Abstract
This investigation reported the isolation and characterization of a potent heavy metal accumulating bacterial strain Enterobacter cloacae B1 from polluted soil at Ghaziabad, India. The minimum inhibitory concentration of the selected bacterial strain was recorded to be 1100 ppm for lead, 900 ppm for cadmium, and 700 ppm for nickel. Bioaccumulation of lead by this bacterial strain was extremely high (95.25 %), followed by cadmium (64.17 %) and nickel (36.77 %). Antioxidant enzymes such as catalase (CAT) and superoxide dismutase (SOD) activity was determined in presence of lead, cadmium, and nickel at a concentration of 400 ppm. To monitor the physiological stress, malondialdehyde (MDA) level was also estimated. In order to optimize the flocculant production, bacterial strain E. cloacae B1 was cultured in specific medium at different incubation period (24 to 72 h), pH (6.0 to 9.0), and temperature (20 to 50 °C). It was observed that surfactant production was maximum at 72 h of incubation period (47.28 %), pH 8.0 (56.63 %), and temperature 40 °C (62.94 %). Protein expression profile in presence of these heavy metals was also interesting. Few proteins were noticed to be overexpressed in presence of these heavy metals.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
In recent year, rapid industrialization throughout the world has made easier for product availability, but it also raises the environmental pollution due to improper infrastructure for waste management (Ahemad and Malik 2011). Heavy metal toxicity in the environment creates different types of health problem (Gupta and Gupta 1998). Few researchers have reported that higher level of lead can cause organ toxicity in human that leads to severe damage in the liver, kidney, and central nervous system, whereas lower level of lead affects heme synthesis and different biochemical processes (Goldstein 1992; WHO 1995), while cadmium cause kidney disease through renal tubule dysfunction (Berohoft 2013). Cadmium is also reported to be a toxic metal for endocrine organ including pituitary (Jiménez-Ortega et al. 2012). On the other hand, nickel cause skin disease (Thyssen et al. 2013) and respiratory problem, and sometimes, it can act as a potent carcinogen (Brüske-Hohlfeld 2009). Different compounds of nickel are also reported to be an inducer for carcinoma formation in rodents (Kasprzak et al. 2003). Though few metals are essential for growth and metabolism in organism (nickel, iron, cobalt, zinc, etc.) and are considered as micronutrient (Beveridge and Doyle 1989; Bruins et al. 2000). However, high level of these metals have adverse effect on environment. Deep and Altalhi (2009) stated that heavy metals are potent inhibitor of normal biodegradation process and cannot be easily removed from environment (Ren et al. 2009). Heavy metal toxicity caused by lead, cadmium, and nickel have been reported by many authors (Chen and Lin 1998; Nath et al. 2012). Reduction of heavy metal toxicity is a great challenge for all developed and developing countries. Due to high cost and competition, industries avoid waste management process which increase heavy metal load on environment. Many naturally occurring bacteria have been found closely associated with the bioremediation process which removes heavy metal from environment and thus reduce toxicity. Due to high adaptive nature and cellular mechanism, bacteria can tolerate and absorb different types of heavy metals (Nies 1999; Etelvina et al. 2005). In recent year, different types of bacterial species have been reported for their bioremediation property (Selvi et al. 2012; Nath et al. 2012; Poornima et al. 2014).
Heavy metal creates different types of physiological stress in bacteria and produce reactive oxygen species (ROS) like O2 −, OH·, and H2O2 (Choudhary et al. 2007). In normal physiological condition, ROS level remains low due to activity of certain protective enzymes like, catalase, superoxide dismutase, and glutathione reductase (Pandey et al. 2013). Lipid peroxydation or malondialdehyde (MDA) production is a major clue for stress response. Among various antioxidant enzymes, catalase (CAT) and superoxide dismutase (SOD) gain special attention which converts the ROS into oxygen and water to maintain cellular integrity. Catalase is a heme-containing protein, widely distributed among various groups including bacteria (Abassi and Kushad 1998; Bailly et al. 2004; Switala and Leuwen 2002). Similarly, SOD is a metalloenzyme that converts highly toxic superoxide into oxygen and less toxic hydrogen peroxide. Thus, for proper evaluation of bioremediation efficiency of the selected bacterial strain, it is necessary to determine the physiological stress in response to heavy metal.
