Abstract
Results of transcriptome analyses suggest that expansin genes play an active role in seed development and yield, but gain- or loss-of-function studies have not yet elucidated the functional role(s) of the expansin gene(s) in these processes. We have overexpressed a sweetpotato expansin gene (IbEXP1) in Arabidopsis under the control of cauliflower mosaic 35S promoter in an attempt to determine the effect of the expansin gene in seed development and yield in heterologous plants. The growth rate was enhanced in IbEXP1-overexpressing (ox) plants relative to wild-type Col-0 plants during early vegetative growth stage. At the reproductive stage, the number of rosette leaves was higher in IbEXP1-ox plants than that in Col-0 plants, and siliques were thicker. IbEXP1-ox plants produced larger seeds, accumulated more protein and starch in each seed, and produced more inflorescence stems and siliques than Col-0 plants, leading to a 2.1–2.5-fold increase in total seed yield per plant. The transcript level of IbEXP1 was up-regulated in response to brassinosteroid (BR) treatment in sweetpotato, and the transcript levels of three BR-responsive genes, fatty acid elongase 3-ketoacyl-CoA synthase 1, HAIKU1 and MINISEED3, were also increased in IbEXP1-ox Arabidopsis plants, suggesting a possible involvement of IbEXP1 in at least one of the BR signaling pathways. Based on these results, we suggest that overexpression of IbEXP1 gene in heterologous plants is effective in increasing seed size and number and, consequently, seed yield.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Seed/crop yield is an important plant trait and the major factor determining the economic value of commercial cultivars of grain crops. This trait mainly refers to two phenotypic characteristics—seed size and seed number. Many efforts have been made to increase seed size and/or number by mutant screening and through the generation of transgenic plants. In maize, Giroux et al. (1996) reported that in vivo site-specific mutagenesis of the large subunit of the starch synthetic enzyme ADP-glucose pyrophosphorylase resulted in an 11–18 % increase in seed weight. A number of studies have shown that seed size is altered when the transcript level of the seed development-related genes is modulated. Downregulation of the expression of the Arabidopsis endosperm development-related genes (MINISEED3, HAIKU1, and HAIKU2) was found to give rise to smaller seeds relative to the wild type (Garcia et al. 2003; Luo et al. 2005; Wang et al. 2010), whereas down regulation of Arabidopsis APETALA2 (AP2) caused an increase in seed mass (Ohto et al. 2005). Increased seed size has also been observed in the cytokinin oxidase-overexpressing (ox) Arabidopsis (Werner et al. 2003), GRAIN INCOMPLETE FILLING 1 (GIF1)-ox rice under the control of its own promoter (Wang et al. 2008), and OsSPL16-ox rice (Wang et al. 2012). However, increases in seed size do not always result in increased seed yield; to the contrary, increases in seed size often result in reduced seed yield through a decrease in seed number (Van Daele et al. 2012). On the other hand, seed yield has generally been found to increase in transgenic plants producing an increased number of seeds. For example, reduced expression of the rice cytokinin oxidase (OsCKX2) or GASA4 gene (plant-specific gibberellic acid-stimulated Arabidopsis gene family) and overexpression of the brassinosteroid (BR) biosynthetic gene, DWARF4, or AtHSD1 (11-β-hydroxysteroid dehydrogenase) were observed to increase seed number, resulting in enhanced seed yield (Choe et al. 2001; Ashikari et al. 2005; Li et al. 2007; Roxrud et al. 2007).
