Abstract
Brazil is the largest sugarcane producer in the world, mainly due to the development of different management strategies. Recently, microbial-plant related studies revealed that bacterial isolates belonging to the genus Burkholderia are mainly associated with this plant and are responsible for a range of physiological activity. In this study, we properly evaluate the physiological activity and genetic diversity of endophytic and rhizospheric Burkholderia spp. isolates from sugarcane roots grown in the field in Brazil. In total, 39 isolates previously identified as Burkholderia spp. were firstly evaluated for the capability to fix nitrogen, produce siderophores, solubilise inorganic phosphates, produce indole-acetic acid and inhibit sugarcane phytopathogens in vitro. These results revealed that all isolates present at least two positive evaluated activities. Furthermore, a phylogenetic study was carried out using 16S rRNA and gyrB genes revealing that most of the isolates were affiliated with the Burkholderia cepacia complex. Hence, a clear separation given by endophytic or rhizospheric niche occupation was not observed. These results presented an overview about Burkholderia spp. isolates from sugarcane roots and supply information about the physiological activity and genetic diversity of this genus, given direction for further studies related to achieve more sustainable cultivation of sugarcane.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sugarcane (Saccharum spp.) occupies a prominent position among the important crops in the world. Brazil is the largest producer of this crop, dominating the market of sugar and ethanol-derived in the last few years (Goldemberg 2007). Therefore, a better understanding of the role played by the bacterial community associated with sugarcane could be a key to improve the performance and sustainability of sugarcane cultivation.
Some endophytic and rhizospheric bacteria have been explored intensively given their beneficial characteristics such as promotion of plant growth and protection against pests and pathogens (Lodewyckx et al. 2002). These bacteria may increase plant fitness by nitrogen fixation, production of phytohormones and antimicrobial substances and induction of systemic resistance. A wide variety of bacteria are able to colonize sugarcane plants; however, the Burkholderia cepacia complex (Bcc) has been shown to be a major bacteria group that interacts with these plants (Mendes et al. 2007). Currently, this genus includes more than 50 species found in various ecological niches, rather in bulk soil and rhizosphere (Coenye and Vandamme 2003).
In this extremely versatile group, certain species are capable of causing disease in plants and humans, while others are very effective in biological control, bioremediation, and promotion of plant growth (Perin et al. 2006b). Such a complex groups has also problems on its taxonomy, where it is possible to observe the grouping of physiological distinct isolates into similar taxonomical branches. It makes necessary the application of robust analysis to accurately classify isolates affiliated to the genus Burkholderia (Mendes et al. 2007). Besides the commonly used gene 16S rRNA, other genes are candidate for Burkholderia isolates grouping and classification. Since 1995, when universal primers for the gene gyrB (which codifies for the DNA gyrase protein) became available (Yamamoto and Harayama 1995), several publications have suggested that gyrB is a suitable gene for bacterial phylogeny, possessing essential attributes such as limited horizontal transfer and presence in all bacterial groups. In Burkholderia spp., this gene demonstrated to be a useful tool to discriminate strains (Tabacchioni et al. 2008).
In the present study, endophytic and rhizospheric Burkholderia spp. were evaluated according to their molecular and physiological characteristics. Isolates belonging the genus Burkholderia were obtained from sugarcane roots (as endophytes and rhizobacteria) and characterized with the aim of investigating niche-related differences based on gyrB and 16S rRNA gene sequences and evaluation of phenotypic characteristics, such as nitrogen fixation, indole-acetic acid and siderophore production, inorganic phosphate solubilization, and inhibition of sugarcane pathogens.
