Abstract
Screening of bacteria from Sambhar lake, an extreme hypersaline environment of India, led to the isolation of 93 haloalkaliphilic bacteria growing optimally in media with 2–25 % salt and 6–12 pH. Based on 16S rRNA gene sequences, 93 isolates were further categorized into 32 groups, with each group representing a different taxa belonging to 3 phyla (Firmicutes, Proteobacteria and Actinobacteria). Majority of the isolates (53.12 %) showed similarity with phylum Firmicutes which was followed by Proteobacteria (40.63 %) and Actinobacteria (6.25 %). The isolates belonging to 32 representative groups were further evaluated for the production of extracellular enzymes viz. amylase, cellulase, protease and xylanase, plant growth promoting attributes and BIOLOG™ substrate usage. Among all the isolates, xylanase producing isolates were in maximum (68 %) as compared to protease (56 %), cellulase (40 %), and amylase (37 %) producing strains. Similarly, among plant growth promoting activities, ammonia producing isolates were highest (56 %) when compared to those producing ACC deaminase (53 %), IAA (50 %), hydrogen cyanide (28 %), siderophore (21 %) and solubilizing P (34 %). Isolates showing enzymatic and PGP activities could be further utilized for promoting plant growth in saline affected area.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Soil salinity is one of the most severe environmental factors limiting the productivity of agricultural crops. Evolving efficient, low cost, easily adaptable methods for management of salinity stress is a major challenge (Venkateswarlu and Shanker 2009). Most of the technologies such as development of salinity tolerant varieties, resource management practices, etc. are cost-intensive. Recent studies indicate that microorganisms can help crops to cope with saline stress (Venkateswarlu and Shanker 2009). Halophilic microorganisms present in hypersaline and alkaline aquatic habitats are adapted to high salinity and require a certain concentration of NaCl for their optimum growth. They have been isolated from various saline environment such as low saline marine environment to hypersaline lakes (e.g., the Dead Sea, the Great Salt Lake) and salt mine drainage. These environments are characterized by large amounts of sodium carbonate, or complexes of this salt, formed by evaporative concentration (Tindall et al. 1980). During the past years, special attention has been focused on the study of microbial diversity present in soda lakes (Scholten et al. 2005; Foti et al. 2007). Several workers have reported that soda lakes contain representatives of prokaryotic diversity, and that they can be considered as responsible component for nutrients cycling (Zavarzin et al. 1999; Jones et al. 1998). The well-studied Soda lakes in the world are located in East African Rift Valley (Rees et al. 2004), in the Libyan Desert (Imhoff et al. 1979) and in North America, i.e., Mono Lake (Humayoun et al. 2003; Scholten et al. 2005). There are also data available on soda lakes in India (Wani et al. 2006).
The microorganisms isolated from soda and saline lakes have tremendous potential in agriculture for improved crop production under saline soils. Besides importance in agriculture, the halophilic microbes are also good source of industrially important enzymes such as amylases, proteases, nucleases, phosphatases, lipases, DNases, pullulanases and xylanases (Rohban et al. 2009). With increasing emphasis on extremozymes during the recent years, the microbial enzymes are being looked to replace chemical catalysts in transformations and manufacturing processes.
Most of the studies on halophilic bacteria from salt lakes, however, have so far focused on phylogenetic analysis of the organisms and only limited information is available on their agricultural and enzymatic potential. Identification of culturable microbial communities from saline lakes and their characterization will not only strengthen database of salt loving microbes for basic studies, but their potential can also be utilized in agriculture or industries if they posses plant growth promoting attributes or have ability to produce salt/alkali tolerant industrially important enzymes. In the present study, we characterized culturable bacterial diversity from extreme salt lake through 16S rRNA gene sequencing. The isolates were further screened for plant growth promoting attributes and production of certain important industrial enzymes.
Materials and methods
Sampling site
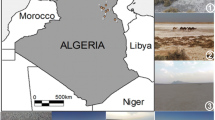
The salterns (Salt beds) present in the Sambhar Salt Lake, Sambhar, Rajasthan, India were selected for the isolation of microorganisms. Sambhar salt lake is the largest inland saline lake of India. It is located in southwest of Jaipur, Rajasthan, India and its catchment lies within the geo coordinates of 26°28′43″–27°33′10″N and 74°35′40″–75°50′28″E, whereas the lake lies within the latitudes of 26°52′–27°02′N and 74°54′–75°14′E (Fig. 1).
