Abstract
Kefir—a traditional beverage whose consumption has been associated with health benefits—is a logical natural product to investigate for new probiotic strains. The aim of the present work was to isolate and identify kefir yeasts and select those with acid and bile tolerance to study their adhesion to epithelial cells and their transit through mouse gut. From 4 milky and 3 sugary kefir grains, 34 yeast strains were isolated and identified by means of classical microbiological and molecular-genetic methods (whole-cell protein pattern, internal-transcribed-spacer amplification, and analysis of restriction-fragment–length polymorphisms). We identified 4 species belonging to 3 genera—Saccharomyces cerevisiae (15 strains), Saccharomyces unisporus (6 strains), Issatchenkia occidentalis (4 strains), and Kluyveromyces marxianus (9 strains)—and selected 13 strains on the basis of resistance to low pH and bile salts. Among the strains selected, Kluyveromyces marxianus CIDCA 8154 and Saccharomyces cerevisiae CIDCA 8112 were further studied. Both strains evidenced the capacity to adhere to epithelial intestine-derived cells in vitro and to survive passage through the gastrointestinal tract of BALB/c mice. The investigation of the potential probiotic features of these kefir-yeast strains should be useful for the development of novel functional foods.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Yeasts are one of the groups of microorganisms most commonly used for animal and human consumption in the food industry worldwide; with Saccharomyces, Candida, and Kluyveromyces being the most representative genera.
Probiotics are defined as: “live microorganisms which, when administered in adequate amounts, exert a beneficial effect on the health of the consumer” (FAO/WHO 2001). Although lactic-acid bacteria and bifidobacteria are the microorganisms most widely studied for probiotic properties, the use of yeast as a probiotic food supplement is gaining relevance (Fleet and Balia 2006). Saccharomyces cerevisiae var. boulardii (Saccharomyces boulardii)—available in various commercial formulations and the probiotic yeast most widely used—is being employed as a therapeutic agent for the treatment of a variety of gut disorders to normalize intestinal flora (Saad et al. 2013; Szajewska et al. 2007; Zanello et al. 2009). Numerous advances in the understanding of the beneficial effects and mechanisms of action of that yeast have been published in recent years (Czerucka et al. 2007). The search for other yeasts with probiotic potential for application to the food industry in additional ways is also an increasing area of investigation. The criteria for the selection of probiotic microorganisms still constitute a topic of controversy, but on consideration of the mechanisms of action two conditions have been widely accepted for the selection: the ability to survive in the gastrointestinal environment and the possession at least one beneficial function (Martins et al. 2005; Morelli 2007).
Kefir is an ancient milk beverage produced by the lactic-acid and alcohol fermentation on the part of mesophilic bacteria and yeasts, respectively. These microorganisms are present in the polysaccharide and protein matrix of the kefir grains that are employed as a starter. The consumption of kefir has been associated with longevity in the people of Caucasus and has been shown to produce health benefits. Kefir has antimicrobial, antihypertensive, anti-inflammatory, anticarcinogenic, antiallergic, and antioxidant activity; participates in immune-system modulation; reduces cholesterol levels; and alleviates lactose intolerance (Ahmed et al. 2013; Farnworth 2005). Another widely disseminated type of fermented beverage is sugary kefir, prepared with kefir grains that ferment sweetened water to produce an effervescent product. Various health benefits have been empirically attributed to sugary kefir, but only its anti-inflammatory activity has been demonstrated conclusively (Moreira et al. 2008).
The beneficial features mentioned here would indicate kefir as being a promising possible source of new microbial strains for the development of functional foods. The aim of the present work was therefore to isolate and identify kefir yeasts that would be suitable as probiotics, to select those strains with acid and bile tolerance, and to study both their adhesion to epithelial cells and their transit through the murine gut.
Materials and methods
Yeast isolates
Yeasts were isolated from 4 milky kefir grains (CIDCA AGK1, CIDCA AGK5, CIDCA AGK7, and CIDCA AGK10) and 3 sugary kefir grains (CIDCA SK1, CIDCA SK2, and CIDCA SK3) that were obtained from Argentine families that traditionally consumed kefir. Kefir grains were ground in a mortar and resuspended in 0.1 % (w/v) triptone in water. Yeasts were isolated by surface—streaking onto yeast-extract–glucose–chloramphenicol (YGC) agar (Merck, D-64271 Darmstadt, Germany) and incubated at 30 °C for 48 h under aerobic conditions. Thirty-four visually different colonies were isolated.
Saccharomyces boulardii was isolated from the commercial probiotic product Floratil® (Lab Biocodex, France).
Identification of yeast isolates
Yeasts were characterized by macroscopic and microscopic morphology, growth in malt extract (Biokar Diagnostics, France), the mode of reproduction, growth kinetics at 37 °C, the ability to hydrolyze urea and ferment sugar (glucose, galactose, sucrose, maltose, lactose), and the assimilation of carbon compounds by means of an API 20C AUX system (BioMerieux, France). Yeasts were identified according to the criteria of Kurtzman and Fell (1998).
