Abstract
Chlorothalonil (2, 4, 5, 6-tetrachloroisophthalonitrile, TPN) has been widely used as a wide-spectrum fungicide in China and other countries, and is considered to be an important soil and water contaminant. Here we report the isolation and characterization of a novel TPN-degrading bacterial strain TP-D1 from a heavily TPN-polluted soil in Henan Province, China, and identified it as a strain of Ochrobactrum lupini based on 16S rRNA gene sequence analysis and its morphological, biochemical, and physiological characteristics. Strain TP-D1 could degrade 90.4 and 99.7% of TPN after 4- and 7-day incubation in mineral salt broth with 50 mg TPN l−1 and in autoclaved soil with 50 μg TPN g−1, respectively. Two new metabolites, methyl 2, 5, 6-trichloro-3-cyano-4-methoxy-benzoate (metabolite C) and methyl 3-cyano-2, 4, 5, 6-tetrachlorobenzoate (metabolite D), were detected besides previously reported 4-hydroxy-2, 5, 6-trichloroisophthalonitrile (TPN-OH, metabolite A). This result suggests that the cyano-group in TPN could be converted into amide groups by strain TP-D1, and reveal the biodegradation mechanism of TPN in soil.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Chlorothalonil (2, 4, 5, 6-tetrachloroisophthalonitrile, TPN) is a widely used wide-spectrum fungicide. It controls plant fungal diseases by reacting with cellular thiols and inhibiting fungal respiration (Godard et al. 1999). According to the US Geological Survey of 1997 and Chlorothalonil-Pesticide Use Statistics of 2008 (http://pesticideinfo.org/Detail_ChemUse.jsp?Rec_Id=PC34550), TPN use was common in the United States; 4.8 × 106 kg TPN was produced each year and applied to more than 50 crops. In China, TPN was usually used to control greenhouse vegetable diseases, and diseases of fruits, rice and wheat. Its production reached 8 × 106 kg each year (New pesticide 2002), but was far from the demand of 2006 and 2007 (Sun 2008). Fungicide residue has been reported in vegetables and fruits (Wang et al. 2005; Xiao et al. 2007), soil and water (Cox 1997), groundwater (Winkler et al. 1996) and in locations as remote as the surface microlayer and fog of the Chukchi Arctic ecosystem (Chernyak et al. 1996). The half-life of TPN in soil is 1–2 months (US Environmental Protection Agency 1986). It may remain in soil for 100 days (Takagi et al. 1991), or one year after repeated application (Motonaga et al. 1996). Though TPN has no acute toxicity, it is still harmful to non-target organisms including soil microbes (Sigler and Turco 2002; Suyama et al. 1993) and terrestrial and aquatic vertebrates (Cox 1997), and has been classified as a “probable human carcinogen” by US EPA (Cox 1997).
TPN can be degraded chemically and microbiologically (Roberts and Hutson 1999). Till now, about 11 kinds of TPN metabolites have been detected (Chaves et al. 2008). In soil and water, the main metabolites were 4-hydroxy-2, 5, 6-trichloroisophthalonitrile (TPN-OH), 1-carbamoyl-3-cyano-4-hydroxy-2, 5, 6-trichlorobenzene (CCHT), and 1, 3-dicarbamoyl-2, 4, 5, 6-tetrachlorobenzene (DTTC) (Roberts and Hutson 1999; Rouchaud and Roucourt 1988), and they were considered to be converted from dechlorination (Sato and Tanaka 1987) and oxidation/hydration of the cyano (–CN) groups in TPN (Putnam et al. 2003; Roberts and Hutson 1999; Rouchaud and Roucourt 1988). Among them, TPN-OH is the most toxic, around thirty times more toxic than its parent compound TPN (Cox 1997; Kenneth and Siegel 1981), more persistent in soil by binding to soil particles (Motonaga et al. 1998), and more readily dissolved in water to lead to second pollution. Therefore, it is important to clarify the biodegradation pathway of TPN to develope new bioremediation strategies for TPN-pollution, especially by a culturable microorganism.
