Abstract
Deschampsia antarctica Desv. is the only gramineae capable of colonizing the Antarctic due to the region’s extreme climate and soil environment. In the present research, bacteria colonizing the rhizospheric soil of D. antarctica were isolated and characterized. The soil studies showed that D. antarctica possesses a wide spectrum of psychrotolerant bacteria with extensive and varied antibiotic resistance, as well as heavy metal tolerance. The bacterial strains isolated from the rhizosphere of D. antarctica also produced a diverse pattern of enzymes. Based on the strain identification with partial characterization of the 16S rRNA gene, the majority of the isolates correspond to different Pseudomonas species, and species of the genus Flavobacterium sp. and Arthrobacter sp. The isolated strains collected from this research constitute a unique collection for future, more detailed taxonomic analysis and physiological characterization, contributing to the search for potential biotechnological uses. These findings and others have great potential for developing new biotechnological products from Antarctic microorganisms.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Polar areas are the coldest regions of the world, containing 99% of the ice on the planet (about 13% in the Arctic and 86% in the Antarctic). The majority of the Antarctic continent is covered by permanent ice and snow, with only 2% available for colonization by plants and animals. Most of the ice and snow-free land is found along the Antarctic Peninsula, its associated islands, and the coastal region around the rest of the Antarctic continent. These are the areas where most of the Antarctic vegetation is found, composed predominantly of lichens and bryophytes. Only two vascular plants, Deschampsia antarctica Desv. and Colobanthus quitensis (Kunth) Bartl., have been able to colonize some of the coastal areas (Casaretto et al. 1994; Alberdi et al. 2002).
Antarctic soils, especially those of the coastal areas, are characterized by a high content of coarse mineral particles and total organic carbon, a low C/N ratio, acidic pH, and frequent nutrient enrichment due to the influence of sea spray and guano from seabirds (Beyer et al. 2000).
Many researchers are interested in D. antarctica due to its ability to live in these extreme conditions, and there is an increasing number of physiological and biochemical studies being undertaken to better understand the mechanisms by which this plant is capable of colonizing the Antarctic environment. Studies have shown that D. antarctica has maximum photosynthetic activity at 13°C and is capable of maintaining 30% of its photosynthetic capacity at 0°C (Edwards and Lewis-Smith 1998). These qualities are among the most important factors contributing to the plant’s survival in the Antarctic.
In contrast to the rather limited diversity of plants and animals found in the Antarctic, the microbial diversity of this continent has been shown to be highly diverse. Apparently barren soil and rock landscapes, as well as the numerous and diverse lakes found at the edges of the continent, harbor a range of prokaryotes, indicating that the extremely low temperatures which prevail throughout the year are no obstacle to microbial colonization (Tindall 2004).
Microorganisms represent the largest reservoir of biodiversity yet to be described, and consequently offer great potential for the discovery of new natural products. It is estimated that Earth sustains between 3 and 30 million species of organisms. Of these, approximately 1.4 million species have been described by science, including virtually all the bird and mammal species.
In contrast, only close to 200,000 of the possible 1.0–1.5 million species of fungi have been characterized. This percentage is even lower for bacteria, with estimates that between 1 and 10% of the probable species have been described (Bull et al. 1992, 2000).
Many of the isolated microorganisms from the Antarctic have been derived from maritime environmental studies of sea ice or sea water, or from the semi-marine lakes of Vestfold Hills, McMurdo Dry Valleys, and Larsemann Hills (Van Trappen et al. 2002). Few studies have isolated bacteria in the soil environment of ice-free areas (Nichols et al. 2002). However, even when only some of the bacterial communities can be isolated (Spring et al. 2000), culturing techniques allow new strains of organisms to be preserved for future detailed analysis, physiological characterization, as well as the search for potential applications such as the production of enzymes adapted to cold, pigments, antibiotics, and other bioactive compounds (Van Trappen et al. 2002).