Whole-cell–protein pattern of yeasts
In order to obtain cell extracts for total-protein analysis, 10 mL of yeast overnight cultures grown at 30 °C in yeast-extract–peptone–dextrose broth (YPD; 1 % yeast extract, 2 % peptone, and 2 % dextrose, w/v) were centrifuged at 10,000×g for 10 min. The pellet was resuspended in 5 mL of distilled water, placed in an ice bath, and sonicated twice for 6 min before recentrifugation at 10,000×g for 10 min. The supernatant was diluted in an equal volume of sample buffer to obtain a final concentration of 123 mM Tris–HCl with 0.1 % SDS, pH 6.8; 5 % 2-mercaptoethanol; 25 % glycerol; and 0.01 % bromophenol blue. Thirty microliters of the soluble protein fraction in sample buffer were applied to the gels.
Protein profiles were analyzed by SDS-polyacrylamide gel electrophoresis (SDS-PAGE) in 12.5 % (w/v) gels according to Laemmli (1970) in a Miniprotean II cell (Biorad Lab, Richmond, CA, USA). Electrophoresis was run at a constant voltage of 120 V and the resulting gels stained in a solution of 10 g Coomasie blue R-250, 400 mL methanol, and 166 mL acetic acid. Distaining was in 250 mL of 40 % (v/v) acetic acid in ethanol with continuous shaking.
The patterns of the stained proteins were analyzed by the Gel-Pro Analyzer software (Media Cybernetics, USA), with the following proteins being used as molecular-weight markers: phosphorylase B, albumin, ovalbumin, carbonic anhydrase, trypsin inhibitor, and α-lactalbumin (94, 67, 43, 30, 20.1, and 14.4 kDa, respectively; Pharmacia Biotech, Uppsala, Sweden). The molecular weight of the proteins in all strains was calculated and the presence or absence of each band recorded. GelCompar II version 4.6 software (Applied Maths) was used for data analysis. A dendrogram was constructed on the basis of the protein profiles by the unweighted-average-linkage method (UPGMA) through the use of the Jaccard correlation coefficient and a 3 % tolerance for band position.
Yeast-DNA isolation
Yeasts were cultured on malt-extract broth for 24 h at 30 °C. The DNA was isolated according to the protocol included with the GFX Genomic Blood DNA Purification Kit (Amersham Biosciences, UK) after a treatment with 2.5 mg/mL litycase (Sigma, USA) and 10 mg/mL proteinase K. DNA electrophoresis was run in 0.8 % (w/v) agarose gels (Promega Corporation, Madison, WI, USA) with TBE buffer (1.08 % [w/v] Tris base, 0.55 % [w/v] boric acid, 0.02 M ethylenediaminetetraacetic acid; pH 8) in a horizontal electrophoresis system (Bio-Rad, Hercules, CA, USA) for 20 min at a constant current of 80 V. Gels were stained with ethidium bromide and visualized under ultraviolet light in a LABNET TM-26 (Edison, NJ, USA) transiluminator.
Yeast ITS-region polymorphism
Yeast DNA was amplified with the primers ITS1 (5′-TCC GTA GGT GAA CCT GCG G-3′) and ITS4 (5′-TCC TCC GCT TAT TGA TAT GC-3′) spanning the internal transcribed sequences (ITS) between the functional 18S, 5.8S, and 28S elements of the ribosomal-RNA gene (Wyder 1998).
The polymerase-chain-reaction (PCR) DNA-sequence amplification was performed in a total reaction volume of 15 μL containing 0.5 μM of each primer, 2.5 U of Taq DNA polymerase (Inbio Highway, Tandil, Argentina), 1 μL of the buffer supplied with the enzyme (100 mM Tris–HCl, 500 mM KCl, pH 9), 1.25 mM MgCl2, 0.2 mM of each dNTP, and 1 μL of the isolated DNA. PCR reactions were performed in a MyCycler Thermal Cycler (Bio-Rad, Hercules, CA, USA). The PCR program consisted of an initial 5-min denaturation at 94 °C, 35 1-min denaturation cycles at 94 °C, a 1-min annealing at 56 °C, and a 2-min extension at 72 °C, followed by a final 10-min extension at 72 °C. Aliquots of the amplification products along with a 100-bp DNA ladder (Inbio Highway, Tandil, Argentina) were analyzed by electrophoresis in horizontal 1.2 % (w/v) agarose gels with TBE buffer for 30 min at a constant current of 70 V. Gels were stained with ethidium bromide and visualized under ultraviolet light. Molecular weights were estimated with Gel-Pro Analyzer software (Media Cybernetics, USA).
Restriction analysis of the ITS region
HinfI, TaqI, and MspI (Fermentas, Life Sciences, USA) restriction endonucleases were used separately to digest the ITS amplification products. Ten microliters of PCR products were digested with 10 U of endonuclease under the conditions recommended for each enzyme. Restriction fragments of the ITS-PCR products were separated by electrophoresis in horizontal 3.0 % (w/v) agarose gels in TBE buffer along with the 100-bp DNA ladder for 40 min at a constant current of 70 V. Gels were stained and visualized as described above.
