Abstract
Wax esters are hydrophobic lipids consisting of a fatty acid moiety linked to a fatty alcohol with an ester bond. Plant-derived wax esters are today of particular concern for their potential as cost-effective and sustainable sources of lubricants. However, this aspect is hampered by the fact that the level of wax esters in plants generally is too low to allow commercial exploitation. To investigate whether wax ester biosynthesis can be increased in plants using transgenic approaches, we have here exploited a fusion between two bacterial genes together encoding a single wax ester-forming enzyme, and targeted the resulting protein to chloroplasts in stably transformed tobacco (Nicotiana benthamiana) plants. Compared to wild-type controls, transgenic plants showed both in leaves and stems a significant increase in the total level of wax esters, being eight-fold at the whole plant level. The profiles of fatty acid methyl ester and fatty alcohol in wax esters were related, and C16 and C18 molecules constituted predominant forms. Strong transformants displayed certain developmental aberrations, such as stunted growth and chlorotic leaves and stems. These negative effects were associated with an accumulation of fatty alcohols, suggesting that an adequate balance between formation and esterification of fatty alcohols is crucial for a high wax ester production. The results show that wax ester engineering in transgenic plants is feasible, and suggest that higher yields may become achieved in the near future.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Increased energy consumption by a growing population, together with an associated intensive usage of petroleum resources, has resulted in undesirable boost of CO2 in the atmosphere. This has led to a search for CO2-neutral energy alternatives to replace petroleum reserves (Durrett et al. 2008). Plants have a high potential as sources of renewable and CO2-neutral bio-products for industrial use, since they can be produced in large quantities, and can be tailor-made using metabolic engineering approaches (Vanhercke et al. 2013). Plant lipids, particularly oils, fatty alcohols and wax esters, are already extensively used in chemical industry, but have in recent years also emerged as candidates for replacing petroleum-derived products for energy purposes. However, due to reasons of insufficient arable land for producing oilseed plants, a production of valuable storage lipid compounds in vegetative tissues rather than seeds, has been proposed as an efficient way for creating alternative oil crops (Carlsson et al. 2011; Chapman et al. 2013; Vanhercke et al. 2013).
Triacylglycerols have, due to their hydrolytic and oxidative instability, limitations in replacing petroleum products, while wax esters have superior resistance properties in this regard (Li et al. 2010). Wax esters are neutral lipids consisting of a fatty acid moiety linked to a fatty alcohol with an ester bond. Wax esters are found in a wide range of organisms, where they serve a number of biological functions, e.g. energy storage in certain bacteria (Fixter et al. 1986; Ishige et al. 2003; Kalscheuer and Steinbuchel 2003), exoskeleton components in insects (Moto et al. 2003), beeswax components (Aichholz and Lorbeer 2000), as well as membrane components in mammals (Cheng and Russell 2004). In terrestrial plants, wax esters cover the aerial surfaces and prevent non-stomatal water loss, but also function as protection against insects, pathogens and UV radiation (PostBeittenmiller 1996). Interestingly, wax esters are also found in considerable concentrations in the spermaceti organs of the sperm whale, where they are believed to influence buoyancy and eco-location. The exceptional properties of spermaceti wax esters have found uses as additives in a number of lubrication, medical and cosmetic applications (Wahlen et al. 2009). After the worldwide ban of sperm whale hunting, jojoba (Simmondsia chinensis) seeds have been exploited as an alternative source of wax esters, as 60 % of the seed dry weight (DW) composes of wax esters (Metz et al. 2000; Miwa 1971). However, the low yield and high cost of cultivating jojoba plants only make the product economically viable for high-value applications. As an alternative, transgenic approaches seem feasible, and biosynthesis of jojoba wax esters in transgenic Arabidopsis seeds by co-expression of two jojoba wax ester-biosynthetic genes has been reported (Lardizabal et al. 2000). It is of interest to refine this technology to translate it into different oilseed crops, as suggested by Carlsson et al. (2011).