Expansins have been characterized as proteins that mediate cell-wall loosening (McQueen-Mason et al. 1992; McQueen-Mason and Cosgrove 1995; Li et al. 2003), and they are believed to be important regulators of wall extension during plant cell growth (Lee et al. 2001; Cosgrove et al. 2002; Li et al. 2003). To date, most studies have focused on the isolation and characterization of expansin genes, and only limited information is available on their precise biological functions in plant growth and development. Gain- and/or loss-of-function approaches have demonstrated that expansins are involved in the ripening process of tomato fruits (Brummell et al. 1999), leaf growth and pedicel abscission in Arabidopsis (Cho and Cosgrove 2000), seedling growth in rice (Choi et al. 2003), petal limb growth and the timing of axillary meristem development in Petunia hybrida (Zenoni et al. 2004, 2011), height and diameter growth in Chinese fir (Cunninghamia lanceolata) (Wang et al. 2011), and root cell division and elongation in Arabidopsis (Guo et al. 2011; Lin et al. 2011). Alternatively, four knock-out mutants of expansin genes generated by homologous recombination showed no detectable phenotype in Physcomitrella patens (Schipper et al. 2002). The authors of a number of very recent studies report conflicting roles for expansin genes under abiotic stress conditions, such as drought and salt stress. Overexpression of a wheat (Triticum aestivum L.) expansin gene, TaEXPB23, in tobacco plants conferred tolerance to salt stress (Han et al. 2012), but ectopic expression of AtEXP3 or AtEXP-β1 enhanced salt sensitivity in transgenic Arabidopsis plants (Kwon et al. 2008). The biological function(s) of expansin genes in seed production has not yet been elucidated. However, Fu et al. (2010) were able to identify genes associated with grain yield and grain dry matter content based on the results of their maize seedling transcriptome analysis, leading these authors to suggest that the α- and β-expansin genes are involved in grain yield. In another study, Lizana et al. (2010) analyzed the temporal and spatial expression profiles of expansin genes during seed development, obtaining results that also led them to suggest that expansin expression is associated with grain size dynamics in wheat.
In an earlier study carried out by our group, we identified cDNAs of three expansin genes (IbEXP1, IbEXP2, and IbEXPL1) from sweetpotato (Ipomoea batatas cv. Jinhongmi) (You et al. 2003) and investigated the transcriptional regulation of these three genes in response to chilling temperature (12–28 °C; Noh et al. 2009). We also investigated the role of IbEXP1 in the formation of the storage root in sweetpotato and found that downregulation of the IbEXP1 gene enhanced storage root development (Noh et al. 2013). In the study reported here, we overexpressed a sweetpotato expansin gene (IbEXP1) in Arabidopsis and observed that leaf growth and axillary meristem formation were enhanced in transgenic IbEXP1-ox Arabidopsis plants, resulting in an increase in seed production.
Materials and methods
Generation of IbEXP1-ox Arabidopsis
The full-size IbEXP1 cDNA was amplified with IbEXP1-specific primers (forward: 5′-gtaggatccCATTCCTCTACCAATTCAACTGAA-3′; reverse: 5′-gatggtaccACTGTCTCCACACTCAGCATT-3′). BamHI and KpnI restriction sites were introduced at the ends of the forward and reverse primers, respectively, in order to facilitate subcloning. The PCR products were then subcloned into the pGEM-T Easy vector (Promega, Madison, WI), digested with the BamHI and KpnI restriction enzymes, and fused to the CaMV 35S promoter in a sense orientation by insertion of the IbEXP1 fragments at the BamHI and KpnI sites of the pMBP1 binary vector. The resulting construct was used to transform Arabidopsis thaliana ecotype Columbia by the floral dip method (Clough and Bent 1998). Transgenic plants were selected on kanamycin-containing agar plates. The transgenic Arabidopsis plants were grown in a culture room at 23–25 °C under a 16/8-h light/dark photoperiod cycle. Wild-type Col-0 and T3 transgenic IbEXP1-ox plants were grown in a relatively larger pot (pot volume for each plant: 330 cm3) or a smaller sized pot (pot volume for each plant: 152 cm3).