Surfactants or flocculants are diverse group of amphipathic molecule that help in alleviating the heavy metal from contaminated soil sediments and are considered to be environment-friendly (Georgiou et al. 1992; Fietcher 1992; Desai and Banat 1997). Chemical surfactants are widely used due to low cost, but these are not environment-friendly material (Kowall et al. 1989; Master et al. 1985). Microbial sources of flocculant are cheap and less toxic. These molecule increases the biodegradation of insoluble pollutant by reducing the surface tension (Hassanshahian 2014). A large number of bacteria, such as Bacillus sp., Pseudomonas sp., Acinetobacter sp., and Arthrobacter sp., are considered to be potent bioflocculant producer (Singh et al. 2014).
Screening of potent heavy-metal-tolerant bacterial strains is necessary. Therefore, the aim of our present investigation were (i) isolation and characterization of heavy-metal-tolerant bacterium, (ii) evaluation of metals removal capacity, (iii) antioxidant profile of this selected bacterial strain, (iv) optimization of surfactant production, and (v) protein expression of the selected bacterial strain upon exposure of these heavy metals.
2 Materials and Methods
2.1 Isolation and Selection of Metal-Tolerant Bacterial Strain
Soil samples were collected from Ghaziabad, Uttar Pradesh, India, and kept in sterilized plastic bag. One gram of soil sample was mixed in 10 ml of sterile distilled water, vortexed carefully, and serial dilution was made up to five times. One hundred microliters of each sample was spread on tryptone soya agar plates (pH 7.0) containing 200 ppm of lead, cadmium, and nickel separately and incubated overnight at 37 °C. Ten different bacterial colonies were primarily selected for further studies. Pure culture of each strain was done by repeated streaking method. In order to check minimum inhibitory concentration (MIC), all the bacterial strains were cultured in tryptone soya broth medium (pH 7.0) with varying concentration of lead, cadmium, and nickel (100–1500 ppm). The bacterial strain B1 showing highest tolerance level was selected for further studies.
2.2 Biochemical Characterization and Identification of the Selected Strain B1
In order to characterize the selected bacterial strain B1, it was cultured in TSA (tryptone soya agar) plate and incubated for 24 h at 37 °C. Primary characterization of the bacterial strain B1 was done by colony morphology (color, surface, margin, and elevation). Different biochemical tests (citrate utilization, nitrate reduction, urease production, oxydase production, etc.) and acid production from glucose, sucrose, lactose, and fructose were carried out according to Mondal et al. (2010). Species level identification of the selected bacterial strain was done by 16S rRNA sequence analysis following the method of Banerjee et al. (2013). In brief, genomic DNA was isolated by lysozyme-SDS method. PCR amplification of the 16S rRNA region was carried out using forward AGAGTTTGATCMTGGCTCAG (B27 F) and reverse GGTTACCTTGT TACGACTT (1492 R) primers. PCR product was sequenced, analyzed, and edited in Mega 4.0. phylogenetic tree was drawn with other close homologous bacterial strains available in NCBI database.
2.3 Quantification of Metal Removal by Atomic Absorption Spectroscopy
In order to quantify the metal removal capacity, the bacterial strain B1 was cultured in nutrient broth (pH 7.0) containing 400 ppm of lead, cadmium, and nickel separately. Removal of heavy metals from the culture medium was determined by following the method of Ahemad and Malik (2011) with few modifications. In brief, 5 ml of culture broth from each sample was taken at 24-h interval for 3 days. Supernatant was collected by centrifugation at 5000 × g for 10 min, mixed 2 ml of concentrated HNO3, and heated at 70 °C until it became transparent. Metals concentration in the supernatant was analyzed by atomic absorption spectroscopy (model Spectra AA55). Nutrient broth without bacterial strain containing the abovementioned metals at concentration of 400 ppm was taken as control.
