Abstract
A thermostable cellulase-producing fungus, HML 0278, was identified as Fusarium chlamydosporum by morphological characteristics and ITS rDNA sequence analysis. HML 0278 produced extracellular cellulases in solid-state fermentation using sugar cane bassage as the carbon source. Native-PAGE analysis demonstrated that this fungus strain was capable of producing the three major components of cellulases and xylanase, with a yield of 281.8 IU/g for CMCase, 182.4 IU/g for cellobiohydrolase, 135.2 IU/g for β-glucosidase, 95.2 IU/g for filter paper activity, and 4,720 IU/g for xylanase. More importantly, the CMCase and β-glucosidase produced by HML 0278 showed stable enzymatic activities within pH 4–9 and pH 4–10, and at temperatures below 70 and 60°C, respectively.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cellulose is the most abundant and renewable biological carbon resource on earth. In nature, cellulases from bacteria and fungi hydrolyze crystalline cellulose into oligosaccharides, which are ultimately hydrolyzed into glucose by the synergistic action of at least three types of cellulolytic enzymes, namely, endo-1,4-β-d-glucanase (EG., EC3.2.1.4), cellobiohydrolase (CBH, EC3.2.1.91), and β-glucosidase (EC3.2.1.21; Bayer et al. 1998; Teeri 1997; Zhang and Lynd 2004).
Xylan is an important component of hemicellulose and it is the second most abundant natural polysaccharide on earth. Xylanase (EC3.2.1.8) refers to a class of enzymes that specifically degrade xylan into oligosacchrides and xylose (Beg et al. 2001; Collins et al. 2005).
Cellulase and xylanase have been widely used in energy, food, animal feed, medical, papermaking, and textile industries (Bhat 2000; Demain et al. 2005; Lynd et al. 2002).
Trichoderma and Aspergillus species are commonly used fungal species in the production of cellulases in industry. Unfortunately, most cellulases produced by these fungi only function effectively at a low temperature within a narrow pH range (Maheshwari et al. 2000). Fusarium species have strong cellulose degradation ability. Fusarium oxysporum produces multiple cellulolytic enzymes such as xylanase and CMCase, and the synergistic action of these enzymes can convert cellulose into ethanol (Gómez-Gómez et al. 2001; Kumar et al. 1991). The present study aimed to isolate new fungal strains that can produce improved thermostable cellulose degradation enzymes.
Here we report the isolation of a new cellulase-producing F. chlamydosporum strain, HML 0278, from the soil beneath the rotten wood. This fungal strain produces all three major components of cellulases as well as a hemicellulase, xylanase. The cellulases produced by this isolate display considerably high enzyme activities. The CMCase and β-glucosidase are thermostable and are active within a broad range of pH, indicating the potential commercial value of this newly isolated fungal strain.
Materials and methods
Isolation of the cellulase-producing fungal strain
The microorganism was isolated from the soil underneath rotten wood in a national nature reserve in Huanjiang Xian, Guangxi, China. The soil sample was diluted and streaked on a Mandels mineral salts agar (Mandels et al. 1976) plate containing 0.3% (W/V) mashed Whatman No. 1 filter paper as the sole carbon source and 1.5% agar (W/V). The plate was incubated at 30°C for 5 days. Single colonies that produced clear hydrolytic zone on the screening plate were selected and re-streaked onto a potato dextrose agar plate (PDA).
Production of cellulases by solid-state fermentation
About 106 spores were inoculated into the cellulase selective medium containing 6 g sugar cane bagasse, 4 g wheat bran, and 30 ml Mandels mineral salts in a 500 ml Erlenmeyer flask then incubated at 30°C for 4 days. The crude enzyme was extracted by adding 200 ml sterile distilled water to the culture and incubating the flask in a 40°C waterbath for 1 h. The culture mixture was centrifuged at 6,000 rpm for 15 min, and the enzyme containing supernatant was collected and stored at 4°C for further investigation.