Materials and methods
Bacterial isolates and growth conditions
The bacteria used in this study were isolated from rhizosphere and root tissues of sugarcane plants (cv. SP80-1842) grown for 3 months in an experimental field located in Piracicaba, Brazil (22°41′S 47°33′W), as previously described by Mendes et al. (2007). In total, 18 individual sugarcane plants were sampled for isolation, generating a wide diversity of bacteria, from which the Burkholderia isolates were selected out for the present study. The isolation procedures were also made according to the protocol previously described by Mendes et al. (2007). Briefly, rhizospheric and root endophytic bacteria were isolated from root segments of different plants collected at a depth of 5–15 cm from the stem base. Isolates were sampled from different host plant, avoiding the selection of clones from the same plant. Rhizosphere bacteria were isolated by vigorously shaking 2 g of roots in PBS buffer (140 mM NaCl, 2.5 mM KCl, 10 mM Na2HPO4, 1.5 mM KH2PO4 pH 7.4) followed by dilution plating on tryptic soy agar (TSA; Difco, Le Pont de Claix, France) supplemented with 50 mg ml−1 of the fungicide imazalil (Agricur, Brazil) and incubation at 28°C. For the isolation of endophytic bacteria, a sterilization process was carried out washing roots in 70% ethanol for 1 min, sodium hypochlorite (2% v/v) for 3 min, and 70% ethanol for 30 s by two rinses with sterilized distilled water. The surface-sterilized root segments were triturated in a mortar with a pestle by using 2 ml of PBS buffer. Aliquots of 100 μl were platted on TSA supplemented with 50 mg ml−1 imazalil. Plates were incubated at 28°C for 10 days. After bacterial growth, colonies showing Burkholderia morphology were sampled for further studies.
Physiological characterization of Burkholderia isolates
The ability to fix atmospheric nitrogen was evaluated according to Döbereiner et al. (1995). Briefly, isolates were inoculated in tubes containing 10 ml of semi-solid NFb medium. Bacterial growth was evaluated after 72 h of incubation at 28°C. The formation of growth disc in the culture medium indicated the ability of the isolates to fix atmospheric nitrogen.
Indole-acetic acid (IAA) production was evaluated using a modification of the qualitative method developed by Bric et al. (1991). Firstly, bacterial isolates were inoculated in 5% tryptone soy broth (TSB) medium supplemented with l-tryptophan (5 mM) and incubated with shaking (150 rpm) at 28°C for 72 h in the dark. Cultures were then centrifuged (5 min, 7,000g, at room temperature), and 400 µl de Salkowski reagent was added to 900 μl of the supernatant. This mixture was incubated for 30 min at room temperature and analyzed by spectrophotometer (530 nm) (Pharmacia Biotech Ultroespec 3000). Absorbance values obtained were interpolated in a standard curve to determine the IAA concentration.
The ability of Burkholderia spp. isolates to solubilise inorganic phosphate was evidenced by a halo obtained after isolate cultivation on culture medium supplemented with inorganic phosphate (Ca3(PO4)2) at 28°C for 72 h, according to Verma et al. (2001). The quantification was done by the estimating of halo size (cm) divided by the colony size (cm), generating a solubilization index (SI).
Siderophore production was evaluated by universal methodology on CAS-Agar medium (Schwyn and Neilands 1987). Isolates were cultivated in CAS-Agar plates at 28°C for 72 h, and the production of siderophore was observed by the presence of a yellow or orange halo around the bacterial colony.
Inhibition of sugarcane phytopathogens
The inhibition potential of Burkholderia isolates against Fusarium verticillioides (Sin: F. moniliforme) and Xanthomonas albilineans was evaluated on one-fifth-strength potato dextrose agar medium and by agar layer methodology (Pugsley and Oudega 1987), respectively.
Genotypic characterization of Burkholderia spp. isolates
Total DNA was extracted from 24-h bacterial cultures grown in 5% TSB medium according to Araujo et al. (2002), and the genotypic diversity of the isolates was evaluated by sequencing the 16S rRNA and gyrB genes. Amplification of the 16S rRNA gene was carried out with an MJ Research PTC-200 thermocycler, using the universal primers for Bacterial Domain, R1387 (Heuer et al. 1997) and PO27F (Lane et al. 1985), resulting in a near-complete 16S rRNA fragment (ca. 1,360 bp). The reaction mixture (50 μl) was preparing containing 3.75 mM of MgCl2, 0.2 mM of each primer, 0.2 mM of each dNTP, 2.5 U of Taq DNA polymerase, 1× Taq buffer and 1 μl of template DNA (5–10 ng). The PCR products were firstly checked for the right size by electrophoresis on agarose gel (0.8% w/v) and purified using a polyethylene glycol (PEG 8000 20%; NaCl 2.5 mM) method, followed by partial sequencing in an automated sequencer (ABI Prism 377, PE Applied Biosystems) using primer R1387.
