Abstract
Inducible Cre–ERT recombinase technology is widely used for gene targeting studies. The second generation of inducible Cre–ERT recombinase, hemizygous B6.129S-Tg(UBC-cre/ERT2)1Ejb/J (hereafter abbreviated as Cre–ERT2), a fusion of a mutated estrogen receptor and Cre recombinase, was engineered to be more efficient and specific than the original Cre–ERT. The putative mechanism of selective Cre-mediated recombination is Cre sequestration in the cytoplasm in the basal state with translocation to the nucleus only in the presence of tamoxifen. We utilized both a reporter mouse (B6.129 (Cg)-Gt(ROSA)26Sor tm4(ACTB-tdTomato,-EGFP)Luo/J) and endothelin converting enzyme-1 floxed transgenic mouse line to evaluate Cre–ERT2 activity. We observed spontaneous Cre activity in both settings. Unintended Cre activity is a confounding factor that has a potentially large impact on data interpretation. Thus, it is important to consider background Cre activity in experimental design.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The development of conditional knockout alleles has enabled the study of gene function in a variety of settings that were previously beyond the scope of simple gene-targeting strategies. Tissue-specific Cre drivers permit the assessment of anatomically restricted gene ablation and examination of embryonically lethal mutations in in post-natal animals. Furthermore, drug regulated Cre strains (Feil et al. 1997; Gossen and Bujard 1992; Johansen et al. 2002; Pluta et al. 2005) can also be used to circumvent developmental phenotypes by delaying gene ablation until a desired time point.
Inducible Cre–ERT recombinase is one of several systems that has allowed researchers to manipulate gene expression in a time dependent manner. The second generation of inducible Cre–ERT recombinase was developed to be more sensitive and specific than the original Cre–ERT (Indra et al. 1999; Jaisser 2000). Hemizygous B6.129S-Tg(UBC-cre/ERT2)1Ejb/J (hereafter abbreviated as Cre–ERT2), a second generation inducible Cre–ERT recombinase, mouse line was generated through lentiviral transgenesis (Lois et al. 2002; Ruzankina et al. 2007). The construct has the common feature of a Cre recombinase-mutated estrogen receptor (Cre–ERT2) protein fusion that is expressed under the control of the Ubiquitin C (UBC) promoter (Ruzankina et al. 2007). Previous in vitro experiments showed that the Cre–ERT2 was limited to the cytoplasm in the absence of tamoxifen (Feil et al. 1997). Past reports have also claimed that Cre–ERT2 is activated exclusively by tamoxifen or 4-hydroxytamoxifen (Feil et al. 1997; Indra et al. 1999; Ruzankina et al. 2007). In the presence of tamoxifen, Cre–ERT2 translocates to the nucleus and mediates recombination (Feil et al. 1997; Indra et al. 1999), but in its absence Cre–ERT2 is limited to the cytoplasm. Background Cre activity can preclude the accurate interpretation of gene function. Therefore, it is important to assess Cre-recombinase activity in the absence of tamoxifen.
Reporter mice are often used to evaluate the spatial and temporal extent of Cre-mediated recombination. In a double fluorescent Cre recombinase reporter mouse B6.129(Cg)-Gt(ROSA)26Sor tm4(ACTB-tdTomato,-EGFP)Luo/J (hereafter abbreviated as mT/mG), cells express membrane targeted tdTomato (mT) prior to Cre excision and membrane targeted enhanced green fluorescent protein (EGFP) (mG) following Cre excision (Muzumdar et al. 2007). The mT/mG cassette is expressed under the control of the CMV β-actin enhanced promoter (pCA). The double fluorescent marker system allows visualization of both recombined and non-recombined cells.
We used the mT/mG mouse line to evaluate Cre–ERT2 activity in vivo. In addition, we also utilized Ece1 flox/flox mice to confirm Cre activity in another targeted gene system. Ece1 encodes for endothelin converting enzyme-1, a key enzyme in the proteolytic processing of big endothelins (big ETs) to mature, active endothelins (ETs). Here, we report that Cre–ERT2 exhibits spontaneous activity in vivo, and that its extent varies among individual animals and among tissues within the same animal.