2.4 Antioxidant Profile of the Bacterial Strain B1
In order to monitor the oxidative stress, bacterial strain B1 was cultured in TSA broth in presence of lead, cadmium, and nickel at 400-ppm concentration. After 24 h of incubation, bacterial cell were harvested by cooling centrifugation (8000 × g, 10 min), washed two times with 50 mM phosphate buffer (pH 7.2) and finally dissolved in same buffer in 2 ml Eppendorf tube. Cytosolic content was collected by disrupting the bacterial cell by ultrasonication in ice bath (30-s pulse and 30-s cooling, for 3 min). Supernatant was collected by centrifugation (8000 × g for 10 min) at 4 °C and kept in −80 °C for further study.
2.4.1 Malondialdehyde Level
MDA production was measured according to the method of Draper and Hadley (1990). Here, 1 ml of sample was carefully mixed with 2 ml of TBA (thiobarbituric acid) reagent (20 % thiobarbituric acid, 0.5 % TBA, and 2.5 N HCl), heated 20 min in boiling water bath, cooled, and supernatant was collected by centrifugation (500 × g, 10 min) at 4 °C. Absorbance of the collected supernatant was measured at 532 nm. MDA equivalent content in the sample was calculated using an extinction coefficient of 1.56 × 105 M−1 cm−1.
2.4.2 Superoxide Dismutase Activity
SOD activity was assayed following the method of Ewing and Janero (1995). Twenty-five microliters of supernatant was taken in a microtiter well and mixed 20 μl of reaction buffer (50 mM phosphate buffer, 0.1 mM EDTA, 98 μM NADH, and 62 μM NBT, pH 7.4). Reaction was initiated by adding 20 μl of an initiating reagent (50 mM phosphate buffer and 33 μM PMS in 0.1 mM EDTA, pH 7.4). Absorbance was measured at 560 nm using the microplate reader (Bio-Rad, Model 680, USA).
2.4.3 Catalase Activity
CAT activity in the bacterial supernatant was measured following the method of Aebi (1984) with little modification. In brief, 20 μl of supernatant was mixed in 980 μl of H2O2 buffer and absorption was measured at 240 nm using spectrophotometer for 60 s at 15-s interval. Activity was calculated by decreased in absorbance of H2O2 at 240 nm.
2.5 Optimization of Flocculant Production
Flocculant production was carried out in the liquid media 0.5 % peptone, 0.5 % yeast extract, 2 % ethanol, 1 % glycerol, 0.05 % K2HPO4, 0.05 % MgSO4·7H2O, 0.2 % NaCl, and 0.2 % CaCO3. Bioflocculating activity was done according to the method of Batta et al. (2013). In brief, 1 ml of cell-free supernatant was added with 9-ml kaolin clay solution (5 g ml−1) and 3 ml of 1 % CaCl2, vortexed, and kept undisturbed for 5 min. The optical density of the upper clear solution was measured at 550 nm by spectrophotometer (Beckman DU 730). Kaolin clay solution without culture supernatant was considered as control. The activity was calculated by the following method:
where X represents the OD of the control at 550 nm, and Y represents the OD of the sample at 550 nm.
Optimization was carried out in different incubation period (24–72 h), pH (6.0–9.0), and temperature (20–50 °C).
2.6 Evaluation of Protein Expression in Stress Condition by SDS-PAGE
In order to check the protein expression upon heavy metal (lead, cadmium, and nickel) exposure, the collected cytosolic content (prepared during antioxidant study) was separated on SDS-PAGE (4–12.5 %). Bacterial culture without metal was taken as control.