Enzyme assays
The CMCase, cellobiohydrolase, filter paper activity (FPase) and β-glucosidase were measured using the methods described by Ghose (1987). One unit of enzyme activity (IU) was defined as the amount of enzyme that produces 1 μmol of reduced sugar (using glucose as a standard) per ml per minute. Xylanase activities were determined according to Bailey et al. (1992). The enzyme activity was expressed in international units and was defined as the amount of enzyme required to release 1 μmol reducing sugar (xylose) per ml per min.
The activities in dry carbon source were expressed in IU/g. The protein content was estimated by Bradford method (1976).
Native-PAGE and activity staining of cellulases
Crude extracellular enzymes were resolved by native polyacrylamide gel electrophoresis (native-PAGE, 8% resolving gel and 4% stacking gel) in a pH 8.3 electrophoresis buffer at a constant voltage of 50 V at 4°C. After the electrophoresis, the gel was cut into several pieces which were then subjected to either silver staining or activity staining by reacting with specific substrates for various cellulases. Cellobiohydrolase activity staining was performed by incubating the gel piece in 0.04% 4-methylumbelliferyl β-d-cellobioside (4-MUC; Sigma) at 30°C for 10 min, followed by the UV light exposure (Reinhold-Hurek et al. 1993). The cellobiohydrolase activity was indicated by the presence of white fluorescence under the UV light. The β-glucosidase activity staining was performed by staining the gel piece with 0.1% esculin (Sigma) and 0.25% ammonium iron (III) citrate (Sigma) at 30°C for 60 min. The presence of the dark yellow product indicated the existence of β-glucosidase (Kwon et al. 1994). The CMCase activity staining was carried out by incubating the gel piece in 1% sodium carboxymethyl cellulose (CMCNa) solution [dissolving CMCNa (Fluka) in 50 mM pH 5.0 sodium acetate buffer] at 30°C for 30 min. The gel was then stained with 0.2% Congo red for 20 min, and destained with 1 mol/l NaCl. CMCase activity was indicated by the presence of red color (Sugimura et al. 2003). The xylanase activity staining was carried out by incubating the gel piece in 1% birch wood xylan (Sigma) solution (birch wood xylan dissolved in pH 5.3 sodium citrate buffer) at 30°C for 60 min, followed by the staining with 0.1% I-KI. Xylanase could hydrolyze birch wood xylan and produce a clear zone which could be subsequently stained blue by I-KI. While the intact birch wood xylan displayed a black color.
Morphological identification of the fungal strain
The morphological characteristics was observed through an Olympus CX41 microscope (Olympus Corporation). The mature culture of the cellulase-producing fungal strain was dried and fixed, and the micromorphology of the cells was observed through a Hitachi S-3400 N scanning electron microscope (SEM; Hitachi High Technologies Inc).
Phylogenetic analysis of the fungal isolate
Freshly cultured mycelia were collected and grinded in a mortar with liquid nitrogen, and the total DNA was extracted following the method described by Michaelon et al. (1998).
The ribosomal DNA ITS regions of the isolate were amplified using the universal primers ITS1 (5′-TCCGTAGGTGAACCTGCGG-3′) and ITS4 (5′-TCCTCCGCTTATTGATATGC-3′). PCR amplification reaction was carried out in a total volume of 50 μl. The reaction mix contained 5 μl of 10xEx Taq buffer (TaKaRa Bio), 2 ng of the total DNA, 1 μl of 10 mmol/l dNTPs (Invitrogene), 0.5 μl of 20 μmol/l ITS1 primer, 0.5 μl of 20 μmol/l ITS4 primer, 2 U of Ex Taq™ Polymerase, and distilled water. The PCR reaction was conducted as following: an initial denaturation at 95°C for 5 min, then 30 cycles of 95°C for 50 s, 52°C for 40 s, and 72°C 50 s, followed by a final extension at 72°C for 10 min.
