Abstract
Benzene has a wide range of industrial applications, but it is also a major source of environmental pollution. The most eco-friendly/cost-effective method of remediation is biodegradation. In the present study, we used a variety of microbial strains in different combinations on a selection of substrate concentrations to determine the most effective degradation processes. Bacterial strains of pure culture (L4, N3, and N6) were isolated from oil sludge in both Luria–Bertani buffer (LB) and nutrient broth media, and identified by 16S-rRNA analysis (≥98% similarity). The degradation experiments were performed using different combinations of bacterial strains (L4, N3, N6, L4 + N3, L4 + N6, N3 + N6, and L4 + N3 + N6) in modified carbon-free media with different concentrations of benzene as a carbon source (60, 100, and 160 mg l−1) at 30 °C. The isolates of L4 (Acc no: FJ686821), N3 (FJ686825) and N6 (FJ868628) were identified as Bacillus spp. using 16S-rRNA gene sequence analysis. All combinations of isolates have the capacity to degrade benzene. However, the L4 + N3 combination was more efficient than the other mixed or single cultures. In the presence of N6 isolate, the degradation rate of benzene decreased, possibly due to inter- and/or intra species interaction amongst the bacteria. The kinetic parameters ‘K m’ of the Lineweaver–Burk regressions conducted as part of this experiment showed that the lower the level of K m was, the better the biodegradation achieved. The results of this study showed that the use of Bacillus strains in benzene decomposition is feasible. In addition, different strain combinations exhibited different degradation patterns, which are attributed to the most efficient mixed cultures of Bacillus spp.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Benzene has a wide range of industrial applications, but it is also a major source of environmental pollution. Accidental leaks and spills of gasoline and other petroleum products from storage tanks and other sources introduce significant quantities of toxic organic chemicals, mainly benzene, toluene, ethyl benzene, and xylene congeners (BTEX) into the environment (Dowd 1984). Biodegradation then plays an important role in their eco-friendly and cost-effective removal from soil and ground water.
Previous research and literature on this issue has mentioned the effectiveness of many bacterial strains with regard to benzene degradation, and the most commonly mentioned one is Pseudomonas spp., especially the Pseudomonas putida. The Pseudomonas putida strains have been examined in a considerable number of studies, some of which we have chosen to list here: P. putida (Shim and Yang 1999, 2002, 2006; Shim et al. 2002, 2005); P. putida 39/D (Kobal et al. 1973; Ziffer et al. 1973); P. putida biotype A (Mallakin and Ward 1996); P. putida BP18 (Baldwin et al. 2000); P. putida F1 (Olsen et al. 1994a, b, 1995; Reardon et al. 2000, 2002; Yu et al. 2001a, b; Gunaseelan and Cowan 2003); P. putida O1 (Oh and Bartha 1997; Oh and Choi 1997); P. putida ML2 (Tan and Mason 1990); P. putida mt2 (Gunaseelan and Cowan 2003), and so on. However, not only P. putida has been proven to be a popular research strain of Pseudomonas, but other strains of Pseudomonas have also been studied in benzene degradation, such as: P. aeruginosa (Marr and Stone 1961; Kim et al. 2003); P. alcaligenes (Oh and Choi 1997); P. fluorescens (Shim and Yang 1999, 2002, 2006; Shim et al. 2002, 2005); P. fragi B1 (Chang et al. 1993); P. maltophilia (Mallakin and Ward 1996); P. mendocina KR (Gunaseelan and Cowan 2003); P. paucimobilis Q1 (Furukawa et al. 1983); P. pickettii PKO1 (Olsen et al. 1995); P. stutzeri (Oh and Bartha 1997; Oh and Choi 1997); P. testosteron (Mallakin and Ward 1996); Pseudomonas sp. (Shirai 1986; Jean et al. 2002, 2008); Pseudomonas sp. ATCC 55595 (Collins and Daugulis 1999a, b); Pseudomonas sp. CFS215 (Alvarez and Vogel 1991; Olsen et al. 1995); Pseudomonas sp. G4 (Olsen et al. 1994a, b, 1995); Pseudomonas sp. GJ31 (Oldenhuis et al. 1989); Pseudomonas sp. W31 (Olsen et al. 1995), as well as other strains.
Although Pseudomonas sp. strains have been popular in benzene-degradation research, other genus strains have also been examined and reported on, such as: Acinectobacter calcoaceticus RJE74 (Winstanley et al. 1987); Acinetobacter johnsonii (Oh and Bartha 1997; Oh and Choi 1997); Alicycliphilus denitrificans strain (Weelink 2008); Arthrobacter sp. strain HCB (Alvarez and Vogel 1991); Aureobacterium esteroaromaticum (Oh and Bartha 1997; Oh and Choi 1997); Burkholderia cepacia G4 (Gunaseelan and Cowan 2003); Mycobacterium rhodochrous (Marr and Stone 1961); Nitrosomonas europea ATCC19718 (Hyman et al. 1985); Rhodococcus sp. strain 33 (Paje et al. 1997); Ralstonia pickettii PKO1 (Gunaseelan and Cowan 2003); Stenotrophomonas maltophilia T3-c (Lee et al. 2002), and so on. Jean et al. (2002) used Pseudomonas spp. for both laboratory experiments and numerical modeling to study the biodegradation and transport of benzene–toluene–xylenes (BTX) in a simulated semi-confined aquifer. Experimental and numerical comparisons indicate that the results by Monod degradation kinetics are more accurate than those by first-order degradation kinetics. Jean et al. (2008) used a glass sand tank in the laboratory to study the effect of inorganic nutrients (sulfate, phosphate, and ammonium chloride) on the aerobic biodegradation of benzene, toluene, and xylene (BTX) by Pseudomonas spp., revealing that the increase in nutrient levels resulted in enhanced bacterial growth and BTX degradation.