D. antarctica offers unusual resistance to the damage caused by stressful environmental factors. Considering the adverse conditions under which D. antarctica survives, identifying and characterizing the bacteria that colonize the soil surrounding its roots is of great interest.
Materials and methods
Soil sampling and processing
Soil samples were collected by the biotechnology company Vitrogen S.A. Sampling was random and taken from four sites, at depths of 0–5 cm, in the area immediately surrounding the roots of D. antarctica. The samples were placed in labeled polyethylene bags and maintained at 4°C. They were then moved to the Laboratorio de Biología Molecular Aplicada at the Universidad de La Frontera, Temuco, Chile. Each sample was homogenized and processed independently, eliminating all components not corresponding to soil.
Isolation of different microorganism groups from the soil
For the microbial analysis, 10 g of soil was placed in 90 ml sterile water agar (0.1%), and agitated for 15 min in a horizontal agitator. A series of dilutions, in base ten, were performed until reaching 10−4. For the total microorganism count, nutritive agar (Merck S.A.) (diluted 1/4) and agar Luria–Bertani (MoBio Laboratories) (LB) were used. The Norris agar media was used to isolate heterotrophic bacteria that set nitrogen in free life (Eckford et al. 2002). The total number of microorganisms developed in each culture medium was determined after incubation at 4 and 15°C during 3, 5, and 10 days. Additionally, each bacterial colony displaying different characteristics was isolated. The pure cultures were maintained at −80°C with glycerol (25%).
Morphological characterization of the principal isolated microorganism groups
The microorganisms were isolated in pure cultures. Gram staining (Bartholomew and Mittwer 1952) was performed for morphological classification. Additionally, cytochrome-c oxidase determinations were performed using Bactident-Oxidase reactive strips (Diagnostica MERCK 13300), and catalase was detected using the Cowan and Steel’s method (1993).
Determination of resistance to antibiotics and heavy metals and tolerance to different temperatures
Antibiotic resistance was determined in Muller agar by adding the respective concentration of each antibiotic employed: gentamycine (10 μg ml−1), kanamycin (10 μg ml−1), streptomycin (10 μg ml−1), actinomycin-D (10 μg ml−1), ampicillin (100 μg ml−1), and tetracycline (30 μg ml−1). The plates were observed after incubation during 24 and 48 h at 4°C.
Resistance to the metal salts cadmium chloride, cobalt chloride, zinc chloride, and mercury chloride was evaluated by adding to LB solid medium the metals in solution in the concentrations 1–2.5–5–10–20–25 and 40 mM of Cd2+, Co2+, Zn2+ and 0.1–0.2–0.4 and 0.5 mM of Hg2+. The plates were incubated for 48 h at 4°C. After incubation, the susceptibility of each isolation was determined by recording growth (+) or growth absence (−).
Tolerance to different temperatures was determined by incubating the bacteria at a known concentration in agar LB at 4, 22, and 37°C. The bacterial growth was determined at 24 and 48 h, and at 14 days of incubation at 37°C.
Genomic DNA extraction
The amplification of the 16S rRNA genes was performed directly on cells lysed by treatment at 80°C for 10 min, followed by 3 min at 100°C.
16S rRNA gene amplification
16S rRNA was selectively amplified from genomic DNA by PCR with oligonucleotide primers designed to anneal to conserved positions in the 3′ and 5′ regions of the 16S rRNA genes. Universal bacterial primers were used, and for all the isolates tested the forward primer 8f (5′-AGAGTTTGATCCTGGCTCAG-3′) and 1492r (5′-GGTTACCTTGTTACGACTT-3′) enabled the amplification of 1,502 bp of the 16S rRNA gene. PCR amplification was performed in a Peltier Thermal Cycler, PTC-200 (M J Research, USA) in 50 μl of reaction containing 2.5 μl reaction buffer 10×; 2.5 μl of MgCl2 25 mM; 2.5 μl of each deoxyribonucleotide 2.5 mM; 2.5 μl of each first 10 μM, and 1 U of Taq DNA polymerase (recombinant, Fermentas).
