Abstract
In Vietnam, the two main viruses that cause disease in rice are the Rice grassy stunt virus (RGSV) and the Rice ragged stunt virus (RRSV). Outbreaks of these two viruses have dramatically decreased rice production in Vietnam. Because natural resistance genes are unknown, an RNAi strategy may be an alternative method to develop resistance to RGSV and RRSV. However, this strategy will be efficient only if putative silencing suppressors encoded by the two viruses are neutralized. To identify these suppressors, we used the classical green fluorescent protein (GFP) agroinfiltration method in Nicotiana benthamiana. Then, we investigated the effects of viral candidate proteins on GFP expression and GFP siRNA accumulation and their interference with the short- or long-range signal of silencing. RGSV genes s2gp1, s5gp2, and s6gp1 and RRSV genes s5gp1, s6gp1, s9gp1, and s10gp1 were selected for viral silencing suppressor investigation according to their small molecular weight, the presence of cysteines, or the presence of a GW motif in related protein products. We confirmed that protein p6 of RRSV displays mild silencing suppressor activity and affects long-range silencing by delaying the systemic silencing signal. In addition, we identified two new silencing suppressors that displayed mild activity: p2 of RGSV and p9 of RRSV.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
RNA silencing is a key defense mechanism against viral infection, and RNA interference (RNAi)-based strategies are now considered as an alternative means to provide an efficient method to control viral diseases by inducing specific resistance in plants [1–3]. RNA silencing is triggered by double-stranded RNAs generated from replicating viral RNAs that are cleaved into duplexes of small interfering RNA (siRNA) of 20–24 nucleotides by members of the RNase III family enzymes called Dicer or Dicer-like (DCL) in plants. One strand of the duplex is subsequently incorporated into an Argonaute (AGO)-containing RNA-induced silencing complex (RISC) to guide the cleavage of homologous viral RNA molecules [1, 2]. However, as a result of the co-evolution between plants and viruses, most viruses encode viral suppressors of RNA silencing (VSRs) to counteract the host RNA silencing pathway [4–6]. One of the most striking features of viral suppressor proteins is their diversity of structures and modes of action. Various types of viral proteins from coat proteins, movement proteins, proteases, and other proteins of previously unknown function have been shown to have silencing suppressor activities [7–10]. Identified VSRs can exhibit different strength levels with respect to suppression activity [11, 12]. They can act at different steps of the plant RNA silencing pathway. Currently, three modes of action for viral silencing suppressors could be distinguished. Firstly, they can inhibit viral siRNA production as it is the case for the helper component-proteinase (HC-Pro) from the Potyvirus which interferes with RNA silencing at a step upstream of the production of siRNA [9, 13, 14]. Secondly, they can act by binding siRNA duplexes, here the example model is the P19 protein from the Tombusvirus [15]. Thirdly, VSR can inactivate RISC by degrading the AGO protein or by inhibiting its cleavage activity as it is the case for the P0 of Polerovirus and the 2b of Cucumovirus, respectively [4].
In Vietnam, the rice cultivation area occupies approximately 4.2 million hectares. The Mekong Delta accounts for 52 % of the total production, which represents 90 % of the export market. However, such intensive production is threatened by various viruses such as Rice ragged stunt virus (RRSV) and Rice grassy stunt virus (RGSV) [16]. The most common way to control viruses is to use natural resistance [17, 18]. Unfortunately, until now, no natural resistance gene against RGSV and RRSV has been identified. An alternative way is to use pesticide for the brown plant hopper (BPH) control, the insect vector for these two viruses. However, such a strategy has severe environmental and human health impacts. Consequently, RNAi-based strategy appears to be an efficient method to control the durable control of RGSV and RRSV.
RGSV belongs to the Tenuivirus genus. This genus consists of 6 members including the reference species Rice stripe virus (RSV) [19]. RGSV particles are thread like, 6–8 nm wide, and mostly circular with a contour length of 200–2400 nm [20]. The RGSV genome contains six single-stranded RNA segments that encode 12 proteins. All segments contain two open reading frames (ORFs), each in an ambisense arrangement. The RNA 1 segment (s1gp1) encodes the p1 protein, whose function is unknown, and the RNA 1 complementary segment (s1gp2) encodes the pC1 protein, which is a putative viral RNA-dependent RNA polymerase (RdRp) [21, 22]. The RNA 5 complementary segment (s5gp2) encodes pC5, a nucleocapsid protein (NP) [23], and the RNA 6 segment (s6gp1) encodes p6, a movement protein (MP) [24]. The functions of the products of the other ORFs present in RNAs 2–6 are unknown. Vietnamese RGSV isolates exhibit low molecular diversity [25].