3 Results
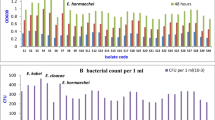
In this present study, heavy-metal-tolerant bacterial strain B1 was isolated from polluted soil (industrial waste disposal area). Morphologically, the selected bacterial strain B1 was creamy white in appearance with smooth surface and irregular margin (Table 1). Biochemical characterization showed that the bacterial strain B1 was positive in case of CAT production, nitrate reduction, ortho-nitrophenol β-galactoside (ONPG) utilization, citrate utilization, and urease production, whereas negative for methyl red, oxidase, and indole production. Carbohydrate utilization ability was also tested for this selected bacterial strain B1. It was observed that it can actively utilized glucose, lactose, and sucrose, but failed to grow in fructose broth. Finally, the bacterial strain B1 was identified as Enterobacter cloacae (Acc. No. KM402758) with the help of 16S rRNA sequence analysis (Fig. 1). Minimum inhibitory concentration of the selected bacterial strain E. cloacae B1 against these three heavy metals lead, cadmium, and nickel is tabulated in Table 2. Nickel was detected to be more toxic (700 ppm), followed by cadmium (900 ppm) and lead (1100 ppm). Metal removal from the culture medium at different time intervals is given in Table 3. Metal absorption was recorded to be highest in case of lead (day 1, 61.45 %, to day 3, 95.25 %). Cadmium absorption ability of the bacterial strain E. cloacae B1 was also quite high (64.17 %, day 3), while nickel removal after day 3 was observed to be 36.77 %.
Antioxidant enzymes activity in presence of these heavy metals is presented in Fig. 2. Bacterial supernatant showed significantly (p < 0.001) higher MDA level in the nickel group and reduced MDA level in the cadmium group, while no such change was detected in the lead group when compared to the control values which indicated that oxidative load was significantly elevated respective nickel and cadmium groups (Fig. 2a). The major antioxidant enzyme CAT activity showed significant (p < 0.001) peak in cadmium group but significantly lower in both nickel and lead groups compare to control one (Fig. 2b). Another ROS scavenging enzyme SOD showed the activity higher in cadmium group, followed by nickel- and lead-treated bacterial samples (Fig. 2c).
Represented antioxidant enzymes activity exhibited by Enterobacter cloacae B1 upon exposure of these heavy metals. a Malondialdehyde (MDA) level at different treatment groups; b catalase (CAT) activity; c superoxide dismutase (SOD) activity. Data were presented as mean ± SEM in vertical bars, n = 3 replicate. Different alphabets (a, b, c, d) indicate significant (p < 0.05) differences in the values of a particular variable in different stress condition whereas, same alphabets denote no significant difference. Data were analyzed using one-way ANOVA along with DMRT analysis for significance
Flocculant production by the bacterial strain E. cloacae B1 at different incubation period, pH, and temperature is given in Fig. 3. Flocculation activity exhibited by E. cloacae B1 was recorded to be highest at 72 h of incubation (Fig. 3a, 47.28 %), pH 8.0 (Fig. 3b, 56.63 %), and temperature 40 °C (Fig. 3c, 62.94 %).
Demonstrated the optimization of flocculant production at different culture parameters. a Effect of incubation period on flocculant production; b optimization of production at different pH; c optimization of production at different temperature. Data were presented as mean ± SEM in vertical bars, n = 3 replicate. Different small letters (a, b, c, d) on the error bars indicate significant (p < 0.05) differences in the values of a particular variable in different stress condition. Data were analyzed using one-way ANOVA along with DMRT analysis for significance
Protein expression of the selected bacterial strain E. cloacae B1 on exposure to lead, cadmium, and nickel is presented in Fig. 4. It was clearly detected that few proteins (a, b, d, e, h, j, and k) were not well expressed on exposure of these heavy metals, whereas protein c was highly expressed in case of lead- and cadmium-treated samples compared to control one. Similarly, protein i was up-regulated in case of cadmium- and nickel-treated samples. Expression of another two proteins f and g were also observed in cadmium-treated sample.