The temperature and cycling conditions were as follows: First, preheating at 94°C for 2 min; then 30 cycles at 94°C for 1 min; 55°C for 1 min; and 72°C for 1.5 min; and a final incubation at 72°C for 10 min. The presence of PCR products and their concentration were checked by electrophoresis of 5 μl products on a 1% agarose gel, stained with ethidium bromide. A molecular weight marker (1 kb DNA ladder, Fermentas) was included. To generate nearly full-length 16S rRNA clones, the PCR product was ligated into the pGEM-Teasy vector (Promega, Madison, Wisconsin, USA) and the ligation reaction was used to transform competent Escherichia coli strain DH5α.
Recombinant colonies were selected on Luria–Bertani agar plates containing 20 μg ml−1 X-gal (5-bromo-4-chloro-3-indolyl-β-d-galactopyranoside), and 0.5 mM IPTG (isopropyl-β-d-thiogalactopyranoside), and 100 μg ml−1 ampicillin. Plates were incubated overnight at 37°C. The presence of inserts was determined by direct PCR on a sample from white (positive) bacterial colonies, using primers flanking the cloning sites on the vector.
Sequencing was done with a Dye Terminator Cycle Sequencing kit and an ABI 3730XL DNASequencer (Applied Biosystems) by Macrogen (Korea). 16S rRNA nucleotide sequences were identified by WWW/BLASTN 2.1.1 (http://www.ncbi.nlm.nih.gov/blast) (Altschul et al. 1990, 1997) using DDBJ/EMBL/GenBank nucleotide sequence databases. The 16S rRNA sequences determined in this study were deposited in the GenBank/EMBL sequence databases.
Enzymatic activity
To determine the enzyme profile, bacterial isolates were inoculated into an APIZYM microbial identification system consisting of 19 substrates (BioMérieux SA, France), and processed according to the manufacturer’s directions. APIZYM enabled the rapid detection of constitutive enzymes based on a colorimetric assay following 16 h incubation at 15°C. The presence of enzyme activity was determined by the intensity of color development following the addition of reagents.
Results and discussion
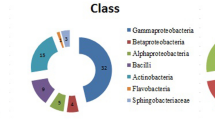
Seventy bacteria were isolated from the soil samples. The majority of the isolated strains presented white-colored colonies and brilliant mucous. Some strains with colored colonies, such as yellow, brown, and purple, were also notable (Fig. 1).
The first colonies appeared after 48 h of incubation at 4°C. The colonies colored primarily purple appeared after 10 days of incubation, where the most abundant counts were found in ¼ diluted nutritive agar rather than in the more rich culture LB medium. Subsequently, the purple-colored colonies developed within 24–48 h at room temperature, and were capable of growing in a more nutrient-rich medium.
Christner et al. (2000) found similar results isolating bacteria immersed in glacier ice, attributing this behavior to the bacteria’s need for time to synthesize enzymes, in order to avoid cellular damage resulting from long exposure to extreme environmental conditions, and to detoxify metabolic products such as hydrogen peroxide and free radicals. The same authors attribute the colored colonies to microorganism defense to avoid lethal DNA damage due to high levels of solar radiation.
The highest density of total cultivable bacteria was found in the sample named D. a. soil I, and it was significantly higher (P < 0.05) than those from the other soil samples (Table 1). Most investigations of bacteria in Antarctic soils have focused on the abundance and diversity of cultivable bacteria in the Dry Valleys, and some investigations indicated that these locations may contain between 106 and 108 prokaryotic cell g−1 (Smith et al. 2006).
From the bacteria collection, twelve were selected for further characterization; these included strains with colored colonies, and a selection of other isolates exhibiting multiple resistance to antibiotics and heavy metals, as well as a variety of morphological and physiological characteristics, as shown in Table 2.