RRSV is the reference species of the Oryzavirus genus in the family Reoviridae. The RRSV genome contains 10 double-stranded RNAs ranging from 1.2 to 3.9 kb [26]. The RRSV virion is an icosahedral particle that consists of a polyhedral core surrounded by flat spikes approximately 20 nm wide and 10 nm high [27]. The RNA 7 (s7gp1) and 10 (s10gp1) segments encode non-structural proteins p7 and p10 with molecular weights (MW) of 68 and 33 kDa, respectively [28], the RNA 5 segment (s5gp) encodes the 91-kDa structural protein p5 [29], and the RNA 8 segment (s8gp1) encodes the 67-kDa structural protein p8, which is endowed with self-aggregation and self-cleavage abilities [30, 31]. The p9 protein encoded by s9gp1 is a 39-kDa protein that contributes largely as a viral spike protein and plays an important role in virus transmission by the insect vector [32, 33].
Resistance against RGSV using artificial siRNA was successfully obtained in different rice varieties [34, 35], but the durability of resistance was not reported. A way to improve the efficiency and durability of artificial siRNA-mediated resistance would be to target the genes that encode VSRs to limit the emergence of virus resistance breakdown. Consequently, before developing an RNAi resistance program, it is crucial to first characterize the VSRs encoded by RGSV and RRSV. In the Oryzavirus genus, p6 of RRSV was reported to be a VSR [36] and suppresses local silencing induced only by sense RNA. P6 might targets an upstream step of the dsRNA formation in the silencing pathway. In the Tenuivirus genus, only the NS3 protein of Rice hoja blanca virus (RHBV) was identified as a VSR [37], this one bind siRNA and miRNA in both plant and insect cells. Here, we selected RGSV and RRSV proteins to test their silencing suppressor activities.
Materials and methods
Transient expression assay using N. benthamiana
Viral genes were amplified from total RNA that was extracted from diseased plants using TriReagent (according to Sigma–Aldrich recommendations, http://www.sigmaaald-rich.com) and then reverse transcribed into complementary DNA (GoScript Reverse Transcription System from Promega, Madison, Wisconsin, USA). The primer pairs used to amplify each viral gene are listed in Table S2. PCR-amplified viral genes were cloned into the pGEM-T vector system (Promega) and sequenced. Viral genes were introduced into the pBin61 binary vector under the control of the 35S promoter and were introduced into Agrobacterium tumefaciens strain C58C1 for protein expression in planta [38]. A. tumefaciens-mediated infiltration (agroinfiltration) into 4-week-old Nicotiana benthamiana leaves was performed as previously described [39]. Silencing against the GFP reporter gene was induced by agroinfiltration of a GFP sequence (35S::GFP). A mixture of the two A. tumefaciens cultures in a 1:1 or 3:1 ratio, one carrying a pBin61 vector with a viral gene (35S::candidate gene) and the other carrying a construct expressing the GFP silencing inducer (35S::GFP) [40], was infiltrated into leaves of a wild-type or transgenic N. benthamiana line constitutively expressing a GFP construct (16c line). For fluorescence quantification experiment, two agroinfiltration spots per leaf of 4-weeks-old N. benthamiana 16c line were performed, one corresponding to the control (GFP plus empty vector) the other one corresponding to the suppressor candidate (GFP plus VSR candidate). Seven days post infiltration, the fresh leaf patches were cut out and scanned using a fluorescent scanner (Typhoon 9400, GE Lifesciences) with the blue laser excitation at 488 nm and a photomultiplier tube voltage at 550 nm. The GFP intensity level was quantified using Image Quant ver. 5.2 software for the control and the suppressor candidate patches. Fluorescence intensity of the leaf background was also evaluated for each leaf. For each leaf, this value was removed from the control and suppressor candidate patch values to reflect the density of the fluorescent only from the infiltrated leaf patches. For each candidate, 2 leaves of 3 plants were considerate and three independent experiments were performed.
RNA filter hybridization
Total RNA was extracted using TriReagent, and the expression levels of the VSR candidate genes were determined using RT-PCR (primers listed in Table S2). All genes were expressed in agroinfiltrated tissues (data not shown). Low molecular weight RNA was enriched from the total RNA samples using polyethylene glycol precipitation, and northern blot hybridizations were performed as described by Lacombe et al. [40]. Hybridizations were performed using a GFP oligonucleotide probe (5′-CTCTTGAAGAAGTCGTGCCGCTTCATATGA-3′) labeled with γ-32P-ATP using T4 polynucleotide kinase (Promega). To verify the equal RNA loading between samples, oligonucleotide complementary to the U6 snRNA (5′-TGTATCGTTCCAATTTTATCGGATGT-3′) was used as endogenous control.