Resistance of yeasts to simulated gastrointestinal conditions
Measurement of bile resistance was performed by a modified ecometric method according to Kociubinski et al. (1999). The method stated in brief: plates of YGC medium (control for growth) and YGC medium containing 1 % (w/v) bile salts (Merck, Darmstadt, Germany) were inoculated with overnight yeast cultures grown at 30 °C in YPD broth and incubated at 30 °C for 48 h. Bile resistance was considered high (+++) when no difference from the control growth was observed.
In order to determine the acid resistance of isolates, yeasts were grown in YPD broth for 24 h at 30 °C, centrifuged, and resuspended at a concentration of 108 CFU/mL in YEGP broth (0.5 % [v/v] yeast extract, 2 % [w/v] glucose, 1 % [w/v] peptone) acidified to pH 2.5 with 3 M HCl. Samples were taken immediately (0 h) and after 3 h at 37 °C, and serial dilutions in 0.1 % (w/v) tryptone in water were plated on YGC agar in order to determine the number of viable cells and calculate the percent survival.
Yeast survival under gastrointestinal conditions in vivo and distribution upon passage through the mouse gut
Six-week-old BALB/c male mice (4 animals per group) were treated with a suspension of 107 CFU/mL of K. marxianus CIDCA 8154, S. cerevisiae CIDCA 8112, or S. boulardii in their ad libitum drinking water for 12 days. Control animals received the normal drinking water without any additions. All groups were fed with standard laboratory mouse chow and housed in a climate-controlled room on a 12-h light–dark cycle. Feces were collected on days 2, 4, 7, 9, 11, 15, and 17; weighed; diluted 100-fold; and resuspended in physiological saline. Serial dilutions were performed in sterile PBS (1.5 mM KH2PO4, 8.1 mM Na2HPO4·2H2O, 0.14 M NaCl, 2.7 mM KCl, pH 7.4) and yeast colony formation counted after seeding on YGC agar. The results were expressed as CFU/g of feces.
To determine the distribution of the yeast within the gastrointestinal tract, another set of experiments was performed in which mice—treated exactly as described above—were sacrificed by cervical dislocation on day 7 of the treatment. One centimeter of each section of the gastrointestinal tract (duodenum, ileum, cecum, and colon) was dissected and the contents scraped from the lumen and resuspended in physiological saline for yeast-colony counts after seeding on YGC agar. The results were expressed as CFU/cm of tissue.
The mice employed were specific pathogen-free, provided by Universidad Nacional de La Plata animal house. All animal experiments were performed according to the guidelines set by the National Institute of Health (NIH publication No. 86-23, 1985 revision).
Yeast adhesion to Caco-2/TC7 cells
Caco-2/TC7 cells, derived from a human epithelial colorectal adenocarcinoma, were routinely grown following the procedure described by Golowczyc et al. (2007). Cells, at subculture passages between 23 and 30, were seeded at a concentration of 2.5 × 105 cells per well in 24-well tissue-culture plates (Corning, NY, USA) and used at postconfluence after 7 days of culture (differentiated cells).
Overnight cultures of K. marxianus CIDCA 8154, S. cerevisiae CIDCA 8112, and S. boulardii on YPD broth were centrifuged and the pellet resuspeded in a sufficient volume of Dulbecco’s Modified Eagle’s Minimal Essential Medium (GIBCO BRL Life Technologies Rockville, USA) to reach an OD600nm = 10 (~108 CFU/mL). Serial dilutions were made in the same medium to concentrations of 107, 106, and 105 CFU/mL. For the adhesion assay, each well was incubated with 0.5 mL of a yeast suspension for 1 h at 37 °C in a 5 %-CO2–95 %-air atmosphere. The monolayer was then washed three times with PBS and lysed in 0.5 mL of sterile distilled water. To determine the number of viable yeast cells associated with the Caco-2 cells, the appropriate dilutions in 0.1 % (w/v) tryptone in water were plated on YGC and colonies were counted. All experiments were performed in triplicate. The percent of yeast adhering to the epithelial cells was calculated with respect to the total number inoculated.
Results and discussion
Yeast isolation and identification
A total of 34 yeast isolates were identified on the basis of physiological and biochemical tests following the criteria of Kurtzman and Fell (1998) and grouped into 4 species belonging to 3 genera. From the 25 lactose-nonfermenting isolates, 15 proved to be Saccharomyces cerevisiae (isolates CIDCA 8112, 8115, 8175, 81102, 81103, 81106, 81108, 81109, 9123, 9124, 9127, 9128, 9132, 9133, and 9136), 6 Saccharomyces unisporus (isolates CIDCA 8111, 8151, 8152, 8155, 81101, and 81107), and 4 Issatchenkia occidentalis (isolates CIDCA 9111, 9125, 91210, and 9131). In addition, 9 lactose-fermenting isolates were identified as Kluyveromyces marxianus (isolates CIDCA 8113, 8116, 8118, 81111, 8153, 8154, 81104, 81105, and 9121).