4 Discussions
Due to global industrialization, heavy metal toxicity in environment is increasing rapidly. Different industries release huge amount of waste containing toxic heavy metals directly into water bodies without processing (Kafilzadeh et al. 2012; Batta et al. 2013; Hookoom and Puchooa 2013). Mechanical removal of these heavy metals is costly and time consuming. Heavy metal accumulation property in bacteria might be an alternative solution. In our study, we have isolated one potent heavy metal-resistant (lead, cadmium, and nickel) bacterial strain B1 from polluted soil at Ghaziabad, India. Morphological, biochemical, and 16S rRNA sequence analysis identified the bacterial isolate as E. cloacae. Recently, Poornima et al. (2014) reported the isolation of chromium-resistant bacterial strain E. coli PS01 from tannery effluent, whereas Pandey et al. (2011) isolated and characterized two lead- and arsenate-tolerant Bacillus sp. from slag deposal site at Burnpur, India. Metal tolerance mechanism in bacteria is well characterized. Bacteria adapted several mechanisms for metal resistance such as different types of transport channel (Nies 1999), production of siderophores (Schalk et al. 2011), and compartmentalization within the cell (Ahemad 2012). Here, minimum inhibitory concentration of the selected bacterial strain E. cloacae B1 against lead was recorded to be quite high (1100 ppm) compared to cadmium and nickel which indicated the high lead absorption ability of the bacterial isolate. Nath et al. (2012) reported the MIC of few bacterial isolates against lead (180 μg ml−1) and cadmium (1200 μg ml−1). Bacterial heavy metal removal property has great application in environmental point of view. Johncy-Rani et al. (2010) isolated three bacterial isolates Bacillus sp., Pseudomonas sp., and Micrococcous sp. and reported their bioaccumulation capacity to be 69.34, 90.41, and 84.27 % in case of copper, cadmium, and lead, respectively. Similarly, Ahemad and Malik (2011) reported the accumulation of different metals like lead, chromium, mercury, and zinc by several bacterial species isolated from agricultural field and wastewater. While in our study, it was observed that lead accumulation capacity of the bacterial strain E. cloacae B1was very high compared to cadmium and nickel.
Bacterial growth in the presence of heavy metals creates physiological stress and produce ROS which is very dangerous for cell. To reduce these oxidative stresses, bacteria produce different types of protective enzymes like CAT and SOD. A significant activity of antioxidant enzyme SOD, glutathione peroxidase (GSHPx), glutathione reductase (GR), and mercury reductase (MR) in rumen bacteria Streptococcus bovis and Selenomonas ruminantium was observed, after exposure to HgCl2 (Lenaèrtovaè et al. 1998), while Pandey et al. (2013) reported the effect of heavy metals on antioxidant enzyme profile in three bacterial species. Our result depicted that addition/ presence of certain heavy metals (lead, nickel, and cadmium) in the culture medium emerged stress condition within the different treated groups as the oxidative stress marker, MDA, was detected to be varying in the treated samples. Moreover, CAT and SOD activities in the cadmium group denoted that their increasing activity has occurred in order to fight back the harmful free radicals (ROS) emerged during oxidative stress to compensate the internal integrity of the cell. These observations were further supported by the MDA level within the cell, whereas in case of both lead- and nickel-treated groups, CAT and SOD activities have reduced as the stress was relatively lower than the cadmium group as indicated by the corresponding MDA level. Result depicted that MDA level was high in control compared to cadmium-treated sample which was quite unusual. In order to determine the oxidative status, we have collected sample after 24 h of incubation period. In Fig. 2b, c, it was clear that CAT and SOD activities were quite high in Cd-treated sample compared to control, as well as Pb- and Ni-treated samples. So, Cd is more toxic for this selected bacterial strain compared to other selected heavy metal. For this reason, MDA level might became elevated sharply and very quickly (far before 24 h of treatment) and again came down (even below control level) after 24 h due to the activity of stress-responsive enzyme system.
Biosurfactant or bioflocculant production from microbial source is cost effective and less hazardous. Several authors reported the production of surfactant from different bacterial species (Dhail 2013; Ugbenyen and Okoh 2013). A study by Bodour et al. (2003) stated that distribution of flocculant-producing bacterial species was dependent on soil conditions, with gram-positive biosurfactant-producing bacteria isolated from heavy-metal-contaminated soil, whereas gram-negative isolates were from hydrocarbon-contaminated soil. In this present study, we have optimized the production of surfactant at different incubation period, pH, and temperature. It was observed that highest production of flocculant was achieved when the incubation period was 72 h, pH was 8.0, and temperature was 40 °C.