Results
Selection of VSR candidate proteins
The RGSV and RRSV candidate proteins were selected first based on their relative low molecular weight, which applies to the majority of VSRs and second based on the presence of cysteines, which can participate in disulfide bridges or zinc finger motifs. Indeed, these two characteristics are possibly involved in silencing suppression mechanisms through a nucleic acid binding function [41–43]. In addition, candidate proteins were also selected for the presence of WG/GW motifs (the nucleotide sequence coding for Glycine—G and Tryptophan—W), which have been demonstrated to be involved in the suppression of silencing via AGO1 sequestration [44]. Based on these criteria, we selected 3 RGSV genes: s2gp1, s5gp2, and s6gp1 coding for proteins p2, pC5, and p6, respectively. For RRSV, proteins p5, p6, p9, and p10 encoded by the s5gp1, s6gp1, s9gp1, and s10gp1 genes, respectively, were selected (Table S1). Interestingly the p2 protein from RGSV belongs to a well-conserved protein superfamily of Tenuivirus_NS3, which includes proteins from RSV (Q01210.1, P26658.1), Maize stripe virus (P27208.1), and RHBV (Q67897.1). All these proteins were previously described as VSRs [45, 46], and the conserved domains characteristic of this superfamily are present in the p2 amino acid sequence (data not shown). The gene encoding protein p6 from RRSV was also selected as it was determined to be the first unique VSR identified thus far within the Oryzavirus genus [36].
Identification of RGSV and RRSV VSRs proteins
To screen for VSR activity, we used the transient A. tumefaciens assays in N. benthamiana that are classically used to identify VSRs [39, 47]. When the genes encoding for silencing suppression (35S::viral suppressor) are co-delivered with the GFP-induced silencing construct, GFP silencing is suppressed, which results in the restoration of GFP fluorescence in the agroinfiltrated patch [40].
The VSR P1Tz3 (from Rice yellow mottle virus (RYMV) Tanzanian isolate) [12] and P19 (from Cymbidium ringspot virus (CymRSV) [39] were used as positive controls of mild and strong VSR activities, respectively. For each tested protein, we performed agroinfiltration in three plants, and three independent experiments were performed.
In the first investigation, agroinfiltrations were performed using 2 leaves from each of 3 wild-type N. benthamiana plants. Leaf patches were visually observed under UV light at 3, 4, 5, 6, 8, and 10 days post infiltration (dpi). The intensity of GFP was compared with negative control patches that were agroinfiltrated without suppressor candidates (patch co-agroinfiltrated with empty pBin61 and a GFP silencing inducer) and positive control patches that were co-infiltrated with P1Tz3 and P19 suppressors (Table 1). From 3 to 6 dpi, the p2-RGSV and p9-RRSV patches displayed GFP expression that was similar to the patches agroinfiltrated with P1Tz3. However, at 8 dpi, the GFP intensity was lower compared with the patches that were agroinfiltrated with P1Tz3. The other candidate proteins did not exert a significant effect on the GFP fluorescence intensity of the agroinfiltrated patches and were similar to the negative control. Surprisingly, p6 from RRSV (previously reported as a VSR [36]) did not show any suppression activity in our assay, and the GFP fluorescence intensity was similar to that of the negative control. Because p6-RRSV was previously identified as a VSR, we repeated the experiment using the same conditions of Wu and colleagues [36]. They used the N. benthamiana line 16c and a 3:1 mixing ratio between the Agrobacterium culture of p6-RRSV and that of the GFP silencing inducer instead of the 1:1 ratio used in the experiment described above. Moreover, in our first investigation, GFP fluorescence intensity was visually estimated under UV light. Assuming that the method might not be sensitive enough to detect minor differences in fluorescence intensity, this intensity was monitored using a fluorescent scanner (Typhoon 9400). Using these optimized conditions, we tested the VSR activity of p6-RRSV and the two new silencing suppressors (p2-RGSV and p9-RRSV) identified using the previous visual method. The pC5-RGSV construct was included as a non-suppressor candidate. In addition, we considered the empty vectors pBin61 and P1Tz3 as negative and positive controls, respectively.
Leaf patches were collected at 7 dpi, and the GFP fluorescence levels were quantified (Figs. 1 and 2). Three independent experiments were performed. Data were analyzed using one-way ANOVA, and comparisons of the mean GFP intensity were contrasted using Duncan’s multiple range test. All statistical analyses were performed at a P value less than 0.05 using Statistica 10 (StatSoft). In these experiments, the mild VSR activity of p2-RGSV and p9-RRSV was confirmed (Fig. 2). In addition, mild VSR activity was detected for p6-RRSV (Figs. 1 and 2), whereas pC5-RGSV and the empty vector pBin61 displayed no detectable VSR activity (Fig. 2).
