Abstract
Capability to produce antilisterial bacteriocins by lactic acid bacteria (LAB) can be explored by the food industry as a tool to increase the safety of foods. Furthermore, probiotic activity of bacteriogenic LAB brings extra advantages to these strains, as they can confer health benefits to the consumer. Beneficial effects depend on the ability of the probiotic strains to maintain viability in the food during shelf-life and to survive the natural defenses of the host and multiply in the gastrointestinal tract (GIT). This study evaluated the probiotic potential of a bacteriocinogenic Lactobacillus plantarum strain (Lb. plantarum ST16Pa) isolated from papaya fruit and studied the effect of encapsulation in alginate on survival in conditions simulating the human GIT. Good growth of Lb. plantarum ST16Pa was recorded in MRS broth with initial pH values between 5.0 and 9.0 and good capability to survive in pH 4.0, 11.0 and 13.0. Lb. plantarum ST16Pa grew well in the presence of oxbile at concentrations ranging from 0.2 to 3.0%. The level of auto-aggregation was 37%, and various degrees of co-aggregation were observed with different strains of Lb. plantarum, Enterococcus spp., Lb. sakei and Listeria, which are important features for probiotic activity. Growth was affected negatively by several medicaments used for human therapy, mainly anti-inflammatory drugs and antibiotics. Adhesion to Caco-2 cells was within the range reported for other probiotic strains, and PCR analysis indicated that the strain harbored the adhesion genes mapA, mub and EF-Tu. Encapsulation in 2, 3 and 4% alginate protected the cells from exposure to 1 or 2% oxbile added to MRS broth. Studies in a model simulating the transit through the GIT indicated that encapsulated cells were protected from the acidic conditions in the stomach but were less resistant when in conditions simulating the duodenum, jejunum, ileum and first section of the colon. To our knowledge, this is the first report on a bacteriocinogenic LAB isolated from papaya that presents application in food biopreservation and may be beneficial to the consumer health due to its potential probiotic characteristics.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lactic acid bacteria (LAB) present many important properties in food manufacturing, such as production of lactic acid, improvement of organoleptic and physical characteristics and reduction of lactose content. In addition, many LAB are used in the food industry, mainly dairy industries, as starter cultures and also as probiotics (Todorov and Dicks 2006; Todorov 2009). Probiotics are defined as live microorganisms which when administered in adequate amounts (106–107 CFU per gram of food) confer health benefits to the host (Bertazzoni-Minelli et al. 2004; FAO/WHO 2001). Probiotic benefits include suppression of growth of pathogens, control of serum cholesterol level, modulation of the immune system, improvement of lactose digestion, synthesis of vitamins, increase in bio-availability of minerals and possible anti-carcinogenic activity (Chan and Zhang 2005; Gomes and Xavier 1999; Kailasapathy and Chin 2000; Shah 2000). Beneficial effects depend on the ability of the probiotic strains to survive the natural defenses of the host and multiply in the gastrointestinal tract (GIT) (Brink et al. 2006; Gilliland 1989; Todorov et al. 2008). Besides these effects, production of antimicrobial compounds such as bacteriocins by probiotic LAB may contribute to colonization of the GIT, providing them competitive advantage over other bacteria (Garriga et al. 1993; Todorov 2009).
Most products currently on the market that contain probiotic bacteria have a very short shelf-life, even when stored at low temperature (0–4°C). Some do not have the required viable cell counts necessary to exert a probiotic effect and thus should not be marketed as probiotic foods (Kailasapathy and Chin 2000). After consumption, the numbers of viable cells lowers significantly due to the acidic environment and bile secretions in the GIT. One method to protect microorganisms from these harsh conditions is their encapsulation in hydrocolloid beads, prepared with alginate, alginate-chitosan or other polymeric compounds (Krasaekoopt et al. 2003). Several studies have shown that microencapsulation of probiotic strains in alginate improved their survival in foods and protected them during passage through the human GIT (Chandramouli et al. 2004; Jankowski et al. 1997; Kailasapathy 2002; Liserre et al. 2007; Mandal et al. 2006; Mollendorff 2008; Sheu and Marshall 1991, 1993; Sheu et al. 1993; Suita-Cruz and Goulet 2001).
This study evaluated the probiotic potential of a bacteriocinogenic Lactobacillus plantarum strain (Lb. plantarum ST16Pa) isolated from papaya fruit (Todorov et al. 2011) and studied the effect of encapsulation of this strain in alginate on survival in conditions simulating the human GIT. These features are important when selecting a starter culture for food biopreservation that also presents potential beneficial properties to the consumer health.
Materials and methods
Cultures
The study was conducted with Lactobacillus plantarum ST16Pa, a bacteriocinogenic strain isolated from papaya fruit (Todorov et al. 2011). Other microbial cultures used in the study were Lb. rhamnosus GG, Lb. plantarum ST202Ch, Lb. plantarum ST69BZ, Enterococcus faecium ST62BZ, Enterococcus faecalis ATCC 19443, Lb. sakei subsp. sakei 2a, Listeria ivanovii subsp. ivanovii ATCC 19119 and Listeria innocua 2030C. Pure cultures were stored at −20°C with 20% glycerol.