Protein expression pattern in the presence of different heavy metals was quite interesting. It was observed that in case of cadmium treatment, several proteins were over-expressed which were not found in lead- and nickel-treated samples. Further research is required to understand the detail mechanism of these stress protein expression and their role in heavy metal tolerance.
5 Conclusion
To the author’s knowledge, the potent heavy-metal-tolerant bacterial species belongs to the genus Pseudomonas sp., Bacillus sp., and Staphylococcus sp. There are very few reports on bioremediation of heavy metals by Enterobacter sp. The lead removal capacity of the isolated bacterial strain E. cloacae B1 was detected to be very high and might be used for bioremediation process. Bioflocculant (a future biological weapon against water pollution) production was also optimized in different culture conditions. It was observed that production rate was maximum at 72 h of incubation period in alkaline condition at 40 °C. Bioflocculant has several advantages over chemical surfactant like activity, degradability, stability, toxicity and cost. So, to reduce the water pollution surfactant producing bacterial species are important in nature. Lastly, different types of proteins are observed to be expressed upon exposure of these heavy metals. Specially, in cadmium treatment, we observed few deep bands which were not so clear in control treatment. Purification and molecular characterization of these stress proteins might be helpful to understand the mechanism of heavy metals remediation by this selected bacterial strain E. cloacae B1.
References
Abassi, N. A., & Kushad, M. M. (1998). Active oxygen-scavenging enzymes activities in developing apple flowers and fruits. Scientia Horticulturae, 74, 183–194.
Aebi, H. (1984). Catalase in vitro. Methods in Enzymology, 105, 121–126.
Ahemad, M. (2012). Implication of bacterial resistance against heavy metals in bioremediation: a review. Institute of Integrative Omics and Applied Biotechnology Journal, 3(3), 39–46.
Ahemad, M., & Malik, A. (2011). Bioaccumulation of heavy metals by zinc resistant bacteria isolated from agricultural soils and irrigated with wastewater. Bacteriology Journal, 2, 12–21.
Bailly, C., Leymarie, J., Lehner, A., Rousseau, S., Come, D., & Corbineau, F. (2004). Catalase activity and expression in developing sunflower seeds as related to drying. Journal of Experimental Botany, 55, 475–483.
Banerjee, G., Ray, A. K., Askarian, F., & Ringø, E. (2013). Characterization and identification of enzyme-producing autochthonous bacteria from the gastrointestinal tract of two Indian air-breathing fish. Beneficial Microbes, 4, 277–284.
Batta, N., Subudhi, S., Lal, B., & Devi, A. (2013). Isolation of lead tolerant novel species, Acinetobacter sp. TL-3: assessment of bioflocculating activity. Indian Journal of Experimental Biology, 51, 1004–1011.
Bernhoft, R. A. (2013). Cadmium toxicity and treatment. The Scientific World Journal. doi:10.1155/2013/394652.
Beveridge, T. J., & Doyle, R. J. (1989). Metal ions and bacteria. New York: Wiley.
Bodour, A. A., Drees, K. P., & Maier, R. M. (2003). Distribution of biosurfactant-producing bacteria in undisturbed and contaminated arid southwestern soils. Applied and Environmental Microbiology, 69(6), 3280–3287.
Bruins, M. R., Kapil, S., & Oehme, F. W. (2000). Microbial resistance to metals in the environment. Ecotoxicology and Environmental Safety, 45, 198–207.
Brüske-Hohlfeld, I. (2009). Environmental and occupational risk factors for lung cancer. Methods in Molecular Biology, 472, 3–23.
Chen, C. Y., & Lin, T. H. (1998). Nickel toxicity to human term placenta: in vitro study on lipid per oxidation. Journal of Toxicology and Environmental Health Part A, 54, 37–47.