The gyrB gene was partially amplified using primers UP-1 (5′-GAA GTC ATC ATG ACC GTT CTG CAY GCN GGN GGN AAR TTY GA-3′) and UP-2r (5′-AGC AGG GTA CGG ATG TGC GAG CCR TCN ACR TCN GCR TCN GTC AT-3′) (Yamamoto and Harayama 1995; Yamamoto and Harayama 1998) (ca. ~1,200 bp). PCR-reaction (50 μl) was performed containing 3.75 mM of MgCl2, 0.2 mM of each primer, 0.2 mM of each dNTP, 1× Taq Buffer, 2.5 U of Taq DNA polymerase and 1 μl of DNA template (5–10 ng). The PCR products were checked in agarose gel, purified using polyethylene glycol (PEG 8000 20%; NaCl 2.5 mM) and partial sequenced with primer UP-1.
Totals sequences obtained from the 16S rRNA and gyrB genes were compared firstly by BLASTn (Basic Local Alignment Search Tool) (Altschul et al. 1997) against the nr/nt database from the NCBI (National Center for Biotechnology Information website). 16S rRNA sequences were also subjected to classification at the Ribosomal Data Project II (RDP) (http://rdp.cme.msu.edu/). Most similar sequences were retrieved from the databases and used for alignment with the sequences obtained in this study. For the 16S rRNA gene, sequences from type strains available in the RDP database were used. Alignment and phylogenetic analyses were conducted using ClustalW algorithm in the software MEGA 4.0.1 (Tamura et al. 2007). Phylogeny was inferred using Neighbor Joining (NJ) (Saitou and Nei 1987) based on a similarity calculated by the parameter of Kimura-2 (Kimura 1980) with a bootstrap of 1,000 replicates (see details on the figure legend).
Results
Phenotypic characterization of Burkholderia isolates
In the present study, a total of 39 bacterial isolates resultant of these isolation processes were previously identified as Burkholderia genus and used for further analysis (Table 1). The ability to fix nitrogen was observed in 87.2% of isolates, with values of 80.0 and 94.7% in endophytic and rhizospheric isolates, respectively (Table 1). In addition, all isolates produced IAA at levels ranging from 3.43 to 19.47 μg ml−1. The IAA production by endophytic isolates ranged from 3.43 to 9.97 μg ml−1, while rhizosphere isolates presented values ranged from 4.40 to 19.47 μg ml−1 (Table 1). The solubilization of inorganic phosphate was detected by radial diffusion in the culture medium and formation of a solid degradation halo (Table 1). The SI values for endophytic and rhizospheric isolates ranged from 1.11 to 6.0 and 2.32–3.75, respectively. With regard to siderophore production, around 35.9% of the isolates grown in CAS-Agar medium revealed the formation of a yellow halo surrounding the bacterial colony. Only 25% of endophytic isolates were able to produce siderophore, while in rhizobacteria this capability was verified for 47.4% of isolates (Table 1). Considering the ability in inhibit the growth of sugarcane phytopathogens, all isolates, except the endophytic isolate CV4.4.3R1, were able to inhibit F. verticillioides growth. In addition, layer agar assays revealed that 65 and 84.2% of Burkholderia isolates from root tissues and rhizosphere, respectively, significantly inhibited the growth of X. albilineans (Table 1).
One of the most striking features of these whole analyses is the percentage of 25.6% of isolates presenting positive correlation for all applied tests, whilst every others isolates (74.4%) presented at least two positive correlations.