Compared to other complex lipids, the biosynthesis of wax esters is moderately simple and has been well investigated (Lardizabal et al. 2000). Fatty acids for wax ester formation are produced through de novo fatty acid synthesis, where primary C16 and C18 acyl chains are formed. This step is catalyzed in the stroma of plastids, after which a further fatty acid elongation is carried out in the cytoplasm by membrane-associated multi-enzyme complexes (Kunst and Samuels 2003). The formation of wax esters consists of at least two sequential enzymatic steps where an activated fatty acid; acyl:coenzyme A or acyl:acyl-carrying protein (acyl:ACP), is first converted to a fatty aldehyde, and then to a fatty alcohol by the action of a fatty acid reductase (FAR). This fatty alcohol is subsequently esterified with an activated fatty acid by a wax synthase (WS) (Khan and Kolattuk 1973; Kolattukudy and Rogers 1978; Sturaro et al. 2005). In prokaryotes, the fatty alcohol formation is catalysed by one or two separate enzymes, whereas this process in eukaryotes is performed by a single bi-functional enzyme, and without the release of an intermediate aldehyde (Hofvander et al. 2011; Metz et al. 2000; Reiser and Somerville 1997).
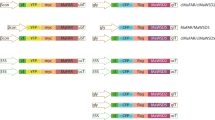
We have recently shown in tobacco (Nicotiana benthamiana), that transient expression of different combinations of Marinobacter genes with a role in the production of wax esters, resulted in accumulation of substantial amounts of wax esters in leaf chloroplasts (Aslan et al. 2014). The activity of the chloroplast-targeted fusion gene tpMaFAR::MhWS was shown to be as effective as a combination of tpMaFAR and tpMhWS when infiltrated separately. Here, we show that an elevated production of wax esters is feasible also in stably transformed N. benthamiana plants. The tpMaFAR::MhWS construct was expressed from the Cauliflower Mosaic Virus 35S promoter (CaMV 35S), and significant amounts of wax esters were produced in both leaves and stems, supporting further development of tobacco as a candidate crop for the production of renewable wax esters in vegetative tissues.
Materials and methods
Plant materials
Nicotiana benthamiana plants were grown in a phytotron unit with a day temperature of 25 °C, and with a relative humidity of 60 %. The photoperiod was 16 h, with light from fluorescent lamps giving a photosynthetic fluence density (PFD) of 320 μmol photons m−2 s−1. Plants were grown in 21 cm × 16 cm dark pots (5 L), and received regular water and fertilization with a commercial fertilizer using common practice.
Sterile plants used for leaf-disc transformations were maintained in growth rooms, with a day temperature of 22 °C and 17 °C at night. The photoperiod was 16 h, with light from fluorescent lamps giving a PFD of 80 µmol photons m−2 s−1. The plants were grown in Murashige–Skoog (MS) medium (Duchefa, RV, Haarlem, The Netherlands) supplemented with 3 % sucrose, and solidified with 0.3 % Gel-rite (Sigma AB, Malmö, Sweden). The tpMaFAR::MhWS gene fusion was made synthetically (GeneWiz Europe, London, UK), and the DNA sequence of the final construct is given in Fig. S1. The tpMaFAR vector used for transformations has been described in Aslan et al. (2014). Genes were cloned in Ti-plasmid pART27, enabling selection for kanamycin resistance in transgenic plants. Transgenic plants were obtained by Agrobacterium-mediated leaf-disc transformation, essentially as described (Horsch et al. 1986).
Regenerated calli were cultivated on shoot-inducing medium, and transferred to fresh medium every 2 weeks for a period of 3 months. Shoots were transferred to small glass jars and further grown until a stem was visible, and then transferred to half-strength MS medium supplemented with 4 µM of the auxin indole-3-butyric acid (Sigma AB, Malmö, Sweden) to promote root formation. Rooted shoots were transferred into soil, and covered with plastic bags until shoot growth became visible. Seeds were collected from self-fertilized primary transformants, and selected for kanamycin (Sigma AB, Malmö, Sweden) resistance on half-strength MS medium supplemented with 125 mg L−1 kanamycin. Homozygous transgenic T3 lines were obtained from self-fertilized T2 transformants having a kanamycin resistance frequency of 3:1.
Chemical analyses
Wax ester and fatty alcohol analyses were performed as described (Aslan et al. 2014).
Chlorophyll levels were measured using a SPAD-502 chlorophyll meter according to the manufacturer (Spectrum Technologies, Thayer Court, Aurora, US).