Whilst Pseudomonas strains have many applications in BTEX bioremediation, other Bacillus strains have been studied in petroleum bioremediation, such as Bacillus subtilis (Mulligan et al. 2001; Medina-Moreno et al. 2005; Toledo et al. 2008); B. subtilis O9 (Cubitto et al. 2004); B. subtilis DM-04 strain (Das and Mukherjee 2007); B. subtilis IFO3335 strain (Yasuda et al. 2003); B. subtilis ATCC 21332 (Whang et al. 2008); B. stearothermophilus (Natarajan et al. 1994); B. cereus (Medina-Moreno et al. 2005), and so on.
A great deal of research has been conducted on the use of different bacteria for BTEX remediation and many studies use Bacillus strains in petroleum carbon-chemical remediation, but there are no reports that specifically mention the use of Bacillus strains for BTEX or single substrate benzene remediation. Therefore, this study screens the dominant bacterial strains for pollutant biodegradation and evaluates their biodegradation efficiency in order to assess their efficacy. While Bacillus spp. was used in this study, benzene degradation and its degradation efficiency with other bacteria were compared. Then a variety of microbial strains were used in different combinations on various substrate concentrations in order to see which combination was most effective in biodegradation.
Materials and methods
Media and bacterial strains
Bacterial strains of pure culture were isolated from the oil sludge of a sewage treatment plant using different media for bacteria culturing, separation, and purification. Some experiments were then conducted to evaluate the potential degradation ability of these strains. Based on past experience, we used media that we have found to be the easiest one to cultivate the bacteria, namely LB (Luria–Bertani buffer) and NB (nutrient broth) media. The composition of the NB medium is beef extract 3 g, peptone 5 g, agar 15 g, and 1 liter of pure, sterile water, with an adjusted pH value of 7.0. The composition of the LB (Luria–Bertani buffer) medium is buffer trypton 5 g, yeast extraction 10 g, NaCl 5 g, and 1 liter of pure, sterile water. We used LB medium for separating and purifying six bacterial strains (LBOSB1~LBOSB6). We also used the NB medium for separating and purifying eight bacterial strains (NBOSB1~NBOSB8), in which the terms LB or NB, OS, and B correspond to the medium, oil sludge, and bacteria strains, respectively. L4, N3, N6, L4 + N3, L4 + N6, N3 + N6, and L4 + N3 + N6 refer to the bacterial isolates number growing in the modified carbon-free media. The non-carbon source medium was modified from a sulfate-reducing one (NaCl, 2.00 g; MgCl2·6H2O, 0.40 g; CaCl2·2H2O, 0.10 g; Na2SO4, 4.00 g; NH4Cl, 0.25 g; KH2PO4, 0.20 g; KCl, 0.50 g; selenite solution, 1.00 ml; sulfide solution, 7.50 ml; trace elements, 1.00 ml). The medium was used for benzene degradation experiments, and carbon source chemicals were removed while retaining the other chemicals with nutritional value.
Organic chemicals
The substrate benzene that was used in this study was reagent grade or better obtained from Fisher Scientific (Fisher Scientific International Company, Loughborough, UK). The properties used for the chemical included dimensionless Henry’s Law coefficients of 0.185 at 20 °C (Turner 1995), and a COD’s (mass of oxygen required to oxidize a unit mass of the compound, g g−1) value of 3.076.
Screening of bacteria
We used oil sludge that was taken from a sewage treatment plant of the China Petroleum Refinery Company in Kaohsiung, Taiwan. The oil sludge was used for culturing the bacteria. Because the bacteria had lived in aromatic substances for a long time, it had evolved the capacity to degrade aromatic organic compounds. First, 5 g of sludge and 100 ml of sterile water were placed into a serum bottle. After 30 min of shaking on a roll bed, we took the supernatant from the upper-part and added 5 ml to a serum bottle that had 250 ml of enriched medium inside. After that, we cultured the bacteria for 3 days at 30 °C constant temperature. We extracted the bacteria in the log growth phase and repeated the steps described above for culturing the bacteria. We repeated the culturing steps at least three times and subsequently isolated the bacteria in an agar plate in log growth phase conditions at 30 °C. The single pure cultures of bacterial strain were isolated by the pour-plate technique (Dubey and Maheshwari 2005; Camper and McFeters 1979). The selected single colony inoculated into a flask that contains a non-carbon medium and benzene, and was cultured separately. Each single strain was put into three sets of flasks and designated a group (n = 3). The strains in the same group that had the lowest benzene residues were considered as the most suitable strains. After strain selection, we sent the strains for identification and preserved them in a refrigerator.
A subculture of the enrichment was used to measure the most dominant bacterial concentration using the pour-plate technique (Camper and McFeters 1979). Three bacterial isolates were identified in our study as the genus Bacillus spp. by the 16S rRNA gene sequence analysis (details are shown, below). The 16S rRNA gene sequences of the three bacterial isolates were aligned with those of the reference strains with more than the ≈99% nucleotide similarity, revealing that the isolates were Bacillus spp. including B. pumilus (Acc no: FJ686821; L4), B. pumilus (Acc no: FJ686825; N3) and B. pumilus (Acc no: FJ868628; N6)
Domesticated strains
The isolated bacteria were added to benzene in a flask and shaken on a roll bed. Subsequently, we cultured the bacteria for 30 days at a constant temperature of 30 °C. After culturing, we refrigerated the culture at 4 °C for preservation.