Genetic diversity and phylogenetic affiliation of Burkholderia isolates
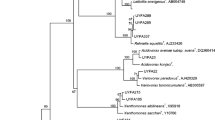
The phylogenetic analysis based on 16S rRNA and gyrB genes allowed an overview of Burkholderia species colonizing sugarcane roots. The isolates analyzed in this work clustered mainly with species associated with the Bcc, except for two isolates that clustered with other species in the genus Burkholderia. Sequence analyses of 16S rRNA gene showed that the evaluated isolates contained a high sequence similarity belonging to the B. cepacia complex (Bcc), more specifically to B. cepacia, B. cenocepacia and B. anthina (Fig. 1). Meanwhile, CV4.2.1R5 and CV3.2.3F3 isolates grouped out of the Bcc, showing similarity with the species B. phytofirmans (Fig. 1a).
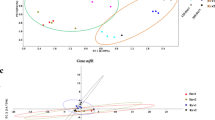
Phylogenetic analysis of bacterial 16S rRNA (a) and gyrB (b) genes retrieved from Burkholderia spp. isolates. The evolutionary relation was inferred using the Neighbor-Joining method (Saitou and Nei 1987). The evolutionary distances were computed using the Kimura 2-parameter method (Kimura 1980). In black it is represented isolates and database sequences belonging to the Burkholderia cepacia complex (Bcc); in gray sequences affiliated to other Burkholderia species. Scale bar indicates the number of base substitutions per site, and bootstrap values above 50% are represented by solid circles next to tree branches (total of 1,000 repetitions). There were a total of 575 and 556 aligned nucleotide positions in the final data set of 16S rRNA and gyrB genes, respectively. Gene sequence of Cupriavidus necator, C. taiwanensis, Ralstonia mannitolilytica and R. solanacearum served as outgroups on both trees. Sequences from isolates are available in the GenBank database under codes GU441604–GU441638 for the 16S rRNA gene and GU441639–GU441677 for the gene gyrB
Sequence analysis of gyrB confirmed the phylogenetic relationship of the isolates with the Bcc complex (Fig. 1b), with the same isolates (CV4.2.1R5 and CV3.2.3F3) with similarity to B. plantarii. Both analyzed genes (16S rRNA and gyrB) seemed to be group isolates independent of niche occupation (endophytic and rhizosphere). However, few indications could be observed like in the 16S rRNA phylogenetic tree two subgroups were formed containing rhizospheric and endophytic isolates (Fig. 1a). Also in the tree resulted from the gyrB gene analysis small groups were observed for part of isolates from each niche (Fig. 1b), but the grouped isolates were different, with exception for the clustering of the isolates CV2.2.2F2 and TC3.4.1F2, which were grouped in both analyses (Fig. 1).
Discussion
The development of plants is constantly influenced by interactions with the microbial community, especially endophytic and rhizospheric bacterial communities. Previous studies have described the importance of the genus Burkholderia in the cultivation of sugarcane (Perin et al. 2006a; Pugsley and Oudega 1987). Recently, Mendes et al. (2007) reported for the first time that bacteria related to Bcc are frequently associated with sugarcane. The understanding of the role of these bacteria on promoting plant growth may allow some practical applications for a more sustainable sugarcane production.
An important issue for a sustainable production of sugarcane is the input of nitrogen, caused by nitrogen-fixing bacteria, an ability commonly found in endophytic bacteria (Oliveira et al. 2002). Such capacity is possibly attributed to the occupation of inner plant root, conferring appropriate conditions for N2 fixation (low levels of O2 and nitrogen) (Steenhoudt and Vanderleyden 2000). Several authors have described new Burkholderia species as nitrogen-fixing rhizobacteria, such as B. unamae (Caballero-Mellado et al. 2004), B. xenovorans (Goris et al. 2004), B. tropica (Reis et al. 2004) and B. silvatlantica (Perin et al. 2006b). Also, rhizobacteria can fix nitrogen when involved in exopolysaccharides, as described by Serrato et al. (2006) for the isolate B. tropica Ppe8. This feature may be widespread throughout the genus Burkholderia since other types of microbial exopolysaccharides have been studied in B. cepacia (Linker et al. 2001). Here we found most of the Burkholderia spp. isolates growing in the NFb medium, as an indicative of the capacity to fix nitrogen, suggesting the importance of this bacterial group for plant growth promotion.