Quantitative real-time PCR
Total RNAs were isolated with PureLink® Plant RNA Reagent (Ambion, Austin, TX) according to the manufacturer’s instructions. A 5-µg sample of total RNA was used for the first-strand cDNA synthesis using the Transcriptor First Strand cDNA Synthesis kit (Roche Diagnostics, Indianapolis, IN) according to the manufacturer’s instructions. The primers for real-time PCR were as follows: IbEXP1-specific primers (5′-GGAGTTGTTCCCGTTGCTTA-3′; 5′-GCCTCCCACGTTGGTTACTA-3′), KCS1-specific primers (5′-GCGTTACGACCGGTTTCGAC-3′; 5′-CCAAACACACATCCATAAATTAATTAAATT-3′), δ-TIP-specific primers (5′-CGGTGGTGGACTTGCCG-3′; 5′-CAGCCATCCATTAAATAATATTCATACAAAT-3′), SHB1-specific primers (5′-GCTGCAATGGTCCTGAATGT-3′; 5′-CCATAAGCGCGACCATAGTT-3′), IKU1-specific primers (5′-ATGTTCCCGTTATCTCCATCC-3′; 5′-TTCTGCTGCGTCTATTCCACT-3′), and MINI3-specific primers (5′-CCCAGATTATCCCTCCGTC-3′; 5′-TCATCATCGCTGCATTGTCT-3′). The quantitative real-time PCR analysis was performed using the LightCycler® 480 Real-Time PCR System (Roche Diagnostics) as described in Noh et al. (2010). For Arabidopsis plants and sweetpotato plants, expression levels were normalized to expression of the elongation factor 1α (EF1α) amplified with the EF1α-specific primers (5′-GCACTGTCATTGATGCTCC-3′; 5′-GTCAAGAGCCTCAAGGAGAG-3′), and to expression of the β-tubulin gene amplified with the β-tubulin-specific primers (5′-CAACTACCAGCCACCAACTGT-3′; 5′-CAGATCCTCACGAGCTTCAC-3′), respectively. The real-time PCR reaction mixture was prepared with 1 × KAPA SYBR® FAST Master mix (Kapa Biosystems, Woburn, MA) in a final reaction volume of 16 μl. The PCR analysis was performed by subjecting the samples to an initial denaturation at 95 °C for 10 min, followed by 45 cycles of 95 °C for 10 s, 60 °C for 20 s, and 72 °C for 30 s.
Protein analysis
One hundred mature dry seeds from Col-0 or IbEXP1-ox transgenic plants were homogenized with 200 μl of extraction buffer [250 mM sucrose, 50 mM Tris HCl, pH 8.0, 2 mM DTT, 2 mM EDTA, SIGMAFAST Protease Inhibitor Cocktail (1 tablet/100 ml)] using a pellet pestle (Sigma, St. Louis, MO). The extracts were centrifuged at 12,000 rpm for 10 min at 4 °C and the supernatants transferred into new tubes. After centrifugation of the homogenate, 2 μl of each extract was loaded on the 12 % sodium dodecyl sulfate–polyacrylamide (SDS/PAGE) gel and electrophoresed; the protein products were stained with Coomassie Brilliant Blue. Protein content was determined using a protein assay kit (Bio-Rad, Hercules, CA).
Determination of starch content
Mature dry seeds (1 g) from Col-0 or IbEXP1-ox transgenic plants were macerated in liquid nitrogen with a pestle and mortar. A sample of the macerated powder was transferred into a flask containing 25 ml distilled water and boiled for 3 min with stirring. The sample was then autoclaved at 135 °C for 1 h for starch digestion, cooled to 60 °C, and distilled water (100 ml) was added. Starch concentrations were determined using a starch assay kit (Sigma) according to the manufacturer’s instructions. Starch content per seed was calculated on the basis of the 1,000-seed weight given in Table 1.
Brassinosteroid treatment
Sweetpotato plantlets bearing a single leaf and petiole (single-leaf plantlets) were collected and incubated in flasks containing distilled water for 3 weeks. After fibrous roots had developed from the distal end of the petiole, the single-leaf plantlets were incubated in various concentrations (0, 100, 200, and 500 μM) of 24-epi-Brassinolide (Sigma) at 25 °C in the dark for 3 h. Following the hormone treatment, total RNA was extracted from the fibrous roots using the RNeasy Plant Mini kit (Qiagen, Hilden, Germany) and used for real time-PCR.