Degradation experiment
After preparing the flasks, we added 200 ml of non-carbon medium and inoculated the pure culture bacteria strain. At the same time, we added benzene as the carbon source. Three different initial concentrations (60, 100, and 160 mg l−1) were used for the bacterial growth and degradation experiments. After that, the flasks were placed on a roll bed and shaken 30 min and then kept at a constant 30 °C for cultivation. Sampling was conducted under air-tight sealed conditions for sample extractions using syringes to sample 1 ml of degraded solution at the sampling times of 0, 6, 12, 18, 24, 33, 42, 51, 63, 75, 87, 99 and 111 h. The degraded solution was then measured for benzene concentration and cell mass. In this study, we selected L4, N3, and N6 strains for single and mixed-culture experiments. These three strains were used to create seven combinations (L4, N3, N6, L4 + N3, L4 + N6, N3 + N6, and L4 + N3 + N6) of benzene degradation experiments. The benzene concentrations were measured by gas chromatography with a flame ionization detector (Varian 3800). To determine the amount of benzene evaporation, a blank test was also performed using the same conditions but without bacterial inoculation.
Analytical methodology
Benzene concentrations were analyzed using the Tekmar 3000 purge and trap concentrator connected to a Varian 3300 gas chromatograph (GC). The GC was equipped with a flame ionization detector (FID) and a 30 cm long EC-WAX capillary column coated with 1–2 μm thick methyl silicon. The initial temperature was maintained at 150 °C for 5 min and then raised to 200 °C at a rate of 10 °C min−1. The temperature was then maintained at 200 °C for 5 min. The total GC sampling time was 15 min. The retention times were 3 min 25 s for benzene in the GC detection system (Experimental error of GC for benzene = ±2.324%). Nitrogen served as the carrier gas, while oxygen and hydrogen served as the fuel gases for the FID. The initial concentration of the benzene was 60, 100 and 15 mg l−1.
Identification of bacteria
DNA isolation
Bacterium DNA was extracted using the DNA/RNA extraction kit (Viogen, Sunnyvale, CA, USA) according to the manufacturer’s instructions. A detailed description of the procedure for DNA isolation is provided in the relevant instruction manual.
Polymerase chain reaction (PCR) amplification
The 16S rRNA gene region of the DNA was amplified by the PCR using a pair of universal primers. The oligonucleotide sequences for the forward primer (16S 8F) and the reverse primer (16S 1510R) were 5′-AGAGTTTGATCCTGGCTCAG-3′ and 5′-GGTTACCTTGTTACGACTT-3′, respectively (Mission Biotech Co. Ltd., San Francisco, CA, USA). Each 50 μl PCR reaction solution contained: 10 μl DNA template, 29.25 μl of double distilled water (11.2 μM), 5 μl 10x reaction buffer, 5 μl MgCl2 (15 mm), 1.25 μl dNTP mix (10 mm), 2 μl of forward primer (10.6 μm), 2 μl of reverse primer (11.2 μm), and 0.5 μl (5 U μl−1) of Taq polymerase (Promega, Madison, WI, USA). The reaction was carried out on a MJ Thermal Cycler (Genetic Research Instruments, Braintree, Essex, UK) programmed to have the sequence of (1) a first cycle of denaturation at 94 °C for 5 min (2) 30 cycles of denaturation at 94 °C for 1 min, annealing at 55 °C for 1 min 30 s, and extension at 72 °C for 2 min, and then (3) 72 °C stabilization of PCR products for 10 min and storage at 4 °C.
Nucleotide sequencing
All PCR products were purified from an agarose gel using the PCR product purification kit (Viogene, Sunnyvale, CA, USA). For each 16S rRNA gene fragment, both strands were cycle-sequenced using the Taq Dye Deoxy Terminator Cycle Sequencing Kit (Applied Biosystem, Foster City, CA, USA). Products of the cycle sequencing reactions were run on the ABI 3730 automated DNA sequencer (Applied Biosystem).
Determination of biodegradation kinetics model parameters
The determination of growth kinetics model parameters is influenced by many factors, especially parameter identifiability, culture history, and the assay procedure used (Grady et al. 1996; Jannasch and Egli 1993; Kova′rova′-Kovar and Egli 1998). Parameter identifiability was determined by establishing a sufficiently low inoculum size and relatively high initial substrate concentration, and culture history effects were examined and found to be minimal (details below). The last factor concerns the mode of cell cultivation (e.g., batch, continuous) that was used to obtain the data to which a model is regressed. Continuous cultivation offers the ability to provide a constant environment to the cells and to maintain cells at low substrate concentrations, and may result in more accurate estimates of affinity constants (e.g., K m in the Monod model) than batch cultivation does. However, estimation of the maximum specific growth rate may be less accurate in continuous cultivations, and the results may be affected by wall growth and mutation over the long cultivation periods required by this technique. Furthermore, it may be difficult to maintain continuous cultures under conditions of strong substrate or product inhibition. For the work presented here, we chose to use batch cultures to measure biodegradation kinetics, evaluate models, and determine model parameters. In particular, we wanted to be able to observe substrate inhibition, patterns of substrate consumption, and related physiological changes. Since each of the three monoaromatics was used as the sole source of carbon and energy and was the limiting growth substrate, all models consisted of two parts: (1) an equation for the specific growth rate, “μ”, as a function of substrate concentration, and (2) an equation for substrate depletion. Various models for μ(S) were tested. The velocity (μ) of enzyme-catalysed reactions is hyperbolically related to the substrate concentration ([S]) through the Michaelis–Menten equation:
where μmax and K m are the kinetic constant characteristics of the particular enzyme and substrate, respectively, μmax is the maximum specific growth rate, and K m is the Monod half-saturation constant.