Similarly, all isolates were able to produce IAA, with higher values for isolates from the rhizosphere, suggesting an association of these isolates with plant growth promotion at roots. In addition, if we consider that IAA has not a known role in bacterial cells, the maintenance of such characteristics must be driven by interactions with plants (Patten and Glick 1996). Another factor involved in plant-bacteria interaction is the ability to solubilise inorganic phosphate (Rodriguez and Fraga 1999) since phosphorus is a commonly limiting nutrient for plant growth in tropical soils. The production and release of organic acids is related to ability of certain bacteria to solubilise inorganic phosphate (Vassilev et al. 2006). We observed that this ability was frequent in both endophytic and rhizospheric Burkholderia spp. isolates, suggesting a role of these bacteria in plant growth.
Bacteria can also enhance plant growth through the synthesis of siderophores, which can take and deliver iron from soil and also limit the growth of pathogenic microorganisms (Siddiqui 2005), especially pathogenic fungi (O’Sullivan and O’Gara 1992). Moreover, siderophores can induce resistance mechanisms in the plant (Schroth and Hancook 1995). In the present study, the rhizospheric isolates were found as major producers of siderophores than endophytes, indicating that this characteristic could increase bacterial competition in the rhizosphere (Copping and Menn 2000).
Burkholderia cepacia complex organisms also produce a number of substances with antagonistic potential for plant pathogens (Heungens and Parke 2000). Some of the isolates used in this study (ESR63, ESR73, ESR100, ESR108) were previously described by Mendes et al. (2007) as producers of pyrrolnitrin, a metabolite with antifungal activity (Ligon et al. 2000). Most of the analyzed isolates were able to inhibit the fungus Fusarium verticillioides (Sin: F. moniliforme) in vitro. Also, Burkholderia spp. isolates from rhizosphere inhibited the growth of the X. albilineans. Therefore, we suggest that Burkholderia spp. living in the rhizosphere compete with other pathogens for nutrients and for occupation of the ecological niche; thus, they must develop tools for intra- and inter-specific competition, such as the production of antibiotics, enzymes and bacteriolytic molecules (Parret et al. 2003). Our results are consistent with the literature and indicate that the evaluated isolates have potential applications in biocontrol programs. It should be noted, however, that although tests conducted in vitro show the abilities presented here, actual promotion of plant growth should be evaluated under field conditions. In previously reports, bacteria classified into the genus Burkholderia that showed nitrogen fixation ability were able to promote the growth of field-cultivated coffee and maize plants (Estrada-De Los Santos et al. 2002).
Another important issue addressed in this study is the genetic diversity and the phylogeny of bacteria belonging to the Burkholderia. Bacteria from this genus have genomes with two or three chromosomes varying in size from 6 to 9 Mb, features that allow great genomic plasticity and adaptability (Michè et al. 2001). Thus, bacteria of this genus can colonize the rhizosphere and inner tissues in a wide range of host plants (Caballero-Mellado et al. 2004; Perin et al. 2006b; Reis et al. 2004). Many of the bacteria in the rhizosphere enter the roots using tissue wounds as the main entrance to the plant, and establish themselves as endophytic populations (Azevedo 1998). Thus, it is assumed that many communities of endophytic bacteria are the result of a process of colonization that began in the external roots surface, the main entry point of endophytic microorganisms (Andreote et al. 2006; Welbaum et al. 2004). The distinction between endophytic and epiphytic bacteria (rhizoplane or phyloplane) is only semantic, and there is no clear boundary, but rather a gradient between groups, as there are bacterial populations that can fluctuate between the colonization of these sites based on environmental factors (Andreote et al. 2009). The results obtained here corroborate with these indications, were a major part of the isolates were groups in phylogenetic analysis independently of the niche occupation, while a minor part was niche-specific, evidencing the mixture of rhizosphere-endophyte populations promoted by the plant colonization.