The PCR product was resolved on a 1% agarose gel and purified using a QIAquick PCR purification kit (Qiagen, Hilden, Germany). After sequencing, the ITS sequence was compared with the NCBI database for homologous sequences using BLAST. DNA sequences highly homologous with HML 0278 ITS were aligned with Vector NTI software, and were also used to construct a phylogenetic tree by the neighbor-joining method with the molecular evolutionary genetics analysis software MEGA4.
Effect of temperature on the activities and stability of CMCase and β-glucosidase
All enzymatic activities were measured by methods described above and expressed as relative activities that were defined as percentages of the highest enzyme activity obtained in each experiment. The optimal temperatures for CMCase and β-glucosidase were determined by evaluating the activity of each enzyme in 50 mM sodium acetate buffer (pH 5.0) at various temperatures ranging 30–90°C. The influence of temperature on the stability of the enzyme was assessed by incubating each enzyme in a waterbath adjusted to various temperatures ranging 40–90°C for 60 min and determining the remaining enzymatic activity at 50°C, thereafter.
Effect of pH on the activities and stability of CMCase and β-glucosidase
To determine the effect of pH on the activities of CMCase and β-glucosidase, the following four buffers were used to achieve the defined pH ranges: 50 mM sodium acetate buffer (pH 3.0–6.0), 50 mM phosphate buffer (pH 6.0–8.0), 50 mM Tris–HCl buffer (pH 8.0–9.0), and 50 mM glycine-sodium hydroxide buffer (pH 9.0–11.0). Under the optimal temperature, enzymes were incubated in these buffers and their pH optima for the highest activities were determined. To evaluate pH stability of CMCase and β-glucosidase, the two enzymes were first stored in buffers with pH values ranging 3.0–11.0 at 4°C for 24 h, and further incubated at 30°C for 3 h. The relative activities of the enzymes were determined at the optimal pH and the optimal temperature. The experiment was repeated three times and the mean relative enzyme activities were calculated.
Results and discussion
Isolation of cellulase-producing fungal strain
The cellulase-degrading activities in certain isolates were shown as transparent halos around the colonies on a Whatman No. 1 filter paper selective agar plate due to the hydrolysis of the filter paper cellulose. The colony HML 0278 produced the largest clear hydrolytic zone and therefore was transferred to a PDA plate for single colony isolation (Fig. 1).
Identification of the fermentation products
HML 0278 grew well in sugar cane baggase and wheat bran. This new isolate produced all three required components of the cellulase enzyme complex. The protein concentrations of crude enzyme produced by HML 0278 was 56 mg/ml. The fermentation products contained 281.8 IU/g of CMCase, 182.4 IU/g of cellobiohydrolase, 135.2 IU/g of β-glucosidase, 95.2 IU/g of filter paper activity, and 4,720 IU/g of xylanase. The cellulase enzyme complex from HML 0278 hydrolyzed crystalline cellulose first into oligosaccharides and ultimately into glucose (Ghose 1987; Teeri 1997; Zhang and Lynd 2004).
The ability of a microorganism to degrade filter paper cellulose depends on its ability to produce cellulases and xylanase. Therefore, using filter paper as the sole carbon source can facilitate the selection of microorganisms that are rich in cellulases. In principle, cellulase-producing microorganisms can be isolated by filter paper cellulose selection followed by enzyme assays to confirm the production of cellulolytic enzymes in the fermentation.
Analysis of the cellulolytic enzymes secreted by HML 0278
Native-PAGE and activity staining showed that the extracellular enzyme mixture of HML 0278 contained major components of cellulases and a hemicellulase, including one type of cellobiohydrolase, two types of β-glucosidases, one type of CMCases, and one type of xylanase, as shown by colored bands on the gel (Fig. 2).