Evaluation of the probiotic potential
Effect of pH and oxbile on growth of Lb. plantarum ST16Pa
This experiment was performed according to Todorov et al. (2008). Lb. plantarum ST16Pa was grown in MRS broth (Difco, USA) adjusted to pH 3.0, 4.0, 5.0, 6.0, 7.0, 9.0, 11.0 and 13.0. Resistance to oxbile salts was tested by cultivating the strain in MRS broth containing 0.2, 0.4, 0.6, 0.8, 1.0, 2.0 and 3.0% (w/v) oxbile (Sigma–Aldrich). All tests were conducted in sterile flat-bottom 96-well micro titer plates (TPP, Switzerland). Each well was filled with 180 μl of tested MRS broth and 20 μl of the culture with OD650nm adjusted to 0.2. Optical density at 600 nm were recorded every hour for 12 h. Cultures grown in MRS broth at pH 7.0 and without oxbile served as controls. Experiments were done in triplicate.
Adhesion of Lb. plantarum ST16Pa to Caco-2 cells
Caco-2 cells were grown in wells of flat-bottom 96-well microtiter plates (TPP, Switzerland) containing 100 μl of Eagle Minimal Essential Medium (MEM) (Merck) supplemented with 10% (v/v) fetal bovine serum (Sigma–Aldrich), 100 U/ml penicillin (Sigma–Aldrich) and 100 U/ml streptomycin (Sigma–Aldrich) for 24 h at 37°C in humidified atmosphere containing 5% CO2 (Todorov et al. 2008). Lb. plantarum ST16Pa was grown in MRS for 24 h at 37°C. The wells containing the Caco-2 cells were inoculated with 100 μl of the bacterial suspension (ca. 1 × 106 CFU/ml) and after 2 h at 37°C the plates centrifuged at 300×g for 2 min at 4°C. The wells were washed twice with 1 ml sterile PBS and the Caco-2 cells lysed by adding 1 ml 0.5% (v/v) Triton X-100. Appropriate decimal dilutions were then plated onto MRS agar. The percentage of adhesion of Lb. plantarum ST16Pa to Caco-2 cells was calculated based on the initial number of viable cells, number of cells in the supernatant and number of cells in the lysate after treatment with Triton X-100. Experiments were done in triplicate. Lb. rhamnosus GG was used as control of adhesion to Caco-2 cells.
Confirmation of the presence of genes encoding MapA and Mub adhesion proteins and EF-Tu elongation factor in Lb. plantarum ST16Pa
DNA from Lb. plantarum ST16Pa was isolated according to Dellaglio et al. (1973). Primers Mub423F (5′-GTA GTT ACT CAG TGA CGA TCA ATG-3′), Mub423R (5′-TAA TTG TAA AGG TAT AAT CGG AGG-3′), Map423F (5′-TGG ATT CTG CTT GAG GTA AG -3′), Map423R (5′-GAC TAG TAA TAA CGC GAC CG-3′), EFTu423F (5′-TTC TGG TCG TAT CGA TCG TG-3′) and EFTu423R (5′-CCA CGT AAT AAC GCA CCA AC-3′) were used for amplification of genes MapA, Mub and EF-Tu by PCR, as described by Todorov and Dicks (2008).
Determination of cell surface hydrophobicity in Lb. plantarum ST16Pa
Cell surface hydrophobicity was measured as described by Doyle and Rosenberg (1995), and modified by Todorov et al. (2008). In summary, Lb. plantarum ST16Pa was grown in MRS broth at 37°C for 18 h. Cells were harvested (6,700×g, 4°C, 6 min), washed twice with sterile saline solution (pH 6.5), re-suspended in the same solution and the optical density (OD580nm) determined. The cell suspension (1.5 ml) was mixed with equal volume of n-hexadecane (Sigma–Aldrich) and vortexed for 2 min. After 30 min at room temperature for separation of the two phases, 1 ml of the aqueous phase was removed and the optical density (OD580nm) determined. The experiment was repeated and the average optical density value determined. The percentage hydrophobicity was calculated as follows: % hydrophobicity = [(OD0−OD30)/OD0] × 100, where OD0 and OD30 refer to the initial OD and OD measured after 30 min, respectively. Experiments were conducted in triplicate.
Effect of commercial drugs and antibiotics on growth of Lb. plantarum ST16Pa
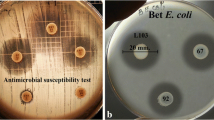
The experiments were performed as described by Carvalho et al. (2009). Commercial drugs listed in Table 1 were purchased from a local drugstore, and solubilized in sterile water to achieve the desired concentration. Lb. plantarum ST16Pa was inoculated into 10 ml MRS broth and incubated at 37°C for 18 h and mixed into MRS soft agar (1.0%, w/v) to achieve a population of 106 CFU/ml. After solidification of the agar, 10 μl of each medicament was spotted onto the surface of the plates, and incubated at 37°C for 24 h. The plates were examined for the presence of inhibition zones around the spotted medicament, and those presenting inhibition zones larger than 2 mm diameter were subjected to the determination of the minimal inhibition concentration (MIC). For this test, serial two-fold dilutions of each medicament were prepared in sterile water and 10 μl spotted onto the surface of MRS soft agar plates, previously inoculated with Lb. plantarum ST16Pa (106 CFU/ml). The plates were incubated at 37°C for 24 h and examined for the presence of inhibition zones around the spots. The MIC corresponded to the highest dilution that resulted in inhibition halos of at least 2 mm diameter.
In a similar experimental approach, the susceptibility of Lb. plantarum ST16Pa to antibiotic disks listed in Table 2 (CEFAR, Sao Paulo, SP, Brazil) was tested. The inhibitory effect of the antibiotics was expressed in millimeters of the inhibition zones (Carvalho et al. 2009).