It has been reported that a carbon supplement was necessary for in vitro TPN degradation by bacteria (Katayama et al. 1991; Sato and Tanaka 1987). However, Flavobacterium NL0-1 (Katayama et al. 1991) and bacterium TB I (Motonaga et al. 1996) have been reported to degrade TPN without nutrient supplement, and two new strains, Ochrobactrum sp. CTN-11 (Liang et al. 2010) and Lysobacter ruishenii CTN-1 (Wang et al. 2010a), also showed the same capacity. TPN metabolites degraded by these bacteria were identified to be TPN-OH (Liang et al. 2010; Motonaga et al. 1996) and methylthiotrichloroisophthalonitrile (Katayama et al. 1997). These two products were considered to be derivatives of TPN with replacement of the chlorine atom at the 4-position (Katayama et al. 1992, 1997). However, detection of CCHT and DTTC in soil (Putnam et al. 2003; Roberts and Hutson 1999; Rouchaud and Roucourt 1988) suggested the possibility of other metabolic pathways to degrade TPN by pure culture. Herein, we report the isolation of a bacterium that could effectively degrade TPN without any nutrient supplement and the identification of two new metabolites resulted from the –cyano group conversion of TPN.
Materials and methods
Chemicals and media
TPN powder (effective component >95.0%) was provided by Ruize Pesticide Company, Henan, China. Standard chlorothalonil (>99.6%) was purchased from National Research Center of China for Certified Reference Materials (Beijing, China). Nutrient agar (NA) (Fang 1996) and mineral salt (MS) agar (1.5 g K2HPO4, 1.0 g NH4NO3, 0.5 g MgSO4·7H2O, 0.5 g (NH4)2SO4, 0.5 g KH2PO4, and 15 g agar per liter water) were used to isolate TPN-degrading bacteria from soil.
Isolation of TPN-degrading bacteria
Soil sample was collected from a field close to a chlorothalonil-producing factory in Henan, China. One gram of soil sample was placed into liquid NA medium supplemented with 50 mg TPN l−1 and incubated at 28°C for 4 days with constant agitation of 180 rev min−1. Five milliliters of the culture was subjected to subculture in the same fresh medium with TPN of 50–600 mg l−1 in 50 mg l−1 increments. One hundred microliters of suspension from each subculture was spread on NA plates containing different concentrations of TPN and incubated at 28°C for 3–7 days. Colonies growing vigorously or producing a clear zone on NA plates were purified and stored at 4°C on NA plate or slant containing 50 mg TPN l−1. In order to identify their modes of utilization of TPN, these isolates were cultured on MS plates (pH 7.0) containing 50 mg TPN l−1. Each strain had triplicate plates. The strain showing the highest TPN-degrading ability, denoted TP-D1, was selected for further testing.
Identification of strain TP-D1
Strain TP-D1 was characterized based on morphological features and physiological and biochemical properties after incubation on NA plates for 48 h according to Bergey’s Manual of Determinative Bacteriology (Holt et al. 1994). Molecular identification of strain TP-D1 was performed as described below. Strain TP-D1 was incubated in Luria-Bertani (LB) medium at 30°C for 24 h, and cells were collected by centrifugation and washed twice with sterile distilled water. Total DNA was extracted following the standard procedure (Sambrook and Russell 1998). Oligonucleotide primers 27F (5′-AGAGTTTGATCCTGGTCAG-3′) and 1492R (5′-TACGGCTACCTTGTTACGACT-3′) (Mincer et al. 2002; Zeng et al. 2008) were employed to amplify the 16S rRNA gene region. PCR reactions were carried out with a thermocycler (PTC-200; MJ Research, Waltham, MA, USA) under the following conditions: 5 min at 95°C, 1 min at 94°C, 31 cycles of 40 s at 95°C, 45 s at 54°C, 2 min at 72°C and one final step of 10 min at 72°C. The PCR product was purified on 1% agarose gel using Gel Extraction Kit (TIANGEN, Beijing, China), ligated into pMD18-T vector (TaKaRa, Ostu, Japan), and transformed into Escherichia coli DH5α competent cells. Positive transformants were screened on LB plates containing 80 mg X-Gal ml−1, 0.5 mM IPTG and 50 mg ampicillin l−1, and sequenced. ClustalX (version 1.8) (Thompson et al. 1997) was used to align the 16S rRNA gene sequence of strain TP-D1 with that of reference species from the GenBank database. A phylogenetic tree was constructed using Mega 4.0 (Tamura et al. 2007).