RNA isolation and transgene expression analysis
Total RNA was extracted from 100 mg leaf materials using the Spectrum™ Plant Total RNA Kit (Sigma-Aldrich, St. Louis, MO, US), and following the protocol of the supplier. All RNA samples were treated with DNase I (Sigma-Aldrich, St. Louis, MO, US) to remove any residual DNA. First-strand cDNA was synthesized using a qScript cDNA Synthesis kit following the manufacturer instructions (Quanta Biosciences, Gaithersburg, MD, US). Quantitative reverse-transcription-polymerase chain reactions (QPCR) were made using a SYBR Green PCR master mix (Applied Biosystems, Life Technologies Europe BV, Stockholm, Sweden) supplemented with cDNA template and 5 μM primers. The primers used for QPCR were: 5′-GGC CTG TGT TCA ACG TGA C-3′ (tpMaFAR::MhWS forward), 5′-CCA ACC TAG CTC CCT CGT AGT-3′ (tpMaFAR::MhWS reverse), 5′- CCT GAC GGA TCG TGT TCT G-3′ (tpMaFAR forward), 5′-GAC TGC GTG GTA TCC AGG TT-3′ (tpMaFAR reverse). PCR was performed in 96-well optical reaction plates (Applied Biosystems, Life Technologies Europe BV, Stockholm, Sweden) as follows: 10 min at 50 °C, 5 min at 95 °C, and 40 cycles of 10 s at 95 °C and 30 s at 60 °C, and finally 1 min at 95 °C. The specificity of the reactions, and the amplicon identities were verified by melting curve analysis. Reaction mixtures without cDNA were used as a negative control. Data were evaluated using the CT method (Livak and Schmittgen 2001), and with correction for the PCR efficiency as determined from the slope of standard curves. Normalization of gene expression levels were made using the ACTIN gene (Liu et al. 2012), and fold-differences in transcript levels, and mean standard error, were calculated as described (Livak and Schmittgen 2001).
Results
Generation of transgenic tobacco plants expressing a 35S:tpMaFAR::MhWS construct
To increase the wax ester biosynthesis in transgenic plants, the Marinobacter genes MaFAR and MhWS were fused to a nucleotide sequence encoding a chloroplast transit peptide, and expressed from the CaMV 35S promoter (Fig. S1). The final 35S:tpMaFAR::MhWS construct was used to transform tobacco leaves, and fourteen primary transformants were generated and transferred to greenhouse conditions. Twelve transformants yielded seeds, while two were unfertile and hence lost. Based on a strong transgene expression in leaves from the second generation of transformants (T2) (Fig. S2a), and detection of the gene construct in nuclear DNA (not shown), the lines #2.10 and #6.1 were chosen for further analyses, and homozygotic lines were obtained from the third generation of transformants (T3). Compared to the lines #2.10 and #6.1, transgene expression in other lines was clearly lower (Fig. S2a).
Transgene expression in the two chosen lines was further assessed by QPCR analysis in T3 leaves collected at three positions (Fig. S3). This showed a gene expression in leaves from all positions. There was in both lines a basipetal increase in transgene expression, and the expression level was slightly higher in line #6.1 than in line #2.10. No significant expression was detected in untransformed wild-type controls.
Wax ester levels in 35S:tpMaFAR::MhWS transformants
For both transgenic lines, an increased level of wax esters was obvious in leaves and stems (Fig. 1). Well in line with the QPCR analysis, the wax ester level in leaves increased basipetally. The highest wax ester level in leaves was 0.28 µmol g−1 fresh weight (FW) in line #2.10 (Fig. 1). Wax ester levels in stems were comparable to those in leaves, but generally highest at the middle position (Fig. 1). As calculated from the total FW of leaves and stems of representative plants, these values corresponded in both transgenic lines to a total wax ester level of ca. 8 mg per plant (85 g FW), or a ca. eight-fold increase compared to the endogenous level in wild-type plants. On a dry weight (DW) basis, the wax ester level in transformants was ca. 0.15 % (Table S1). Wax ester levels in other lines was lower, well in line with a lower degree of transgene expression in these lines (Fig. S2b).
Wax ester levels in wild-type (Wt) tobacco plants (white), and in the 35S:tpMaFAR::MhWS transgenic lines #2.10 (grey), and #6.1 (black). Total wax esters were extracted from leaves (a) and stems (b) at the top, middle, and basal positions, and quantified by gas chromatography. Leaf and stem materials were collected from plants at the age of 8-weeks. Mean value ± range of two separate extractions of one sample made from six pooled plants per genotype
The wax esters in both leaves and stems were composed mainly of the fatty alcohols 16:0-OH (palmitoyl alcohol) and 18:0-OH (stearoyl alcohol) (Fig. 2a, b). The corresponding composition of the fatty acid moieties was broader, and prominent compounds were 16:0 (palmitic acid), 18:0 (steraric acid), 18:2 (linoleic acid), 20:0 (arachidic acid), and 22:0 (behenic acid) (Fig. 2c, d). Corresponding absolute levels are shown in Fig S4.