Aggregation properties of Lb. plantarum ST16Pa
The experiment was performed according to Basson et al. (2008) with modifications (Todorov et al. 2008). Lb. plantarum ST16Pa was grown in MRS broth for 24 h at 37°C, harvested (7,000×g, 10 min, 20°C), washed, re-suspended in sterile saline solution (pH 6.5) and diluted to OD660nm = 0.3. One ml of the cell suspension was transferred to a 2 ml sterile plastic cuvette and the OD660nm recorded over 60 min by using a spectrophotometer. After 60 min the cell suspension was centrifuged (300×g, 2 min, 20°C) and the OD60 of the supernatant determined.
Auto-aggregation was determined using the following equation: % Auto-aggregation = [(OD0−OD60)/OD0] × 100, where OD0 and OD60 refer to the initial OD and OD measured after 60 min, respectively.
For evaluation of co-aggregation of Lb. plantarum ST16Pa with Lb. plantarum ST202Ch, Lb. plantarum ST69BZ, E. faecium ST62BZ, E. faecalis ATCC 19443, Lb. sakei subsp. sakei 2a, L. ivanovii subsp. ivanovii ATCC 19119 and L. innocua 2030C, the lactobacilli were grown in MRS broth for 24 h at 37°C and the other cultures were prepared in BHI for 24 h at 37°C. The experimental protocol for study of co-aggregation was the same used for auto-aggregation, except that one milliliter of Lb. plantarum ST16Pa and one milliliter of the co-aggregation partner were coupled.
Effect of encapsulation on survival of Lb. plantarum ST16Pa in conditions simulating the gastrointestinal tract
Encapsulation of Lb. plantarum ST16Pa
The encapsulation method of Sheu et al. (1993) was used. Lb. plantarum ST16Pa was grown in 10 ml MRS broth to an optical density (OD600nm) of 3.0 (approximately 1 × 109 CFU/ml) and harvested (2,000×g, 10 min, 4°C). The cells were washed twice with 10 ml sterile peptone water (0.1%, w/v) and resuspended in 5 ml of the same solution. For encapsulation, the Lb. plantarum ST16Pa cell suspension was mixed with 25 ml of sterile 2, 3 and 4% sodium alginate solutions prepared in distilled water. After vortexed to homogeneity, the mixtures were transferred to sterile syringes fitted with a 0.45 mm diameter needle and slowly ejected and allowed to drop into sterile 100 ml 0.05 M CaCl2 supplemented with 0.1% (v/v) Tween 80. After 30 min for stabilization, the beads were harvested (350×g, 10 min, 4°C), washed three times with sterile peptone water and collected by filtration through sterile Grade 1 Whatman filter paper. The average diameter of the alginate beads was 1 mm. The beads were stored in sterile peptone water at 4°C for a maximum of 2 days.
Effect of pH and oxbile on survival of encapsulated Lb. plantarum ST16Pa
One gram of alginate beads containing entrapped cells of Lb. plantarum ST16Pa was transferred to 10 ml of 0.08 M HCl supplemented with 0.2% (w/v) NaCl (pH 1.6), gently mixed and incubated at 37°C in an orbital shaker (60 rpm). After 1 and 3 h of incubation, the beads were harvested by filtration through sterile Grade 1 Whatman filter paper and mixed with 10 ml phosphate buffer (0.1 M KH2PO4, pH 7.4) for depolymerization and release of cells. The cell suspension was submitted to ten-fold dilutions in sterile distilled water and plated onto MRS agar and incubated at 37°C for 48 h. The experiment was repeated by transferring one gram of alginate beads to 10 ml of 0.1 M KH2PO4 (pH 7.4). Samples were taken after 30 min, 1, 2 and 3 h and viable counts determined as described before. In both experiments, viable cell numbers were also determined immediately after encapsulation. The same experiments were carried out with non-encapsulated Lb. plantarum ST16Pa. The experiments were performed in triplicate.
The effect of oxbile on survival of encapsulated Lb. plantarum ST16Pa was measured adding one gram of the beads to 10 ml 1 and 2% (w/v) oxbile (Oxoid). After 0, 3, 6 and 12 h of incubation at 37°C, the beads were depolymerized and the number of viable cells determined by plating onto MRS agar, as described before. The experiments were also carried out with non-encapsulated Lb. plantarum ST16Pa, adding free cells to the oxbile solutions. The experiments were performed in triplicate.
Release of encapsulated Lb. plantarum ST16Pa at conditions simulating different sections of the gastrointestinal tract
Ten grams of the alginate beads containing Lb. plantarum ST16Pa were transferred to 250 ml of a sterile solution simulating the stomach condition (12.5 g MRS, 0.78 g NaCl, 0.28 g KCl, 0.03 g CaCl2 and 0.15 g NaHCO3 per liter, pH adjusted to 4.0 with 1 M HCl), prepared according to Mollendorff (2008). After 2.5 h at 37°C, the pH of the solution was adjusted to 3.0 with sterile 1 M HCl and the number of viable Lb. plantarum ST16Pa determined as described before. The beads were then removed from this solution, washed twice with sterile 0.1% (m/v) peptone water and transferred to a mixture prepared with 250 ml of the stomach simulating solution and 125 ml of sterile pancreatic solution [1.5 g NaHCO3, 0.11 g Pancreatin (Sigma–Aldrich), 0.08 g Oxgall (Oxoid)], prepared according to Mollendorff (2008). The pH of the mixture was adjusted to 6.5 with sterile 1 N NaOH. After 4 h at this pH, simulating the conditions in the duodenum and ileum, the pH was reduced to 6.0 with sterile 1 N HCl and kept at this value for 2.5 h to simulate conditions in the jejunum and first section of the colon. All incubations were performed at 37°C. Samples (1 ml) were withdrawn after 2 h (simulating the duodenum), 4 h (simulating the jejunum), 6 h (simulating the ileum) and 6.5 h (simulating the first section of the colon), and submitted to counts of Lb. plantarum ST16Pa. The percentage of cell release from the beads and the percentage of free cells survival were calculated as described before. The experiment was repeated with non-encapsulated cells of Lb. plantarum ST16Pa to serve as control. All experiments were performed in triplicates.