Enrichment of samples for isolation of bacteria
Water and sediment samples were collected from four different sampling sites and used for isolation of bacteria following an enrichment strategy. To 10 ml of liquid Horikoshi Medium I with 20 % NaCl and pH 9, 1 g sediment or 1 ml water sample was inoculated and incubated for 7 days at 37 °C. An aliquot of 100 μl from each enriched sample was spread plated on three different media viz. Horikoshi I, Horikoshi II (Joshi et al. 2008) and nutrient agar with modifications and incubated at 37 °C for 48 h. The media were modified by incorporating them with 10 and 20 % NaCl and pH was maintained at 9.0 using 1 N NaOH. Following incubation, the isolates were purified through repeated streaking and all purified isolates were preserved in 35 % glycerol for further utilization.
DNA isolation and PCR
Extraction of total genomic DNA from 93 isolates was carried out using Gen Elute Bacterial Genomic DNA Kit (Sigma, USA). Amplification of 16S rRNA gene was carried out by using universal primers pA (5′-AGTTTGATCCTGGCTAG-3′) and pH (5′-AGGAGGTGATCCAGCCGCA-3′) (Edwards et al. 1989). The amplification conditions were used as described earlier (Sahay et al. 2011).
Sequencing of 16S rRNA gene and phylogenetic analysis
The PCR amplified 16S rRNA genes were purified with a Quiaquick purification kit (Qiagen). The nucleotide sequences were di-deoxy cycle sequenced with fluorescent terminators (Big Dye, Applied Biosystems) and run in 3130xl Applied Biosystems ABI prism automated DNA sequencer. The rRNA gene sequence was checked by sequencing both strands using primers pA and pH for forward and reverse reaction. The partial 16S rRNA gene sequences of the isolated strains were compared with those available in the databases. Identification to the species level was determined as a 16S rRNA gene sequence similarity of ≥97 % with that of a prototype strain sequence in the GenBank. The phylogenetic tree was constructed on the aligned datasets using the neighbour-joining method implemented in the program MEGA 4.0.2 (Tamura et al. 2007). Bootstrap analysis was performed on 1,000 random samples taken from the multiple alignments.
Nucleotide sequence accession numbers
All 16S rRNA gene sequences of the strains (grouped) were deposited in GenBank and accession numbers (JN645842–JN645873) were obtained.
Screening of isolates for tolerance to NaCl and pH
For assessing the tolerance level of NaCl, isolates were inoculated in different wells of micro-titer plates having nutrient broth with different salt concentration (2–30 %) in increment of 1 %. Micro-titer plates were incubated at 37 °C in a temperature-controlled Automated Microbiology Growth Analysis System (Oy Growth Curve Ab Ltd, Finland), where growth in terms of optical density at 600 nm was measured in each well at regular interval. All the observations were taken in triplicate. Similarly, for studying pH tolerance, the isolates were spot inoculated on nutrient agar (NA) medium with pH ranging from 6 to 13, in an increment of pH 1. The pH in the medium was maintained by addition of 50 mM each of phosphate buffer (pH 6.0–8.0), bicarbonate buffer (pH 9.2–10.6) or sodium hydroxide/potassium chloride buffer (pH 12.0–13.0) to the growth medium.
Screening of isolates for extracellular enzyme production
All the thirty two isolates were tested qualitatively for production of four extracellular enzymes. The production of amylase (Manachini et al. 1988), protease (Ramesh et al. 2001), xylanase (Sanghi et al. 2007) and cellulase (Zvereva et al. 2006) by all the isolates at NaCl concentration supporting optimum growth was estimated.