Whole-cell–protein profiles were suitable for distinguishing the 4 yeast species isolated from the kefir grains (Fig. 1). Protein profiles of all the isolates were obtained and a dendrogram of the relationship among them constructed (Fig. 2). Cluster analysis confirmed the determinations made from the microscopic characteristics along with the assimilation and fermentation of sugars since the isolates corresponding to different species grouped in separate clusters. The lactose-fermenting K. marxianus grouped in one cluster, while the lactose-nonfermenting yeasts grouped in another. Within the cluster of the latter, with 50.47 % similarity, one subcluster corresponds to the genus Saccharomyces and the other to Issatchenkia. Among the Saccharomyces yeasts, strains that belonged to S. cerevisiae had 72.20 % similarity and those of S. unisporus 90.91 %. The I. occidentalis isolates grouped at a 90.48 % degree of similarity. Finally, within the cluster of the species K. marxianus, the isolates shared a 72.42 % similarity. The application of whole-cell–protein electrophoresis has been successfully used by others for different yeast-classification purposes (Noumi et al. 2012; Paramithiotis et al. 2000; Vancanneyt et al. 1992). Our results indicate that analysis by SDS-PAGE enables the facile and prompt acquisition of species-specific patterns that could be used to differentiate yeasts present in kefir.
A length polymorphism of ITS1–ITS4 region was observed in the PCR products among the species with the sizes ranging between 420 and 915 bp (Table 1; Fig. 3). I. occidentalis had the shortest amplified sequence at about 435 bp, followed by K. marxianus and S. unisporus at around 740 and 770 bp, respectively, and finally by S. cerevisiae at roughly 880 bp. These lengths were in agreement with previously reported values (Bockelmann et al. 2008; Clemente Jimenez et al. 2004; Coton et al. 2006; Esteve Zarzoso et al. 1999; Latorre García et al. 2007; Fernandez-Espinar et al. 2000; Orberá Ratón 2004). The ITS electrophoretic analysis thus allowed the differentiation of S. cerevisiae and I. occidentalis from the other kefir isolates but not K. marxianus from S. unisporus since the ITS fragments of those species were too similar in length. This circumstance is in agreement with earlier findings, where the size of the amplified ITS1-ITS4 fragment did not constitute sufficient information for differentiating among species. We thus decided to perform a restriction-length–polymorphism analysis on the ITS fragments in order to explore the sequence heterogeneity present.
Agarose-gel electrophoresis of ITS1–ITS4 PCR products before (a) and after (b) their digestion with TaqI, HinfI, or MspI. Lane 1, Kluyveromyces marxianus CIDCA 8154; lane 2, K. marxianus CIDCA 81104; lane 3, Saccharomyces cerevisiae CIDCA 8112; lane 4, S. cerevisiae CIDCA 9127; lane 5, Saccharomyces unisporus CIDCA 81107; lane 6, Issatchenkia occidentalis CIDCA 9131; lane 7, I. occidentalis CIDCA 9125. MW molecular weight, bp base pairs
Restriction analysis of the ITS1-ITS4 PCR products was accordingly performed with the enzymes TaqI, HinfI, and MspI. Figure 3 shows the restriction profiles obtained for representative isolates from each species. The fragment sizes of the restriction products obtained for each yeast isolate are listed in Table 1. Treatment of the ITS1–ITS4 region with the enzyme MspI produced different fragments from I. occidentalis (about 240 and 110 bp) and S. cerevisiae (about 650 and 115 bp), whereas for K. marxianus and S. unisporus fragments of about 680 and 700 bp, respectively, were obtained (Fig. 3). For K. marxianus and S. unisporus the difference between the intact ITS1–ITS4 regions and the MspI restriction-fragment sizes was therefore about 60–70 bp. Cleavage fragments with those small sizes were not detected in these runs, probably because fragments shorter than 100 bp are often not visualized on gels. Regardless of kefir-grain origin, the size of the PCR-amplification products combined with the restriction analysis of the ITS1–ITS4 region with the endonucleases TaqI or HinfI allowed a differentiation of all the species analyzed (Table 1). Those restriction profiles were accordingly similar among the strains that belonged to the same species and furthermore had comparable lengths to those that had been previously published (Clemente Jimenez et al. 2004; Esteve Zarzoso et al. 1999; Fernandez-Espinar et al. 2000; Latorre García et al. 2007; Orberá Ratón 2004).
We wish to emphasize that both the physiological tests and the molecular analyses contribute to the identification of isolates. Although the characterization of the isolates by physiological testing is laborious, that method allows the unequivocal identification of each species. By contrast, the analysis of the total-protein profile is a simpler and faster method; but, like the restriction analysis of the ITS1–ITS4 region, needs the inclusion of accurate reference strains for a definitive assignment of identity. These conclusions are also in agreement with the statements by other authors that although molecular attributes form the basis of the current taxonomy of yeasts, physiological tests are still necessary for identification (Senses-Ergul et al. 2006).
