, Liu Y, Yuan J, Du J, Wang LP, Sun F, Shao Z (2014) Multilocus Sequence analysis for assessment of phylogenetic diversity and biogeography in Thalassospira bacteria from diverse marine environments. PLoS One 9:e106353
Lee CS, Li X, Shi W, Cheung SC, Thornton I (2006) Metal contamination in urban, suburban, and country park soils of Hong Kong: a study based on GIS and multivariate statistics. Sci Total Environ 356:45–61
Martins A, Tenreiro T, Andrade G, Gadanho M, Chaves S, Abrantes M, Calado P, Tenreiro R, Vieira H (2013) Photoprotective bioactivity present in a unique marine bacteria collection from Portuguese deep sea hydrothermal vents. Mar Drugs 11:1506–1523
Mason OU, Di Meo-Savoie CA, Van Nostrand JD, Zhou JZ, Fisk MR, Giovannoni SJ (2009) Prokaryotic diversity, distribution, and insights into their role in biogeochemical cycling in marine basalts. ISME J 3:231–242
Mino S, Kudo H, Arai T, Sawabe T, Takai K, Nakagawa S (2014) Sulfurovum aggregans sp. nov., a hydrogen-oxidizing, thiosulfate-reducing chemolithoautotroph within the Epsilonproteobacteria isolated from a deep-sea hydrothermal vent chimney, and an emended description of the genus Sulfurovum. Int J Syst Evol Microbiol 64:3195–3201
Mohandass C, Rajasabapathy R, Ravindran C, Colaco A, Santos RS, Meena RM (2012) Bacterial diversity and their adaptations in the shallow water hydrothermal vent at D. Joao de Castro Seamount (DJCS), Azores, Portugal. Cah Biol Mar 53:65–76
Mouloud G, Daoud H, Bassem J, Atef I, Hani B (2013) New bacteriocin from Bacillus clausii StrainGM17: purification, characterization, and biological activity. Appl Biochem Biotechnol 171:2186–2200
Moyer CL, Dobbs FC, Karl DM (1995) Phylogenetic diversity of the bacterial community from a microbial mat at an active, hydrothermal vent system, Loihi Seamount, Hawaii. Appl Environ Microbiol 61:1555–1562
Nakagawa S, Inagaki F, Takai K, Horikoshi K, Sako Y (2005a) Thioreductor micantisoli gen. nov., sp nov., a novel mesophilic, sulfur-reducing chemolithoautotroph within the epsilon-Proteobacteria isolated from hydrothermal sediments in the mid-Okinawa Trough. Int J Syst Evol Microbiol 55:599–605
Nakagawa S, Takai K, Inagaki F, Chiba H, Ishibashi J, Kataoka S, Hirayama H, Nunoura T, Horikoshi K, Sako Y (2005b) Variability in microbial community and venting chemistry in a sediment-hosted backarc hydrothermal system: impacts of subseafloor phase-separation. FEMS Microbiol Ecol 54:141–155
Nakagawa S, Takai K, Inagaki F, Hirayama H, Nunoura T, Horikoshi K, Sako Y (2005c) Distribution, phylogenetic diversity and physiological characteristics of epsilon-Proteobacteria in a deep-sea hydrothermal field. Environ Microbiol 7:1619–1632
Nakamura K, Toki T, Mochizuki N, Asada M, Ishibashi J, Nogi Y, Yoshikawa S, Miyazaki J, Okino K (2013) Discovery of a new hydrothermal vent based on an underwater, high-resolution geophysical survey. Deep Sea Res Part I Oceanogr Res Pap 74:1–10
Nishijima M, Lindsay DJ, Hata J, Nakamura A, Kasai H, Ise Y, Fisher CR, Fujiwara Y, Kawato M, Maruyama T (2010) Association of thioautotrophic bacteria with deep-sea sponges. Mar Biotechnol 12:253–260
Orcutt BN, Sylvan JB, Knab NJ, Edwards KJ (2011) Microbial ecology of the dark ocean above, at, and below the Seafloor. Microbiol Mol Biol Rev 75:361–422
Pena A, Busquets A, Gomila M, Bosch R, Nogales B, García-Valdés E, Lalucat J, Bennasar A (2012) Draft genome of Pseudomonas stutzeri Strain ZoBell (CCUG 16156), a marine isolate and model organism for denitrification studies. J Bacteriol 194:1277–1278
Perez-Rodriguez I, Bohnert KA, Cuebas M, Keddis R, Vetriani C (2013) Detection and phylogenetic analysis of the membrane-bound nitrate reductase (Nar) in pure cultures and microbial communities from deep-sea hydrothermal vents. FEMS Microbiol Ecol 86:256–267
Plouviez S, Faure B, Le Guen D, Lallier FH, Bierne N, Jollivet D (2013) A new barrier to dispersal trapped old genetic clines that escaped the easter microplate tension zone of the Pacific vent mussels. PLoS One 8:e81555
Polz MF, Cavanaugh CM (1995) Dominance of one bacterial phylotype at a mid-Atlantic ridge hydrothermal vent site. Proc Natl Acad Sci USA 92:7232–7236
Price RE, Lesniewski R, Nitzsche KS, Meyerdierks A, Saltikov C, Pichler T, Amend JP (2013) Archaeal and bacterial diversity in an arsenic-rich shallow-sea hydrothermal system undergoing phase separation. Front Microbiol 4:158
Raghukumar C, Mohandass C, Cardigos F, D’Costa PM, Santos RS, Colaco A (2008) Assemblage of benthic diatoms and culturable heterotrophs in shallow-water hydrothermal vent of the D. Joao de Castro Seamount, Azores in the Atlantic Ocean. Curr Sci 95:1715–1723