Effect of p6 from RRSV on GFP silencing in N. benthamiana 16c lines. P1Tz3 or p6 and the GFP silencing inducer constructs were co-infiltrated into GFP overexpression N. benthamiana 16c line leaves. The suppressor and GFP Agrobacterium cultures (OD600 = 0.6) were mixed in a 3:1 (v/v) ratio and used for leaf agroinfiltration. GFP intensity detection using a fluorescent scanner (Typhoon 9400) was measured in leaf patches after 7 dpi. The scanning parameters were Blue 2 (488 nm), PMT 550. a GFP fluorescence is shown as a dark color. The intensity is correlated with the darkness of the patches. b GFP intensity was quantified using Image Quant ver. 5.2 software (negative control: pBIN61 empty vector). Two leaves from each of three plants were analyzed, and three independent experiments were performed. Results are averages. The differences observed between values noted a and b were significant at p value < 0.05
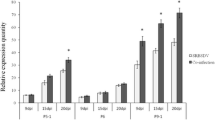
Effects of the viral protein candidates on GFP silencing in N benthamiana 16c lines. Leaves of GFP overexpression N. benthamiana 16c lines were co-infiltrated with viral candidate genes and GFP inducer constructs. The ratio between the candidate proteins and GFP inducer constructs was 3:1 (v:v). P1Tz3 (Rice yellow mottle virus) was used as a positive control for the mild silencing suppressor protein. Leaf patches collected at 7 dpi were scanned using the fluorescent scanner Typhoon 9400. The GFP intensity level was quantified using Image Quant ver. 5.2 software. Two leaves from each of three plants were analyzed, and three independent experiments were performed. Results are averages. The differences observed between values noted a and b were significant at p value < 0.05
P19 from CymRSP and P1 from RYMV suppress the silencing pathway by interacting with either one or both 21- and 24-nt siRNAs [39, 40, 48], leading to a strong decrease in siRNA accumulation. To determine whether the viral candidate proteins affect the accumulation of the GFP siRNA in agroinfiltrated leaves, we measured the accumulation of GFP siRNA using northern blot analysis. Using either 1:1 or 3:1 ratio (suppressor: inducer) in the wild type and 16c lines of N. benthamiana, respectively, agroinfiltrated leaf patches were collected after 7 dpi. We used P19 and P1Tz3 as strong and mild suppressor positive controls, respectively. The data are presented in Fig. 3A and 3B. As expected, the accumulation of GFP 21- and 24-nt siRNA was reduced in the presence of P19 [39, 48], while the quantity of GFP 24-nt siRNA was only specifically reduced in the presence of P1Tz3 [40]. However, for other candidate proteins, the level of accumulation of the 21- and 24-nt GFP siRNA was not dramatically affected compared to the negative control (empty pBin61 and GFP inducer). Same observations were made whatever the suppressor:inducer ratio used (Fig. 3a, b). These results reveal that viral candidate proteins from RGSV/RRSV do not severally modify GFP siRNA accumulation.
Effects of the candidate viral proteins from RGSV and RRSV on GFP siRNA accumulation. Leaves of wild-type N. benthamiana were co-infiltrated with viral candidate genes and a GFP inducer either in a 1:1 (v/v) ratio, (a) or in a 3:1 (v:v) ratio (b). Leaf patches were collected at 7 dpi, and RNA was extracted as described by Lacombe et al. (2010). The GFP and U6 probes were end-labeled with [γ-32P] ATP using T4 polynucleotide kinase (Promega, http://www.promega.com), and northern blots were performed as described by Lacombe et al. [32]. U6 hybridization provides a control for RNA loading of the gels
Effect of p2-RGSV, p6-RRSV, and p9-RRSV on short- and long-range silencing signals
In plants, the silencing signal spreads between cells through a short-distance movement of 10–15 cells (short-range silencing movement) [49, 50]. In leaves of the N. benthamiana 16c line displaying transiently induced GFP silencing patches, the short-distance movement was well visualized as a thin red border at the margin of the agroinfiltrated zone [40]. To investigate the effects of RGSV/RRSV proteins on this limited movement, the silenced borders of the infiltration zones expressing P19, viral candidate proteins from RGSV and RRSV, or no suppressor were compared. In addition, in plants, the long-distance movement of the silencing signal can spread to the upper leaves, leading to systemic silencing (long-range silencing movement) [50]. The effects of the viral candidate proteins on long-range silencing movement were also investigated. The progression of GFP silencing in the newly emerging leaves of three N. benthamiana (line 16c) plants that were agroinfiltrated with the different viral proteins was examined over time (Table 2). In the negative control (empty pBin61), short- and long-range silencing movements were observed. By contrast, P19 completely blocked both the short- and the long-range systemic silencing movements as previously reported [49]. None of the viral candidate proteins affected the short-range silencing movement. Among all tested proteins, only p6-RRSV displayed an inhibitory effect (from 9 to 17 dpi) on the long-range silencing movement (Table 2). This was confirmed in another experiment in which the number of agroinfiltrated plants was increased from three to fifteen. These results were presented in Fig. 4 and confirmed that the long-range silencing signal movement was delayed by p6-RRSV. This was particularly clear until 10 dpi, when approximately 50 % of the plants that were agroinfiltrated with the empty pBIN61 vector showed systemic long-range silencing of GFP in the emerging leaves, while no systemic silencing was observed in the plants agroinfiltrated with p6-RRSV construct. However, from 10 dpi, the systemic signal inhibition by p6-RRSV was not maintained, and some plants displayed GFP silencing in the emerging leaves at this time. From 10 to 24 dpi, the number of plants exhibiting GFP silencing in the emerging leaves progressively reached approximately 30 %. This proportion was ~50 % of the plants that were agroinfiltrated with the empty pBin61 vector.