Materials and methods
Targeted mouse line production
The generation of mT/mG and Cre–ERT2 mice has been described (Lois et al. 2002; Muzumdar et al. 2007; Ruzankina et al. 2007). Both mT/mG and Cre–ERT2 mice were purchased from Jackson Laboratory. mT/mG and Cre–ERT2 mice were intercrossed to generate mT/mG and Cre–ERT2 mT/mG mice (Fig. 1a). By 8 ± 1 weeks, the mice were either left untreated, or treated with vehicle (98% corn oil, 2% ethanol) or tamoxifen (MP Biomedicals) at a concentration of 0.5 µM/g of body weight dissolved in 98% corn oil and 2% ethanol once daily for 5 days via intraperitoneal (IP) injection. Both mT/mG and Cre–ERT2 mT/mG were maintained to an age of 10 ± 1 weeks.
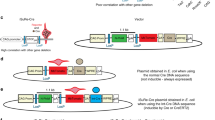
Generation of Cre–ERT2 mT/mG mice and inducible Ece1 Knockout (KO) mice. a A schematic showing the generation of Cre–ERT2 mT/mG mice. Mice harboring mT/mG gene were treated with tamoxifen. Mice harboring Cre–ERT2 mT/mG gene were left untreated or treated with tamoxifen or vehicle at 8 ± 1 weeks of age. b A schematic showing the generation of inducible Ece1 KO mice. T Mice harboring Ece1 flox/flox allele were intercrossed with Cre–ERT2 Ece1 +/ Δ mice to generate the experimental animals. These mice were then treated with either tamoxifen or vehicle at 8 ± 1 weeks of age
The conditional Ece1 knockout mouse construct was purchased from the knockout mouse project (KOMP). The construct was made on a C57BL/6 N background. Four generation backcrossing with C57BL/6 J were performed to generate Ece1 KOMP mice with C57BL/6 J background. The Ece1 KOMP mice were then initially bred to transgenic mice harboring FLP recombinase (B6.Cg–Tg (ACTFLPe)9205Dym/J, purchased from Jackson laboratory) and subsequently bred to EIIA Cre recombinase (B6.FVB–Tg(EIIa-cre)C5379Lmgd/J, purchased from Jackson laboratory) mice to generate the Ece1 flox and Ece1 Δ alleles, respectively. Ece1 +/flox mice were intercrossed to generate Ece1 flox/flox mice. Ece1 +/ Δ mice were then intercrossed with Cre–ERT2 mice to generate Cre–ERT2 Ece1 +/ Δ mice. Ece1 flox/flox and Cre–ERT2 Ece1 +/ Δ mice were intercrossed to generate Ece1 +/flox, Ece1 Δ/flox, Cre–ERT2 Ece1 +/flox and Cre–ERT2 Ece1 Δ/flox mice (Fig. 1b). All experimental animals were left untreated or injected with either vehicle or tamoxifen at 8 ± 1 weeks of age.
Mice were housed 2–5 mice/500 cm2 cage with a 12 h light–dark cycle, given acidified water, and fed laboratory rodent chow 5001 (PMI Nutrition International, Richmond, IN, USA) ad libitum according to the guidelines set forth by “The Guide for the Care and Use of Laboratory Animals”. All animal procedures were approved by the William S. Middleton VA Hospital Institutional Animal Care and Use Committee (IACUC).
Genotyping
DNA was isolated from tail snips and tissue samples using the IBI genomic DNA mini kit (Peosta, IA USA) following manufacturer’s protocol. For the PCR reaction, we used Failsafe PCR system (Epicentre Technologies, Madison WI) with PreMix J for the (mT/mG transgene) and Premix H for Ece1 flox allele genotyping.