Results
Vegetative growth was enhanced in IbEXP1-ox plants
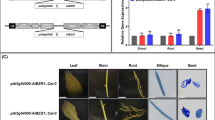
A sweetpotato IbEXP1 cDNA was overexpressed in Arabidopsis with the aim of addressing the efficacy of IbEXP1 expression in vegetative growth and seed production. Transgenic IbEXP1-ox Arabidopsis plants were generated under the control of the CaMV 35S promoter, and 25 independent transgenic lines were obtained, of which three (#1, #4, and #22) were randomly selected for further analysis. The qRT-PCR analysis with RNAs extracted from rosette leaf tissue revealed that IbEXP1 was overexpressed in all three IbEXP1-ox lines but at different levels: the transcript level of IbEXP1 was relatively high in transgenic line #1 and relatively low in #22 (Fig. 1a). As an indicator of vegetative growth, we measured the size of the fourth rosette leaf to emerge from the T3 homozygous plants of the three transgenic lines and Col-0. Under normal growth conditions, the growth of the fourth rosette leaf in IbEXP1-ox plants exceeded that of its counterpart in Col-0 plants; by 3 weeks post-planting, both the length and width of the fourth rosette leaf were greater in IbEXP1-ox plants than in Col-0 plants (Figs. 1b, S1), suggesting that the growth of rosette leaves was accelerated in IbEXP1-ox plants. At the reproductive stage (6 weeks after planting), however, the fourth rosette leaf was of a similar size in both IbEXP1-ox and Col-0 plants (Fig. S2), indicating that although the initial growth rate was enhanced in IbEXP1-ox plants during the early vegetative growth stage, the ultimate size of the rosette leaf was unaltered from that of Col-0 plants. In contrast, the number of rosette leaves at the reproductive stage markedly increased in IbEXP1-ox plants compared to Col-0 plants (Figs. 1c, S3). Rosette leaf development ceased earlier in Col-0 plants, whereas it continued even after silique development in IbEXP1-ox plants, resulting in a higher number of rosette leaves in the latter plants. Taken together, these results indicate that vegetative growth was markedly enhanced in IbEXP1-ox plants.
Vegetative growth in IbEXP1-ox Arabidopsis plants. a Transcript levels of IbEXP1 in IbEXP1-ox plants. Total RNA was extracted from rosette leaves of IbEXP1-ox plants. Real-time RT-PCR data were normalized to those for elongation factor 1α (EF1α) expression. Error bars indicate the standard deviation between three technical replicates measured on Arabidopsis rosette leaves collected from at least three different plants and subsequently pooled for analysis. b Vegetative growth in IbEXP1-ox plants at 3 weeks after planting. E IbExp1-ox line #1; C Col-0. c Number of rosette leaves at reproductive stage. Small rosette leaves that were not yet fully expanded were also counted. Data were collected at 6 weeks after planting. Mean values of >12 plants with standard deviations are shown. Different letters above the bars indicate significantly different means (P < 0.05) as analyzed by Duncan’s multiple range test using the IBM SPSS Statistics 21 program. a–c Exp-1, 4, and 22 represent IbEXP1-ox lines #1, #4, and #22, respectively
Seed size was increased in IbEXP1-ox plants
Reproductive growth was also distinctly different in IbEXP1-ox and Col-0 plants and was especially evident for silique size (Fig. S4a). Siliques were significantly thicker in IbEXP1-ox plants than in Col-0 plants, whereas they did not significantly differ in length (increase in length in IbEXP1-ox plants was within the range of standard deviation; Figs. S4b, c). At harvest, IbEXP1-ox seeds were larger than Col-0 seeds in terms of both length and width (Fig. 2). This increase in seed size was observed in all three IbEXP1-ox lines (#1, #4, #22). Compared to Col-0 plants, the weight per 1,000 seeds in IbEXP1-ox lines #1, #4, and #22 was 1.4-, 1.4-, and 1.3-fold higher, respectively (Table 1).