Table 2 demonstrates that a large number of bacteria are resistant to antibiotics at the given doses. Only four of the 70 isolated strains did not show resistance to any of the antibiotics studied (data not shown). Of the 70 tested isolates, 76% were resistant to ampicillin (100 μg ml−1), actinomycin-D (10 μg ml−1) and gentamycin (10 μg ml−1), while 66% were resistant to tetracycline (30 μg ml−1), 51% to streptomycin (10 μg ml−1), and 36% to kanamycin (10 μg ml−1). Multiple resistances (to six antibiotics) were exhibited by 5% of the bacteria. Likewise, 29% of the isolates showed multiple resistances to metals (data not shown).
Resistance to ampicillin was very pronounced in all our isolates, a phenomenon also reported from other sources (Zemelman et al. 1980; De Souza et al. 2006). Transfer of virulence, conjugative, and/or antibiotic-resistance phenotypes may result in the increased dissemination of antibiotic-resistance and/or virulence determinants to bacteria in the environment.
We speculate that the resistance to several antibiotics shown by these Antarctic bacteria is due to the presence of guano from migratory birds which nest in Antarctica and spend much of their time in areas where vegetation is present.
With respect to the bacteria’s heavy metal tolerance, the evaluated strains presented low Cd tolerance, intermediate Zn and Hg tolerance, and high Co tolerance.
Even though some heavy metals are essential trace elements, many can be toxic to living beings, including bacteria, at high concentrations, due to the formation of complex compounds inside the cell. Microorganisms have developed several mechanisms to tolerate heavy metals including energy dependent metal effluxes (Purchase et al. 1997; Nies et al. 1998; Peitzsch et al. 1998; Goldberg et al. 1999; Grass et al. 2000; Franke et al. 2001); metal precipitation as insoluble salts (Blake et al. 1993); alteration of the membrane’s permeability to metal (Levine and Marzluf 1989); immobilization of metal within the cell wall (Cervantes and Gutiérrez-Corona 1994); chelating agent production (Silver and Phung 1996); and biochemical transformation of the metallic ion (Williams and Silver 1984).
Mergeay et al. (1985) evaluated the minimum inhibitory concentrations of several metallic ions on E. coli, in agar media, finding that the least toxic metal was manganese (20 mM) and the most toxic was mercury (2 mM). The heavy metals Zn, Co, and Cd presented minimum inhibitory concentrations of 1, 1, and 0.5 mM, respectively.
Additionally, Calomiris et al. (1984), Baquero et al. (1998), and Lawrence (2000) have demonstrated a correlation between metal tolerance and antibiotic resistance in bacteria because both genes are probably located close to each other in the same plasmid, and consequently, it is likely they are transferred together into the environment.
With respect to optimal growth temperature, the strains evaluated correspond to psychrotolerant strains, presenting a growth range up to 22°C, and growth absence at 37°C.
Nichols et al. (2002) indicate that microbial adaptation to permanently cold environments includes the optimization of basic cellular processes that are necessary for growth and survival. These can be summarized in three categories: enzyme functioning, nutrient transportation, and cellular membrane functions. The adaptation of the cellular processes in each one of these areas represents potential biotechnological products. Two examples obtained from Antarctic ice bacteria are polyunsaturated fatty acid (PUFA) production, and production of enzymes active in cold temperatures.
Based on the strain identification results (Table 2), the majority of the colonies that were whitish with mucoid features corresponded to different Pseudomonas species grouped in the γ-proteobacteria subdivision (strains numbered Da-bac 9, Da-bac 1-6, Da-bac 1-8, Da-bac 1-12, Da-bac 10, Da-bac 7-4, Da-bac 7-9, Da-bac 11, Da-bac 6-1, and Da-bac 12). Among the colored colonies, we found species from the Cytophaga–Flavobacteria complex, including species from the genus Flavobacterium sp. (Da-bac 3-9) and isolates from the Gram positive groups with high G + C content, such as Arthrobacter sp. (Da-bac 3–6) (Table 3).