Considering the taxonomic affiliation of the isolates, the major part (37 out of 39) were assigned to the Bcc, 16S rRNA sequences grouped them into three species: B. cepacia, B. cenocepacia and B. anthina. Interestingly, similar analysis based on gyrB sequences limited the classification to the B. cepacia and B. cenocepacia species, corroborating with data from previous studies, which have shown the differential topology of trees for gyrB and 16S rRNA genes (Coenye et al. 2001). We agree on the statement that the taxonomy of the Bcc is not yet fully resolved (Coenye et al. 2003), since the presence of various phenotypic-distinct isolates within species such as B. cepacia and B. cenocepacia represent the need of an additional level of taxonomic resolution. However, these results allow us to conclude that the Bcc is the main source of Burkholderia spp. in association with sugarcane plants, corroborating previous results (Mendes et al. 2007). The other two isolates (CV4.2.1R5 and CV3.2.3F3) were identified by the gene 16S rRNA as B. phytofirmans. However, by assessing gyrB gene sequences, these two isolates were clustered with sequences from the species B. plantarii, initially described as Pseudomonas plantarii, a rice phytopathogen (Azegami et al. 1988). Hence, it should be remarked that the gyrB sequence for B. phytofirmans was not available at database and the bootstrap values for this classifications were low (<50) (Fig. 1). Furthermore, in both sequence analyses, many isolates were not clearly grouped with any sequence from databases, indicating the possible existence of a yet non-described subset of the Burkholderia genus. To complete the description of new species of the genus Burkholderia, however, detailed sequencing studies should be conducted.
In summary, our results have demonstrated that most of sugarcane-associated bacteria of the genus Burkholderia are related to the Bcc, harboring besides the sequence similarities a number of phenotypes which can be targeted for the use as plant growth promoters and as biocontrol agents, especially for sugarcane plants.
References
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Andreote FD, Lacava PT, Gai CS, Araújo WL, Maccheroni W Jr, van Overbeek LS, van Elsas JD, Azevedo JL (2006) Model plants for studying the interaction between Methylobacterium mesophilicum and Xylella fastidiosa. Can J Microbiol 52:419–426
Andreote FD, Azevedo JL, Araújo WL (2009) Assessing the diversity of bacteria communities associated with plants. Braz J Microbiol 40:417–432
Araujo WL, Marcon J, Maccheroni W Jr, van Elsas JD, van Vuurde JWL, Azevedo JL (2002) Diversity of endophytic bacterial populations and their interaction with Xylella fastidiosa in citrus plants. Appl Environ Microbiol 68:4906–4914
Azegami K, Nishiyama K, Tabei H (1988) Infection courts of rice seedlings with Pseudomonas plantarii and Pseudomonas glumae. J Plant Dis Prot 54:337–341
Azevedo JL (1998) Biodiversidade microbiana e potencial Biotecnológico. In: Melo IS, Azevedo JL (ed) Ecologia microbiana. EMBRAPA—Meio Ambiente, Jaguariúna, pp 445–461
Bric JM, Bostock RM, Silverstone S (1991) Rapid in situ assay for indoleacetic acid production by bacteria immobilized on a nitrocellulose membrane. Appl Environ Microbiol 57:535–538
Caballero-Mellado J, Martinez-Aguilar L, Paredes-Valdez G, Estrada-De los Santos P (2004) Burkholderia unamae sp. nov., an N2-fixing rhizospheric and endophytic species. Int J Syst Evol Microbiol 54:1165–1172
Coenye T, Vandamme P (2003) Diversity and significance of Burkholderia species occupying diverse ecological niches. Environ Microbiol 5:719–729