RRSV-P6 delays long-range silencing movement. Leaves of transgenic 16c N. benthamiana were co-infiltrated with p6 and GFP silencing inducer. The strong silencing suppressor P19 from Cymbidium ringspot virus was used as a positive control; P19 blocks short- and long-range signal movements. pBin61 empty vector was used as a negative control. A. tumefaciens cultures (OD600 = 0.6) were mixed at a 1:1 (v:v) ratio. Fifteen plants were used for each construction, and agroinfiltration was performed on 2 leaves from each plant. The plants were observed under UV light at 7, 10, 12, 15, 18, 24, 25, and 30 dpi, and the percentage of plants displaying silencing of systemic tissues is indicated
Discussion
It was previously determined that p6-RRSV is a VSR. In addition, we determined that p2-RGSV and p9-RRSV exhibited VSR activity. These three proteins suppressed the RNA silencing pathway when co-infiltrated with the GFP silencing inducer. A comparison with the mild suppressor P1Tz3 [40] suggests that these proteins may also have mild silencing suppression activity. As described for p6-RRSV [36], this mild activity was revealed by the 3:1 (suppressor:inducer, v:v) ratio rather than the 1:1 ratio, which suggests that the ratio between the VSR and the silencing inducer in the transient expression assay in N. benthamiana may be important to identify VSRs with mild activity. In addition, p6-RRSV appeared to act differently than p2-RGSV and p9-RRSV. Indeed, although p2-RGSV, p6-RRSV, and p9-RRSV did not block the short-range silencing signal movement, the long-range silencing signal was delayed by p6-RRSV.
Interestingly, p6-RRSV and p2-RGSV, and p1-RGSV are predicted to be probable VSR using the VSR prediction web tool, plant VsupPred recently developed (http://bioinfo.icgeb.res.in/pvsup/submit.html) [51]. Here, we functionally confirmed these predictions for two of them, p6-RRSV and p2-RGSV, the third one p1-RGSV was not tested here.
The expression of these three VSRs did not dramatically modify the accumulation of GFP siRNA. A previous work reported that p6-RRSV would affect GFP siRNA accumulation in N. benthamiana. However, the reduction observed by northern blot was quite moderated compared to the effect of TAV2b VRS control [36]. We can speculate that this reduction in siRNA accumulation induced by p6-RRSV would not be important enough to be detected by northern blot performed in our conditions. Because of the absence of detectable effect on siRNA accumulation, we propose that p2-RGSV, p6-RRSV, and p9-RRSV, which displayed mild VSR activity in our experiments, may interact directly or indirectly with key components of the silencing pathway other than siRNA, as has been described for P38 from Turnip crinkle virus and P0 from Beet western yellow virus, which both inhibit AGO1 [11], and P2 from RSV, which interacts with OsSGS3 [52, 53]. The means by which these VSRs suppress the silencing pathway needs to be further investigated.
Finally, we have shown that, similar to previous reports, many viruses encode more than one VSR [54]. Here, we confirmed p6-RRSV as a mild VSR, and we identified p9-RRSV as a new mild VSR for RRSV. Because we did not investigate all the proteins encoded by the two viruses studied here, we cannot rule out the possibility that other VSRs might be encoded by RRSV and RGSV. In addition, because these two viruses are often found together in the same plants, at least 3 VSRs may act synergistically to initiate infection throughout the plant. Because of their silencing suppressor activities and potential trans effects in rescuing silenced VSR activity, these three VSRs should be targeted together (possibly with the other remaining VSRs encoded by RRSV and RGSV) to achieve efficient RNAi resistance against RRSV and RGSV.