Results and discussion
A previous study indicated that Lb. plantarum ST16Pa produces a large spectrum bacteriocin capable of inhibiting many food spoilage bacteria and foodborne pathogens, including Listeria monocytogenes, a psychrotrophic foodborne pathogen of increasing importance (Todorov et al. 2011). Further research evaluating potential probiotic characteristics of this strain is important as this additional beneficial property would lead to a broader application of the strain in foods, both for improvement of quality and safety of food products and for conferring beneficial health effects to the consumer.
Effect of pH and oxbile on growth of Lb. plantarum ST16Pa
Capability to grow and survive at low pH and high concentrations of oxbile salts, as those encountered in the human GIT, is a major characteristic of a strain with potential probiotic activity (Vinderola and Reinheimer 2003). Results reported in this study indicate that Lb. plantarum ST16Pa grew well in MRS broth with initial pH values of 5.0, 6.0, 7.0 and 9.0 (Fig. 1A), however, at pH 4.0 the growth rate was much lower. Previous studies indicated that pH affects the growth of bacteriocinogenic Lactobacillus spp. in different degrees. Growth of several strains of Lb. plantarum, Lb. rhamnosus, Lb. pentosus and Lb. paracasei was suppressed at pH 3.0 and 4.0, and variable results were recorded for pH of 11.0 and 13.0, but poor growth recorded for strains Lb. paracasei ST242BZ and ST284BZ, Lb. rhamnosus ST462BZ, Lb. plantarum ST664BZ and Lb. pentosus ST712BZ at pH 13.0 (Todorov et al. 2008).
Lactobacillus plantarum ST16Pa grew well in the presence of oxbile at concentrations ranging from 0.2 to 3.0% (Fig. 1B), suggesting that this strain would be able to survive in the human GIT. Bile salts have different effects in different Lactobacillus strains. Growth of strains Lb. plantarum ST194BZ, ST414BZ and ST664BZ, Lb. rhamnosus ST462BZ, and Lb. pentosus ST712BZ were less affected by the presence of 0.3% bile, compared to strains Lb. paracasei ST242BZ and ST284BZ and Lb. rhamnosus ST461BZ (Todorov et al. 2008). Bile at 0.6% (w/v) suppressed the growth of Lb. plantarum ST194BZ and ST441BZ, Lb. paracasei ST242BZ and ST284BZ and Lb. rhamnosus ST462BZ, but not Lb. rhamosus ST461BZ, Lb. plantarum ST664BZ and Lb. pentosus ST712BZ. Bile levels above 0.6% (w/v) suppressed the growth of all tested strains (Lb. plantarum ST194BZ, ST441BZ and ST664BZ, Lb. paracasei ST242BZ and ST284BZ, Lb. rhamnosus ST461BZ and ST462BZ and Lb. pentosus ST712BZ) (Todorov et al. 2008).
Similar effects of oxbile and pH on Lb. plantarum 423, Lb. salivarius 241 and Lb. curvatus DF38 were reported by Brink et al. (2006). In another study, Haller et al. (2001), reported that as many as 10% of Lb. plantarum cells, but less than 0.001% of Lb. sakei and Lb. paracasei cells, survived when exposed to HCl (pH 2.0) and bile salts.
Adhesion of Lb. plantarum ST16Pa to Caco-2 cells
Adhesion of probiotic strains to intestinal cells such as Caco-2 cells is believed to be a critical factor to increase the possibility of colonizing the GIT and survive in this hostile environment (Tuomola and Salminen 1998). Lb. plantarum ST16Pa adhered to Caco-2 cells at a rate similar to that recorded for the well-known probiotic reference strain Lb. rhamnosus GG (9.5 and 11.3%, respectively) (data not shown). These values are similar to those reported by Todorov et al. (2008) for several bacteriocins producing LAB isolated from boza (0.26–9.0%) and by Tuomola and Salminen (1998) for some probiotic and dairy Lactobacillus strains (3.2–14.4%). Bertazzoni-Minelli et al. (2004) reported much lower adhesion rates for probiotic Lactobacillus casei strains (0.08–0.74%).
Identification of genes encoding MapA and Mub adhesion proteins and EF-Tu elongation factor in Lb. plantarum ST16Pa
The expression of mucus adhesion proteins, such as those encoded by the mub and mapA genes, and of GTP-binding EF-Tu protein has been shown to be critical in adhesion of probiotic strains to human intestine cells (Ramiah et al. 2008). Results of PCR amplification using specific primers and sequence analysis of the PCR amplicons indicated that Lb. plantarum ST16Pa contains the mub, mapA and EF-Tu genes (Fig. 2). Most probably presence of these genes is species dependent and not related with the ecological origin of the strain/s. However, same genes were recorded in other Lb. plantarum strains isolated from plants (Ramiah et al., 2008). The presence of these adhesion genes in this Lb. plantarum strain is not surprising due to the high adhesive properties of this microbial species. Whole genome sequence data of Lb. plantarum WCFS1, a human isolate, revealed the presence 223 extracellular proteins, most of them involved in the adhesion of the cell to its environment (Kleerebezem et al. 2003). The domain composition of Lb. plantarum WCFS1 proteins predicted to be associated with adhesion indicated the presence of seven proteins to be involved with mucus. Some of these possible mucus binding proteins contain the Mub domain, a trait unique to LAB (Boekhorst et al. 2006). The presence of these adhesive proteins in lactic acid bacteria is imperative for probiotic strains to colonize the GIT.