Despite many years of study, the microbiological composition of kefir grains is still not yet fully known. The food standards of the Food and Agriculture Organization of the United Nations state that kefir possesses lactose-fermenting yeasts (K. marxianus) and nonlactose-fermenting (S. unisporus, S. cerevisiae, and S. exiguus), but the yeast composition has not been fully defined since other authors have found additional species (Mainville et al. 2006). Among the yeasts registered in kefir grains from different origins the genera Zygosaccharomyces, Candida, Kluyveromyces, Saccharomyces, Torulaspora, Issatchenkia, Pichia, and Debaryomyces may be mentioned (Ahmed et al. 2013); with Kluyveromyces, Saccharomyces, and Issatchenkia being isolated in the present work from the grains of the CIDCA collection. The composition of kefir microflora strongly depends on the origin of the grains, on the local conditions of culture, and on the storage and elaboration processes (Garrote et al. 2001, 2010). With respect to yeast composition, three of the milky-kefir grains analyzed here—CIDCA AGK1, AGK5, and AGK10—were found to contain S. cerevisiae, S. unisporus, and K. marxianus; whereas only S. cerevisiae was isolated from CIDCA AGK7 grains. Similarly, in a previous study the same three species were detected in kefir grains CIDCA AGK10 by means of an independent culture method (Londero et al. 2012). In contrast, the 3 sugary kefir grains studied—CIDCA SK1, SK2, and SK3—contained, in addition to the three above-mentioned species, yeasts belonging to the genus Issatchenkia.
Survival of yeasts after passage through the gastrointestinal tract as assayed in vitro and in vivo
Survival to the passage through the gastrointestinal tract is a desirable characteristic in the choice of probiotic microorganisms since viability plays a significant role in certain of their beneficial properties (Romanin et al. 2010; Saad et al. 2013). The potential ability of the identified isolates to survive under the conditions of transit through the gastrointestinal tract as assayed indirectly in vitro is demonstrated by the results presented in Table 2. All the species tested contained strains that were highly resistant to bile salts and an acidic environment.
As to the bile resistance, all the yeasts were able to grow in the presence of 0.5 % (w/v) bile salts in the culture medium (data not shown), while 20 isolates grew well (+++ and ++) even when the concentration was as high as 1 % (w/v). That a strain of Saccharomyces boulardii isolated from a commercial probiotic product and included in the present study was more sensitive to bile salts than several strains isolated from kefir is furthermore highly noteworthy.
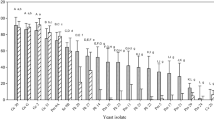
With respect to acid tolerance, the highest percentage of survival was observed for S. cerevisiae CIDCA 8175, 81103, 81106, and 9133 along with I. occidentalis CIDCA 9125. Fourteen isolates exhibited survivals between 50 % and 90 % after 3 h, whereas with the remaining strains the viability decreased by more than 50 %.
That the food matrix could exert an additional protective action against acid damage deserves consideration. Based on the capacities of both acid and bile resistance, 13 yeast isolates (shown in bold in Table 2) were preferred as probiotic candidates. Among them, K. marxianus CIDCA 8154 and S. cerevisiae CIDCA 8112 were selected as representatives of each of the principal genera found in CIDCA kefir grains for the following studies both on the basis of their immunomodulatory capacity (Romanin et al. 2010) and on their capability to reduce the cytotoxic action of Clostridium difficile and frost resistance (Diosma 2010). In addition, the aforementioned commercial probiotic strain of S. boulardii was included in the studies as a reference organism.
The adhesive capacity of selected yeasts was examined in vitro with the Caco-2/TC7 intestine-derived cell line. Regardless of the yeast concentration added to Caco-2/TC7 cells (e. g., 108, 107, 106, or 105 CFU/mL), the percent adhesion was 3.0 ± 0.9 % for K. marxianus CIDCA 8154, 1.5 ± 0.4 % for S. boulardii, and 0.5 ± 0.1 % for S. cerevisiae CIDCA 8112. In agreement with these results, Kumura et al. (2004) had reported a 4 % adhesion to Caco-2 cells for a strain of K. marxianus isolated from kefir. Different percentages of adhesion to intestinal cells have been reported for S. cerevisiae varying from 0.6 to 6.2 % for 6 strains (Klingberg et al. 2008), from 0.2 to 40 % for 4 strains (Kumura et al. 2004), and from 1.9 to 16.8 % for 18 strains (van der Aa Kühle et al. 2005). These last authors described that the adhesion of 8 S. boulardii strains to the neonatal-piglet mid-jejunal-epithelium cell line (IPEC-J2) varied from 1.1 to 28.0 %. Therefore, compared to the studies cited here, the S. cerevisiae CIDCA 8112 and S. boulardii used in the present work adhered poorly to epithelial cells.