The activity staining of cellulases secreted from HML 0278 on the native-PAGE gel. Lane 1 cellobiohydrolase activity staining, lane 2 and 3 β-glucosidase activity staining, lane 4 and 5 CMCase activity staining, lane 6 and 7 xylanase activity staining, Xylanase hydrolyzed birch wood xylan and produced a clear zone which was subsequently stained blue by I-KI, while the intact birch wood xylan displayed a black color, lane 8 and 9 silver stained extracellular enzymes, lane M the Serva native-PAGE protein marker (SERVA Electrophoresis GmbH, Germany) showing (1) 160 kDa aldolase rabbit (pI 8.2–9.1); (2) 240 kDa catalase bovine (pI 5.5–6); (3) 17.8 kDa myoglobin equine (pI 6.9–7.4); and (4) 67 kDa albumin bovine (pI 4.7–4.9) as indicated by the numbers beside the protein bands. The arrows point to the location of the corresponding enzymes
The colony morphology and microscopic features of HML 0278
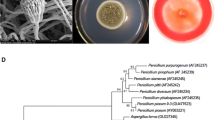
The colony of HML 0278 was white and fluffy. It reached 5 cm in diameter when cultured on a PDA plate for 4 days (Fig. 3). HML 0278 produced branched hyphae with septa (Fig. 4a). Thick-walled conidia were paragenic or terminal (Fig. 4b). It contained two types of conidia. Branching conidiophore-bearing microconidia were spindle-shaped, single or in chain on the conidiophores, and scattering among mycelia (Fig. 4c). Macroconidia were sickle-shaped and septate, single or tuft (Fig. 4d). The colony morphology and microscopic characteristics of HML 0278 were consistent with those of Fusarium chlamydosporum (Dai 1987; Wei 1979).
Phylogenetic analysis of the fungal isolate
The ITS regions of the nuclear rDNA unit are often highly conserved within species and therefore are considered valuable genetic markers for species identification (Bruns et al. 1991; Gardes and Bruns 1993). HML 0278 DNA was successfully extracted and the ITS region was successfully amplified. Sequence homology analysis and the phylogenetic tree identified HML 0278 as F. chlamydosporum (99% sequence homology of ITS regions; Fig. 5). Based on colony morphology, microscopic features, and ITS sequence analysis, HML 0278 was classified into F. chlamydosporum.
Evaluation of enzyme activity and native-PAGE staining further identified that the extracellular enzymes secreted by HML 0278 contained xylanase and the three major cellulases, namely CMCase, avicelase, and β-glucosidase. Our study, for the first time, reported that an F. chlamydosporum, HML 0278, produced complete components of the cellulases as well as xylanase.
Effect of temperature on the activities and stability of CMCase and β-glucosidase
The optimal temperature for CMCase activity was 60°C at pH 5. The enzyme showed considerable satility at temperatures below 70°C, and it retained 82% of its original activity even after 1 h incubation at 70°C. The optimal temperature for β-glucosidase activity was 55°C. The enzyme remained stable below 60°C, and it still maintained 80% of its initial activity even after 1 h incubation at 60°C. The CMCase and β-glucosidase of HML 0278 can be considered thermophilic (Fig. 6).
The effect of pH on the stability and activities of CMCase and β-glucosidase
CMCase was relatively stable at pH 4–9, and the maximal enzyme activity was obtained at pH 6. The β-glucosidase was relatively stable within pH 4–10, and its maximal activity was achieved at pH 5 (Fig. 7). The ability of an enzyme to retain high activity at elevated pH is a potentially useful property in processes employing alkaline delignification (Alani et al. 2008).
The CMCase produced by Fusarium oxysporum was stable under 55°C (Christakopoulos et al. 1995; Christakopoulos et al. 1989), the activities of F. oxysporum that degraded carboxymethylcellulose (CMC), filter paper, and Avicel were very low (Christakopoulos et al. 1996). The CMCase of Fusarium moniliforme was stable at pH 3–8, and the optimum temperature was found to be 60°C (Matsumoto et al. 1974). The CMCase of Fusarium solani was stable at pH 5–7 (Wood 1969). Our data showed that the CMCase produced by HML 0278 tolerates higher pH and higher temperature than the Cellulase produced by other Fusarium species.