References
S.W. Ding, O. Voinnet, Antiviral immunity directed by small RNAs. Cell 130, 413–426 (2007)
S.W. Ding, RNA-based antiviral immunity. Nat. Rev. Immunol. 10, 632–644 (2010)
B. Tabassum, I.A. Nasir, U. Aslam, T. Husnain, How RNA Interference Combat Viruses in Plants, Functional Genomics, Dr. Germana Meroni (Ed.), ISBN: 978-953-51-0727-9, InTech,DOI:10.5772/51870.Availablefrom:http://www.intechopen.com/books/functional-genomics/how-rna-interference-combat-viruses-in-plants (2012)
M. Incarbone, P. Dunoyer, RNA silencing and its suppression: novel insights from in planta analyses. Trends Plant Sci. 18, 382–392 (2013)
L. Lakatos, T. Csorba, V. Pantaleo, E.J. Chapman, J.C. Carrington, Y.P. Liu, V.V. Dolja, L.F. Calvino, J.J. Lopez-Moya, J. Burgyan, Small RNA binding is a common strategy to suppress RNA silencing by several viral suppressors. EMBO J. 25, 2768–2780 (2006)
Z. Merai, Z. Kerenyi, S. Kertesz, M. Magna, L. Lakatos, D. Silhavy, Double-stranded RNA binding may be a general plant RNA viral strategy to suppress RNA silencing. J. Virol. 80, 5747–5756 (2006)
J.A. Díaz-Pendón, S.W. Ding, Direct and indirect roles of viral suppressors of RNA silencing in pathogenesis. Annu. Rev. Phytopathol. 46, 303–326 (2008)
H.S. Guo, S.W. Ding, A viral protein inhibits the long range signaling activity of the gene silencing signal. EMBO J. 21, 398–407 (2002)
C. Llave, K.D. Kasschau, J.C. Carrington, Virus-encoded suppressor of posttranscriptional gene silencing targets a maintenance step in the silencing pathway. Proc. Natl. Acad. Sci. U.S.A. 97, 13401–13406 (2000)
B.M. Roth, G.J. Pruss, V.B. Vance, Plant viral suppressors of RNA silencing. Virus Res. 102, 97–108 (2004)
J. Lin, C. Wei, Y. Li, Viral suppression of RNA silencing. Sci. China Life Sci. 55, 109–118 (2012)
C. Siré, M. Bangratz-Reyser, D. Fargette, C. Brugidou, Genetic diversity and silencing suppression effects of rice yellow mottle virus and the P1 protein. Virol. J. 5, 1–12 (2008)
G. Brigneti, O. Voinnet, W.X. Li, L.H. Ji, S.W. Ding, D.C. Baulcombe, Viral pathogenicity determinants are suppressors of transgene silencing in Nicotiana benthamiana. EMBO J. 17, 6739–6746 (1998)
C. Mallory, L. Ely, T.H. Smith, R. Marathe, R. Anandalakshmi, M. Fagard, H. Vaucheret, G. Pruss, L. Bowman, V.B. Vance, HC-Pro suppression of transgene silencing eliminates the small RNAs but not transgene methylation or the mobile signal. Plant Cell 13, 571–583 (2001)
K. Ye, L. Malinina, D.J. Patel, Recognition of small interfering RNA by a viral suppressor of RNA silencing. Nature 426, 874–878 (2003)
N.V. Vien, H.M. Trung, Overview about rice virus and Rice grassy stunt virus, rice ragged stunt virus protection in Mekong Delta. J. Agric. Agric. Dev. 18, 9–13 (2006)
P. Cournoyer, S.P. Dinesh-Kumar, NB-LRR immune receptors in plant virus defence, in Recent Advances in Plant Virology, ed. by C. Caranta, M.A. Aranda, M. Tepfer, J.J. Lopez-Moya (Caister Academic Press, Norfolk, 2011), pp. 149–176
P. Gomez, A. Rodriguez-Hernandez, B. Moury, M. Aranda, Genetic resistance for the sustainable control of plant virus diseases: breeding, mechanisms and durability. Eur. J. Plant Pathol. 125, 1–22 (2009)
M.A. Mayo, J.E. De Miranda, B.W. Falk, R. Goldbach, A.L. Haenni, S. Toriyama, Genus Tenuivirus, in Virus Taxonomy, ed. by M.H. van Regenmortel, C.M. Fauquet, D.H.L. Bishop, E.B. Carstens, M.K. Estes, S.M. Lemon, J. Maniloff, M.A. Mayo, D.J. McGeoch, C.R. Pringle, R.B. Wickner (Academic Press, NewYork, 2000), pp. 904–908
H. Hibino, T. Usugi, T. Omura, T. Tsuchizaki, K. Shohara, M. Iwasaki, Rice grassy stunt virus: a planthopper-borne circular filament. Phytophatology 75, 894–898 (1985)
G.J. Miranda, O. Azzam, Y. Shirako, Comparison of nucleotide sequences between northern and southern philippine isolates of rice grassy stunt virus indicates occurrence of natural genetic reassortment. Virology 266, 26–32 (2000)
S. Toriyama, T. Kimishima, M. Takahashi, T. Shimizu, N. Minaka, K. Akutsu, The complete nucleotide sequence of the rice grassy stunt virus genome and genomic comparisons with viruses of the genus Tenuivirus. J. Gen. Virol. 79, 2051–2058 (1998)