The ability to adhere to intestinal epithelium has been widely used as a criterion to select and characterize probiotic yeasts (Maccaferri et al. 2012; Kourelis et al. 2010; Kumura et al. 2004; van der Aa Kühle et al. 2005). Nevertheless, the relevance of adhesivity to the probiotic capacity of yeast is still unclear. The ability to modulate the immune system is generally considered a highly desirable characteristic of probiotic strains (Collado et al. 2009), and the contact between the yeast and the epithelial cells seems to be involved in the down-regulation of the epithelial inflammatory response induced by flagellin (Romanin et al. 2010). In contrast, for other yeast mechanisms of action—such as coaggregation with intestinal pathogens (Tiago et al. 2012) or toxin binding (Brandão et al. 1998; Shetty and Jespersen 2006)—a prompt transit through the gastrointestinal tract may be advantageous. In addition, the beneficial effects of yeasts with low adhesion capacities have been described. Tasteyre et al. (2002) demonstrated that a nonadhesive S. boulardii strain inhibited the adhesion of Clostridium difficile to Vero green-monkey-kidney cells through proteolytic activity as well as through steric hindrance. van der Aa Kühle et al. (2005) described that the expression of IL-1α decreased in IPEC-J2 cells exposed to a Shiga-like toxin 2 when the cells were either pre- or coincubated with S. boulardii even though this yeast strain was of low adhesion, thus suggesting that adhesivity is not necessarily a mandatory prerequisite for probiotic effects. For these reasons, a further investigation of the influence of adhesion on yeast mechanisms of action would be of great usefulness in order to clarify whether or not adhesion to intestinal epithelium is a relevant criterion for the selection of probiotic yeasts.
Other considerations necessary for the choice of probiotic-yeast candidates are their survival, persistence, and distribution through the gastrointestinal tract. These aspects were studied here for the three selected strains in an in vivo murine model. All the strains tested tolerated passage through the gastrointestinal tract of inoculated Balb/c mice since they were subsequently detected in the feces during the entire period of probiotic administration (Fig. 4a). After 4 days, the yeast concentration was 104–106 CFU/g feces for all the mice that had consumed the probiotic. A maximum concentration of 107 CFU/g feces was detected on Day 7 and was maintained until the last determination on Day 11 before the end of administration on Day 12. No significant differences in yeast counts among the treatments were detected. That kefir-isolated yeasts showed the same capacity to survive and persist in the mouse intestinal tract as the S. boulardii commercial probiotic strain is of extreme relevance here. In all instances, the yeast could be detected in feces for up to 3 days after the termination of administration. These results are in agreement with other pharmacokinetic studies performed in man and in rats demonstrating that S. boulardii organisms were cleared from the stools by 2–5 days after discontinuation of ingestion (Edwards-Ingram et al. 2007; Elmer et al. 1999; Pecquet et al. 1991).
Yeast counts from feces (a) or from different portions of gastrointestinal tract (b) of mice treated during 12 or 7 days, respectively, with Kluyveromyces marxianus CIDCA 8154 (filled bar), Saccharomyces cerevisiae CIDCA 8112 (cross lined bar) or Saccharomyces boulardii (empty bar) present in drinking water at concentrations of 107 CFU/mL. The results are expressed as mean ± SE of means
To characterize the distribution of the yeast in the gastrointestinal tract, Balb/c mice were treated with K. marxianus CIDCA 8154 or S. boulardii for 7 days (Fig. 4b). The highest concentration of yeast was found in the distal portions of the gut, reaching a mean density of 104 CFU/cm in the colon and of 105 CFU/cm in the cecum. Similar results had been found by Edwards-Ingram et al. (2007), who described that, at 1 and 3 h after gastric gavage administration of 108 CFU/mL of 2 S. cerevisiae strains and 1 S. boulardii strain, a majority of the yeasts were present in the colon and the cecum in a viable state. Since only little information about the survival and distribution of probiotic yeasts in the gastrointestinal tract is available, we need to emphasize that the findings here may be useful for the design of further probiotic-based strategies aimed at treating different gastrointestinal-tract disorders.
Conclusions
In the present study 15 strains of S. cerevisiae, 6 strains of S. unisporus, 4 strains of I. occidentalis, and 9 strains of K. marxianus were isolated from the complex microbial consortium of kefir grains. Among them, 13 isolates have high acid- and bile-resistance phenotypes and are thus potentially suitable for probiotic purposes. K. marxianus CIDCA 8154 and S. cerevisiae CIDCA 8112 also exhibited a capacity to adhere to intestinal epithelial cells in vitro, survive passage through the gastrointestinal tract of Balb/c mice, and during that intestinal residence spread all along the tract with higher abundance throughout the distal portions. At the end of the administration both yeast strains become cleared from the gut within 5 days. Although further studies about the health benefits from and mechanisms involved in these findings are still required, the results obtained here should nevertheless be useful for the development of new probiotic products based on the different strains of yeast isolated from kefir.