The bacteria most commonly reported in cold environments are Gram negative microorganisms from the α, β, and γ-proteobacterias (Pseudomonas spp. and Vibrio spp.), and the phylum Cytophaga–Flavobacterium–Bacteroides. Among Gram positive bacteria, the most common are coryneformes, such as Arthrobacter and Micrococcus sp. These genera usually dominate in number and species diversity over Archaeas, although in deep sea waters they are found in equal numbers with Methanogenium and Methanococcus, among the genera most cited. Among Cyanobacteria that have been identified in Antarctica, the most common are Oscillatoria, Phormidium, and Nostoc (Pandey et al. 2004). The psycrophile yeasts, particularly Cryptococcus spp., have been repeatedly isolated from soil samples, and some researchers describe it as the most important adaptation in desert soils (Vishniac and Klinger 1986).
Aislabie et al. (2006), studying Antarctic soils using a combination of methods based on culturing and non-culturing techniques, found that the dominant groups correspond to the bacterial divisions Bacteroidetes, Actinobacteria, Proteobacteria, Thermus-Deinococcus, Acidobacteria, Firmicutes and Cyanobacteria, which are among the divisions commonly reported in soils.
It is now widely acknowledged that culture-based community studies inevitably induce a high degree of bias, whereas important groups of organisms that may be fastidious, co-culture dependent, or in a viable but non-culturable state, may be unrepresented. The isolated strains maintained in the collection from the present study constitute a unique collection for future, more detailed taxonomic analysis and physiological characterization, as part of the search for potential biotechnological uses.
Indeed, Bowman et al. (1999) indicate growing interest in developing research on prokaryotes in the Antarctic in order to isolate new bacteria, create culture collections, search for bioactivity, taxonomy, polyunsaturated fatty acid (PUFA) production, enzymes with activity in cold, and bioremediation. The research performed to date confirms the isolation of new bacteria from this extreme environment.
The bacterial strains isolated from the rhizosphere of D. antarctica produced diverse patterns of enzymes as shown in Table 4, where the diversity of enzymatic activities presented by the colored bacterial strains is especially notable. The strains, identified as different species of Pseudomonas, produced alkaline and acid phosphatases, esterase (C4), esterase lipase (C8), leucine and valine arylamidase, phosphohydrolase, and α-glucosidase. The genera Flavobacterium produced alkaline and acid phosphatases, esterase lipase (C8), leucine and valine arylamidase, phosphohydrolase, α-glucosidase, β-galactosidase, N-acetyl-β-glucosaminidase and α-mannosidase. Isolates identified as Arthrobacter sp. produced the most diverse array of enzymes.
The enzyme patterns varied with species of bacteria, but in general the Gram positive isolates produced peptidases, while the Gram negative isolates produced carbohydrases. Phosphatases, esterases, and lipases were produced by both Gram positive and Gram negative isolates.
Some of the isolates obtained in this investigation excreted extracellular enzymes such as lipases, proteases and phosphatases, which possess potential biotechnological applications due to their adaptation to cold (Wery et al. 2003).
In addition, the results obtained in the present study suggest that the bacterial strains isolated from rhizospheric soil in the Antarctic could serve as a model to study the physiological, biochemical, and molecular mechanisms of heavy metal tolerance and antibiotic resistance in cold climates, and elucidate the actual way in which these bacteria are becoming resistant to multiple antibiotics.
These and other findings have great potential for developing new biotechnological products from Antarctic microorganisms.