These constants are estimated in experiments in which [S] is varied and μ measured. They are usually extracted by some linear transformation of the data, such as the Lineweaver–Burk or Hanes plots. However, these transformations are unsatisfactory, as they can result in artificial weighting of the data leading to erroneous estimates of μmax and K m. A better approach is to fit the data directly to the best possible hyperbola. This is achieved by using non-linear regression analysis. Lineweaver–Burk or Hanes plots, and the kinetic constants derived from them are also provided for comparison.
The linear plot equations are as follows:
Hanes Plot:
Lineweaver–Burk Plot:
For any pair of μ and [S] values there is an infinite number of μmax and K m values which will satisfy the Michaelis–Menten equation. In the parameter space plot, μmax and K m are treated as the variables. For each data pair, a line is drawn connecting μ on the ordinate axis and [S] on the abscissa. For n data pairs there will be n such lines and these will have n(n − 1)/2 intersections. The μ and [S] values at the intersections correspond respectively to the μmax and K m values satisfying the Michaelis–Menten equation for the corresponding two pairs of [S]/μ data. The best estimates of μmax and K m for the data set as a whole are taken as the medians of the points of intersection.
Results and discussion
Isolates and identification
All three isolates (L4, N3 and N6) were identified as Bacillus spp. (Accession no: FJ686821, FJ686825 and FJ868628 respectively) using 16S rRNA gene sequence analysis. All combinations of isolates that are attributed to the Bacillus genus can degrade benzene. All L4, N3 and N6 Bacillus species are aerobic, rod-shaped, Gram-positive bacteria which are ubiquitous in nature. Gram-stained cells, ≈1 μm wide, ≈1.5–2.5 μm long, are arranged singly or in short chains. 16S rRNA sequence analysis is a fast and accurate method to identify the L4, N3 and N6 phylogeny position. In this study, full-length (about 1500 bp) 16S rRNA genes were sequenced and used to construct a phylogenetic tree (Fig. 1a–c). The results showed that the tested strain L4 was classified in the Bacillus genera, and the nucleotide similarities between L4 (Accession no: FJ686821) and B. pumilus (EU660356), B. pumilus (FJ234439), and B. pumilus (EU869282) were 99.0, 99.0 and 99.0, respectively (Fig. 1a). Therefore, L4 (FJ686821) is closely related to B. pumilus, and was not further identified, but designated as Bacillus spp. The isolated strain N3 was classified in the Bacillus genera; the nucleotide similarities between N3 (Accession no: FJ686825) and B. pumilus (EU596537), B. subtilis (EF563825) and B. altitudinis (AJ831842) were 99.0, 98.0, and 97.5% respectively. Therefore, N3 (FJ686825) is closely related to B. pumilus (Fig. 1b). The results found that the tested strain N6 was classified in the Bacillus genera; the nucleotide similarities between N6 (Accession no: FJ868628) and B. pumilus (EU869273), B. subtilis (EF563825), B. subtilis (EU931563) were 99.5, 99.4 and 98.5%, respectively. Therefore, N6 (FJ868628) is closely related to B. pumilus (Fig. 1c).
Phylogenic relationship of three different isolates belonging to Bacillus pumilus: a L4 (Acc no: FJ686821) b N3 (Acc no: FJ686825) and c N6 (Acc no: FJ868628). The calculation of evolutionary distance and classification of phylogenitic relationship were determined using the Jukes–Cantor distance and neighbor-joining algorithm
Degradation and growth results
The biodegradations of benzene are shown in Fig. 2a–g and the bacterial growth curves with different initial concentrations (60, 100, and 160 mg l−1) are shown in Fig. 2a–c. The degradation results reflecting the relationship between incubation time and benzene concentration reveal that single-strain cultures L4 and N3 could degrade benzene to a low-concentration in a fast time, while N6 degraded benzene slowly (Fig. 2a–c). The 100 mg l−1 benzene was completely degraded by the N3 and N6 isolates within 75 h incubation time, but was completely degraded within 63 h by the L4 isolate. At a low concentration, such as 60 mg l−1, the complete degradation of benzene was found to be similar for all selected isolates. However, at a higher concentration (160 mg l−1), the N3 isolate was more efficient with regard to degrading the benzene than the other two single isolates (L4 and N6). In addition, the isolate L4 was more efficient at degrading the benzene at the initial concentration of 100 mg l−1 than N3 and N6 do. Similarly, N3 is more efficient with regard to degrading the benzene at an initial concentration of 160 mg l−1 when compared to other isolates, such as L4 and N3.