Screening of isolates for plant growth promoting (PGP) attributes
Isolates were also screened for their ability to exhibit plant growth promoting attributes through assessing phosphate solubilization and production of siderophore, ammonia, hydrogen cyanide, ACC-deaminase and indolic compounds at their optimum NaCl tolerance concentration. All the isolates were screened for phosphate solubilisation on Pikovskaya’s agar plates (Pikovskaya 1948). Pikovskaya’s medium containing tri-calcium phosphate was prepared and poured into sterilized petriplate. Isolates of halophilic bacteria were spot inoculated on the plates and incubated at 28 °C for 4–7 days. The plates were observed for clearing or solubilization zones around the colonies. Bacterial isolates were assayed for siderophore production on the Chrome azurol S agar medium as described by Schwyn and Neilands (1987). Chrome azurol S agar plates were spot inoculated with test organism and incubated at 28 °C for 48–72 h. Development of yellow–orange halo around the growth was considered as positive for siderophore production. Bacterial isolates were tested for the production of ammonia in peptone water. Freshly grown cultures were inoculated in 10 ml peptone water in each tube and incubated for at 24–72 h. Nessler’s reagent (0.5 ml) was added in each tube. Development of brown to yellow colour was a positive test for ammonia production (Cappuccino and Sherman 1992). Single isolates were streaked onto KB medium supplemented with 4.4 g glycine L−1′ to screen for cyanide production. Thereafter the Petri dishes were inverted. A piece of filter paper impregnated with 0.5 % picric acid (yellow) and 2.0 % sodium carbonate was placed in the lid of each Petri dish. The Petri dishes were sealed with parafilm and held at 20 °C for 48–96 h. Discoloration of the filter paper to orange-brown after incubation indicates microbial production of cyanide (Bakker and Schippers 1987). ACC (1-Aminocyclopropane-1-carboxylic) deaminase activity was analyzed by using DF Salt minimal medium with ACC as sole source of nitrogen (Penrose and Glick 2003). Indole-3-aceticacid (IAA) production was estimated colorimetrically (Glickmann and Dessaux 1995).
Assessment of bacterial carbon source utilization
Carbon substrate usage by isolates was measured using the GP2 (Gram Positive) and GN2 (Gram negative) MicroPlate™ (Biolog, Inc., Hayward, California, USA), each 96-well microtiterplate has 95 wells that contain a single substrate per well with one water negative control well, for a total of 96 substrates. Sample preparation and analysis was performed following the directions of the manufacturer (Biolog). Each plate was inoculated with a bacterial suspension (150 μl/well) and incubated for 3–4 days at 37 °C. The resulting utilization patterns were read against a substrate blank well at A590 nm with an automated plate reader (Molecular Devices Corporation, USA). Overall color development in BIOLOG plates was expressed as average well color development (AWCD). AWCD of the plate was calculated using the formula (C − R)/{[P(C − R)/95]}, where R is the value of the control well and C is the values of wells containing sole carbon sources.
Results
Isolation of bacteria from enriched samples
The total bacterial count in the sediment samples ranged from 32 to 59 × 105 cfu g−1 whereas in water samples it ranged from 14 to 38 × 105 cfu ml−1. A total of 140 bacteria were isolated from the water and sediment samples of Sambhar lake. All the isolates were screened for salt tolerance at graded concentrations of NaCl (2–30 %) with regular increment of 1 %. Among 140 culturable isolates that exhibit distinct colony characteristics, a total of 93 isolates showing tolerance to ≥5 % NaCl were selected for further study. These isolates were purified and used for phylogenetic analysis.
Sequencing of 16S rRNA gene and phylogenetic analysis
The 16S rRNA gene of all the 93 isolates was amplified and a single amplicon of about 1,545 bp was obtained for each isolate. Based on near complete sequence analysis of 16S rRNA gene sequences and BLAST search, the 93 isolates were identified and categorized into 32 groups. Taxonomic and phylogenetic analysis of these 32 bacteria categorized them into three phyla viz. Firmicutes, Actinobacteria and Proteobacteria (Fig. 2; Table 1). Of the 32 groups, 19 were Gram positive and 13 were Gram negative. Amongst the 19 Gram positive isolates, 17 belonged to the phylum Firmicutes (Low G + C) and 2 belonged to phylum Actinobacteria (High G + C) (Table 1). All the 13 Gram negative isolates belonged to the phylum Proteobacteria with predominance of γ-proteobacteria (12) and only one β-proteobacteria (Alcaligenes). The phylogenetic tree constructed to determine their affiliations are shown in Fig. 2.