Conclusions
In this study, a novel thermostable cellulase-producing fungus, HML 0278, has been investigated. Based on the colony morphology, microscopic features, and ITS sequence analysis, HML 0278 was classified into F. chlamydosporum.
F. chlamydosporum HML 0278 produced extracellular cellulases in solid-state fermentation using a mixture of sugar cane bassage and wheat bran as the carbon source. Using sugar cane baggase as a carbon source has many advantages. For example, sugar cane baggase and wheat bran are rich in nutrients, especially multiple oligosaccharides. Thus they are able to promote the rapid growth of cells. In addition to easy availability and cost-effectiveness, these two types of carbon sources are also a good inducer for the production of cellulases and xylanase by fungal cells (Matthew et al. 2000; Muniswaran and Charyulu 1994).
Native-PAGE analysis demonstrated that this fungal strain was capable of producing the three major components of cellulases and xylanase, with a yield of 281.8 IU/g for CMCase, 182.4 IU/g for cellobiohydrolase, 135.2 IU/g for β-glucosidase, 95.2 IU/g for filter paper activity, and 4,720 IU/g for xylanase. The CMCase exhibits stability within pH 4–9, and at temperatures below 70°C. The β-glucosidase displays stability within pH 4–10 and at temperatures below 60°C. As the most abundant enzymes produced by this strain, β-glucosidase and CMCase display superior thermo and pH stability. The newly isolated F. chlamydosporum HML 0278 provides an improved cellulose hydrolytic efficacy and therefore it has potential applications in industries such as fuel ethanol industry, and paper and agricultural industries that process cellulose-derived materials (Alani et al. 2008; Maheshwari et al. 2000).
References
Alani F, Anderson WA, Moo-Young M (2008) New isolate of Streptomyces sp. with novel thermoalkalotolerant cellulases. Biotechnol Lett 30:123–126
Bailey MJ, Biely P, Poutanen K (1992) Interlaboratory testing of methods for assay of xylanase activity. J Biotechnol 23:257–270
Bayer EA, Chanzy H, Lamed R, Shoham Y (1998) Cellulose, cellulases and cellulosomes. Curr Opin Struct Biol 8(5):548–557
Beg QK, Kapoor M, Mahajan L, Hoondal GS (2001) Microbial xylanases and their industrial applications: a review. Appl Microbiol Biotechnol 56:326–338
Bhat MK (2000) Cellulases and related enzymes in biotechnology. Biotechnol Adv 18:355–383
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Bruns TD, White TJ, Taylor JW (1991) Fungal molecular systematics. Annu Rev Ecol Syst 22:525–564
Christakopoulos P, Macris BJ, Kekos D (1989) Direct fermentation of cellulose to ethanol by Fusarium oxysporum. Enzyme Microb Tech 11:236–239
Christakopoulos P, Kekos D, Macris BJ, Claeyssens M, Bhat MK (1995) Purification and characterization of a less randomly acting endo-1, 4-beta-d-glucanase from the culture filtrates of Fusarium oxysporum. Arch Biochem Biophys 316(1):428–433
Christakopoulos P, Kekos D, Macris BJ, Claeyssens M, Bhat MK (1996) Purification and characterisation of a major xylanase with cellulase and transferase activities from Fusarium oxysporum. Carbohydr Res 289:91–104
Collins T, Gerday C, Feller G (2005) Xylanases, xylanase families and extremophilic xylanases. FEMS Microbiol Rev 29:3–23
Dai FL (1987) Morphology and classification of fungi, 1st edn. Scientific & Technological Press, Beijing, pp 297–314
Demain AL, Newcomb M, Wu JHD (2005) Cellulase, clostridia, and ethanol. Microbiol Mol Biol Rev 69:124–154