Chemical composition of wax esters in leaves and stems collected at three positions (top, middle, basal) in transgenic 35S:tpMaFAR::MhWS tobacco plants. Composition of the wax ester fatty alcohol moieties in leaves (a), and in stems (b), and of the fatty acid moieties in leaves (c), and stems (d). Mean value ± range from two separate extractions of one sample made from six pooled plants per genotype
Phenotype of transgenic plants
The growth and development of the transgenic lines was somewhat different from that of wild-type controls (Fig. 3). Lines #2.10 and #6.1 displayed a stunted growth, with chlorotic leaves and stems (Fig. S5). A reduced level of chlorophyll in chlorotic leaves was verified (Fig. S6).
General appearance of wild-type tobacco (Nicotiana benthaminiana) plants, and of representative 35S:tpMaFAR::MhWS transformants. (a) Wild-type plant, and transgenic lines #1.2 (b), #2.10 (c), #6.1 (d), #9.4 (e), #10.4 (f), #11.1 (g), #12.2 (h). Bars are equal to 50 cm. Plants were depicted at nine weeks from sowing
The chlorotic leaf phenotype indicated that the biosynthesis of wax esters became associated with the production of a substance with harmful effects. This substance might be one or several wax esters, or alternatively, an intermediate in the wax ester biosynthesis, such as free fatty alcohols, i.e. fatty alcohols not incorporated into wax esters. Analysis of the latter showed that the level of free fatty alcohols was significant in both leaves and stems, and comparable to the amount of fatty alcohols bound in wax esters. The highest free alcohol level was measured in leaves at the middle position, being ca. 0.4 µmol g−1 FW in both lines (Fig. 4a). High levels of free alcohols were also measured in stem tissues, where they showed an acropetal increase (Fig. 4b), indicating an inverse relationship between the level of free fatty alcohols and wax esters. Moreover, the chemical composition of free alcohols was similar to that bound in wax esters, and consisted almost exclusively of stearoyl and palmitoyl alcohols (Fig. 4c, d).
Level and composition of free fatty alcohols in wild-type (Wt) tobacco plants, and in 35S:tpMaFAR::MhWS transgenic lines. Total level of free fatty alcohols in leaves (a), and in stems (b) of wild-type plants (white), and of transgenic lines #2.10 (grey), and #6.1 (black). Chemical composition of free alcohols in leaves (c), and stems (d). Total free alcohols and their composition was measured at the plant top, middle, and basal positions. Mean value ± range of two separate extractions of one sample made from six pooled plants per genotype
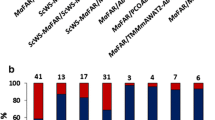
To further investigate the potentially negative effects of free fatty alcohols, the single tpMaFAR gene was expressed from the CaMV 35S promoter in transgenic tobacco plants, thus generating ten 35S:tpMaFAR primary transformants. At the flowering stage, the transformants displayed a reduced fertility and pollen production, although all transformants could be rescued by hand-pollination. However, almost all (80 %) of the 35S:tpMaFAR lines showed a lethal phenotype in the T2 generation, with only a few plants surviving per line (Fig. 5a). The lethal phenotype was obvious both on selective as well as on non-selective media, and during normal as well as reduced light conditions. None of the wild-type plants or control transformants showed this phenotype in parallel analyses. QPCR analysis of surviving 35S:tpMaFAR plants on non-selective media demonstrated that the plants expressed the transgene to a considerable extent (Fig. 5b). However, a corresponding analysis of the dead seedlings was not feasible, likely due to a poor RNA quality when isolated from the dead materials. On the other hand, free fatty alcohols could be extracted from both phenotypes, and was shown to be more than six-fold higher in dead seedlings compared to viable ones (Fig. 5c). Similar to 35S:tpMaFAR::MhWS lines, the surviving 35S:tpMaFAR seedlings displayed in the T2 generation an aberrant phenotype during later stages of development, including necrotic leaves, reduced pollen function, and reduced fertility (Fig. 5d, e). Taken together, the results strongly suggest that the negative effects on growth observed in 35S:tpMaFAR::MhWS plants, at least in part were due to an accumulation of free fatty alcohols.