Degradation of TPN in MS broth
Cells of strain TP-D1 were collected from NA plates after incubation at 30°C for 1–2 days and suspended in 10 ml MS broth (pH 6.8). The density of the bacterial suspension was adjusted to approximately 1 × 109 c.f.u. ml−1. Bacterial suspension was diluted 100 times into 50 ml MS broth containing 50 mg TPN l−1, and incubated at 30°C and 180 rev min−1 for 4 days. To determine the degrading ability of strain TP-D1 in MS broth, samples were collected at days 1, 2, 3 and 4, respectively, and the amount of TPN was determined by gas chromatography (GC) with an electron capture detector (ECD). Fifty milliliters of MS broth was poured into a 250-ml flask to evaporate acetone and water using a Rotavapor R-215 (Buchi, Postfach, Switzerland) under reduced pressure. The residue was dissolved in 25 ml m-xylene, and 1 ml of the solution was diluted with acetone to an appropriate concentration for GC analysis. Column DB-5 (30 m × 0.25 mm × 0.25 μm) was used, and the operation conditions were as follows: the oven was heated up to 100°C and maintained for 2 min, then increased to 210°C at a rate of 20°C min−1; the temperature of injection port and detector port were maintained at 240 and 250°C, respectively; nitrogen carrier gas flow rate was 29 ml min−1; and injection volume was 1.5 μl. The recovery rate was 90.7–108.8%. Bacteria-free MS broth containing TPN was used as blank control and the experiment was carried out in triplicate. The degrading ability of strain TP-D1 was determined as the percentage of TPN degraded.
Degradation of TPN in soil
The degrading ability of strain TP-D1 in soil was also determined. Dry natural soil was passed through a 60-micron sieve and autoclaved at 160°C for 1.5 h. TPN solution was sprayed onto 10 g of the soil in a Petri dish to the final concentration of 50 μg g−1 dry soil. A bacterial suspension of strain TP-D1 was mixed with the TPN-containing soil at 107 c.f.u. g−1 dry soil. Plates were incubated at 28°C in darkness for 7 days, and soil moisture was maintained at 20% by adding distilled water when needed. The amounts of TPN in soil samples were monitored at day 0, 1, 3, and 7, respectively. Each treatment had triplicates, and bacteria-free soil with TPN was used as control.
Extraction and analysis of TPN in soil was conducted following the protocol of Motonaga et al. (1996) with some modifications. The concentration of NaCl and the volume of hexane were 5% and 100-ml, respectively. The residue of hexane evaporation was dissolved in 10 ml 100% methanol and sonicated for 3 min. Two milliliters of the extracts were analysed by HPLC under the following conditions: column, C18 reversed phase Shim-Pack CLC-ODS (150 × 4.6 mm); mobile phase, acetonitrile and 0.1% acetic acid buffer at 65:35 (vol/vol); flow rate, 1 ml min−1; injection volume, 50 μl; room temperature for column performance; and u.v. wavelength at 232 nm. Recovery rate of TPN from the soil was from 83.5 to 101.2%.
Strain growth and metabolite production in modified MS broth
In order to avoid the possible utilization of acetone as carbon by strain TP-D1, TPN was pre-dissolved in acetone at 50 mg l−1, and the acetone in the flasks was evaporated before adding the modified MS broth. The modified MS broth containing 1.5 g K2HPO4, and 0.5 g KH2PO4 per liter water (pH 6.8) was used to avoid the influence of chloride anion and other inorganic ions in MS broth. The bacterial suspension of strain TP-D1 mentioned above (1 × 109 c.f.u. ml−1) was diluted 100 times into 50 ml modified MS broth and incubated at 30°C and 180 rev min−1 for 3 days. Samples were taken at 0, 12, 24, 48, 56, 72 h of incubation, and the amount of cells were counted as c.f.u. ml−1. TPN-free modified MS broth was used as blank control and the experiment was carried out in triplicates.
To detect the metabolites, TPN was pre-dissolved in acetone, bacterial suspension of 107–108 c.f.u. ml−1 was incubated in the modified MS broth at 30°C and 180 rev min−1 in darkness for 6 days, and samples were taken at day 0, 2, 4 and 6 for TPN metabolite analysis. Each sample had triplicates. After methanol extraction, an aliquot of 10 ml sample was extracted using solid-phase extraction disks (OASIS® HLB; Waters Corporation, Milford, MA, USA) after pH adjustment as described previously (Chaves et al. 2008). A total of 1 ml extract was used for analysis by liquid chromatography-electrospray ionization mass spectrometry (LC-ESI-MS) and liquid chromatography-atmospheric pressure chemical ionization mass spectrometry (LC-APCI-MS). HPLC-MS (2695-ZQ4000; Waters) equipped with a u.v. detector (2487; Waters) and electrospray ion source was used to detect the TPN metabolites under the following conditions. LC-MS conditions were: column, Xterra C18 (2.1 mm × 100 mm × 3.5 μm); column temperature, 35°C; u.v. detector, 230 nm; mobile phase, acetonitrile (added 0.1% formic acid) 10% (0–0.5 min)–95% (0.5–12 min); and flow rate, 0.3 ml min−1. MS conditions were: mass scanning range (m/z), 80–600; mass detector, a quadrupole mass spectrometer; source temperature, 100°C; cone voltage, 30 V; capillary voltage, 3,000 V; desolvation gas flow rate, 500 l h−1; and desolvation temperature, 300°C.