Coenye T, Vandamme P, Govan JRW, Lipuma JJ (2001) Minireview: taxonomy and identification of the Burkholderia cepacia complex. J Clin Microbiol 39:3427–3436
Copping LG, Menn JJ (2000) Biopesticides: a review of their action, applications and efficacy. Pest Manag Sci 56:651–676
Döbereiner J, Baldani VLD, Baldani JL (1995) Como isolar e identificar bactérias diazotróficas de plantas não-leguminosas. Embrapa-SPI, Brasília
Estrada-De Los Santos P, Mavingui P, Cournoyer B, Fontaine F, Balandreau J, Caballero-Mellado J (2002) A N2-fixing endophytic Burkholderia sp. associated with maize plants cultivated in Mexico. Can J Microbiol 48:285–294
Goldemberg J (2007) Ethanol for a sustainable energy future. Science 315:808–810
Goris J, de Vos P, Caballero-Mellado J, Park JH, Falsen E, Quensen JF III, Tiedje JM, Vandamme P (2004) Classification of the PCB- and biphenyl-degrading strain LB400 and relatives as Burkholderia xenovorans sp. nov. Int J Syst Evol Microbiol 54:1677–1681
Heuer H, Krsek M, Baker P, Smalla K, Wellington EMH (1997) Analysis of actinomycete communities by specific amplification of genes encoding 16S rRNA and gel-electrophoretic separation in denaturing gradients. Appl Environ Microbiol 63:3233–3241
Heungens K, Parke JL (2000) Zoospore homing and infection events: effects of the biocontrol bacterium Burkholderia cepacia AMMDR1 on two oomycete pathogens of pea (Pisum sativum L.). Appl Environ Microbiol 66:5192–5200
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide-sequences. J Mol Evol 16:111–120
Lane DJ, Pace B, Olsen GJ, Stahl DA, Sogin ML, Pace NR (1985) Rapid-determination of 16S ribosomal RNA sequences for phylogenetic analyses. Proc Natl Acad Sci USA 82:6955–6959
Ligon JM, Hill DS, Hammer PE, Torkewitz NR, Hans-Joachim DH, van Pée KH (2000) Natural products with antifungal activity from Pseudomonas biocontrol bacteria. Pest Manag Sci 56:688–695
Linker A, Evans LR, Impallomeni G (2001) The structure of a polysaccharide from infectious strains of Burkholderia cepacia. Carbohydr Res 335:45–54
Lodewyckx C, Vangronsveld J, Porteous F, Moore ERB, Taghavi S, Mezgeay M, Der Lelie D (2002) Endophytic bacteria and their potential applications. Crit Rev Plant Sci 21:583–606
Mendes R, Pizzirani-Kleiner AA, Araújo WL, Raaijmakers JM (2007) Endophytic bacteria from sugarcane: genetic and biochemical characterization of Burkholderia cepacia complex isolates. Appl Environ Microbiol 73:7259–7267
Michè L, Faure D, Blot M, Cabanne-GiuliI E, Balandreau J (2001) Detection and activity of insertion sequences in environmental strains of Burkholderia. Environ Microbiol 3:766–773
O’Sullivan DJ, O’Gara F (1992) Traits of fluorescent Pseudomonas spp. involved in suppression of plant root pathogens. Microbiol Mol Biol Rev 56:662–676
Oliveira ALM, Urquiaga S, Döbereiner J, Baldani JI (2002) The effect of inoculating endophytic N2-fixing bacteria on micropropagated sugarcane plants. Plant Soil 242:205–215
Parret AHA, Schoofs G, Proost R, De Mot R (2003) Plant lectin-like bacteriocin from rhizosphere-colonizing Pseudomonas isolate. J Bacteriol 185:897–908
Patten CL, Glick BR (1996) Bacterial biosynthesis of indole-3-acetic acid. Can J Microbiol 42:207–220
Perin L, Martínes-Aguilar L, Paredes-Valdez G, Baldani JI, Estrada-De-Los-Santos P, Reis VM, Caballero-Mellado J (2006a) Burkholderia silvatlantica sp. nov., a diazotrophic bacterium associated with sugarcane and maize. Int J Syst Evol Microbiol 56:1931–1937
Perin L, Martínes-Aguilar L, Castro-Gonzáles R, Estrada-De-Los-Santos P, Cabellos-Avelar T, Guedes HV, Reis VM, Caballero-Mellado J (2006b) Diazotrophic Burkholderia species associated with field-grown maize and sugarcane. Appl Environ Microbiol 72:3103–3110