Lethal phenotype of 35S:tpMaFAR transformants. Seedlings of wild-type (Wt) tobacco (a) and a representative 35S:tpMaFAR line (b) grown on MS medium without kanamycin selection. (c) Relative tpMaFAR transgene expression in the 35S:tpMaFAR line. (d) Total level of free fatty alcohols in wild-type plants, and in viable or dead 35S:tpMaFAR transformants. Mean value ± range of duplicate extractions. (e) Aberrant phenotype of 35S:tpMaFAR transformants, and a close-up (f)
Discussion
The use of plants as bioreactors for production of valuable compounds such as lipids may provide an alternative to conventional oil resources. Towards this end, we have recently evaluated genes important to wax ester biosynthesis using transient expression in N. benthamiana leaves (Aslan et al. 2014). This identified the 35S:tpMaFAR::MhWS gene fusion as one of the constructs capable of increasing wax esters, and led us to here evaluate its potential for wax ester production in vegetative parts of stably transformed tobacco plants.
Results showed a clearly increased wax ester content in 35S:tpMaFAR::MhWS transformants compared to the wild type (Figs. 1 and S2), being about eight-fold higher as measured at the whole plant level. Within the plant, wax ester levels increased in both leaves and stems, and the pattern was well correlated with the transgene expression (Figs. 1, S2, and S3), indicating sufficient amounts of biosynthetic substrates in all parts of the tobacco plant. The positive correlation between transgene expression and wax ester levels suggests that increased transgene expression, for instance by using stronger promoters or minimizing transgene post-transcriptional silencing, might be one way of increasing wax ester production even further. However, the total level of wax esters in 35S:tpMaFAR::MhWS young leaves, 0.08 % (DW) (Table S1), was clearly lower than the 0.2 % (DW) reported in equivalent leaf materials transiently expressing the same gene construct (Aslan et al. 2014). Moreover, wax ester levels were in all parts of the stably transformed plants also lower than the between 2 and 4 % reported for transgenic Arabidopsis seeds (Heilmann et al. 2012). Considering that 35S:tpMaFAR::MhWS transformants displayed certain negative effects on growth and development (Fig. 3), we hypothesize that the stably transformed plants already at the present degree of transgene expression from the CaMV 35S promoter might have been counter-selected for a strong transgene expression during the regeneration process, resulting in a sub-optimal production in the final transformants. The results suggest high levels of free fatty alcohols to be a factor hampering growth and a high wax ester production (Fig. 5). Transient co-expression of separate 35S:tpMaFAR and 35S:tpMhWS genes led to a doubling of wax ester levels compared to the 35S:tpMaFAR::MhWS gene fusion, but also to a doubling of fatty alcohols (Aslan et al. 2014). This indicates that the high levels of fatty alcohols observed in our transformants at least in part are due to catalytic limitations of the WS enzyme as such, rather than artifacts due to the gene fusion. Hence, rather than maximizing the overall transgene expression, future strategies might instead focus on minimizing the level of un-incorporated fatty alcohols. This might be done by an increased expression or activity of the wax synthase step, for instance by using a broader substrate range wax ester synthase enzyme with higher specificity for the predominant 16:0-OH and 18:0-OH alcohols. Sequential transformation of WS transformants with FAR constructs, or WS × FAR transformant crosses, might be alternative strategies. Moreover, further investigations should also aim at optimizing the balance between target organ and plant biomass, organ-specific product yield, and total plant production per cultivated surface area.
The major components of wax esters in transgenic lines, 16:0-OH (palmitoyl alcohol) and 18:0-OH (stearoyl alcohol) (Fig. 2) were well in line with those reported earlier (Aslan et al. 2014) and with analyses of MaFAR substrates in vitro (Hofvander et al. 2011). In addition to this, MhWS was thought to use substrates shorter than 16 carbons (Barney et al. 2012), although this did not seem to be the case in the current study, where a broad wax ester profile was detected (Fig. 2). The fatty acid content of the wax esters was composed mainly of 16:0 and 18:0 fatty acids, which indicates that the MhWS enzyme preferentially accepts C16 and C18 acyl-ACP substrates in vivo. In addition, wax esters in the current study contained also the very long chain fatty acids 20:0 and 22:0 at considerable levels, both in stem and leaf tissues (Fig. 2c, d). It is known that very long chain fatty acid biosynthesis occurs at the endoplasmic reticulum in plant cells (Samuels et al. 2008). This suggests that a part of the wax ester biosynthesis in the current transgenic lines might have occurred outside the plastids, where wax esters accumulated in our previous study (Aslan et al. 2014). The mechanism behind this result should be further investigated for optimization of the tobacco transformation in the future.