The degradation of benzene and bacterial growth at 60, 100 and 160 mg l−1 of initial benzene concentrations using single and mixed culture of Bacillus pumilus: a L4, b N3, c N6, d L4 + N3, e L4 + N6, f N3 + N6, g L4 + N3 + N6 and h blank experiment. Error bars are represented as mean ± SE of n = 3; p < 0.05
The mixed cultures L4 + N3 exhibited greater efficacy in degrading benzene than any other mixture or single culture examined (Fig. 2a–g) and achieved total degradation (100%) at the initial concentrations of 60, 100 and 160 mg l−1 after 42, 51 and 75 h, respectively. It is evident that in this case a mixture of two bacterial strains has improved the efficacy of benzene degradation, and that in some cases (i.e., L4 + N6, N3 + N6, L4 + N3 + N6), the benzene degradation rates actually declined over single strain comparisons (Fig. 2e–g). It seems that N6 has some type of inhibitory effect on benzene degradation, perhaps due to inter- and/or intra-species interaction of the bacteria. After 111 h of incubation time, the degradation percentage was 100% for single and mixed cultures of various combinations, except L4 + N3 + N6 (Fig. 2g). However, the mixed-culture combination L4 + N3 + N6 had the best degradation efficiency, up to 100% after 51 h at a low initial concentration of 60 mg l−1. For the combinations L4, N3, N6, and L4 + N3, when the initial concentration increased, the degradation percentage also rose. In contrast, For L4 + N6, N3 + N6 and L4 + N3 + N6, when the initial concentration increased, the degradation percentage slightly decreased. For the single-strain cultures L4, N3, and N6, the higher initial concentration had the larger degradation percentage. Similar results were found from the mixed culture of L4 + N3. For the mixed cultures of L4 + N6, N3 + N6, L4 + N3 + N6, the higher the initial concentration, the lower the degradation percentage. This result shows that inhibition caused by N6 is more obvious in a high concentration environment. The blank tests for benzene evaporation showed that the substrate concentrations did not change significantly during the experiments (Fig. 2h).
Degradation patterns
The initial concentration of benzene and rate of degradation results revealed that these seven combinations could be grouped into three types of degradation pattern. L4, N3, and L4 + N3 comprise the first category, while N6 alone is the second category, and L4 + N6, N3 + N6 as well as L4 + N3 + N6 are the third category (Fig. 2a–g).
The degradation patterns for the first category (L4, N3, and L4 + N3) are characterized as: (1) When the initial concentrations of benzene increased within the range of 160 mg l−1, the amount of benzene degradation (mg l−1 of benzene concentration was degraded by bacteria) also increased efficiently (complete degradation, 100%). (2) Bacteria strains in this category had the ability to degrade benzene to low concentrations within a short period of time, and their degradation curves were concave downward. (3) Bacterial growth lagged in the beginning and increased rapidly. L4, N3 and L4 + N3 are attributed to the first category. Degradation behavior of this type reflects the gradual increase in degradation efficiency (%) (i.e., the percentage of benzene that was degraded by the bacteria compared to the initial concentration of benzene in mg l−1) with increased concentration. For the low concentration (60 mg l−1) of benzene substrate, complete degradation (100%) occurred within 35–40 h incubation, whereas at higher initial concentrations, such as 100 and 160 mg l−1, the complete degradation (100%) occurred at 50–75 h and 75–100 h, respectively. In a comparison of L4, N3, and L4 + N3, the results show that the degradation efficiency of the mixed culture L4 + N3 was better than that of a single culture of L4 or N3. This could indicate that L4 and N3 exhibit some level of co-operative behavior. The degradation patterns of L4, N3, and L4 + N3 show that these single and mixed strains can degrade benzene very rapidly, and their degradation curves are concave downward. The bacterial cultures in the first category demonstrate excellent growth results. From the bacteria growth curve, we found that bacteria growth initially exhibits a lag time and then exhibits a rapid growth spurt. For the bacterial cultures in the first category, greater concentration results in better growth conditions for the bacteria.
The characteristics of the second category such as the N6 isolate, were that: (1) When the initial concentration increased, the actual amount of benzene degraded also rose. However, the rate of degradation was slightly less than that of first category. (2) The bacterial isolate degraded benzene gradually to low concentrations, and the degradation curve was flatter than that of the first category. (3) The bacterial growth pattern was such that the growth during the initial lag time was slow, and then proceeded rapidly. Degradation behavior for this type of bacteria is similar to that of the first category at low concentration. For the initial concentration of benzene substrate at 60 mg l−1, complete degradation (100%) was noticed with 40 h incubation, whereas for concentrations of 100 and 160 mg l−1 the complete degradations (100%) were noticed after 80 and 110 h, respectively. The degradation curve of N6 presents a slow degradation rate and a relatively flat curve. The bacterial growth pattern, however, exhibited a relatively short initial lag time and subsequently rapid growth, revealing an excellent growth rate. A greater substrate concentration resulted in better growth of the bacteria.