Results
Isolation of TPN-degrading bacteria
Two strains had ability to degrade TPN on NA plates. Growth tests on MS plates indicated that TPN could be degraded without other nutrient supplement. Strain TP-D1 showing higher TPN-degrading ability (with a clear zone of around 2 mm-in-diameter on MS plates) was purified and used for further testing.
Identification of strain TP-D1
Morphological and biochemical tests indicated that the cells of strain TP-D1 were rod-shaped, 1.6 × 0.75 μm in dimension, motile with one or two polar flagella, non-spore-forming, aerobic, gram-negative, and non-fluorescent. Colonies on NA plate were circular, smooth with regular margins, milk-white and mucilaginous with 2.0 mm in diameter after 3 days of incubation. The isolate was positive to oxidase and catalase reactions. It can utilize arginine, ornithine, lysine, citrate, urea, sucrose, fructose, xylose, mannitol, inositol and glucose. It was negative to esculin hydrolysis, the Kovacs Indole, ONPG, H2S, Voges-Proskauer, methyl red, and nitrate reduction tests.
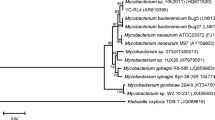
The full-length 16S rRNA gene sequence of strain TP-D1 (1,404 bp, GenBank accession no. EF998851) was compared with the related sequences in the GenBank, and its phylogenetic tree was constructed. Strain TP-D1 exhibited high sequence similarity with bacteria belonging to the genus Ochrobactrum (Fig. 1). Similarity calculations indicated that strain TP-D1 was closely related to O. lupini (99.8%), O. anthropi (99.8%), and O. tritici (98.9%).
Phylogenetic tree based on 16S rRNA gene sequences showing the relationship between strain TP-D1 and representative species of the genus Ochrobactrum and related genera. This tree was constructed with the neighbor-joining method using MEGA 4.0. GenBank accession numbers are shown in parentheses. The significance of each branch is indicated by a bootstrap value for 1,000 subsets. Bar, 2 nt substitutions per 100 nt
Comparison of the representative physiological and biochemical characteristics of strain TP-D1 with other close relatives (Table 1) indicated that strain TP-D1 was more similar to O. lupini except for the contrary reaction of esculin. Thus strain TP-D1 was identified as a strain of O. lupini.
Degrading ability of strain TP-D1 in MS broth and soil
TPN in MS broth was detected by GC and showed a peak at 9.3 min. Calculation based on peak areas showed that the degrading efficacy increased when incubation was prolonged and reached 90.4% after 4-day incubation (Fig. 2a). The degrading efficiency of strain TP-D1 in autoclaved soil was determined by HPLC. TPN was degraded of 95.0% after 3-day incubation and 99.7% after 7 days (Fig. 2b).
Strain growth and metabolite production in modified MS broth
The population of strain TP-D1 in modified MS broth increased gradually from 0 to 60 h, but decreased at 72 h (Fig. 3).
Four TPN metabolites were detected by HPLC after a 2-day incubation. Among them, peak A had a retention time at 6.74 min (Fig. 4a, a-1) and was detected by LC-ESI-MS at negative mode (Fig. 4a, a-3). Peak B, C and D were detected at 7.75, 8.12 and 8.55 min (Fig. 4a, a-1), respectively, at positive mode (Fig. 4a, a-2). The m/z of each metabolite was listed in Table 2. During the degradation process, the range of metabolites varied a lot. At day 0, only TPN (Peak E) was detected at negative mode. At day 2, four metabolites (peak A, B, C, and D) were detected. At day 6, only peak A was detected either by LC-ESI-MS or by LC-APCI-MS. In the sample of blank control only TPN was detected.
HPLC, ion spectra and mass spectra of TPN metabolites degraded by strain TP-D1 in MS broth after incubation at 30°C for 2 days. Metabolites were extracted with methanol (V/V = 1:1), followed by SPE procedure and LC-ESI-MS analysis. a HPLC (a−1) and ion spectra of TPN and its metabolites detected with LC-ESI-MS (a−2, a−3). b Mass spectra of metabolites in (a)
The metabolite represented by peak A was the major degradation product of TPN. The mass spectra of three chlorine compounds at m/z 244.5, m/z 248.6, and m/z 250.6 (Fig. 4b, b-1) were identical to TPN-OH based on the analysis of LC-ESI-MS and LC-APCI-MS at negative mode. Due to the easy substitution of the para-chlorine of TPN, metabolite A might also be TPN-OH. Peak B represented a minor metabolite (Fig. 4b, b-2). From its isotope clusters for four chlorine atoms, peak B should be a cyano derivative of TPN. In combination with its m/z, it was deduced to be 3-methoxycarbonyl-2, 4, 5, 6-tetrachlorobenzeneacetamide. Metabolite C was deduced to be methyl 2, 5, 6-trichloro-3-cyano-4-methoxy-benzoate from its isotope clusters of three chlorine atoms and its m/z (Fig. 4b, b-3). Metabolite D was assumed to be converted from hydration of the cyano- group of TPN, and might be methyl 3-cyano-2, 4, 5, 6-tetrachlorobenzoate (Fig. 4b, b-4). And metabolites C and D might be esterified from CCHT and CTB in the presence of formic acid during the analysis procedure.