Taken together, we have here shown that wax esters can be produced at high levels in vegetative tissues of stably transformed tobacco plants, reaching approximately 0.15 % (DW) per plant. It will now be of interest to optimize the production, e.g. by sequential transformations, transformant crosses, inducible expression, or optimizing substrate preferences, for obtaining wax ester levels of commercial interest.
References
Aichholz R, Lorbeer E (2000) Investigation of combwax of honeybees with high-temperature gas chromatography and high-temperature gas chromatography-chemical ionization mass spectrometry II: high-temperature gas chromatography-chemical ionization mass spectrometry. J Chromatogr A 883:75–88. doi:10.1016/s0021-9673(00)00386-1
Aslan S et al (2014) Wax esters of different compositions produced via engineering of leaf chloroplast metabolism in Nicotiana benthamiana. Metab Eng 25C:103–112. doi:10.1016/j.ymben.2014.07.001
Barney BM, Wahlen BD, Garner E, Wei J, Seefeldt LC (2012) Differences in substrate specificities of five bacterial wax ester synthases. Appl Environ Microbiol 78:5734–5745. doi:10.1128/AEM.00534-12
Carlsson AS, Yilmaz JL, Green AG, Stymne S, Hofvander P (2011) Replacing fossil oil with fresh oil—with what and for what? Eur J Lipid Sci Technol 113:812–831. doi:10.1002/ejlt.201100032
Chapman KD, Dyer JM, Mullen RT (2013) Commentary: why don’t plant leaves get fat? Plant Sci Int J Exp Plant Biol 207:128–134. doi:10.1016/j.plantsci.2013.03.003
Cheng JB, Russell DW (2004) Mammalian wax biosynthesis. I. Identification of two fatty acyl-Coenzyme A reductases with different substrate specificities and tissue distributions. J Biol Chem 279:37789–37797. doi:10.1074/jbc.M406225200
Durrett TP, Benning C, Ohlrogge J (2008) Plant triacylglycerols as feedstocks for the production of biofuels. Plant J Cell Mol Biol 54:593–607. doi:10.1111/j.1365-313X.2008.03442.x
Fixter LM, Nagi MN, McCormack JG, Fewson CA (1986) Structure, distribution and function of wax esters in acinetobacter-calcoaceticus. J Gen Microbiol 132:3147–3157
Heilmann M, Iven T, Ahmann K, Hornung E, Stymne S, Feussner I (2012) Production of wax esters in plant seed oils by oleosomal cotargeting of biosynthetic enzymes. J Lipid Res 53:2153–2161. doi:10.1194/jlr.M029512
Hofvander P, Doan TT, Hamberg M (2011) A prokaryotic acyl-CoA reductase performing reduction of fatty acyl-CoA to fatty alcohol. FEBS Lett 585:3538–3543. doi:10.1016/j.febslet.2011.10.016
Horsch RB, Klee HJ, Stachel S, Winans SC, Nester EW, Rogers SG, Fraley RT (1986) Analysis of agrobacterium-tumefaciens virulence mutants in leaf-disks. Proc Natl Acad Sci USA 83:2571–2575. doi:10.1073/pnas.83.8.2571
Ishige T, Tani A, Sakai Y, Kato N (2003) Wax ester production by bacteria. Curr Opin Microbiol 6:244–250. doi:10.1016/s1369-5274(03)00053-5
Kalscheuer R, Steinbuchel A (2003) A novel bifunctional wax ester synthase/acyl-CoA:diacylglycerol acyltransferase mediates wax ester and triacylglycerol biosynthesis in Acinetobacter calcoaceticus ADP1. J Biol Chem 278:8075–8082. doi:10.1074/jbc.M210533200
Khan AA, Kolattuk P (1973) Control of synthesis and distribution of acyl moieties in etiolated euglena-gracilis. Biochemistry 12:1939–1948. doi:10.1021/bi00734a017
Kolattukudy PE, Rogers L (1978) Biosynthesis of fatty alcohols, alkane-1,2-diols and wax esters in particulate preparations from uropygial glands of white-crowned sparrows (zonotrichia-leucophrys). Arch Biochem Biophys 191:244–258. doi:10.1016/0003-9861(78)90087-5
Kunst L, Samuels AL (2003) Biosynthesis and secretion of plant cuticular wax. Prog Lipid Res 42:51–80. doi:10.1016/s0163-7827(02)00045-0