Agarose gels showing amplified fragments for genes MapA (lanes 1, 2 and 3), Mub (lanes 4, 5 and 6) and EF-Tu (lanes 7, 8 and 9) amplified by PCR. Lanes 2, 5 and 8 correspond to Lactobacillus plantarum ST16Pa and lanes 1, 4 and 7 to Enterococcus faecium ST88Ch. Lanes 3, 6 and 9 contain no DNA and lane M corresponds to 100 bp molecular weight marker
Determination of cell surface hydrophobicity in Lb. plantarum ST16Pa
Bacterial cells with a high hydrophobicity usually form strong interactions with mucosal cells. The interactions between microbial and host cells are non-specific, but in probiotic strains, there is a good correlation between surface hydrophobicity and the ability to adhere to the intestinal mucosa (Wadström et al. 1987). The initial interaction may be weak, is often reversible and precedes subsequent adhesion processes mediated by more specific mechanisms involving cell-surface proteins and lipoteichoic acids (Granato et al. 1999; Rojas et al. 2002; Roos and Jonsson 2002). Hydrophobicity varies among even genetically closely related species and even among strains of the same species (Schar-Zammaretti and Ubbink 2003). Although hydrophobicity may assist in adhesion, it is not a prerequisite for strong adhesion to human intestinal cells.
The hydrophobicity recorded for Lb. plantarum ST16Pa was higher than that recorded for the reference probiotic Lb. rhamnosus GG strain (68.7 and 53.3%, respectively). Similar results were observed by Todorov et al. (2008), for strains Lb. rhamnosus ST461BZ, Lb. rhamnosus ST462BZ and Lb. plantarum ST664BZ (75–80%) and Lb. rhamnosus GG (55%).
Effect of commercial drugs and antibiotics on growth of Lb. plantarum ST16Pa
Probiotic consumers may be under treatment for a variety of illnesses, and the beneficial effects of the probiotic strain may be hampered by possible interactions with the medicaments used by these consumers. As shown in Table 1, there were several drugs that reduced the growth of Lb. plantarum ST16Pa, especially non-steroidal anti-inflammatory drugs that contain potassium diclofenac or ibuprofen arginine, promethazine hydrochloride (antihistaminic), orphenadrine citrate and sodium metamizole (analgesic), paroxetine (antidepressant) and amiodarone (antiarrhrythmic). The negative effect of sodium diclofenac on growth of potential probiotic strains was also observed in other studies (Botes et al. 2008; Carvalho et al. 2009; Todorov et al. 2007, 2008; Todorov and Dicks 2008). Dimenhydrinate inhibited the growth of Lb. rhamnosus ST462BZ and Lb. plantarum ST664BZ (Todorov et al. 2008).
The majority of the antibiotics tested in this study inhibited the growth of Lb. plantarum ST16Pa (Table 2). The strain was resistant to amicacin, tobramicin, vancomicin, oxacilin, kanamicin, metronidazol and nalidixic acid. Resistance of potential probiotic LAB to antibiotics is a controversial subject, as these strains may be reservoirs of antibiotic resistance genes, and can be transferred horizontally to other in the human GIT (Dicks et al. 2009). Resistance may be inherent to a bacterial genus or species, but may also be acquired through exchange of genetic material, mutations and the incorporation of new genes (Ammor et al. 2007; Levy and Marshall 2004; Salyers et al. 2004; Teuber 1999).
It should be emphasized that the interaction between medicaments or antibiotics and probiotic bacteria in the GIT depends on their concentration in this environment (Todorov et al. 2007, 2008), so that the Minimal Inhibitory Concentration (MIC) values play an important for the proper evaluation of these interactions. As an example, spidufen reduced the growth of Lb. plantarum ST16Pa in MRS, but the MIC for this drug was as high as 15.0 mg/ml (Table 1), indicating that the recommended daily dose (600 mg) will hardly affect the survival of Lb. plantarum ST16Pa in the GIT. More important are the drugs for treatment of chronic diseases, such as atlansil, an anti-arrhytmic drug normally used in long course treatments and arotin, a drug from the group of the anti-depressants with neuroleptic effect, also used in prolonged treatments, which presented MIC of 1.25 and 1.0 mg/ml, respectively. Due to their long-term application, they may accumulate in the gastrointestinal tract and affect the viability of Lb. plantarum ST16Pa.
Aggregation properties of Lb. plantarum ST16Pa
Aggregation is an important feature for biofilm formation. Lb. plantarum has a number of genes encoding for surface proteins responsible for recognition of or binding to components present in the environment. Several of these genes are homologous to proteins with predicted functions, such as mucus-binding, aggregation-promotion and intracellular adhesion (Kleerebezem et al. 2003).
Aggregation properties need to be discussed taking in account the bacteriocin spectrum of activity. If the bacteriocin produced by Lb. plantarum ST16Pa does not inhibit the co-aggregation partner, the two strains can co-exist and facilitate biofilm formation. In opposite, if the bacteriocin is active against the co-aggregation partner, high co-aggregation will facilitate the bactericidal mode of action and easier elimination of the co-aggregation partner from the system.