The characteristics of the third category were that: (1) When the initial concentration increased, the amount of benzene degradation increased, although the rate of degradation was significantly less than for the other two categories; (2) The bacterial growth and the benzene degradation were less noticeable in both single and mixed cultures, and the degradation curve was flatter than that for the first and second categories. (3) The bacterial growth was inhibited perhaps due to intra- and/or inter-bacterial species interaction. The mixed cultures of isolates, such as L4 + N6, N3 + N6, and L4 + N3 + N6, are attributed to the third category. The degradation behavior of this type showed a gradual decrease in efficiency as the initial concentration increased. For 60 mg l−1 benzene substrate, complete degradation (100%) was achieved within 50 h for all mixed cultures. For 100 and 160 mg l−1, the incubation times were increased from 75 to 100 h and >100 h, respectively. The degradation curves for L4 + N6, N3 + N6, and L4 + N3 + N6 are flatter and thus show that the degradation rate was slower. The degradation curves of the different concentrations of the three mixed cultures had obvious differences. Bacterial growth and the degradation rate were slightly lower for the third category compared to the previous two. In comparing the three types we found some differences, such as the degradation efficiency of the first category was good, whereas while the degradation rate and bacterial growth of the second category were both good, they were less so than those of first. In addition, the degradation efficiency of the third category was lower than that of the other two, and the degradation rate and bacterial growth were also slower. It is noted that a co-operative relationship between the L4 and N3 isolates could enhance the degradation efficiency of benzene and bacterial growth. In addition, N6 to L4 or N3 exhibited a competitive relationship, suggesting that competition between N6 and L4 or N3 could inhibit the degradation efficiency of benzene and also bacterial growth.
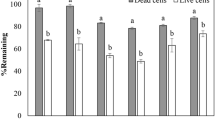
The Box-and-Whisker plots of the measured benzene clearly indicate the biodegradation rates by bacteria at different levels of nutrient concentration. The results indicate that an increase in the concentration of nutrients would promote benzene degradation (Fig. 3a–b). The bacterial isolates of a single culture, such as L4 or N3, are more efficient than N6 (Fig. 3a). Similarly, the mixed culture of L4 + N3 is more efficient than that of L4 + N6, L4 + N3 + N6, N3 + N6 (Fig. 3b). In a comparison of each single and mixed culture, the mixed cultures other than N3 + N6, are more efficient than each single culture, except for N3.
Biodegradation of benzene with kinetics model parameters
The benzene biodegradation experiments are shown in Fig. 2a–g. The Monod model was used to describe the growth kinetics and to model the consumption (biodegradation) of benzene. The determination of the growth kinetics model parameters was influenced by many factors, especially parameter identifiability, culture history, and the assay procedure used. Parameter identifiability was set up by establishing a sufficiently low inoculum size and relatively high initial substrate concentration, and culture history effects were examined and found to be minimal. The last factor concerns the mode of cell cultivation (e.g., batch, continuous). Continuous cultivation offers the ability to provide a constant environment to the cells and to maintain them at low substrate concentrations, thereby resulting in more accurate estimates of affinity constants (e.g., K m in the Monod model) than is possible with batch cultivation. For the work presented here, we chose to use batch cultures to measure biodegradation kinetics, to evaluate models, and to determine model parameters. We used the degradation data for Lineweaver–Burk regression analysis and obtained the bacterial kinetic parameters μmax and K m. The K m value (Table 1) of N3 (K m = 10.11 mg l−1) was greater than that for two other single cultures, namely L4 (K m = 4.26 mg l−1) and N6 (K m = 1.618 mg l−1). The K m values for mixed-culture combinations of L4 + N3 (K m = 8.158 mg l−1) and L4 + N6 (K m = 5.169 mg l−1) were greater than those of the single and mixed cultures L4, N3, N6, N3 + N6 (K m = 0.645 mg l−1) and the L4 + N3 + N6 (K m = 2.251 mg l−1) combination, which had the lowest K m. Lineweaver–Burk plot showed that the mixed-culture isolates had a better degradation ability for benzene than the single isolates did, while inhibition also occurred with N3 + N6. Also Lineweaver–Burk regression analysis indicated that the benzene degradation in all combinations of mixed cultures could be inhibited in the presence of the N6 isolate. Comparing these results with those in the existing studies in kinetic constant characteristic of the particular substrate (K m), P. putida F1 had a K m value of 0.73 mg l−1, while P. putida O1 and P. fragi B1 had K m values of 0.44 and 0.335 mg l−1, respectively. The K m values from Bacillus spp. of this study are the same or greater than those shown above, suggesting that Bacillus spp. also has good degradation ability for benzene.
Conclusions
The results of this study show that the use of Bacillus strains for benzene degradation is feasible. Different strain combinations have been examined in terms of their degradation patterns. The results show that mixed cultures of Bacillus spp. exhibit the greatest efficacy in benzene degradation. All combinations of three isolates, namely L4 (Acc no: FJ686821), N3 (Acc no: FJ686825), and N6 (Acc no: FJ868628), have the capacity to degrade benzene. However, the L4 + N3 combination is more efficient than the other mixed or single cultures. The degradation efficiencies of benzene were in the order of N3 > L4 + N3 > L4 + N6 > L4 > L4 + N3 + N6 > N6 > N3 + N6. In the presence of N3 isolate, the degradation of benzene increases. However, in the presence of N6 isolate, the degradation rate of benzene decreases, possibly due to inter- and/or intra-species interaction amongst the bacteria. The results presented in this study of a mixed culture combination of two bacterial strains isolated from a benzene-contaminated site might provide data that will be of practical use with regard to the degradation of benzene in real field sites. In future work, it would be useful to evaluate the possibility of utilizing mixed bacterial strains rather than a pure cultured strain for the bioremediation of benzene-contaminated soils and waters.