In conclusion, the metabolites of TPN degraded by strain TP-D1 in modified MS broth might be TPN-OH, methyl 2, 5, 6-trichloro-3-cyano-4-methoxy-benzoate, and methyl 3-cyano-2, 4, 5, 6-tetrachlorobenzoate. It was proposed that TPN was degraded by strain TP-D1 via direct hydroxylation of its chloro- group (Pathway I), hydration of its cyano- group (Pathway II), or both (Pathway III) (Fig. 5).
Discussion
Bacterial degradation of TPN has been reported previously (Katayama et al. 1991; Sato and Tanaka 1987). TPN metabolites, DTTC and CCHT that were probably bio-converted from the oxidation of the cyano-group of TPN, have been detected in soil, suggesting the presence of some bacteria responsible for the oxidation process (Roberts and Hutson 1999; Rouchaud and Roucourt 1988). Herein we report on the isolation and characterization of a bacterial strain, denoted TP-D1, which can degade TPN without carbon supplement and produce two new metabolites besides TPN-OH. Compared with the reported TPN-degrading bacteria, strain TP-D1 had similar TPN-degrading ability with Ochrobactrum sp. CTN-11 (Liang et al. 2010), but stronger than that of Flavobacterium NL0-1 (Katayama et al. 1991) and TB I (Motonaga et al. 1996).
TPN-OH (Motonaga et al. 1996; Liang et al. 2010) and methylthiotrichloroisophthalonitrile (Katayama et al. 1997) have been reported to be TPN metabolites. In this study, we detected two hitherto unreported metabolites, methyl 2, 5, 6-trichloro-3-cyano-4-methoxy-benzoate (metabolite C) and methyl 3-cyano-2, 4, 5, 6-tetrachlorobenzoate (metabolite D). Metabolite B was not identified due to the limited information about the conversion of –CONH2 group in TPN into –CH2CONH2 group. The –COOCH3 group in metabolites B, C, and D might be esterified in the presence of formic acid during the analysis procedure instead of during the extraction process because the reaction conditions we were improper. During the detection procedure, we set up a blank control to avoid TPN conversion by chemicals during the extraction and analysis procedure, but no other metabolite except for TPN was detected (Table 2). It indicated that metabolites C and D derived from TPN degradation. We assume that metabolites C and D come from CCHT (Rouchaud and Roucourt 1988) and 3-cyano-2,4,5,6-tetrachlorobenzamide (CTB) (Szalkowski and Stallard 1977), respectively. CTB has been detected as a chemical hydrolysis product of TPN at pH 9.0 (Szalkowski and Stallard 1977); however, it is a former product of DTTC. DTTC and CCHT have been recovered from soil under field conditions, and are thought to be the biodegradation products under mild conditions (Rouchaud and Roucourt 1988). Although we did not detect CCHT and CTB in this study, detection of its downstream metabolites C and D suggested that TPN could be degraded by a specific bacterium through conversion of cyano- group of TPN, and CTB could be degraded microbiologically. Considering the presence of metabolite C, we proposed another degradation pathway besides the well-known TPN degradation pathways I and II (Putnam et al. 2003; Regitano et al. 2001; Roberts and Hutson 1999) (Fig. 5). In pathway II and III, the conversion of –cyano group of TPN was an important step, but no report has described the –cyano group breaking or substitution by degrading enzymes. Hitherto, only chlorine substitution of TPN by a hydrolytic dehalogenase (Wang et al. 2010b) and glutathione S-transferase (Kim et al. 2004) has been described. Isolation of strain TP-D1 and establishing its degradation pathway benefit our understanding of the TPN degradation mechanism.