Salinity and pH tolerance
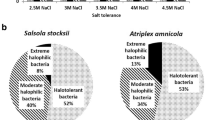
Of the 32 isolates, 4 isolates (SL26, SL125, SL128, and SL139) could tolerate 25 % NaCl, 8 isolates (SL34, SL141, SL10, SL36, SL145, SL121, SL131 and SL35) were able to tolerate maximum of 20–24 % NaCl concentration whereas the remaining 21 isolates could tolerate maximum of 10–19 % NaCl. Similarly, variation in pH tolerance was also observed. Some of the isolates like SL21, SL26, SL131 and SL19 were able to grow in a pH range of 6.0–12.0 (Table 1). Isolates (SL3, SL13, SL15, SL20, SL35, SL37, SL40, SL101, SL107, SL109, SL111, SL125, SL128, SL136, SL139, SL141, and SL145) capable of growing at higher NaCl concentration of >5 % and pH >9 can be grouped as halo-alkali-tolerant strains (Table 2).
Extracellular enzymes and plant growth promotion
The ability to produce four different enzymes was tested among the isolates. Out of 32 isolates 18, 12, 13 and 22 isolates were found to produce amylase, protease, cellulase and xylanase, respectively. Among all the isolates, xylanase producing isolates were highest (68 %) as compared to protease (56 %), cellulase (40 %), and amylase (37 %) producing strains. Four isolates viz., SL10, SL26, SL35 and SL131 showed all four enzyme activities. Highest amylase activity (52.30 IU ml−1 min−1) was presented by isolate SL109, protease activity (58.80 IU ml−1 min−1) by SL23, cellulase activity (52.94 IU ml−1 min−1) by SL136 and xylanase activity (51.83 IU ml−1 min−1) by SL101. Further screening of these isolates for the production of six PGP traits at high salt concentrations revealed that 6 isolates solublized phosphorus, 16 isolates produced IAA, 7 produced siderophore, 12 produced ACC deaminase, 18 produced ammonia, and 9 showed the production of HCN (Table 2). Similarly, among plant growth promoting activities, ammonia producing strains were highest (56 %) when compared to ACC deaminase (53 %), IAA production (50 %), P-solubilization (34 %), hydrogen cyanide (28 %), and siderophore (21 %) producing strain. Isolate SL35 solublized highest amount of phosphorus (981.32 μg PO4 ml−1) and isolate SL109 showed highest IAA production of 570.98 μg mg−1protein. Interestingly, one isolates SL23 exhibited all six PGP activities along with three enzyme activities viz., amylase, protease and xylanase.
Carbon utilization pattern
A total of 32 representative isolates were selected for carbon substrate utilization at 5 % NaCl concentration. The BIOLOG metabolic profiles of thirty two bacteria (19 Gram-positive and 13 Gram-negative) belonging to Firmicutes, Actinomycetes and Proteobacteria based on use/non-use of 95 substrates is shown in Fig. 3a, b. At 52 % similarity, 13 clusters could be made for Gram-positive. Isolate SL17 could utilize highest number of the sugars tested (48 %) whereas SL19 utilized the lowest number of sugars (11 out of 95 sugars). Gram negative bacteria produced 11 clusters at 46 % similarity level and isolate SL121 could utilize highest number of the sugars tested (67/95; 65.26 %) whereas SL125 utilized the lowest number of sugars (12 out of 95 sugars). The variability in utilization of carbon sources by Halomonas strains was very high (67/95) as compared to Firmicutes and Actinobacteria (46/95) and shows elevated carbon utilization unevenness throughout UPGMA dendrogram. The utilization of l-arabinose, d-mannose, pyruvic acid methyl ester, succinic acid mono-methyl-ester, d-glucuronic acid, β-hydroxybutyric acid were different significantly among halomonas strains.