S. Toriyama, T. Kimishima, M. Takahashi, The proteins encoded by rice grassy stunt virus RNA5 and RNA6 are only distantly related to the corresponding proteins of other members of the genus Tenuivirus. J. Gen. Virol. 78, 2355–2363 (1997)
A. Hiraguri, O. Netsu, T. Shimizu, T. Uehara-Ichiki, T. Omura, N. Sasaki, H. Nyunoya, T. Sasaya, The nonstructural protein pC6 of rice grassy stunt virus trans-complements the cell-to-cell spread of a movement-defective tomato mosaic virus. Arch. Virol. 156, 911–916 (2011)
H.A. Ta, D.P. Nguyen, S. Causse, T.D. Nguyen, V.V. Ngo, E. Hebrard, Molecular diversityof Rice grassy stunt virus in Vietnam. Virus Genes 46, 383–386 (2013)
T. Omura, Y. Minobe, I. Kimura, H. Hibino, T. Tsuchizaki, Y. Saito, Improved purification procedure and RNA segments of Rice ragged stunt virus. Ann. Phytopathol. Soc. Jpn. 49, 670–675 (1988)
K. Hagiwara, Y. Minobe, Y. Nozu, H. Hibino, I. Kimura, T. Omura, Component proteins and structures of rice ragged stunt virus. J. Gen. Virol. 67, 1711–1715 (1986)
N.M. Upadhyaya, K. Ramm, J.A. Gellatly, Z. Li, W. Kositratana, P.M. Waterhouse, Rice ragged stunt oryzavirus genome segments S7 and S10 encode non-structural proteins of M(r) 68,025 (Pns7) and M(r) 32,364 (Pns10). Arch. Virol. 142, 1719–1726 (1997)
Z. Li, N.M. Upadhyaya, W. Kositratana, A.J. Gibbs, P.M. Waterhouse, Genome segment 5 of rice ragged stunt virus encodes a virion protein. J. Gen. Virol. 77, 3155–3160 (1996)
H.H. Lu, Z.X. Gong, T.Q. Cao, Studies on the genomic coding assignments of rice ragged stunt virus. Chin. J. Virol. 6, 167–172 (1990)
N.M. Upadhyaya, E. Zinkowsky, W. Kositratana, P.M. Waterhouse, The Mr 43 K major capsid protein of rice ragged stunt Oryzavirus is a posttranslationally processed product of a Mr 67,348 polypeptide encoded by genome segment 8. Arch. Virol. 141, 1689–1701 (1996)
C.G. Shao, H.J. Lü, J.H. Wu, Z.X. Gong, Nucleic acid binding activity of Pns6 encoded by genome segment 6 of rice ragged stunt oryzavirus. Acta Biochim. Biophys. Sin. 36, 457–466 (2004)
G.Y. Zhou, X.B. Lu, H.J. Lu, J.L. Lei, S.X. Chen, Z.X. Gong, Rice ragged stunt oryzavirus: role of the viral spike protein in transmission by the insect vector. Ann Appl. Biol. 135, 573–578 (1999)
T. Shimizu, E. Nakazono-Nagaoka, T. Uehara-Ichiki, T. Sasaya, T. Omura, Targeting specific genes for RNA interference is crucial to the development of strong resistance to Rice stripe virus. Plant Biotechnol. J. 9, 503–512 (2011)
T. Shimizu, T. Ogamino, A. Hiraguri, E. Nakazono-Nagaoka, T. Uehara-Ichiki, M. Nakajima, K. Akutsu, T. Omura, T. Sasaya, Strong Resistance Against Rice grassy stunt virus is induced in transgenic rice plants expressing double-stranded RNA of the viral genes for nucleocapsid or movement proteins as targets for RNA Interference. Virology 103, 513–519 (2013)
J. Wu, Z. Du, C. Wang, L. Cai, M. Hu, Q. Lin, Z. Wu, Y. Li, L. Xie, Identification of Pns6, a putative movement protein of RRSV, as a silencing suppressor. Virol. J. 7, 335–341 (2010)
E. Bucher, T. Sijen, P. de Haan, R. Goldbach, M. Prins, Negative-strand tospoviruses and tenuiviruses carry a gene for a suppressor of gene silencing at analogous genomic positions. J. Virol. 77, 1329–1336 (2003)
C.M. Hamilton, A. Frary, C. Lewis, S.D. Tanksley, Stable transfer of intact high molecular weight DNA into plant chromosomes. Proc. Natl. Acad. Sci. U.S.A. 93, 9975–9979 (1996)
A. Hamilton, O. Voinnet, L. Chappell, D. Baulcombe, Two classes of short interfering RNA in RNA silencing. EMBO J. 21, 4671–4679 (2002)
S. Lacombe, M. Bangratz, F. Vignols, C. Brugidou, The rice yellow mottle virus P1 protein exhibits dual functions to suppress and activate gene silencing. Plant J. 61, 371–382 (2010)
F.X. Gillet, D.I. Cattoni, S. Petitot-Becard, F. Delalande, V. Poignavent, J.P. Brizard, Y. Bessin, A. Van Dorsselaer, N. Declerck, S. Sanglier-Cianferani, C. Brugidou, F. Vignols, The RYMV-encoded viral suppressor of RNA silencing P1 is a Zinc-binding protein with redox-dependent flexibility. J. Mol. Biol. 425, 2423–2435 (2013)