The mT/mG transgene and Cre–ERT2 allele were identified by PCR from tail DNA preparations. Ece1 KOMP, Ece1 flox and Ece1 Δ alleles were genotyped by PCR using tail and tissue DNA preparations. The list of primers and PCR setting is summarized in Table 1.
The transgene containing the Cre allele was identified by quantitative Real-time PCR reactions, performed in a StepOne RT-PCR instrument (Life Technologies, Carlsbad, CA, USA) using TaqMan assay (Life Technologies, Carlsbad, CA, USA) following manufacturer’s recommended protocol. The TaqMan copy number assay is Mr00635245_cn (Life Technologies, Carlsbad, CA, USA).
Tissue preparation and histology
Mice were euthanized using isoflurane followed by cervical dislocation. For cryosection preparation, kidney, liver, heart and lung were perfused with 4% cold paraformaldehyde in 0.1 M phosphate buffered saline (PBS), fixed for 6–24 h at 4 °C, cryoprotected in 30% sucrose overnight at 4 °C and embedded in OCT, following a previously described method (Muzumdar et al. 2007). Samples were stored at −80 °C until use. Ten micron sections were obtained using a Leica CM1800 cryostat, washed three times with PBS, counterstained with DAPI, washed an additional three times and mounted.
Data and image analysis
Two-dimensional and z-stack confocal images were obtained with a Nikon C1 confocal microscope and EZ-C1 software (v. 3.91) in 1.0 μm slices with a 400× objective. Images were processed with Image J (FIJI).
Results
UBC Cre–ERT2 activity in mT/mG mice
Reporter mice that only carried the mT/mG allele (n = 6) did not emit mG fluorescence in the green channel in any of the tissues examined (Fig. 2), indicating that mG labeling was Cre dependent as previously shown (Muzumdar et al. 2007). In the tamoxifen treated Cre–ERT2 mT/mG mice (n = 6), we observed approximately ~50–90% of mG positive cells in all tissues examined (Fig. 2). Both vehicle (n = 4) and untreated Cre–ERT2 mT/mG (n = 4) mice also unexpectedly showed >50 and 10–80% mG positive cells, respectively, in the kidney and liver (Figs. 2, 3). In the kidney, mG positive cells were observed primarily in the mesangium and tubules. In the liver, mG labeling was observed in the hepatocytes. In heart and lung, there were ~5–30% of mG positive cells observed in untreated groups for both tissues (Figs. 2, 3). Moreover, mG labeling was mainly observed in the cardiomyocytes and in the alveoli and bronchi of the lung. There was also a small degree of overlap in the localization of mT and mG fluorescence, suggesting co-localization of recombined and non-recombined cells. The degree of Cre efficiency in the double transgenic progeny of mT/mG and Cre–ERT2 mice varied among animals. Spontaneous Cre activity was detected in all tissues examined in untreated Cre–ERT2 mT/mG mice (Figs. 2, 3). mG positive cells were detected in the untreated and vehicle treated Cre–ERT2 mT/mG mice indicating that Cre-mediated recombination occurred in the absence of tamoxifen. Furthermore, mG expression in the untreated group excludes unrecognized estrogenic activity of the corn oil/ethanol vehicle as the basis for Cre-mediated recombination in the absence of tamoxifen.