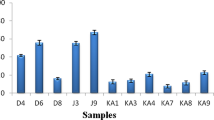
Auto-aggregation is a strain-specific trait (Todorov and Dicks 2008). Lb. plantarum ST16Pa was determined to have an auto-aggregation level of 37.05% (Fig. 3), which is lower than the levels reported by Todorov et al. (2008) for other lactobacilli: Lb. pentosus ST712BZ (67%) and Lb. paracasei ST284BZ (99%). The co-aggregation of Lb. plantarum ST16Pa with Lb. plantarum ST202Ch, Lb. plantarum ST69BZ, E. faecium ST62BZ, E. faecalis ATCC 19443, Lb. sakei subsp. sakei 2a, L. ivanovii subsp. ivanovii ATCC 19119 and L. innocua 2030C also varied according to the strain (Fig. 3), but the lowest levels were observed for Listeria and Enterococcus strains. Low levels of co-aggregation may play an important role in preventing the formation of biofilms, and in this way preventing the persistence of pathogenic species in the GIT.
Auto-aggregation and co-aggregation of Lactobacillus plantarum ST16Pa with Lactobacillus plantarum ST202Ch, Lactobacillus plantarum ST69BZ, Enterococcus faecium ST62BZ, Enterococcus faecalis ATCC 19443, Lactobacillus sakei subsp. sakei 2a, Listeria ivanovii subsp. ivanovii ATCC 19119 and Listeria innocua 2030c. Each result represents an average of three experiments
Effect of pH and oxbile on survival of encapsulated Lb. plantarum ST16Pa
Encapsulation of Lb. plantarum ST16Pa in alginate protected the cells from the acidic pH, regardless the concentration of alginate used in the encapsulation process. The counts of Lb. plantarum ST16Pa after exposure to simulated gastric conditions (pH 1.6) for 3 h were 1.2 log lower than the initial count, while for non-encapsulated cell a five-log reduction was observed (data not shown).
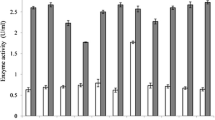
The effects of oxbile on the survival of encapsulated and free Lb. plantarum ST16Pa are shown in Figs. 4a–c and 5, respectively. In the absence of oxbile, survival of encapsulated (Fig. 4a) and non-encapsulated Lb. plantarum ST16Pa (Fig. 5) in MRS up to 12 h was similar, regardless the percentage of alginate used for encapsulation. However, exposure to 1 and 2% oxbile for 12 h caused less than two log reduction in the counts of Lb. plantarum ST16Pa when encapsulated in 2, 3 or 4% alginate (Fig. 4b, c), while the number of non-encapsulated cells decreased five logs (Fig. 5). There was not a direct correlation between the concentration of alginate in the encapsulation process and the number of viable cells after exposure to 1 and 2% oxbile. Similar results were reported by Mollendorff (2008) in encapsulation of two strains of Lb. plantarum. However, Mandal et al. (2006) observed an increase in viability of Lb. casei NCDC-298 with the increase in alginate concentration, similarly to Lee and Heo (2000), who reported a decrease in the death rate of cells of Bifidobacterium longum with the increase in alginate concentration. Strain specific interactions between bacterial species and alginate or variations in the encapsulation methodology could explain these differing results.
a Survival of encapsulated L. plantarum ST16Pa in MRS with no added oxbile (encapsulation in 2% alginate—light grey bars, in 3% alginate—dark grey bars and in 4% alginate—black bars). b Survival of encapsulated L. plantarum ST16Pa in MRS containing 1% oxbile (encapsulation in 2% alginate—light grey bars, in 3% alginate—dark grey bars and in 4% alginate—black bars). c Survival of encapsulated L. plantarum ST16Pa in MRS containing 2% oxbile (encapsulation in 2% alginate—light grey bars, in 3% alginate—dark grey bars and in 4% alginate—black bars)
Release of encapsulated Lb. plantarum ST16Pa at conditions simulating different sections of the gastrointestinal tract
Counts of free and encapsulated Lb. plantarum ST16Pa after exposure to conditions simulating the transit through the GIT are shown in Fig. 6. After 2.5 h in the solution simulating the stomach (pH 3.0), no counts of Lb. plantarum ST16Pa were obtained when they were encapsulated in alginate, indicating that the cells remained inside the capsules and not affected by the low pH. In counterpart, the counts of free cells after the same treatment were 2 log lower than the initial counts. When transferred from the solution simulating the stomach due to low pH to the new stomach solution added of pancreatic components (digestive enzymes and bile salts), the protection given by the encapsulation weakened along exposure to this new environment as the number of released cells increased gradually with time. However, the number of viable cells obtained for non-encapsulated Lb. plantarum ST16Pa was always lower than that recorded for their encapsulated counterparts. This result indicates that protection given by encapsulation is more evident in conditions simulating the stomach than in conditions simulating the duodenum, jejunum, ileum and the first part of the large intestine. Similar results were reported for whey protein-based microencapsulated cells of bifidobacteria when tested at simulated intestinal conditions (Picot and Lacroix 2004). In contrast, Trindade and Grosso (2000) reported a decrease in viability of immobilized Bifidobacterium bifidum and Lb. acidophilus after exposure to 2% (w/v) and 4% (w/v) bile salts.
Based on the results reported in this study, Lb. plantarum ST16Pa presents high potential of application by the food industry as probiotic since it not only produces a bacteriocin, but also possesses many characteristics responsible for survival and possible reproduction in the harsh conditions of the gastrointestinal tract. Studies run with Lb. plantarum ST16Pa encapsulated in alginate indicate that encapsulation can be used to provide extra protection to the cells. Further studies on survival of the encapsulated bacteria in foods and on the delivery of viable cells to the consumer are necessary to elucidate the complimentary probiotic activity of this strain. To our knowledge, this is the first report on a bacteriocinogenic LAB isolated from fruit that presents application in food biopreservation and may be beneficial to the consumer health due to its potential probiotic characteristics.