References
Aislabie JM, Chhour KL, Saul DJ, Miyauchi S, Ayton J, Paetzold RF, Balks MR (2006) Dominant bacteria in soils of marble Point and Wright Valley, Victoria Land, Antarctica. Soil Biol Biochem 38:3041–3056
Alberdi M, Bravo L, Gutiérrez A, Gidekel M, Corcuera L (2002) Ecophysiology of Antarctic vascular plants. Physiol Plantarum 115:479–486
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402. Available at http://www.ncbi.nlm.nih.gov/BLAST/
Baquero F, Negri MC, Morosini MI, Blázquez J (1998) Antibiotic-selective environments. Clin Infect Dis 27:5–11
Bartholomew JW, Mittwer T (1952) The Gram stain. Bacteriol Rev 16:1–29
Beyer L, Bölter M, Seppelt RD (2000) Nutrient and thermal regime, microbial biomass and vegetation of Antarctic soils in the Windmill Islands Region of east Antarctica (Wilkes Land). Arct Antarct Alp Res 32:30–39
Blake RC, Choate DM, Bardhan S, Revis N, Barton L, Zocco TG (1993) Chemical transformation of toxic metals by a Pseudomonas strain from a toxic waste site. Environ Toxicol Chem 12:1365–1376
Bowman JP, McCammon SA, Brown MV, Nichols DS, McMeekin TA (1999) Diversity and association of psychrophilic bacteria in Antarctic sea ice. Appl Environ Microbiol 63:3068–3078
Bull AT, Goodfellow M, Slater JH (1992) Biodiversity as a source of innovation in biotechnology. Ann Rev Microbiol 46:219–252
Bull AT, Ward AC, Goodfellow M (2000) Search and discovery strategies for biotechnology: the paradigm shift. Microbiol Mol Biol Rev 46:573–606
Calormiris J, Armstrong JL, Seidler RJ (1984) Association of metal tolerance with multiple antibiotic resistance of bacteria isolated from drinking water. Appl Environ Microbiol 47:1238–1242
Casaretto JA, Serey I, Zúñiga G (1994) Size structure of a population of Deschampsia antarctica (Desv.) in Robert Island, Maritime Antarctica. Serie Científica INACH 44:61–66
Cervantes C, Gutiérrez-Corona F (1994) Cooper resistance mechanisms in bacteria and fungi. FEMS Microbiol Rev 14:121–137
Christner BC, Mosley-Thompson E, Thompson LG, Zagorodnov V, Sandman K, Reeve J (2000) Recovery and identification of viable bacteria immured in glacial ice. Icarus 144:479–485
Cowan S, Steel S (1993) Manual for the identification of medical bacteria, 7th edn. Cambridge University Press
De Souza MJ, Nair S, Loka Bharathi PA, Chandramohan D (2006) Metal and antibiotic-resistance in psychrotrophic bacteria from Antarctic Marine waters. Ecotoxicology 15:379–384
Eckford R, Cook FD, Saul D, Aislabie J, Foght J (2002) Free-living heterotrophic nitrogen-fixing bacteria isolated from fuel-contaminated Antarctic soils. Appl Environ Microbiol 68:5181–5185
Edwards J, Lewis-Smith R (1998) Photosynthesis and respiration of Colobanthus quitensis and Deschampsia antarctica Desv. from the marine Antarctic. Brit Antarct Surv Bull 81:43–63
Franke S, Gregor G, Nies DH (2001) The product of the ybdE gene of the Escherichia coli chromosome is involved in detoxification of silver ions. Microbiology 147:965–972
Goldberg M, Pribyl T, Juhnke S, Nies DH (1999) Energetics and topology of a CzcA, a cation/proton antiporter of the resistance-nodulation-cell division protein family. J Biol Chem 274:26065–26070
Grass G, Große C, Nies DH (2000) Regulation of the cnr cobalt and nickel resistance determinant from Ralstonia sp. strain CH34. J Bacteriol 182:1390–1398
Lawrence JG (2000) Clustering of antibiotic resistance genes: beyond the selfish operon. Am Soc Microbiol News 66:281–286
Levine WB, Marzluf GA (1989) Isolation and characterization of a cadmium-resistant mutant of Neurospora crassa. Can J Microbiol 35:359–365