Gardes M, Bruns TD (1993) ITS primers with enhanced specificity for basidiomycetes-application to the identification of mycorrhizae and rusts. Mol Ecol 2:113–118
Ghose TK (1987) Measurement of cellulose activities. Pure Appl Chem 59:257–260
Gómez-Gómez E, Isabel M, Roncero G, Di Pietro A, Hera C (2001) Molecular characterization of a novel endo-beta-1, 4-xylanase gene from the vascular wilt fungus Fusarium oxysporum. Curr Genet 40:268–275
Kumar PKR, Singh A, Schuegerl K (1991) Fed-batch culture for the direct conversion of cellulosic substrates to acetic acid/ethanol by Fusarium oxysporum. Process Biochem 26:209–216
Kwon KS, Lee J, Kang HG, Hah YC (1994) Detection of beta-glucosidase activity in polyacrylamide gels with esculin as substrate. Appl Environ Microbiol 60:4584–4586
Lynd LR, Weimer PJ, van Zyl WH, Pretorius IS (2002) Microbial cellulose utilization: fundamentals and biotechnology. Microbiol Mol Biol Rev 66:506–577
Maheshwari R, Bharadwaj G, Bhat MK (2000) Thermophilic fungi: their physiology and enzymes. Microbiol Mol Biol Rev 64:461–488
Mandels M, Andreotti RE, Roche C (1976) Measurement of saccharifying cellulose. Biotechnol Bioeng Symp 6:21–33
Matsumoto K, Endo Y, Tamiya N, Kano M, Miyauchi K (1974) Studies on cellulase produced by the phytopathogens. Purification and enzymatic properties of cellulase of Fusarium moniliforme. J Biochem 76:563–572
Matthew TH, David AM, Tony H (2000) Approach to designing rotating drum bioreactors for solid-state fermentation on the basis of dimensionless design factors. Biotechnol Bioeng 67(3):274–282
Michaelon LV, Lazarus CM, Griffiths G, Napier JA, Stobart AK (1998) Isolation of a delta5-fatty acid desaturase gene from Mortierella alpina. J Biol Chem 273:19055–19059
Muniswaran A, Charyulu N (1994) Solid substrate fermentation of coconut coir pith for cellulase production. Enzyme Microb Technol 16:436–440
Reinhold-Hurek B, Hurek T, Claeyssens M, Van MM (1993) Cloning, expression in Escherichia coli and characterization of cellulolytic enzymes of Azoarcus sp., a root-invading diazotroph. J Bacteriol 175:7056–7065
Sugimura M, Watanabe H, Lo N, Saito H (2003) Purification, characterization, cDNA cloning and nucleotide sequencing of a cellulase from the yellow-spotted longicorn beetle, Psacothea hilaris. Eur J Biochem 270:3455–3460
Teeri TT (1997) Crystalline cellulose degradation: new insight into the function of cellobiohydrolase. Trends Biotechnol 15:160–167
Wei JC (1979) Fungi identification guide, 1st edn. Shanghai Science & Technology Press, Shanghai, pp 405–614
Wood TM (1969) The cellulase of Fusarium solani. Resolution of the enzyme complex. Biochem J 115(3):457–464
Zhang YH, Lynd LR (2004) Toward an aggregated understanding of enzymatic hydrolysis of cellulose: noncomplexed cellulase systems. Biotechnol Bioeng 88:797–824
Acknowledgments
This work was supported in part by the National Natural Science Foundation of China (20066001), Guangxi Natural Science Foundation (2010GXNSFA013103), and by the Open Fund of Guangxi Key Laboratory of Subtropical Bioresource Conservation and Utilization (SB0607).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Qin, Y., He, H., Li, N. et al. Isolation and characterization of a thermostable cellulase-producing Fusarium chlamydosporum . World J Microbiol Biotechnol 26, 1991–1997 (2010). https://doi.org/10.1007/s11274-010-0383-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11274-010-0383-x