Choudhary, M., Jetley, U. K., Khan, M. A., Zutshi, S., & Fatma, T. (2007). Effect of heavy metal stress on Proline, Malondialdehyde and Superoxide Dismutase activity in the Cyanobacterium spirulina platensis-S5. Ecotoxicology and Environmental Safety, 66, 204–209.
Deeb, B. E., & Altalhi, A. D. (2009). Degradative plasmid and heavy metal resistance plasmid naturally coexist in phenol and cyanide assimilating bacteria. American Journal of Biochemistry and Biotechnology, 5(2), 84–93.
Desai, J. D., & Banat, I. M. (1997). Microbial production of surfactants and their commercial potential. Microbiology and Molecular Biology Reviews, 61, 47–64.
Dhail, S. (2013). Microbial enhanced oil recovery using potent biosurfactant produced by Pseudomonas sp. Arabian sea, Mumbai. Journal of Petroleum and Gas Engineering, 4(3), 57–60.
Draper, H. H., & Hadley, M. (1990). Malondialdehyde determination as index of lipid peroxidation. Methods in Enzymology, 186, 421–423.
Etelvina, M. A. P. F., Ana Isabel, G. L., & Sofia Isabel, A. P. (2005). Cadmium tolerance plasticity in Rhizobium leguminoserum bv. Viciae: glutathione as a detoxifying agent. Canadian Journal of Microbiology, 51, 7–14.
Ewing, J. F., & Janero, D. R. (1995). Microplate superoxide dismutase assay employing a nonenzymatic superoxide generator. Analytical Biochemistry, 232, 243–248.
Fietcher, A. (1992). Biosurfactant: moving toward industrial application. Trends in Biotechnology, 10, 208–217.
Georgiou, G., Lin, S. C., & Sharma, M. M. (1992). Surface active compounds from microorganisms. Biotechnology, 10(1), 60–65.
Goldstein, G. W. (1992). Neurological concepts of lead poisoning in children. Pediatric Annals, 21(6), 384–388.
Gupta, U. C., & Gupta, S. C. (1998). Trace element toxicity relationships to crop production and livestock and human health: implications for management. Communications in Soil Science and Plant Analysis, 29, 1491–1522.
Hassanshahian, M. (2014). Isolation and characterization of biosurfactant producing bacteria from Persian Gulf (Bushehr provenance), Marine Pollution Bulletin (www.elsevier.com/locate/marpolbul).
Hookoom, M., & Puchooa, D. (2013). Isolation and identification of heavy metal tolerant bacteria from industrial and agricultural areas in Mauritius. Current Research in Microbiology and Biotechnology, 1(3), 119–123.
Jiménez-Ortega, V., Barquilla, P. C., Fernández-Mateos, P., Cardinali, D. P., & Esquifino, A. I. (2012). Cadmium as an endocrine disruptor: correlation with anterior pituitary redox and circadian clock mechanisms and prevention by melatonin. Free Radical Biology and Medicine, 53(12), 2287–2297.
Johncy-Rani, M., Hemambika, B., Hemapriya, J., & Rajeshkannan, V. (2010). Comparative assessment of heavy metal removal by immobilized and dead bacterial cells: a biosorption approach. Global Journal of Environmental Research, 4(1), 23–30.
Kafilzadeh, F., Afrough, R., Johari, H., & Tahery, Y. (2012). Range determination for resistance/tolerance and growth kinetic of indigenous bacteria isolated from lead contaminated soils near gas stations (Iran). European Journal of Experimental Biology, 2(1), 62–69.
Kasprzak, K. S., Bal, W., & Karaczyn, A. A. (2003). The role of chromatin damage in nickel-induced carcinogenesis. A review of recent developments. Journal of Environmental Monitoring, 5, 183–187.
Kowall, N. W., Pendleury, W. W., & Kessler, J. B. (1989). Aluminium-induced neurofibrillary degeneration affects a subset of neurons in rabbit cerebral cortex, basal forebrain and upper brain stem. Neuroscience, 29, 329–337.