References
Ahmed Z, Wang Y, Ahmad A, Khan ST, Nisa M, Ahmad H, Afreen A (2013) Kefir and health: a contemporary perspective. Crit Rev Food Sci 53:422–434
Bockelmann W, Heller M, Heller KJ (2008) Identification of yeasts of dairy origin by amplified ribosomal DNA restriction analysis (ARDRA). Int Dairy J 18:1066–1071
Brandão RL, Castro IM, Bambirra EA, Amaral SC, Fietto LG, Tropia MJM, Neves MJ, Dos Santos RG, Gomes NCM, Nicoli JR (1998) Intracellular signal triggered by cholera toxin in Saccharomyces boulardii and Saccharomyces cerevisiae. Appl Environ Microbiol 64:564–568
Clemente Jimenez JM, Mingorence Cazorla L, Martínez Rodriguez S, Las Heras Vázquez FJ, Rodríguez Vico F (2004) Molecular characterization and oenological properties of wine yeasts isolated during spontaneous fermentation of six varieties of grape must. Food Microbiol 21:149–155
Collado MC, Isolauri E, Salminen S, Sanz Y (2009) The impact of probiotic on gut health. Curr Drug Metab 10:68–78
Coton E, Coton M, Levert D, Casaregola S, Sohier D (2006) Yeast ecology in French cider and black olive natural fermentations. Int J Food Microbiol 108:130–135
Czerucka D, Piche T, Rampal P (2007) Review article: yeast as probiotics—Saccharomyces boulardii. Alim Pharmacol Therap 26:767–778
Diosma G (2010) Estudio y selección de levaduras con propiedades probióticas. Magister Thesis. Facultad de Ciencias Agrarias y Forestales. Universidad Nacional de La Plata, Argentina
Edwards-Ingram L, Gitsham P, Burton N, Warhurst G, Clarke I, Hoyle D, Oliver SG, Stateva L (2007) Genotypic and physiological characterization of saccharomyces boulardii, the probiotic strain of Saccharomyces cerevisiae. Appl Environ Microbiol 73:2458–2467
Elmer GW, McFarland LV, Surawicz CM, Danko L, Greenberg RN (1999) Behaviour of Saccharomyces boulardii in recurrent Clostridium difficile disease patients. Aliment Pharmacol Ther 13(12):1663–1668
Esteve Zarzoso B, Belloch C, Uruburu F, Querol A (1999) Identification of yeasts by RFLP analysis of the 5.8S rRNA gene and the two ribosomal internal transcribed spacers. Int J Syst Bacteriol 49:329–337
FAO/WHO (2001) Joint FAO/WHO expert consultation on evaluation of health and nutritional properties of probiotics in food including powder milk with live lactic acid bacteria. Cordoba, Argentina 1–4 October 2001. Available at: http://www.who.int/foodsafety/publications/fs_management/
Farnworth ER (2005) Kefir—a complex probiotic. Food Sci Technol Bull Funct food 2:1–17
Fernandez-Espinar TM, Esteve Zarzoso B, Querol A, Barrio E (2000) RFLP analysis of the internal transcribed spacers and the 5,8S rRNA gene región of the genus Saccharomyces: a fast method for species identification and the differentiation of flor yeasts. Anton Leeuw Int J G 78:87–97
Fleet GH, Balia R (2006) The Public Health and Probiotic Significance of Yeasts in Foods and Beverages. In: Querol A, Fleet G (eds) The yeast handbook. Yeasts in food and beverages. Springer, Heidelberg, pp 381–397
Garrote GL, Abraham AG, De Antoni G (2001) Chemical and microbiological characterization of kefir grains. J Dairy Res 68(4):639–652
Garrote GL, Abraham AG, De Antoni GL (2010) Microbial Interactions in Kefir: a natural probiotic drink. In: Mozzi F, Raya RR, Vignolo GM (eds) Biotechnology of lactic acid bacteria: novel applications. ISBN: 978-0-8138-1583-1. Wiley-Blackwell, Ames, IO, USA, pp 327–340
Golowczyc MA, Mobili P, Garrote GL, Abraham AG, De Antoni GL (2007) Protective action of Lactobacillus kefir carrying S-layer protein against Salmonella enterica serovar Enteritidis. Int J Food Microbiol 118(3):264–273
Klingberg TD, Lesnik U, Arneborg N, Raspor P, Jespersen L (2008) Comparison of Saccharomyces cerevisiae strains of clinical and nonclinical origin by molecular typing and determination of putative virulence traits. FEMS Yeast Res 8(4):631–640
Kociubinski G, Pérez P, De Antoni G (1999) Screening of bile resistance and bile precipitation in lactic acid bacteria and bifidobacteria. J Food Protect 62:905–912
Kourelis A, Kotzamanidis C, Litopoulou-Tzanetaki E, Scouras ZG, Tzanetakis N, Yiangou M (2010) Preliminary probiotic selection of dairy and human yeast strains. J Biol Res-Thessalon 13:93–104
Kumura H, Tanoue Y, Tsukahara M, Tanaka T, Shimazaki K (2004) Screening of dairy yeast strains for probiotic applications. J Dairy Sci 87:4050–4056
Kurtzman CP, Fell JW (1998) The yeasts a taxonomic study, 4th edn. Elsevier, Amsterdam
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Latorre García L, Castillo Agudo L, Polaina J (2007) Taxonomical classification of yeasts isolated from kefir based on the sequence of their ribosomal RNA genes. World J Microbiol Biotechnol 23:785–791
Londero A, Hamet F, De Antoni GL, Garrote GL, Abraham AG (2012) Kefir grains as a starter for whey fermentation at different temperatures: chemical and microbiological characterization. J Dairy Res 79(3):262–271