mT/mG labeling of mT/mG and Cre–ERT2 mT/mG mice. Fixed tissue sections of mT/mG mice (n = 6) did not show mG labeling for all tissues examined. Fixed tissue sections of Cre–ERT2 mT/mG mice showed mosaic pattern of mT and mG labeling in the kidney, liver, heart and lung tissue sections. Tamoxifen treated Cre–ERT2 mT/mG mice (n = 6) showed >50% Cre efficiency as represented by the percent of mG labeling in all tissues examined. mG labeling was observed in tissue sections of Cre–ERT2 mT/mG untreated (n = 4) and vehicle (n = 4) treated groups, indicating spontaneous Cre activity in the absence of tamoxifen
Cre activity in untreated 10 ± 1 weeks old Cre–ERT2 mT/mG mice. These are representative images of tissue sections from two different animal samples (I, II) to show the range of the spontaneous Cre activity. Spontaneous Cre activity was observed in kidney, liver, heart, and lung tissue sections. Additionally, the Cre activity appears to be animal dependent
Cre–ERT2 activity in Ece1 flox/flox mice
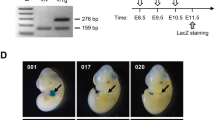
To verify our findings on the Cre activity of Cre–ERT2 mice, we used an Ece1 flox/flox targeted mouse line. We monitored for the presence of the various Ece1 ∆ alleles under various conditions. We compared tail DNA samples of untreated Cre–ERT2 Ece1 +/flox (n = 3) mice with Ece1 +/flox (n = 3) and Cre–ERT2 Ece1 ∆/flox (n = 3) at 3 weeks of age. Using 5 µl PCR reaction samples, we did not detect the Ece1 ∆ allele band (~300 bp) in Cre–ERT2 Ece1 +/flox and Ece1 +/flox DNA (Fig. 4a). However, when 20 µl PCR reaction samples were electrophoresed, the Ece1 ∆ allele band was detected in some of Cre–ERT2 Ece1 +/flox DNA samples (Fig. 4b). We also observed a diminishing Ece1 flox allele band (~1100 bp) and a more prominent Ece1 ∆ allele band in the 17-week old vehicle treated Cre–ERT2 Ece1 ∆/flox mice (n = 2) in both the liver and kidney using 5 µl PCR reaction volumes (Fig. 4c). Collectively, these results indicated spontaneous Cre activity in the absence of tamoxifen and suggest continued Cre activity over time.
Cre activity analysis in Ece1 floxed mice. St standard, Ece1 flox = ~1100 bp, Ece1 + = ~1000 bp, Ece1 Δ = ~300 bp, a using a 5 µl PCR reaction volume, genomic DNA from 3 weeks old tail preparations showed an undetectable Ece1 Δ band in the untreated Cre–ERT2 Ece1 +/flox (n = 3) and Ece1 +/flox (n = 3) samples, while a weaker Ece1 flox band and a more intense Ece1 Δ band were observed in the untreated Cre–ERT2 Ece1 Δ/flox samples (n = 3). b Using a 20 µl PCR reaction volume, genomic DNA from untreated 3 weeks old tail preparation showed Ece1 Δ band detected in Cre–ERT2 Ece1 +/flox mice (n = 3), while none was observed in the Ece1 +/flox samples (n = 3). c Genomic DNA from 17 weeks old mice kidney and liver showed a less intense Ece1 flox band and a more intense Ece1 Δ band for vehicle treated Cre–ERT2 Ece1 Δ/flox
Discussion
Using the mT/mG reporter mice, our data show that the UBC Cre–ERT2 mouse line has substantial tamoxifen-independent Cre activity. The Cre-recombinase activity also varies among individuals, tissues and cell types. There are several possible explanations for these findings. Some organs may have higher expression of the reporter gene, higher expression of Cre-recombinase or a combination of the two. The Cre may have higher efficiency in specific cell types because the levels of circulating tamoxifen or endogenous estrogen are higher in certain tissues. There are also differences in cell turnover rates and blood flow in different organ systems that need to be considered. Regardless of the factors that may contribute to the variability of the Cre activity, our study demonstrates that Cre–ERT2 is active in the absence of tamoxifen. It also further confirms previous finding on tamoxifen independent UBC-driven Cre–ERT2 activity in vivo (Seime et al. 2015). Sanger sequencing of the UBC-driven Cre–ERT2 gene did not show any mutations that could potentially render the Cre–ERT2 perpetually active (data not shown). Many of the Cre–ERT2 lines have been reported to have spontaneous Cre recombination regardless of its specific promoter (Leone et al. 2003; Mishra et al. 2015; Papoutsi et al. 2015; Seime et al. 2015), which demonstrate that it is an inherent property of the fusion protein. The ubiquitin C promoter is widely and highly expressed, which enabled us to detect ligand independent Cre-recombinase activity in many tissues at surprisingly high levels. Our findings differ from those reported previously in showing more generalized and more extensive Cre-recombinase activity. Thus, investigators should consider the impact of spontaneous Cre activity on their studies.