Discussion
The isolation and characterization of halotolerant and halophilic bacteria from hypersaline environment is of practical importance because these bacteria have biotechnological potential with regard to the production of useful biomolecules, such as osmolytes (compatible solutes), hydrolytic enzymes and exopolysaccharides (Margesin and Schinner 2001). In the present investigation, we have isolated 93 bacteria from hypersaline condition of Sambhar Lake, India. Phylogenetic analysis of 16S rRNA gene sequences categorized them into 32 groups belonging to three phyla i.e., Firmicutes, Proteobacteria and Actinobacteria. In terms of diversity, the majority of isolates were from the Bacillus group within the firmicutes (53 %) and gamma-proteobacteria (40 %). Similar results were reported from the haloalkaline lake Elmenteita, Kenya but the abundance of gamma-proteobacteria was higher as compared to firmicutes (Mwirichia et al. 2010). Deshmukh et al. (2011) also reported majority of bacteria isolated from Lonar soda lake of Maharashtra state, India belonging to phylum firmicutes and few to proteobacteria. In another study on the bacterial diversity of Zabuye salt lake in Tibet region using culture independent approach, the phylogenetic tree showed that some clones (57.14 % of total clones) belonged to 23 genera in gamma-proteobacteria, alpha-proteobacteria, delta-proteobacteria, bacteroidetes, firmicutes, and verrucomicrobia and the rest clones were quite different (Zhang and Kong 2010). Several workers have reported the abundance of firmicutes in different lakes such as Wadi An Natrun Mono Lake, CA, USA (Mesbah et al. 2007), Mongolian Baer Soda Lake (Ma et al. 2004), Kenyan Soda Lake (Vargas et al. 2004) and brackish water Pulicat lake (Sahay et al. 2011), Chaka lake Tibetan plateau (Jiang et al. 2007). In the present study, the firmicutes were represented mostly by members of the diverse Bacillus spectrum and included Bacillus, Halobacillus, Thalassobacillus, Virgibacillus, Sediminibacillus, Oceanobacillus and Amphibacillus. Other genera included were Exiguobacterium and Alkalibacterium. In general the abundance of alkaliphilic, endospore forming Gram positive bacteria belonging to the genus Bacillus and other Bacillus derived genera have been reported from haloalkaline lakes (Mwirichia et al. 2010) and brackish water lake (Sahay et al. 2011). Of the 13 members of the proteobacteria, 12 belonged to gamma-proteobacteria and only 1 belonged to beta-proteobacteria (Alcaligenes sp.). Dominance of proteobacteria was also reported earlier by various workers while working on marine sediments solar salterns (Yeon et al. 2005), brackish water (Sahay et al. 2011) and saline-alkaline lakes (Joshi et al. 2008). Amongst the gamma-proteobacteria, the halomonads were most dominant as has been reported from other soda lake (Mwirichia et al. 2010). The other 3 isolates identified as Marinobacter hydrocarbonoclasticus, Salicola marasensis and Nitrincola sp. have also been isolated from saline habitats. The abundance of Halomonas in these habitats is due to their ability to tolerate or require high salt concentration for growth (Jiang et al. 2007).
The variations in the diversity pattern obtained from Sambhar lake and other salt or soda lakes could be due to edaphic or environmental factors; and flora of the region. Most of the permanent vegetation around the Sambhar lake is xerophytic in nature. The main tree species growing in the catchment area are Acacia senegal, A. nilotica, Prosopis cineraria. The main grasses are Cenchrus pennisetiformis, C. cilliaris and Chloris dolichostachya. Another soda lake from India, Lonar lake is surrounded by forest where principal tree species are teak, tamarind and babul.
The distribution and abundance of microbial population to a permanently saline environment includes optimization of basic cell processes, enzyme modulation, nutrient transport and cell membrane function. The bacterial isolates from hypersaline condition also showed the ability to tolerate a wide range of salinity, pH and temperatures, and presented combined hydrolytic activity, which provides the advantage of uses in various industrial processes (Mellado and Ventosa 2003). In the present investigation, xylanase producing isolates were dominant as compared to amylase, cellulase and protease. The observed discrepancy could be attributed to the fact that extremely halophilic/halotolerant and Gram-positive halophilic bacteria produced more extracellular enzymes such as (amylase, protease, cellulase and xylanase (Table 2) in comparison to moderate halophilic and Gram-negative isolates. Similar variations in the production of industrial enzymes were reported among the isolates from Howz Soltan lake, Iran and Pulicat lake, India (Rohban et al. 2009; Sahay et al. 2011).