Strain TP-D1 and strain TB I (Motonaga et al. 1996) could degrade TPN and grow slightly in MS broth. The reason might be that the medium was not purified enough and contained some carbon sources. However, their degrading metabolites were different, strain TB I only degraded TPN into TPN-OH. This difference might be ascribed to the detection method besides their different degrading ability. In this study, we used SPE, LC-APCI-MS and LC-ESI-MS techniques, which facilitate the analysis of amide degradation products (Chaves et al. 2008). During the whole degradation process, metabolites B, C and D could be detected at day 2, but were absent at day 6, suggesting that these metabolites were intermediates. However, no other metabolites were detected, probably due to the unsuitable detection method or the low concentrations of these metabolites. In this study, we also detected TPN-OH as a major metabolite (Table 2), and its presence might kill the cells and inhibit the growth of strain TP-D1 (Fig. 3) (Motonaga et al. 1996), consequently limiting the continuous conversion of cyano- group of TPN. Therefore, strain TP-D1 is not a safe bacterium to be used directly for bioremediation of TPN-polluted soil.
Strain TP-D1 was classified to be O. lupini according to its 16S rRNA gene sequence and biochemical and physiological characteristics. Several Ochrobactrum strains have been reported to have the ability to degrade pesticide and organic chemicals, including TPN (Liang et al. 2010a), triazophos (Dai et al. 2005), parathionmethyl (Bai et al. 2004; Qiu et al. 2006) and aniline (Wei et al. 1998). However, even the same bacterial strain has different degrading ability to chemicals with similar groups. For example, Ochrobactrum strain mp_4 could degrade triazophos, methylparathion, phoxim, parathion and malathion, but not methamidophos (Dai et al. 2005). In order to know the degrading specificity of strain TP-D1, we tested it against several chemicals including pentachloronitrobenzene (PCNB), hexachlorocyclohexane (BHC), dichloro-diphenyl-trichloroethane (DDT), triazophos, parathionmethyl and aniline on MS agar plates at a concentration of 5 mg l−1, and no bacterial growth or clear zone around the colonies was detected (unpublished data). This indicated that strain TP-D1 could not metabolize the tested chemicals as sole carbon source, or the ability of strain TP-D1 to degrade these chemicals was too weak to be observed. Further research on the degradation specificity of strain TP-D1 should be conducted in liquid broth.
TPN could be degraded by a hydrolytic dehalogenase, but the enzyme is specific. For example, the hydrolytic dehalogenase isolated from Pseudomonas sp. CTN-3 had no capacity to degrade some chloroaromatics including PCNB (Wang et al. 2010b). Further research on the degrading enzymes of strain TP-D1 should be conducted. Genes coding for these enzymes will be cloned, and their functions will be verified in future.
Conclusions
A novel TPN-degrading strain, O. lupini TP-D1, was isolated from heavily TPN-polluted soil. Strain TP-D1 had a strong ability to degrade TPN without nutrient supplement, and produced two new metabolites besides TPN-OH. We propose that strain TP-D1 could degrade TPN via another new metabolic pathway related to the conversion of –cyano group in TPN. Because TPN-OH was still the main metabolite of TPN degraded by strain TP-D1, our future objectives are to isolate other strains that are devoid of TPN-OH production, or to engineer strain TP-D1 not to produce TPN-OH.
Abbreviations
- BHC:
-
Hexachlorcyclonhexane
- CCHT:
-
1-Carbamoyl-3-cyano-4-hydroxy-2, 5, 6-trichlorobenzene
- c.f.u.:
-
Colony forming units
- CTB:
-
3-Cyano-2, 4, 5, 6-tetrachlorobenzamide
- DDT:
-
Dichloro-diphenyl-trichloroethane
- DTTC:
-
1, 3-Dicarbamoyl-2, 4, 5, 6-tetrachlorobenzene
- ECD:
-
Electron capture detector
- GC:
-
Gas chromatography
- HPLC:
-
High performance liquid chromatography
- LB:
-
Luria-Bertani
- LC-ESI-MS:
-
Liquid chromatography-electrospray ionization mass spectrometry
- LC-APCI-MS:
-
Liquid chromatography-atmospheric pressure chemical ionization mass spectrometry