To address the potential difference in Cre-recombination efficiency between the reporter construct and a specific floxed gene, we performed genomic DNA genotyping for the Ece1 ∆ allele using our Ece1 floxed mouse constructs. In the untreated Cre–ERT2 Ece1 +/flox mice, the Ece1 Δ allele was observed at 3 weeks of age, before tamoxifen treatment. Collectively, these findings demonstrate the need for caution of unintended recombination activity in many inducible Cre strains. Although the use of tamoxifen increases the Cre efficiency, the results stress the importance on the use of appropriate controls and the inclusion of both sexes when studying conditional knockout alleles.
Spontaneous Cre activity can be a confounding factor in analyzing gene function and must be considered during experimental design. Without the use of appropriate controls, meaningful results can be overlooked. In many cases, the use of only a vehicle treated control group may not be sufficient to characterize the phenotypic outcomes of homozygous gene deletion. Due to the unintended intrinsic Cre activity, vehicle treated controls may have a comparable degree of gene deletion to tamoxifen treated animals, masking significant genotype-dependent effects. As a result, some changes in phenotypes may be understated. Littermates that do not harbor the Cre transgene are valuable controls that should be included in tamoxifen inducible gene ablation studies.
These experiments have both strengths and limitations. The strengths include the use of both reporter mice and inducible Ece1 knockout mice to evaluate Cre activity in two targeted gene systems. We also examined the Cre-recombinase activity in multiple tissues allowing for comparison of the Cre efficiency in different organ systems. We excluded vehicle as a possible inducer of Cre activity. A study limitation is that we did not perform quantitative genotyping for Ece1 ∆ alleles; thus, we were unable to confirm the effect of sexual dimorphism on the Cre–ERT2 activity. Further experiments are required to quantitatively identify possible sexual dimorphism of spontaneous Cre-recombinase activity. Additionally, we only examined Cre activity over a small range of times. Therefore, we were unable to address the accumulation of recombined cells beyond young adulthood. Recent findings show that cross contamination of exogenous tamoxifen can occur in animals that are housed together through exposure to the general environment and/or coprophagous behavior (Brake et al. 2004). In our study, there were untreated and tamoxifen treated mice that were housed in the same cage. Thus, we were unable to exclude the possibility of tamoxifen cross contamination may contribute to the Cre activity observed in the untreated animals. However, the detection of the Ece1 ∆ allele in 3 weeks old mice, prior to drug treatment, demonstrates that coprophagia is not a sufficient explanation for spontaneous Cre recombinase activity.
Despite the shortcomings of some Cre constructs, the Cre-recombinase system has been proven to be a very useful tool in biological studies. The Cre–ERT2 system has been used to deduce many gene functions over the years. Although new approaches may be required to refine this powerful tool, fully understanding the extent of Cre activity will help researchers better characterize the outcomes of their studies.