We have also evaluated halophilic bacteria for six plant growth promoting attributes. In our study, we found that out of thirty two, six isolates were positive for P-solubilization and isolate SL35 showed highest solubilization of phosphorous (981.32 μg PO4 ml−1). Phosphate-solubilizing (PSB) γ-proteobacteria brings about mobilization of insoluble phosphates in the soil and increase plant growth under conditions of poor phosphorus availability (Rodriguez and Fraga 1999). Use of these PSB as bioinoculants will increase the available P in soil, helps to minimize the P-fertilizer application, reduces environmental pollution and promotes sustainable agriculture in future.
It has been reported that indolic compounds (indole-3-acetic acid, indolepyruvic acid, and indoleacetamide) have positive effect on root growth and causing rapid establishment of roots beneficial for young seedlings as it increase their capacity to anchor themselves to the soil and to obtain water and nutrients from their environment (Vessey 2003). We have also tested the ability of isolates to produce IAA. Of the total, 16 (50 %) isolates were tested positive. Similarly, good proportions of isolates (53.12 %) were also found positive for ACC deaminase. Cheng et al. (2007) reported that ACC deaminase bacteria conferred salt tolerance onto plants by lowering the synthesis of salt induced stress and promoted the growth of canola in saline environment. One of the most important strain (Bacillus sp. SL23) which showed three enzyme activity and all six PGP attributes can be exploited biotechnologically and as well as agriculturally. The strain can be used in various industrial areas such as the detergent industry, leather industry (Birbir and Ilgaz 1996) and food preservation (Mellado and Ventosa 2003; Ventosa and Nieto 1995).The enzymes derived from halophiles display optimal activity in the presence of NaCl and maintain stability over a wide pH range (pH 6–10). This makes them excellent additives for laundry detergent, baking industries and for biopolishing of fabrics and production of stonewashed denims (Setati 2010). PGP attributes can be utilized as bioinoculant to improve the production of crop under saline condition.
The Biolog metabolic profiling analysis demonstrated to be highly effective method for differentiating among the species groups by cluster analysis (Dawson et al. 2002). However, this method has also been adapted and used to characterize the functional potential of the bacteria (Garland and Mills 1991). No positive correlations between taxonomic and metabolic diversity were found, which is in accordance with previous study by O’Donnell et al. (1993). They suggested that factors other than community structure, such as soil pH, were more important in regulating metabolic activity. In our study it was observed that high salt and pH tolerating bacteria utilized more carbon as compared to low salt tolerating bacteria. The possible reason behind this may be attributed to the fact that bacteria require a sufficient supply of carbon to feed their metabolic pathways under stress condition.
In saline environment, organisms encounter limited amounts of complex mixtures of carbon sources that are often present at low concentrations. As a result, microbial cells have developed multiple systems to utilize a wide array of different substrates as carbon sources resulting increase in metabolic activity.
Conclusion
The above study has demonstrated that Sambhar lake harbors a wealth of diverse microorganisms, especially high salt tolerant bacteria, having multiple PGP attributes and different enzyme activity. The enzymes are generally haloalkaliphilic which make them suitable to be employed in different industrial processes. These bacteria can also be exploited in developing microbe based formulations to improve the productivity of crop plants grown in saline soils.