Meargeay M, Nies D, Schlegel HD, Gerits J, Charles P, Van Gijsegem F (1985) Alcaligenes eutrophus CH34 is a facultative chemolithotroph with plasmid-borne resistance to heavy metals. J Bacteriol 162:328–334
Nichols DS, Sanderson K, Buia A, Van de Kamp J, Holloway J, Bowman JP, Smith M, Mancuso Nichols C, Nichols PD, McMeekin TA (2002) Bioprospecting and biotechnology in Antarctica. In: Jabour-Green J, Haward M (eds) The Antarctic: past, present and future. Antarctic Cooperative Research Centre, Research Report #28, Hobart, pp 85–103
Nies DH, Koch S, Wachi S, Peitzsch N, Saier MH (1998) CHR of prokaryotic proton motive force-driven transporters probably contains chromate/sulphate antiporters. J Bacteriol 180:5799–5802
Pandey KD, Shukla SP, Shukla PN, Giri DD, Singh JS, Singh P, Kashyap AK (2004) Cyanobacteria in Antarctica: ecology, physiology and cold adaptation. Cell Mol Biol (Noisy-le-Grand) 50:575–584
Peitzsch N, Eberz G, Nies DH (1998) Alcaligenes eutrophus as a bacterial chromate sensor. Appl Environ Microbiol 64:453–458
Purchase D, Miles RJ, Young TWK (1997) Cadmium uptake and nitrogen fixing ability in heavy-metal-resistant laboratory and field strains of Rhizobium leguminosarum biovar trifolii. FEMS Microbiol Ecol 22:85–93
Silver S, Phung LT (1996) Bacterial heavy metal resistance: new surprises. Annu Rev Microbiol 50: 753–789
Smith JJ, Ah Tow L, Stafford W, Cary C, Cowan D (2006) Bacterial diversity in three different Antarctic cold Desert mineral soils. Microb Ecol 51:413–421
Spring S, Schulze R, Overmann J, Schleifer KH (2000) Identification and characterization of ecologically significant prokaryotes in the sediment of freshwater lakes: molecular and cultivation techniques. FEMS Microbiol Rev 24:573–590
Tindall BJ (2004) Prokaryotic diversity in the Antarctic: the tip of the iceberg. Microb Ecol 47:271–283
Van Trappen S, Mergaert J, Van Eygen S, Dawyndt P, Cnockaert MC, Swings J (2002) Diversity of 746 heterotrophic bacteria isolated from microbial mats from ten Antarctic lakes. Syst Appl Microbiol 25:603–610
Vishniac H, Klinger J (1986) Yeasts in the Antarctic deserts. In: Megusar F, Gantar M (eds) Perspectives in microbial ecology. Proceedings of the 4th ISME, Slovene Society for Microbiology, Ljubljana, Slovenia, pp 46–51
Wery N, Gerike U, Sharman A, Chaudhuri JB, Hough DW, Danson MJ (2003) Use of packed-column bioreactor for isolation of diverse protease-producing bacteria from Antarctic soil. Appl Environ Microbiol 69:1457–1464
Williams JW, Silver S (1984) Bacterial resistance and detoxification of heavy metals. Enzyme Microb Technol 6:530–537
Zemelman R, Silva J, Henríquez M (1980) Antibiotic resistant bacteria in seawater from Concepción Bay. Arch Biol Exp 13:121
Zúñiga GE, Alberdi M, Fernández J, Montiel P, Corcuera LJ (1994) Lipid content in leaves of Deschampsia antarctica from the maritime Antarctic. Phytochemistry 37:669–672
Acknowledgements
The authors are grateful for Grants INACH No. 03-01 and Grant DIUBB 055209 2/R 2005; Convenio Desempeño II 2007 Universidad de La Frontera; to the Instituto Antártico Chileno, Ministerio de Relaciones Exteriores 2005–2007 for logistical support; and the biotechnology company Vitrogen S.A.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Barrientos-Díaz, L., Gidekel, M. & Gutiérrez-Moraga, A. Characterization of rhizospheric bacteria isolated from Deschampsia antarctica Desv.. World J Microbiol Biotechnol 24, 2289–2296 (2008). https://doi.org/10.1007/s11274-008-9743-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11274-008-9743-1