Maccaferri S, Klinder A, Brigidi P, Cavina P, Costabile A (2012) Potential probiotic kluyveromyces marxianus B0399 modulates the immune response in caco-2 cells and peripheral blood mononuclear cells and impacts the human gut microbiota in an in vitro colonic model system. Appl Environ Microbiol 78(4):956–964
Mainville I, Robert N, Lee B, Farnworth ER (2006) Polyphasic characterization of the lactic acid bacteria in kefir. Syst Appl Microbiol 29:59–68
Martins FS, Nardi RM, Arantes RM, Rosa CA, Neves MJ, Nicoli JR (2005) Screening of yeasts as probiotic based on capacities to colonize the gastrointestinal tract and to protect against enteropathogen challenge in mice. J Gen Appl Microbiol 51:83–92
Moreira MEC, Santos MHD, Zolini GPP, Wouters ATB, Carvalho JCT, Schneedorf JM (2008) Anti-inflammatory and cicatrizing activities of a carbohydrate fraction isolated from sugary kefir. J Med Food 11(2):356–361
Morelli L (2007) In vitro assessment of probiotic bacteria: from survival to functionality. Int Dairy J 17:1278–1283
Noumi E, Snoussi M, Marcilla Diaz A, Bakhrouf A, Valentin E (2012) Numerical analysis of whole-cell and cell wall proteins’ profiles of human oral cavity Candida isolates. Afr J Microbiol Res 6(21):4503–4511
Orberá Ratón T (2004) Molecular identification methods of yeasts of biotechnological interest. Rev Iberoam Micol 21:15–19
Paramithiotis S, Müller MRA, Ehrmann MA, Tsakalidou E, Seiler H, Vogel R, Kalantzopoulos G (2000) Polyphasic identification of wild yeast strains isolated from Greek sourdoughs. Syst Appl Microbiol 23(1):156–164
Pecquet S, Guillaumin D, Tancrede C, Andremont A (1991) Kinetics of saccharomyces cerevisiae elimination from the intestines of human volunteers and effect of this yeast on resistance to microbial colonization in gnotobiotic mice. Appl Environ Microbiol 57(10):3049–3051
Romanin D, Serradell M, González Maciel D, Lausada N, Garrote GL, Rumbo M (2010) Downregulation of intestinal epithelial innate response by probiotic yeasts isolated fron kefir. Int J Food Microbiol 140:102–108
Saad N, Delattre C, Urdaci M, Schmitter JM, Bressollier P (2013) An overview of the last advances in probiotic and prebiotic field. LWT Food Sci Technol 50:1–16
Senses-Ergul S, Agoston R, Belak A, Deak T (2006) Characterization of some yeasts isolated from foods by traditional and molecular tests. Int J Food Microbiol 108:120–124
Shetty PH, Jespersen L (2006) Saccharomyces cerevisiae and lactic acid bacteria as potential mycotoxin decontaminating agents. Trends Food Sci Tech 17(2):48–55
Szajewska H, Skorka A, Dylan M (2007) Meta-analysis: Saccharomyces boulardii for treating acute diarrhoea in children. Alim Pharmacol Therap 25:257–264
Tasteyre A, Barc M, Karjalainen T, Bourlioux P, Collignon A (2002) Inhibition of in vitro cell adherence of Clostridium difficile by Saccharomyces boulardii. Microb Pathogenesis 32(5):219–225
Tiago FCP, Martins FS, Souza ELS, Pimenta PFP, Araujo HRC, Castro IM, Brandão RL, Nicoli JR (2012) Adhesion to the yeast cell surface as a mechanism for trapping pathogenic bacteria by Saccharomyces probiotics. J Med Microbiol 61(PART 9):1194–1207
van der Aa Kühle A, Skovgaard K, Jespersen L (2005) In vitro screening of probiotic properties of Saccharomyces cerevisiae var. boulardii and food-borne Saccharomyces cerevisiae strains. J Food Microbiol 101:29–39
Vancanneyt M, Van Lerberge E, Berny JF, Hennebert GL, Kersters K (1992) The application of whole-cell protein electrophoresis for the classification and identification of basidiomycetous yeast species. Antonie Van Leeuwenhoek 61(1):69–78
Wyder MT (1998) Identification and characterization of the yeast flora in kefir and smear ripened cheese—contribution of selected yeasts to cheese ripening. PhD Thesis, Swiss Federal Institute of Technology, Zurich
Zanello G, Meurens F, Berri M, Salmon H (2009) Saccharomyces boulardii effects on gastrointestinal diseases. Curr Issues Mol Biol 11:47–58
Acknowledgments
G. Diosma is researcher at the Facultad de Ciencias Agrarias y Forestales of Universidad Nacional de La Plata; A. Londero, F. Rey-Burusco, and D. E. Romanin are fellows; and G. L. Garrote is a researcher of the Consejo Nacional de Investigaciones Científicas y Técnicas (CONICET). This work was supported by Agencia Nacional de Promoción Científica y Tecnológica (ANPCyT), CONICET and UNLP. Dr. Donald F. Haggerty, a retired career investigator and native English speaker, edited the final version of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Diosma, G., Romanin, D.E., Rey-Burusco, M.F. et al. Yeasts from kefir grains: isolation, identification, and probiotic characterization. World J Microbiol Biotechnol 30, 43–53 (2014). https://doi.org/10.1007/s11274-013-1419-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11274-013-1419-9