- MS:
-
Mineral salt
- NA:
-
Nutrient agar
- PCNB:
-
Pentachloronitrobenzene
- SPE:
-
Solid-phase extraction
- TPN:
-
Chlorothalonil; 2, 4, 5, 6-tetrachloroisophthalonitrile
- TPN-OH:
-
4-Hydroxy-2, 5, 6-trichloroisophthalonitrile
References
Bai WQ, Li M, Qiu XH, He FQ, Leng XF (2004) Degrative pathway of parathion-methyl by Ochrobacterum sp. B2. Chin J Pest Sci 6:48–54 in Chinese
Chaves A, Shea D, Danehower D (2008) Analysis of chlorothalonil and degradation products in soil and water by GC/MS and LC/MS. Chemosphere 71:629–638
Chernyak SM, Rice CP, McConnell LL (1996) Evidence of currently-used pesticides in air, ice, fog, seawater and surface microlayer in the Bering and Chukchi Sesas. Mar Pollut Bull 32:410–419
Chlorothalonil—Pesticide use statistics for 2008 (2010). PAN pesticides database—California pesticide use. http://pesticideinfo.org/Detail_ChemUse.jsp?Rec_Id=PC34550
Cox C (1997) Chlorothalonil. J Pest Reform Winter 17:14–20
Dai QH, Zhang RF, Jiang JD, Gu LF, Li SP (2005) Isolation, identification and characterization of Trizopgos degrading bacterium mp_4. Acta Pedol Sin 42:110–115 in Chinese
Fang ZD (1996) Basic methods on plant diseases research. 3rd edn. Chinese Agricultural Press. P182
Godard T, Fessard V, Huet S, Mourot A, Deslandes E, Pottier D et al (1999) Comparative in vitro and in vivo assessment of genotoxic effects of etoposide and chlorothalonil by the comet assay. Mutat Res 444:103–116
Holmes B, Popoff M, Kiridjian M, Kersters K (1988) Ochrobactrum anthropi gen. nov., sp. nov. from human clinical specimens and previously known as Group Vd. Int J Syst Bacteriol 38:406–416
Holt JG, Krieg NR, Sneath PHA, Staley JT, Williams ST (1994) Bergey’s manual of determinative bacteriology, 9th edn. Williams & Wilkins, Battimore
Katayama A, Isemura H, Kuwatsuka S (1991) Population change and characteristics of chlorothalonil degrading bacteria in soil. J Pestic Sci 16:239–245
Katayama A, Ukai T, Nomura K, Kuwatsuka S (1992) Formation of a methylthiolated metabolite from the fungicide chlorothalonil by soil bacteria. Biosci Biotech Biochem 56:1520–1521
Katayama A, Itou T, Ukai T (1997) Ubiquitous capability to substitute chlorine atoms of chlorothalonil in bacteria. J Pest Sci 22:12–16
Kenneth AR, Siegel MR (1981) Mechanism of action and fate of the fungicide chlorothalonil (2, 4, 5, 6-Tetrachloroisophthalonotrile) in biological systems 3. Interaction with mammalian DNA, histones, and isolated rat liver nuclei. Pestic Biochem Physiol 16:120–128
Kim YM, Park K, Joo GJ, Jeong EM, Kim JE, Rhee IK (2004) Glutathione-dependent biotransformation of the fungicide chlorothalonil. J Agric Food Chem 52:4192–4196
Lebuhn M, Achouak W, Schloter M, Berge O, Meier H, Barakat M et al (2000) Taxonomic characterization of Ochrobactrum sp. isolates from soil samples and wheat roots, and description of Ochrobactrum tritici sp. nov. and Ochrobactrum grignonense sp. nov. Int J Syst Evol Microbiol 50:2207–2223
Liang B, Li R, Jiang D, Sun JQ, Qiu JG, Zhao YF, Li SP, Jiang JD (2010) Hydrolytic dechlorination of chlorothalonil by Ochrobactrum sp. CTN-11 isolated from a chlorothalonil-contaminated soil. Curr Microbiol 61:226–233
Mincer TJ, Jensen PR, Kauffman CA, Fenical W (2002) Widespread and persistent populations of a major new marine actinomycete taxon in ocean sediments. Appl Environ Microbiol 68:5005–5011
Motonaga K, Takagi K, Matumoto S (1996) Biodegradation of chlorothalonil in soil after suppression of degradation. Biol Fertil Soils 23:340–345
Motonaga K, Takagi K, Matumoto S (1998) Suppression of chlorothalonil degradation in soil after repeated application. Environ Toxicol Chem 17:1469–1472
New pesticide (in Chinese) (2002) Productivity of chlorothalonil reached 8000 t in China. 2–3:9
Putnam RA, Nelson JO, Clark JM (2003) The persistence and degradation of chlorothalonil and chlorpyrifos in a cranberry bog. J Agric Food Chem 51:170–176
Qiu XH, Bai WQ, Zhong QZ, Li M, He FQ, Li BT (2006) Isolation and characterization of a bacterial strain of the genus Ochrobactrum with methyl parathion mineralizing activity. J Appl Microbiol 10:986–994