Raguenes G, Cambon-Bonavita MA, Lohier JF, Boisset C, Guezennec J (2003) A novel, highly viscous polysaccharide excreted by an Alteromonas isolated from a deep-sea hydrothermal vent shrimp. Curr Microbiol 46:448–452
Rajasabapathy R, Mohandass C, Colaco A, Dastager SG, Santos RS, Meena RM (2014) Culturable bacterial phylogeny from a shallow water hydrothermal vent of Espalamaca (Faial, Azores) reveals a variety of novel taxa. Curr Sci 106:58–69
Rogers JS (1997) On the consistency of maximum likelihood estimation of phylogenetic trees from nucleotide sequences. Syst Biol 46:354–357
Takai K, Komatsu T, Inagaki F, Horikoshi K (2001) Distribution of archaea in a black smoker chimney structure. Appl Environ Microbiol 67:3618–3629
Takai K, Inagaki F, Nakagawa S, Hirayama H, Nunoura T, Sako Y, Nealson KH, Horikoshi K (2003) Isolation and phylogenetic diversity of members of previously uncultivated epsilon-proteobacteria in deep-sea hydrothermal fields. FEMS Microbiol Lett 218:167–174
Takai K, Oida H, Suzuki Y, Hirayama H, Nakagawa S, Nunoura T, Inagaki F, Nealson KH, Horikoshi K (2004) Spatial distribution of marine Crenarchaeota group I in the vicinity of deep-sea hydrothermal systems. Appl Environ Microbiol 70:2404–2413
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Tasiemski A, Jung S, Boidin-Wichlacz C, Jollivet D, Cuvillier-Hot V, Pradillon F, Vetriani C, Hecht O, Sönnichsen FD, Gelhaus C, Hung CW, Tholey A, Leippe M, Grötzinger J, Gaill F (2014) Characterization and function of the first antibiotic isolated from a vent organism: the extremophile metazoan Alvinella pompejana. PLoS One 9:e95737
Taylor PW (2013) Alternative natural sources for a new generation of antibacterial agents. Int J Antimicrob Agents 42:195–201
Thankappan B, Ramesh D, Ramkumar S, Natarajaseenivasan K, Anbarasu K (2014) Characterization of Bacillus spp. from the gastrointestinal tract of Labeo rohita-towards to identify novel probiotics against fish pathogens. Appl Biochem. doi:10.1007/s12010-014-1270-y
Thornburg CC, Zabriskie TM, McPhail KL (2010) Deep-sea hydrothermal vents: potential hot spots for natural products discovery? J Nat Prod 73:489–499
Tollefson J (2014) China plunges into ocean research. Nature 506:276
Valiente Moro C, Tran FH, Raharimalala FN, Ravelonandro P, Mavingui P (2013) Diversity of culturable bacteria including Pantoea in wild mosquito Aedes albopictus. BMC Microbiol 13:70
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267
Watsuji T, Nakagawa S, Tsuchida S, Toki T, Hirota A, Tsunogai U, Takai K (2010) Diversity and function of epibiotic microbial communities on the galatheid crab, Shinkaia crosnieri. Microbes Environ 25:288–294
Yin Q, Fu BB, Li BY, Shi XC, Inagaki F, Zhang XH (2013) Spatial variations in microbial community composition in surface seawater from the ultra-oligotrophic center to rim of the south Pacific Gyre. PLoS One 8:e55148
Zhao W, Zeng X, Xiao X (2014) Thermococcus eurythermalis sp. nov., a conditional piezophilic hyperthermophilic archaeon with a wide temperature range isolated from an oil-immersed chimney in the Guaymas Basin. Int J Syst Evol Microbiol. doi:10.1099/ijs.0.067942-0
Zhou HY, Li JT, Peng XT, Meng J, Wang FP, Ai Y (2009a) Microbial diversity of a sulfide black smoker in main endeavour hydrothermal vent field, Juan de Fuca Ridge. J Microbiol 47:235–247
Zhou MY, Chen XL, Zhao HL, Dang HY, Luan XW, Zhang XY, He HL, Zhou BC, Zhang YZ (2009b) Diversity of both the cultivable protease-producing bacteria and their extracellular proteases in the sediments of the South China Sea. Microb Ecol 58:582–590
Zhou MY, Wang GL, Li D, Zhao DL, Qin QL, Chen XL, Chen B, Zhou BC, Zhang XY, Zhang YZ (2013) Diversity of both the cultivable protease-producing bacteria and bacterial extracellular proteases in the coastal sediments of King George Island, Antarctica. PLoS One 8:e79668
Acknowledgments
This work was supported by the grants from the Strategic Priority Research Program of the Chinese Academy of Sciences (XDA11030401) and the Taishan Scholar Program of Shandong Province. We thank the WPOS Sample Center for providing the samples.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflict of interest to declare.
Rights and permissions
About this article
Cite this article
Sun, Ql., Wang, Mq. & Sun, L. Characteristics of the cultivable bacteria from sediments associated with two deep-sea hydrothermal vents in Okinawa Trough. World J Microbiol Biotechnol 31, 2025–2037 (2015). https://doi.org/10.1007/s11274-015-1953-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11274-015-1953-8