References
Bakker A, Schippers B (1987) Microbial cyanide production in the rhizosphere in relation to potato yield reduction and Pseudomonas spp-mediated plant growth stimulation. Soil Biol Biochem 19:451–457
Birbir M, Ilgaz A (1996) Isolation and identification of bacteria adversely affecting hide and leather quality. J Soc Leather Technol Chem 80:147–153
Cappuccino JC, Sherman N (1992) Microbiology: a laboratory manual, 3rd edn. Benjamin/Cumming Pub Co, New York (NH3 test)
Cheng Z, Park E, Glick BR (2007) 1-Aminocyclopropane-1-carboxylate (ACC) deaminase from Pseudomonas putida UW4 facilitates the growth of canola in the presence of salt. Can J Microbiol 53:912–918
Dawson SL, Fry JC, Dancer BN (2002) A comparative evaluation of five typing techniques for determining the diversity of fluorescent Pseudomonads. J Microbiol Meth 50:9–22
Deshmukh KB, Pathak AP, Karuppayil MS (2011) Bacterial diversity of Lonar Soda Lake of India. Ind J Microbiol 51:107–111
Edwards U, Rogall T, Blocker H, Emde M, Bottger EC (1989) Isolation and direct complete nucleotide determination of entire genes. Characterization of a gene coding for 16S ribosomal RNA. Nucleic Acids Res 17:7843–7853
Foti M, Sorokin DY, Lomans B, Mussmann M, Zacharova EE, Pimenov NV, Kuenen JG, Muyzer G (2007) Diversity, activity and abundance of sulfate-reducing bacteria in saline and hypersaline soda lakes. Appl Environ Microbiol 73:2093–2100
Garland JL, Mills AL (1991) Classification and characterization of heterotrophic microbial communities on the basis of patterns of community level sole-carbon-source utilization. Appl Environ Microbiol 57:2351–2359
Glickmann E, Dessaux Y (1995) A critical examination of the specificity of the salkowski reagent for indolic compound produced by phytopathogenic bacteria. Appl Environ Microbiol 61:793–796
Humayoun SB, Bano N, Hollibaugh JT (2003) Depth distribution of microbial diversity in Mono Lake, a meromictic soda lake in California. Appl Environ Microbiol 69:1030–1042
Imhoff JF, Sahl HG, Soliman GSH, Truper HG (1979) The Wadi Natrun, chemical composition and microbial mass developments in alkaline brines of eutrophic desert lakes. Geomicrobiol J 1:219–234
Jiang H, Dong H, Yu B, Liu X, Li Y, Ji S, Zhang CL (2007) Microbial response to salinity change in Lake Chaka, a hypersaline lake on Tibetan plateau. Environ Microbiol 10:2603–2621
Jones BE, Grant WD, Duckworth AW, Owenson GG (1998) Microbial diversity of soda lakes. Extremophiles 2:191–200
Joshi AA, Kanekar PP, Kelkar AS, Shouche YS, Vani AA, Borgave SB, Sarnaik SS (2008) Cultivable bacterial diversity of alkaline Lonar lake. India Microb Ecol 55:163–172
Ma Y, Weizhou Z, Xue Y, Zhon P, Ventosa A, Grant WD (2004) Bacterial diversity of the inner Mongolian Baer Soda lake as revealed by 16S rRNA gene sequence analysis. Extremophiles 8:45–51
Manachini PL, Fortina MG, Parini C (1988) Alkaline protease produced by Bacillus thermoruber—a new species of Bacillus. Appl Microbiol Biotechnol 28:409–413
Margesin R, Schinner F (2001) Potential of halotolerant and halophilic microorganisms for biotechnology. Extremophiles 5:73–83
Mellado ME, Ventosa A (2003) Biotechnological potential of moderately and extremely halophilic microorganisms. In: Barredo JL (ed) Microorganisms for health care, food and enzyme production. Research Signpost, Kerala, pp 233–256
Mesbah NM, Aboum-El-Ela SH, Wiegel J (2007) Novel and unexpected prokaryotic diversity in water and sediments of the alkaline, hypersaline lakes of the Wadi An Natrun, Egypt. Microb Ecol 54:598–617
Mwirichia R, Muigai AW, Tindall B, Boga HI, Stackebrandt E (2010) Isolation and characterisation of bacteria from the haloalkaline Lake Elmenteita, Kenya. Extremophiles 14:339–348
O’Donnell AG, Falconer C, Goodfellow M, Ward AC, Williams E (1993) Biosystematics and diversity amongst novel carboxydotrophic actinomycetes. Antonie Van Leeuwenhoek 64:325–340
Penrose DM, Glick BR (2003) Methods for isolating and characterizing ACC deaminase-containing plant growth-promoting rhizobacteria. Physiol Plant 118:10–15