Pugsley AP, Oudega B (1987) Methods for studying colicins and their plasmids. In: Hardy KG (ed) Plasmids: a practical approach. IRL Press, Oxford, pp 105–161
Reis VM, Estrada-De los Santos P, Tenorio-Salgado S, Vogel J, Stoffels M, Guyon S, Mavingui P, Baldani VLD, Schmid M, Baldani JI, Balandreau J, Hartmann A, Caballero-Mellado J (2004) Burkholderia tropica sp. nov., a novel nitrogenfixing, plant-associated bacterium. Int J Syst Evol Microbiol 54:2155–2162
Rodriguez H, Fraga R (1999) Phosphate solubilizing bacteria and their role in plant growth promotion. Biotechnol Adv 17:319–339
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Schroth MN, Hancook GH (1995) Disease suppressive soil and root colonizing bacteria science. Soil Biol Biochem 24:539–542
Schwyn B, Neilands JB (1987) Universal chemical assay for the detection and determination of siderophores. Anal Biochem 160:47–56
Serrato RV, Sassaki GL, Cruz LM, Pedrosa FO, Gorin PAJ, Lacomini M (2006) Culture conditions for the production of an acidic exopolysaccharide by the nitrogen-fixing bacterium Burkholderia tropica. Can J Microbiol 52:489–493
Siddiqui ZA (2005) PGPR: prospective biocontrol agents of plant pathogens. In: Siddiqui ZA (ed) PGPR: biocontrol and biofertilization. Springer, The Netherlands, pp 111–142
Steenhoudt O, Vanderleyden J (2000) Azospirillum, a freeliving nitrogen-fixing bacterium closely associated with grasses: genetic, biochemical and ecological aspects. FEMS Microbiol Rev 24:487–506
Tabacchioni S, Ferri L, Manno G, Mentasti M, Cocchi P, Campana S, Ravenni N, Taccetti G, Dalmastri C, Chiarini L, Bevivino A, Fani R (2008) Use of the gyrB gene to discriminate among species of the Burkholderia cepacia complex. FEMS Microbiol Lett 281:175–182
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) Software Version 4.0. Mol Biol Evol 24:596–1599
Vassilev N, Vassileva M, Nikolaeva I (2006) Simultaneous P-solubilizing and biocontrol activity of microorganisms: potentials and future trends. Appl Microbiol Biotechnol 71:137–144
Verma SC, Ladha JK, Tripathi AK (2001) Evaluation of plant growth promoting and colonization ability of endophytic diazotrophs from deep water rice. J Biotechnol 91:127–141
Welbaum G, Sturz AV, Dong Z, Nowak J (2004) Fertilizing soil microorganisms to improve productivity of agroecosystems. Crit Rev Plant Sci 23:175–193
Yamamoto S, Harayama S (1995) PCR amplification and direct sequencing of gyrB genes with universal primers and their application to the detection and taxonomic analysis of Pseudomonas putida strains. Appl Environ Microbiol 61:1104–1109
Yamamoto S, Harayama S (1998) Phylogenetic relationships of Pseudomonas putida strains deduced from the nucleotide sequences of gyrB, rpoD and 16S rRNA genes. Int J Syst Bacteriol 48:813–819
Acknowledgments
This work was supported by FAPESP (Foundation for Research Assistance of São Paulo State, Brazil) (Proc. 08/52407-9) and CNPq (National Council of Research, Brazil) awarded the fellowship to D. M. Luvizotto. We also thank Armando C. F. Dias and Maria C. Quecine and José A. Silva for technical supplying and critical discussions.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Luvizotto, D.M., Marcon, J., Andreote, F.D. et al. Genetic diversity and plant-growth related features of Burkholderia spp. from sugarcane roots. World J Microbiol Biotechnol 26, 1829–1836 (2010). https://doi.org/10.1007/s11274-010-0364-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11274-010-0364-0