Lenaèrtovaè, V., Holovskaè, K., & Javorsky, P. (1998). The influence of mercury on the antioxidant enzyme activity of rumen bacteria Streptococcus bovis and Selenomonas ruminantium. FEMS Microbiology Ecology, 27, 319–325.
Master, C., Multhaup, G., & Simms, G. (1985). Neuronal origin of a cerebral amyloid: neurofibrillary tangles of Alzheimer’s disease contain the same protein as the amyloid of plaque cores and blood vessels. EMBO Journal, 4, 2757–2763.
Mondal, S., Roy, T., & Ray, A. K. (2010). Characterization and identification of enzyme-producing bacteria isolated from the digestive tract of bata, Labeo bata. Journal of the World Aquaculture Society, 41, 369–377.
Nath, S., Deb, B., & Sharma, I. (2012). Isolation and characterization of cadmium and lead resistant bacteria. Global Advance Research Journal of Microbiology, 1(11), 194–198.
Nies, D. (1999). Microbial heavy-metal resistance. Applied Microbiology and Biotechnology, 51, 730–750.
Pandey, S., Saha, P., Biswas, S., & Maiti, T. K. (2011). Characterization of two metals resistant Bacillus strains isolated from slag deposal site at Burnpur, India. Journal of Environmental Biology, 32, 773–779.
Pandey, S., Barai, P. K., & Maiti, T. K. (2013). Influence of heavy metals on the activity of antioxidant enzymes in the metal resistant strains of Ochrobactrum and Bacillus sp. Journal of Environmental Biology, 34, 1033–1037.
Poornima, M., Kumar, R. S., & Thomas, P. D. (2014). Isolation and molecular characterization of bacterial Strains from tannery effluent and reduction of chromium. International Journal of Current Microbiology and Applied Science, 3(4), 530–538.
Ren, W. X., Li, P. J., Geng, Y., & Li, X. J. (2009). Biological leaching of heavy metals from a contaminated soil by Aspergillus niger. Journal of Hazardous Materials, 167(1–3), 164–169.
Schalk, I. J., Hannauer, M., & Braud, A. (2011). New roles of bacterial siderophores in metal transport and tolerance. Environmental Microbiology, 13, 2844–2854.
Selvi, A. T., Anjugam, E., Devi, R. A., Madhan, B., Kannappan, S., & Chandrasekaran, B. (2012). Isolation and characterization of bacteria from tannery effluent treatment plant and their tolerance to heavy metals and antibiotics. Asian Journal of Experimental Biological Sciences, 3(1), 34–41.
Singh, A. K., Dhanjal, S., & Cameotra, S. S. (2014). Surfactin restores and enhances swarming motility under heavy metal stress. Colloids and Surfaces B: Biointerfaces, 116, 26–31.
Switala, J., & Leuwen, P. C. (2002). Diversity of properties among catalases. Archives of Biochemistry and Biophysics, 401, 145–154.
Thyssen, J. P., Gawkrodger, D. J., White, I. R., Julander, A., Menné, T., & Lidén, C. (2013). Coin exposure may cause allergic nickel dermatitis: a review. Contact Dermatitis, 68, 3–14.
Ugbenyen, A. M., & Okoh, A. I. (2013). Characteristics of a bioflocculant produced by a consortium of Cobetia and Bacillus species and its application in the treatment of waste waters. Water SA, 40(1), 139–1144.
World Health Organization. (1995). Inorganic lead. Geneva: Environmental Health Criteria. No. 165.
Acknowledgments
We are grateful to Helix Biogenesis for providing laboratory facilities and financial supports.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Banerjee, G., Pandey, S., Ray, A.K. et al. Bioremediation of Heavy Metals by a Novel Bacterial Strain Enterobacter cloacae and Its Antioxidant Enzyme Activity, Flocculant Production, and Protein Expression in Presence of Lead, Cadmium, and Nickel. Water Air Soil Pollut 226, 91 (2015). https://doi.org/10.1007/s11270-015-2359-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11270-015-2359-9