References
Ammor MS, Flóres AB, Mayo B (2007) Antibiotic resistance in not-enterococcal lactic acid bacteria and bifidobacteria. Food Microbiol 24:559–570
Basson A, Flemming LA, Chenia HY (2008) Evaluation of adherence, hydrophobicity, aggregation characteristics and biofilm development of Flavobacterium johnsoniae-like isolates. Microb Ecol 55:1–14
Bertazzoni-Minelli E, Benini A, Marzotto M, Sbarbati A, Ruzzenente O, Ferrario R, Hendriks H, Dellaglio F (2004) Assessment of novel probiotic Lactobacillus casei strains for the production of functional foods. Int Dairy J 14:723–736
Boekhorst J, Wells M, Kleerebezem M, Siezen RJ (2006) The predicted secretome of Lactobacillus plantarum WCFS1 sheds light on interactions with its environment. Microbiology 152:3175–3183
Botes M, Van Reenen CA, Dicks LMT (2008) Evaluation of Enterococcus mundtii ST4SA and Lactobacillus plantarum 423 as probiotics by using a gastro-intestinal model with infant milk formulations as substrate. Int J Food Microbiol 128:362–370
Brink M, Todorov SD, Martin JH, Senekal M, Dicks LMT (2006) The effect of prebiotics on production of antimicrobial compounds, resistance to growth at low pH and in the presence of bile, and adhesion of probiotic cells to intestinal mucus. J Appl Microbiol 100:813–820
Carvalho KG, Kruger MF, Furtado DN, Todorov SD, Franco BDGM (2009) Evaluation of the role of environmental factors in the human gastrointestinal tract on the behaviour of probiotic cultures Lactobacillus casei Shirota and Lactobacillus casei LC01 by the use of a semi-dynamic in vitro model. Ann Microbiol 59:439–445
Chan ES, Zhang Z (2005) Bioencapsulation by compression coating of probiotic bacteria for their protection in an acidic medium. Process Biochem 40:3346–3351
Chandramouli V, Kailasapathy K, Peiris P, Jones M (2004) An improved method of microencapsulation and its evaluation to protect Lactobacillus spp. in simulated gastric conditions. J Microbiol Meth 56:27–35
Dellaglio F, Bottazzi V, Troatelli LD (1973) Deoxyribonucleic acid homology and vase composition in some thermophylic lactobacilli. J Gen Microbiol 74:289–297
Dicks LMT, Todorov SD, Franco BDGM (2009) Current status of antibiotic resistance in lactic acid bacteria. In: Bonilla AR, Muniz KP (eds) Antibiotic resistance: causes and risk factors, mechanisms and alternatives. Pharmacology—research, safety testing and regulation. Nova Publisher, New York, pp 379–425
Doyle RJ, Rosenberg M (1995) Measurement of microbial adhesion to hydrophobic substrates. Meth Enzymol 253:542–550
FAO/WHO (2001) Evaluation of health and nutritional properties of powder milk and live lactic acid bacteria. In: Food and agriculture organization of the United Nations and World Health Organization expert consultation report. FAO, Rome
Garriga M, Hugas M, Aymerich T, Monfort JM (1993) Bacteriocinogenic activity of Lactobacillli from fermented susages. J Appl Bacteriol 75:142–148
Gilliland SE (1989) Acidophilus milk products: A review of potential benefits to consumers. J Dairy Sci 72:2483–2494
Gomes AMP, Xavier MF (1999) Bifidobacterium spp. and Lactobacillus acidophilus: biological, biochemical, technological and therapeutical properties relevant for use as probiotics. Trends Food Sci Technol 10:139–157
Granato D, Perotti F, Masserey I, Rouvet M, Golliard M, Servin A, Brassart D (1999) Cell surface-associated lipoteichoic acid acts as an adhesion factor for attachment of Lactobacillus johnsonii La1 to human enterocyte-like Caco-2 cells. Appl Environ Microbiol 65:1071–1077
Haller D, Colbus H, Ganzle MG, Scherenbacher P, Bode C, Hammes WP (2001) Metabolic and functional properties of lactic acid bacteria in the gastro-intestinal ecosystem: A comparative in vitro study between bacteria of intestinal and fermented food origin. Syst Appl Microbiol 24:218–226
Jankowski T, Zielinska M, Wysakowska A (1997) Encapsulation of lactic acid bacteria with alginate/starch capsules. Biotechnol Techn 11:31–34
Kailasapathy K (2002) Microencapsulation of probiotic bacteria: technology and potential applications. Curr Issues Intest Microbiol 3:39–48
Kailasapathy K, Chin J (2000) Survival and therapeutic potential of probiotic organisms with reference to Lactobacillus acidophilus and Bifidobacteria spp. Immunol Cell Biol 78:80–88
Kleerebezem M, Boekhorst J, van Kranenburg R, Molenaar D, Kuipers OP, Leer R, Tarchini R, Peters SA, Sandbrink HM, Fiers MWEJ, Stiekema W, Lankhorst RMK, Bron PA, Hoffer SM, Groot MNN, Kerkhoven R, de Vries M, Ursing B, de Vos WM, Siezen RJ (2003) Competitive genome sequence of Lactobacillus plantarum WCFS1. Proc Nat Acad Sci USA 100:1990–1995
Krasaekoopt W, Bhandari B, Deeth H (2003) Evaluation of encapsulation techniques of probiotics for yoghurt. Int Dairy J 13:3–13
Lee KY, Heo TR (2000) Survival of Bifidobacterium longum immobilized in calcium alginate beads in simulated gastric juices and bile salt solution. Appl Environ Microbiol 66:869–873
Levy SB, Marshall B (2004) Antibacterial resistance worldwide: causes, challenges and responses. Nat Med Rev 10:S122–S129