References
Brake RL, Simmons PJ, Begley CG (2004) Cross-contamination with tamoxifen induces transgene expression in non-exposed inducible transgenic mice. Genet Mol Res GMR 3:456–462
Feil R, Wagner J, Metzger D, Chambon P (1997) Regulation of Cre recombinase activity by mutated estrogen receptor ligand-binding domains. Biochem Biophys Res Commun 237:752–757
Gossen M, Bujard H (1992) Tight control of gene expression in mammalian cells by tetracycline-responsive promoters. Proc Natl Acad Sci USA 89:5547–5551
Indra AK, Warot X, Brocard J, Bornert JM, Xiao JH, Chambon P, Metzger D (1999) Temporally-controlled site-specific mutagenesis in the basal layer of the epidermis: comparison of the recombinase activity of the tamoxifen-inducible Cre–ER(T) and Cre–ER(T2) recombinases. Nucleic Acids Res 27:4324–4327
Jaisser F (2000) Inducible gene expression and gene modification in transgenic mice. J Am Soc Nephrol JASN 11(Suppl 16):S95–S100
Johansen J, Rosenblad C, Andsberg K, Moller A, Lundberg C, Bjorlund A, Johansen TE (2002) Evaluation of Tet-on system to avoid transgene down-regulation in ex vivo gene transfer to the CNS. Gene Ther 9:1291–1301
Leone DP, Genoud S, Atanasoski S, Grausenburger R, Berger P, Metzger D, Macklin WB, Chambon P, Suter U (2003) Tamoxifen-inducible glia-specific Cre mice for somatic mutagenesis in oligodendrocytes and Schwann cells. Mol Cell Neurosci 22:430–440
Lois C, Hong EJ, Pease S, Brown EJ, Baltimore D (2002) Germline transmission and tissue-specific expression of transgenes delivered by lentiviral vectors. Science 295:868–872
Mishra P, Varuzhanyan G, Pham AH, Chan DC (2015) Mitochondrial dynamics is a distinguishing feature of skeletal muscle fiber types and regulates organellar compartmentalization. Cell Metab 22:1033–1044
Muzumdar MD, Tasic B, Miyamichi K, Li L, Luo L (2007) A global double-fluorescent Cre reporter mouse. Genesis (New York, NY 2000) 45:593–605
Papoutsi T, Odelin G, Moore-Morris T, Puceat M, de la Pompa JL, Robert B, Zaffran S (2015) Msx1CreERT2 knock-In allele: a useful tool to target embryonic and adult cardiac valves. Genesis (New York, NY 2000) 53:337–345
Pluta K, Luce MJ, Bao L, Agha-Mohammadi S, Reiser J (2005) Tight control of transgene expression by lentivirus vectors containing second-generation tetracycline-responsive promoters. J Gene Med 7:803–817
Ruzankina Y, Pinzon-Guzman C, Asare A, Ong T, Pontano L, Cotsarelis G, Zediak VP, Velez M, Bhandoola A, Brown EJ (2007) Deletion of the developmentally essential gene ATR in adult mice leads to age-related phenotypes and stem cell loss. Cell Stem Cell 1:113–126
Seime T, Kolind M, Mikulec K, Summers MA, Cantrill L, Little DG, Schindeler A (2015) Inducible cell labeling and lineage tracking during fracture repair. Dev Growth Differ 57:10–23
Acknowledgements
We would like to thank Baozhi Yuan and Harmut Weiler for helpful discussions, and Aron Geurts and Michael Grzybowski for Sanger Sequencing and analysis. This material is based on work supported in part by award I21 RX1440, by the Office of Research and Development, Rehabilitation Research and Development Service, and a Merit Review Award from the Office of Research and Development, Biomedical and Laboratory Research and Development Service, the Department of Veterans Affairs, and performed in the Geriatrics Research, Education, and Clinical Center at the William S. Middleton Memorial Veterans Hospital. The views expressed here are those of the authors, and the contents do not represent the views of the US Department of Veterans Affairs or the United States Government. Research reported in this publication was supported in part by National Institute of Arthritis and Musculoskeletal and Skin Diseases of the National Institute of Health under Award Number NIH AR54753. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institute of Health.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kristianto, J., Johnson, M.G., Zastrow, R.K. et al. Spontaneous recombinase activity of Cre–ERT2 in vivo. Transgenic Res 26, 411–417 (2017). https://doi.org/10.1007/s11248-017-0018-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11248-017-0018-1