Pikovskaya RI (1948) Mobilization of phosphorus in soil connection with the vital activity of some microbial species. Microbiologiya 17:362–370
Ramesh B, Reddy PRM, Seenayya G, Reddy G (2001) Effect of various flours on the production of thermostable β-amylase and pullulanase by Clostridium thermosulfurogenes SV2. Bioresour Technol 76:169–171
Rees HC, Grant WD, Jones BE, Heaphy S (2004) Diversity of Kenyan soda lake alkaliphiles assesses by molecular methods. Extremophiles 8:63–71
Rodriguez H, Fraga R (1999) Phosphate solubilizing bacteria and their role in plant growth promotion. Biotechnol Adv 17:319–339
Rohban R, Ali MA, Ventosa A (2009) Screening and isolation of halophilic bacteria producing extracellular hydrolyses from Howz Soltan Lake Iran. J Ind Microbiol Biotechnol 36:333–340
Sahay H, Singh S, Kaushik R, Saxena AK, Arora DK (2011) Characterization of halophilic bacteria from environmental samples of Pulicat brackish water lake, India. Biologia 66:741–747
Sanghi A, Garg N, Sharma J, Kuhar K, Kuhad RC, Gupta VK (2007) Optimization of xylanase production using inexpensive agroresidues by alkaliphilic Bacillus subtilis ASH in solid state fermentation. World J Microbiol Biotechnol 24:633–640
Scholten JCM, Joye SB, Hollibaugh JT, Murrell JC (2005) Molecular analysis of the sulfate-reducing and archaeal community in a meromictic soda lake (mono Lake, California) by targeting 16SrRNA, mcrA, apsA and dsrAB genes. Microb Ecol 50:29–39
Schwyn B, Neilands JB (1987) Universal chemical assay for the detection and determination of siderophores. Anal Biochem 160:47–56
Setati ME (2010) Diversity and industrial potential of hydrolase producing halophilic/halotolerant eubacteria. Afr J Biotechnol 9:1555–1560
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4, molecular evolutionary genetics analysis (MEGA) software version 4.0.2. Mol Biol Evol 24:1596–1599
Tindall BJ, Mills AA, Grant WD (1980) An alkalophilic red halophilic bacterium with low magnesium requirement from Kenyan soda lake. J Gen Microbiol 116:257–260
Vargas VA, Delgado OD, Hatti-Kaul R, Mattiasson B (2004) Lipase producing microorganisms from a Kenyan alkaline soda lake. Biotechnol Lett 26:81–86
Venkateswarlu B, Shanker AK (2009) Climate change and agriculture, adaptation and mitigation strategies. Indian J Agron 54:226–230
Ventosa A, Nieto JJ (1995) Biotechnological applications and potentialities of halophilic microorganisms. World J Microbiol Biotechnol 11:85–94
Vessey JK (2003) Plant growth promoting rhizobacteria as biofertilizers. Plant Soil 255:571–586
Wani AA, Surakasi VP, Siddharth J, Raghavan RG, Patole MS, Ranade D, Shouche YS (2006) Molecular analysis of microbial diversity associated with the Lonar soda lake in India, an impact crater in a basalt area. Res Microbiol 157:928–937
Yeon SH, Jeong WJ, Park JS (2005) The diversity of culturable organotrophic bacteria from local solar salterns. J Microbiol 43:1–10
Zavarzin GA, Zhilina TN, Kevbrin VV (1999) The alkaliphilic microbial community and its functional diversity. Microbiol 63:503–521
Zhang X, Kong F (2010) Bacterial diversity in Zabuye Salt Lake Tibet by culture-independent approaches. Acta Microbiologica Sinica 50:334–341
Zvereva EA, Fedorova TV, Kevbrin VV, Zhilina TN, Rabinovich ML (2006) Cellulase activity of a haloalkaliphilic anaerobic bacterium, strain Z-7026. Extremophiles 10:53–60
Acknowledgments
This work was supported by Indian Council of Agricultural Research (ICAR) Network Project on Application of Microorganisms in Agriculture and Allied Sectors. Authors are thankful to National Bureau of Agriculturally Important Microorganisms (NBAIM) for providing all the facilities.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sahay, H., Mahfooz, S., Singh, A.K. et al. Exploration and characterization of agriculturally and industrially important haloalkaliphilic bacteria from environmental samples of hypersaline Sambhar lake, India. World J Microbiol Biotechnol 28, 3207–3217 (2012). https://doi.org/10.1007/s11274-012-1131-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11274-012-1131-1