Liserre AM, Re MI, Franco BDGM (2007) Microencapsulation of Bifidobacterium animalis subsp. lactis in modified alginate-chitosan beads and evaluation of survival in simulated gastrointestinal conditions. Food Biotechnol 21:1–16
Mandal S, Puniya AK, Singh K (2006) Effect of alginate concentrations on survival of microencapsulated Lactobacillus casei NCDC-298. Int Dairy J 16:1190–1195
Mollendorff JW (2008) Characterization of bacteriocins produced by lactic acid bacteria from fermented beverages and optimization of starter cultures. McS Thesis, Stellenbosch University, Stellenbopsch, South Africa
Picot A, Lacroix C (2004) Encapsulation of bifidobacteria in whey protein-based microcapsules and survival in simulated gastrointestinal conditions and in yoghurt. Int Dairy J 14:505–515
Ramiah K, Van Reenen CA, Dicks LMT (2008) Surface-bound proteins of Lactobacillus plantarum 423 that contribute to adhesion of Caco-2 cells and their role in competitive exclusion and displacement of Clostridium sporogenes and Enterococcus faecalis EF-Tu protein binds to GTP. Res Microbiol 159:470–475
Rojas M, Ascencio F, Conway PL (2002) Purification and characterization of a surface protein from Lactobacillus fermentum 104R that binds to porcine small intestinal mucus and gastric mucin. Appl Environ Microbiol 68:2330–2336
Roos S, Jonsson H (2002) A high-molecular-mass cell-surface protein from Lactobacillus reuteri 1063 adheres to mucus components. Microbiology 148:2481–2489
Salyers AA, Gupta A, Wang Y (2004) Human intestinal bacteria as reservoirs for antibiotic resistance genes. Trends Microbiol 12:412–416
Schar-Zammaretti P, Ubbink J (2003) The cell wall of lactic acid bacteria: surface constituents and macromolecular conformations. Biophys J 85:4076–4092
Shah NP (2000) Probiotic bacteria: selective enumeration and survival in dairy foods. J Dairy Sci 83:894–910
Sheu TY, Marshall RT (1991) Improving culture viability in frozen dairy desserts by microencapsulation. J Dairy Sci 74:107–112
Sheu TY, Marshall RT (1993) Microencapsulation of lactobacilli in calcium alginate gels. J Food Sci 58:557–561
Sheu TY, Marshall RT, Heymann H (1993) Improving survival of culture bacteria in frozen desserts by microentrapment. J Dairy Sci 76:1902–1907
Suita-Cruz P, Goulet J (2001) Improving probiotic survival rates. Food Technol 55:36–42
Teuber M (1999) Spread of antibiotic resistance with food-borne pathogens. Cell Mol Life Sci 56:755–763
Todorov SD (2009) Bacteriocins from Lactobacillus plantarum–production, genetic organization and mode of action. A review. Braz J Microbiol 40:209–221
Todorov SD, Dicks LMT (2006) Screening for bacteriocin producer lactic acid bacteria from boza, a traditional cereal beverage from Bulgaria. Characterization of produced bacteriocins. Process Biochem 41:11–19
Todorov SD, Dicks LMT (2008) Evaluation of lactic acid bacteria from kefir, molasses and olive brine as possible probiotics based on physiological properties. Ann Microbiol 58:661–670
Todorov SD, Botes M, Danova ST, Dicks LMT (2007) Probiotic properties of Lactococcus lactis subsp. lactis HV219, isolated from human vaginal secretions. J Appl Microbiol 103:629–639
Todorov SD, Botes M, Guigas C, Schillinger U, Wiid I, Wachsman MB, Holzapfel WH, Dicks LMT (2008) Boza, a natural source of probiotic lactic acid bacteria. J Appl Microbiol 104:465–477
Todorov SD, Prévost H, Lebois M, Dousset J, LeBlanc JG, Franco BDGM (2011) Bacteriocinogenic Lactobacillus plantarum ST16Pa isolated from papaya (Carica papaya)—from isolation to application: Characterization of a bacteriocin. Food Res Int 44:1351–1363
Trindade CSF, Grosso CRF (2000) The effect of the immobilization of Lactobacillus acidophilus and Bifidobacterium lactis in alginate on their tolerance to gastrointestinal secretions. Milchwissenschaft 55:496–499
Tuomola EM, Salminen S (1998) Adhesion of some probiotic and dairy Lactobacillus strains to Caco-2 cell cultures. Int J Food Microbiol 41:45–51
Vinderola CG, Reinheimer JA (2003) Lactic acid starter and probiotic bacteria: a comparative in vitro study of probiotic characteristics and biological barrier resistance. Food Res Int 36:895–904
Wadström T, Andersson K, Sydow M, Axelsson L, Lindgren S, Gullmar B (1987) Surface properties of lactobacilli isolated from the small intestine of pigs. J Appl Bacteriol 62:513–520
Acknowledgments
The authors would like to thank Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES, Brasil), Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq, Brasil), and Consejo Nacional de Investigaciones Científicas y Técnicas (CONICET, Argentina) for financial support.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Todorov, S.D., LeBlanc, J.G. & Franco, B.D.G.M. Evaluation of the probiotic potential and effect of encapsulation on survival for Lactobacillus plantarum ST16Pa isolated from papaya. World J Microbiol Biotechnol 28, 973–984 (2012). https://doi.org/10.1007/s11274-011-0895-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11274-011-0895-z