Abbreviations
- μ:
-
Specific growth rate
- μmax :
-
The maximum specific growth rate
- [S]:
-
Substrate concentration (mg l−1)
- K m :
-
Kinetic constant characteristic of the particular enzyme and substrate
References
Alvarez PJJ, Vogel TM (1991) Substrate interactions of benzene, toluene, and para-xylene during microbial degradation by pure cultures and mixed culture aquifer slurries. Appl Environ Microbiol 57:2981–2985
Baldwin BR, Mesarcit MB, Nies L (2000) Broad substrate specificity of naphthalene- and biphenyl-utilizing bacteria. Appl Microbiol Biotechnol 53:748–753
Camper AK, McFeters GA (1979) Chlorine injury and the enumeration of waterborne coliform bacteria. Appl Environ Microbiol 37:633–641
Chang MK, Voice TC, Criddle CS (1993) Kinetics of competitive inhibition and cometabolism in the biodegradation of benzene, toluene, and pxylene by two pseudornonas isolates. Biotechnol Bioeng 41:1057–1065
Collins LD, Daugulis AJ (1999a) Benzene/toluene/p-xylene degradation. Part I. Solvent selection and toluene degradation in a two-phase partitioning bioreactor. Appl Microbial Biotechnol 52:354–359
Collins LD, Daugulis AJ (1999b) Benzene/toluene/p-xylene degradation. Part II. Effect of substrate interactions and feeding strategies in toluene/benzene and toluene/p-xylene fermentations in a partitioning bioreactor. Appl Microbiol Biotechnol 52:360–365
Cubitto MA, Morán AC, Commendatore M, Chiarello MN, Baldini MD, Siñeriz F (2004) Effects of Bacillus subtilis O9 biosurfactant on the bioremediation of crude oil-polluted soils. Biodegradation 15:281–287
Das K, Mukherjee AK (2007) Crude petroleum-oil biodegradation efficiency of bacillus subtilis and Pseudomonas aeruginosa strains isolated from a petroleum-oil contaminated soil from north-east India. Bioresour Technol 98:1339–1345
Dowd RM (1984) Leaking underground storage tanKm. Environ Sci Technol 18:309A
Dubey RC, Maheshwari DK (2005) Practical microbiology. 1st edn (Reprint). S. Chand and Company Ltd., New Delhi, pp 10–25
Furukawa K, Simon JR, Chakrabarty AM (1983) Common induction and regulation of biphenyl, xylene/toluene, and salicylate catabolism in Pseudomonas paucimobilis. J Bacteriol 154:1356–1362
Grady CPL Jr, Smets BF, Barbeau DS (1996) Variability in kinetic parameter estimates: a review of possible causes and a proposed terminology. Water Res 30:742–748
Gunaseelan A, Cowan R (2003) Substrate inhibition kinetics for toluene and benzene degrading pure cultures and a method for collection and analysis of respirometric data for strongly inhibited cultures. Biotechnol Bioeng 83:798–809
Hyman MR, Sansome-Schmith AW, Shears JH, Wood PM (1985) A kinetic study of benzene oxidation to phenol by whole cells of Nitrosomonas europea and evidence for the further oxidation of phenol to hydroquinone. Arch Microbiol 143:302–306
Jannasch H, Egli T (1993) Microbial growth kinetics: a historical perspective. Antonie van Leeuwenhoek 63:213–224
Jean J-S, Tsai C-L, Ju S-H, Tsao C-W, Wang S-M (2002) Biodegradation and transport of benzene, toluene, and xylenes in a simulated aquifer: comparison of modeled and experimental results. Hydrol Process 16:3151–3168
Jean J-S, Lee M-K, Wang S-M, Chattopadhyay P, Maity JP (2008) Effects of inorganic nutrient levels on the biodegradation of benzene, toluene, and xylene (BTX) by Pseudomonas spp. in a laboratory porous media sand aquifer model. Bioresour Technol 99:7807–7815
Kim SB, Park CH, Kim DJ, Jury WA (2003) Kinetics of benzene biodegradation by Pseudomonas aeruginosa: parameter estimation. Environ Toxicol Chem 22:1038–1045
Kobal VM, Gibson DT, Davis RE, Garza A (1973) X-ray determination of the absolute stereochemtstry of the initial oxidation product formed from toluene by Pseudomonas putida 39/D. J Am Chem Soc 95:4420–4421
Kova′rova′-Kovar K, Egli T (1998) Growth kinetics of suspended microbial cells: from single-substrate-controlled growth to mixed-substrate kinetics. Microbiol Mol Biol Rev 62:646–666
Lee EY, Jun YS, Cho KM, Ryu HW (2002) Degradation characteristics of toluene, benzene, ethylbenzene, and xylene by Stenotrophomonas maltophilia T3-c. J Air Waste Manage Assoc 52:400–406
Mallakin A, Ward O (1996) Degradation of BTEX compounds in liquid media and in peat biofilters. J Ind Microbiol 16:309–318
Marr EK, Stone RW (1961) Bacterial oxidation of benzene. J Bacteriol 18:425–430
Medina-Moreno SA, Huerta-Ochoa S, Gutierrez-Rojas M (2005) Hydrocarbon biodegradation in oxygen-limited sequential batch reactors by consortium from weathered, oil-contaminated soil. Can J Microbiol 51:231–239
Mulligan CN, Yong RN, Gibbs BF (2001) Surfactant-enhanced remediation of contaminated soil: a review. Eng Geol 60:371–380
Natarajan MR, Lu Z, Oriel P (1994) Cloning and expression of a pathway for benzene and toluene from Bacillus stearothermophilus. Biodegradation 5:77–82