A.C. Jamieson, S.H. Kim, J.A. Wells, In vitro selection of zinc fingers with altered DNA-binding specificity. Biochemistry 33, 5689–5695 (1994)
V.R. Wezel, H. Liu, Z. Wu, J. Stanley, Y. Hong, Contribution of the zinc finger to zinc and DNA binding by a suppressor of posttranscriptional gene silencing. J. Virol. 77, 696–700 (2003)
H. Jin, J.K. Zhu, A viral suppressor protein inhibits host RNA silencing by hooking up with Argonautes. Genes Dev. 24, 853–856 (2010)
H. Hemmes, L. Lakatos, R. Goldbach, J. Burgyan, M. Prins, The NS3 protein of Rice hoja blanca tenuivirus suppresses RNA silencing in plant and insect hosts by efficiently binding both siRNAs and miRNAs. RNA 13, 1079–1089 (2007)
G. Wu, J. Wang, Y. Yang, B. Dong, Y. Wang, G. Sun, C. Yan, F. Yan, J. Chen, Transgenic rice expressing rice stripe virus NS3 protein, a suppressor of RNA silencing, shows resistance to rice blast disease. Virus Genes 48, 566–569 (2014)
O. Voinnet, D.C. Baulcombe, Systemic signalling in gene silencing. Nature 389, 553 (1997)
D. Silhavy, A. Molnar, A. Lucioli, G. Szittya, C. Hornyik, M. Tavazza, J. Burgyan, A viral protein suppresses RNA silencing and binds silencing-generated 21-to 25-nucleotide double-stranded RNAs. EMBO J. 21, 3070–3080 (2002)
C. Himber, P. Dunoyer, G. Moissiard, C. Ritzenthaler, O. Voinnet, Transitivity-dependent and-independent cell-to-cell movement of RNA silencing. EMBO J. 22, 4523–4533 (2003)
P. Dunoyer, C. Himber, O. Voinnet, DICER LIKE 4 is required for RNA interference and produces the 21 nucleotide small interfering RNA component of the plant cell to cell silencing signal. Nat. Genet. 37, 1356–1360 (2005)
Z. Jagga, D. Gupta, Supervised learning classification models for prediction of plant virus encoded RNA silencing suppressor. PLoS ONE 9, e97446 (2014)
Z. Du, D. Xiao, J. Wu, D. Jia, Z. Yuan, Y. Liu, L. Hu, Z. Han, T. Wei, Q. Lin, Z. Wu, L. Xie, P2 of Rice stripe virus (RSV) interacts with OsSGS3 and is a silencing suppressor. Mol. Plant. Pathol. 12, 808–814 (2011)
E. Glick, A. Zrachya, Y. Levy, A. Mett, D. Gidoni, E. Belausov, V. Citovsky, Y. Gafni, Interaction with host SGS3 is required for suppression of RNA silencing by tomato yellow leaf curl virus V2 protein. Proc. Natl. Acad. Sci. U.S.A. 105, 157–161 (2008)
R. Vanitharani, P. Chellappan, J.S. Pita, C.M. Fauquet, Differential roles of AC2 and AC4 of cassava geminiviruses in mediating synergism and suppression of posttranscriptional gene silencing. J. Virol. 78, 9487–9498 (2004)
Acknowledgments
We greatly thank Prof. David Baulcombe, the Gatsby Charitable Foundation and PBL (Norwich), for providing the N. benthamiana 16c line. We also thank Dr. Thomas Kroj, INRA for very useful suggestion regarding GFP scanning technique. We are very grateful to Dr. D Sérémé for statistical analysis. TDN was supported by an IRD ARTS PhD fellowship.
Author contributions
SL and CB designed the research; TN, SL, and MB performed viral genes cloning, siRNA expression and transient expression of 35S::candidate gene, and the silencing inducer 35S::GFP in N. benthamiana. TN performed experimentation described in Fig. 4 and Tables 1 and 2. HT, DV, and PG contributed for analysis data and discussion. TN, SL, CB, and PG have written the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by Thomas Hohn.
Thanh Duc Nguyen and Séverine Lacombe contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11262_2015_1229_MOESM1_ESM.docx
Supplemental data Table S1: RGSV and RRSV viral proteins. The ones selected for analysis are noted in bold.Supplemental data Table S2: List of primers used for virus gene amplification. Supplementary material 1 (DOCX 21 kb)
Rights and permissions
About this article
Cite this article
Nguyen, T.D., Lacombe, S., Bangratz, M. et al. p2 of Rice grassy stunt virus (RGSV) and p6 and p9 of Rice ragged stunt virus (RRSV) isolates from Vietnam exert suppressor activity on the RNA silencing pathway. Virus Genes 51, 267–275 (2015). https://doi.org/10.1007/s11262-015-1229-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-015-1229-2