Lardizabal KD, Metz JG, Sakamoto T, Hutton WC, Pollard MR, Lassner MW (2000) Purification of a jojoba embryo wax synthase, cloning of its cDNA, and production of high levels of wax in seeds of transgenic Arabidopsis. Plant Physiol 122:645–655. doi:10.1104/pp.122.3.645
Li W et al (2010) Green waxes, adhesives and lubricants. Philos Trans Ser A Math Phys Eng Sci 368:4869–4890. doi:10.1098/rsta.2010.0197
Liu D, Shi L, Han C, Yu J, Li D, Zhang Y (2012) Validation of reference genes for gene expression studies in virus-infected Nicotiana benthamiana using quantitative real-time PCR. PLoS ONE 7:e46451. doi:10.1371/journal.pone.0046451
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25:402–408. doi:10.1006/meth.2001.1262
Metz JG, Pollard MR, Anderson L, Hayes TR, Lassner MW (2000) Purification of a jojoba embryo fatty acyl-coenzyme A reductase and expression of its cDNA in high erucic acid rapeseed. Plant Physiol 122:635–644. doi:10.1104/pp.122.3.635
Miwa TK (1971) Jojoba oil wax esters and derived fatty acids and alcohols: gas chromatographic analyses. J Am Oil Chem Soc 48:259–264. doi:10.1007/bf02638458
Moto K et al (2003) Pheromone gland-specific fatty-acyl reductase of the silkmoth, Bombyx mori. Proc Natl Acad Sci USA 100:9156–9161. doi:10.1073/pnas.1531993100
PostBeittenmiller D (1996) Biochemistry and molecular biology of wax production in plants. Ann Rev Plant Physiol Plant Mol Biol 47:405–430. doi:10.1146/annurev.arplant.47.1.405
Reiser S, Somerville C (1997) Isolation of mutants of Acinetobacter calcoaceticus deficient in wax ester synthesis and complementation of one mutation with a gene encoding a fatty acyl coenzyme a reductase. J Bacteriol 179:2969–2975
Samuels L, Kunst L, Jetter R (2008) Sealing plant surfaces: cuticular wax formation by epidermal cells. Annu Rev Plant Biol 59:683–707. doi:10.1146/annurev.arplant.59.103006.093219
Sturaro M, Hartings H, Schmelzer E, Velasco R, Salamini F, Motto M (2005) Cloning and characterization of GLOSSY1, a maize gene involved in cuticle membrane and wax production. Plant Physiol 138:478–489. doi:10.1104/pp.104.058164
Vanhercke T, Wood CC, Stymne S, Singh SP, Green AG (2013) Metabolic engineering of plant oils and waxes for use as industrial feedstocks. Plant Biotechnol J 11:197–210. doi:10.1111/pbi.12023
Wahlen BD, Oswald WS, Seefeldt LC, Barney BM (2009) Purification, characterization, and potential bacterial wax production role of an NADPH-dependent fatty aldehyde reductase from Marinobacter aquaeolei VT8. Appl Environ Microbiol 75:2758–2764. doi:10.1128/AEM.02578-08
Acknowledgments
We thank Prof. Sten Stymne (Dept. of Plant Breeding, SLU, Alnarp) for support and advice, and Drs. Sarosh Bejai (Dept. of Plant Biology, SLU, Uppsala) and Frédéric Domergue (Laboratoire de Biogenèse Membranaire, CNRS, Univ. Bordeaux Ségalen, Bordeaux, France) for advice concerning QPCR data analysis and DNA constructions, respectively. The work was supported by EU FP7 project “ICON”, the Swedish Research Council Formas, and the Swedish Governmental Agency for Innovation Systems, VINNOVA.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare they have no competing interests.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Aslan, S., Hofvander, P., Dutta, P. et al. Increased production of wax esters in transgenic tobacco plants by expression of a fatty acid reductase:wax synthase gene fusion. Transgenic Res 24, 945–953 (2015). https://doi.org/10.1007/s11248-015-9893-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11248-015-9893-5