Oh YS, Bartha R (1997) Construction of a bacterial consortium for the biofiltration of benzene, toluene and xylene emissions. World J Microbiol Biotechnol 13:627–632
Oh YS, Choi SC (1997) Characterization of BTX-degrading bacteria and identification of substrate interactions during their degradation. J Microbiol 35:193–199
Oldenhuis R, Kuijk L, Lammers A, Janssen DB, Witholt B (1989) Degradation of chlorinated and non-chlorinated aromatic solvents in soil suspensions by pure bacterial cultures. Appl Microbiol Biotechnol 30:211–217
Olsen RH, Kukor JJ, Kaphammer B (1994a) A novel toluene-3-monooxygenase pathway cloned from Pseudornonas pickettii PKO1. J Bacteriol 176:3749–3756
Olsen RH, Mikesell MD, Kukor JJ (1994b) Enumeration and characterization of BTEX-degrading bacteria from hypoxic environments functional with mixed electron acceptors. Res Microbiol 145:47–49
Olsen RH, Mikesell MD, Kukor JJ, Byrne AM (1995) Physiological attributes of microbial BTEX degradation in oxygen-limited environments. Environ Health Perspect 103:49–51
Paje MLF, Neilan BA, Couperwhite L (1997) A Rhodococcus species that thrives on medium saturated with liquid benzene. Microbiology 143:2975–2981
Reardon KF, Mosteller DC, Rogers JB (2000) Biodegradation kinetics of benzene, toluene, and phenol as single and mixed substrates for Pseudomonas putida F1. Biotechnol Bioeng 69:385–400
Reardon KF, Mosteller DC, Rogers JB, DuTeau NM, Kim K-H (2002) Biodegradation kinetics of aromatic hydrocarbon mixtures by pure and mixed bacterial cultures. Environ Health Perspect 110:1005–1011
Shim H, Yang ST (1999) Biodegradation of benzene, toluene, ethylbenzene, and o-xylene by a coculture of pseudomonas putida and pseudomonas fluorescens immobilized in a fibrous-bed bioreactor. J Biotechnol 67:99–112
Shim H, Yang ST (2002) BTEX removal from contaminated groundwater by a co-culture of pseudomonas putida and pseudomonas fluorescens immobilized in a continuous fibrous-bed bioreactor. J Chem Technol Biotechnol 77:1308–1315
Shim H, Yang ST (2006) Kinetics of BTEX biodegradation by a coculture of Pseudomonas putida and Pseudomonas fluorescens. J Environ Eng Manage 16:309–317
Shim H, Shin E, Yang ST (2002) A continuous fibrous-bed bioreactor for BTEX biodegradation by a co-culture of Pseudomonas putida and Pseudomonas fluorescens. Adv Environ Res 7:203–216
Shim H, Hwang B, Lee SS, Kong SH (2005) Kinetics of BTEX biodegradation by a coculture of Pseudomonas putida and Pseudomonas fluorescens under hypoxic conditions. Biodegradation 16:319–327
Shirai K (1986) Screening microorganisms for catechol production from benzene. Agric Biol Chem 50:2875–2880
Tan HM, Mason JR (1990) Cloning and expression of the plasmid-encoded benzene dioxygenase genes from P. putida ML2. Fems Microbtol Lett 72:259–264
Toledo FL, Gonzalez-Lopez J, Calvo C (2008) Production of bioemulsifier by Bacillus subtilis, Alcaligenes faecalis and Enterobacter species in liquid culture. Bioresour Technol 99:8470–8475
Turner LH (1995) Vapor-liquid equilibrium of highly non-ideal aqueous organic system. Ph.D. thesis, Rutgers university, New jersey, USA
Weelink SAB (2008) Degradation of benzene and other aromatic hydrocarbons by anaerobic bacteria. Ph.D. thesis, Wageningen university, Wageningen, Netherlands
Whang LM, Liu PW, Ma CC, Cheng SS (2008) Application of biosurfactants, rhamnolipid, and surfactin, for enhanced biodegradation of diesel-contaminated water and soil. J Hazard Mater 151:155–163
Winstanley C, Taylor SC, Williams PA (1987) pWW174: a large plasmid from Acinetobacter calcoaceticus encoding benzene catabolism by the ß-ketoadipate pathway. Mol Microbiol 1:219–227
Yasuda Y, Katoh S, Inoue Y, Shiomi N (2003) A bioremediation method based on self-immobilization of cells in shallow layer of soil. J Chem Eng Jpn 36:216–219
Yu H, Kim BJ, Rittmann BE (2001a) The roles of intermediates in biodegradation of benzene, toluene, and p-xylene by Pseudomonas putida F1. Biodegradation 12:455–463
Yu H, Kim BJ, Rittmann BE (2001b) A two-step model for the kinetics of BTX degradation and intermediate formation by Pseudomonas putida F1. Biodegradation 12:465–475
Ziffer H, Jerina DM, Gibsox DT, Kobal Val M (1973) Absolute stoichiochemistry of the (+)-cts-l, 2-dihydroxy-3-methylcyclohexa-3.5-diene produced from toluene by Pseudomonas putida. J Am Chem Soc 95:4048–4049
Acknowledgments
This work was supported by the National Science Council in Taiwan (NSC No. NSC 96-2116-M-006-003).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Liu, JH., Maity, J.P., Jean, JS. et al. Biodegradation of benzene by pure and mixed cultures of Bacillus spp.. World J Microbiol Biotechnol 26, 1557–1567 (2010). https://doi.org/10.1007/s11274-010-0331-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11274-010-0331-9