Regitano JB, Tornisielo VL, Lavorenti A, Pacovsky RS (2001) Transformation pathways of 14C-Chlorothalonil in tropical soils. Arch Environ Contam Toxicol 40:295–302
Roberts T, Hutson D (1999) Metabolic pathways of agrochemicals. Part 2: insecticides and fungicides. The Royal Society of Chemistry, Cambridge, United Kingdom, P1380–P1384
Rouchaud J, Roucourt P (1988) Hydrolytic biodegradation of chlorothalonil in the soil and in cabbage crops. Toxicol Environ Chem 17:59–68
Sambrook J, Russell DW (1998) Molecular cloning: a laboratory manual. 3rd edn. Cold Spring Harbor Laboratory Press
Sato K, Tanaka H (1987) Degradation and metabolites of a fungicide, 2, 4, 5, 6-tetra-chloroisophthalonitrile (TPN) in soil. Biol Fertil Soils 3:205–209
Sigler WV, Turco RF (2002) The impact of chlorothalonil application on soil bacterial and fungal populations as assessed by denaturing gradient gel electrophoresis. Appl Soil Ecol 21:107–118
Sun W (2008) Export value of Dacheng pesticide break through one hundred million. Shandong Chem Ind 1:41
Suyama K, Yamamoto H, Tatsuyama K, Komada H (1993) Effect of long-term application of a fungicide, chlorothalonil, on cellulose decomposition and microflora in soil under upland conditions. J Pestic Sci 18:225–230
Szalkowski MB, Stallard DE (1977) Effect of pH on the hydrolysis of chlorothalonil. J Agric Food Chem 25:208–210
Takagi K, Wada H, Yamazaki S (1991) Effect of long-term application of a fungicide, chlorothalonil (TPN) on upland ecosystem. Soil Sci Plant Nutr 37:583–590
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Trujillo ME, Willems A, Abril A, Planchuelo A, Rivas R, Ludeña D et al (2005) Nodulation of Lupinus albus by strains of Ochrobactrum lupini sp. nov. Appl Environ Microbiol 71:1318–1327
US Environmental Protection Agency (US EPA) (1986) Pesticide fact sheet: chlorothalonil. No. 36. Washington, DC, 30 Sept
US Geological Survey (1997) National pesticide synthesis project. Summary (online). Available at http://water.Wr.Usgs.gov/pnsp/ (verified & October 2001)
Wang J, Xue M, Chen XX, Wang YP, Li XR, Xie YS et al (2005) Chlorothalonil residues in greenhouse growing vegetables and corresponding impact resulted from different treatments. J Safety Environ 5:79–82 in Chinese
Wang GL, Wang L, Chen HH, Shen B, Li SP, Jiang JD (2010a) Lysobacter ruishenii sp. nov., a chlorothalonil-degrading bacterium isolated from a long-term chlorothalonil-contaminated soil in China. Int J Syst Evol Microbiol doi:10.1099/ijs.0.020990-0
Wang GL, Li R, Li SP, Jiang JD (2010b) A novel hydrolytic dehalogenase for the chlorinated aromatic compound chlorothalonil. J Bacteriol 192:2737–2745
Wei CH, Ren Y, Wu CF (1998) The characteristics of aniline biodegradation by Ochroba ctrum anthropi. Environ Sci 19:22–24 in Chinese
Winkler ES, Potter TL, Veneman PLM (1996) Chlorothalonil binding to aquatic humic substances assessed from gas purge studies. J Environ Sci Health B 31:1155–1170
Xiao Y, Zhang ZY, Sun SL, Wang JZ, Yang BD, Qiao FQ (2007) Residual dynamics of chlorothalonil on shed strawberry. Agrochemicals 46:548–550 in Chinese
Zeng LP, Huang JF, Zhang YF, Qiu GZ, Tong JB, Chen D et al (2008) An effective method of DNA extraction for bioleaching bacteria from acid mine drainage. Appl Microbiol Biotechnol 79:881–888
Acknowledgments
This study was supported by the National Hi-Tech R & D Program of China (Grant No. 2008AA10Z403) and the Japanese International Cooperation Agency. Thanks are given to Ms Xiuli Shen and Ms Baoli Sun for their kind assistance in TPN analysis, and to Dr. Conghui Wang for his help in analysis of TPN metabolites, and to Dr. Zhiqiang An for his critical review of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Xiu-Zhen Shi and Rong-Jun Guo contributed equally to this paper
Rights and permissions
About this article
Cite this article
Shi, XZ., Guo, RJ., Takagi, K. et al. Chlorothalonil degradation by Ochrobactrum lupini strain TP-D1 and identification of its metabolites. World J Microbiol Biotechnol 27, 1755–1764 (2011). https://doi.org/10.1007/s11274-010-0631-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11274-010-0631-0