Seed development in IbEXP1-ox plants. a Seed phenotype in IbEXP1-ox plants. Seeds were harvested at 10 weeks after planting and air-dried for more than 2 weeks. b Seed size comparison between IbEXP1-ox plants and Col-0 plants. Mean values of >50 seeds from at least six different plants with standard deviations are shown. Different letters above the bars indicate significantly different means (P < 0.05) as analyzed by Duncan’s multiple range test using the IBM SPSS Statistics 21 program. a, b Exp-1, 4, and 22 represent IbEXP1-ox lines #1, #4, and #22, respectively
To determine whether the amount of storage reserves had also increased in the larger IbEXP1-ox seeds, we analyzed protein and starch levels. Total proteins were extracted from identical numbers of Col-0 and IbEXP1-ox seeds, and fractionation of equal portions of the extracts by SDS/PAGE revealed differences in protein levels. The levels of the major seed storage proteins in Arabidopsis (12S) (Fig. 3a) as well as total protein levels per seed (Table S1) were consistently higher in IbEXP1-ox seeds from lines #1, #4, and #22 than in Col-0 seeds. Starch content was also distinctly different in IbEXP1-ox and Col-0 seeds, with the level of starch per seed markedly increasing in IbEXP1-ox seeds from lines #1, #4, and #22 compared to Col-0 seeds (Fig. 3b; Table S1). These results indicate that the larger IbEXP1-ox seeds also accumulated more storage reserves than the Col-0 seeds.
Accumulation of protein and starch in the seeds from IbEXP1-ox plants. a Protein content in seeds from IbEXP1-ox plants. Protein extracts from an equal number of Col-0 and IbEXP1-ox lines seeds were fractionated on a 12 % SDS/polyacrylamide gel and stained. b Starch content in each seed from Col-0 and IbEXP1-ox lines. Starch content per seed was calculated on the basis of the 1,000-seed weight in Table 1. a, b Exp-1, 4, and 22 represent IbEXP1-ox lines #1, #4, and #22, respectively
Seed yield was increased in IbEXP1-ox plants
IbEXP1-ox plants and Col-0 plants differed in terms of the numbers of primary and secondary inflorescence stems (Table 1; Fig. S5). Relative to Col-0 plants, the IbEXP1-ox plants had 1.4- to 1.5-fold more primary inflorescence stems. The increase in the number of secondary inflorescence stems was even greater, with IbEXP1-ox plants having 1.8–2.0-fold more secondary inflorescence stems than Col-0 plants. This increase in the number of primary and secondary inflorescence stems resulted in an increase in the total number of siliques in the IbEXP1-ox plants, which increased by 1.8–2.0-fold in the IbEXP1-ox plants. Consequently, seed yield per plant was also increased in IbEXP1-ox plants relative to Col-0 plants (Fig. 4; Table 1). Total seed weight per individual plant was increased by 2.5-, 2.2-, and 2.1-fold in IbEXP1-ox lines #1, #4, and #22, respectively. This increase (2.1- to 2.5-fold) in total seed yield exceeded the increase in both seed weight (1.3- to 1.4-fold) and silique number (1.8- to 2.0-fold). These results suggest that the increase in both seed weight and number likely affected total seed yield in an additive manner. In addition, the extent of the increase in seed yield in the IbEXP1-ox lines (2.5-fold in #1, 2.2-fold in #4 and 2.1-fold in #22) correlated with the elevated levels of IbEXP1 transcripts, suggesting that the increase in seed yield resulted from an overexpression of IbEXP1.
Seed production in IbEXP1-ox plants. a Comparison of total seed yield between IbEXP1-ox line #1 (Exp-1) and Col-0. Total seeds produced in an IbEXP1-ox line #1 or a Col-0 plant were collected in a 1.5-ml Eppendorf tube. Red line indicates the upper most position of the harvested seeds. b Branching phenotype in IbEXP1-ox line #1 (Exp-1) plants. Picture was taken prior to seed harvest at 10 weeks after planting
Transcript levels of the brassinosteroid-responsive genes were altered in IbEXP1-ox plants
Overexpression of the BR biosynthetic gene (DWARF4) resulted in an increase in vegetative growth and in the number of branches and siliques in Arabidopsis (Choe et al. 2001), which is a similar phenotype to that of the IbEXP1-ox plants in our study. Similarly, the transcript level of AtEXP5 in Arabidopsis was found to be upregulated in BR-treated Arabidopsis plants (Müssig et al. 2002; Coll-Garcia et al. 2004; Park et al. 2010). These findings led us to examine transcriptional regulation of IbEXP1 in response to BR treatment in sweetpotato. Based on the findings of Noh et al. (2009) which showed that IbEXP1 is expressed in the leaf, petiole, and fibrous root of sweetpotato, we analyzed the regulation of IbEXP1 transcription in the fibrous root as this plant organ is the optimal tissue for BR treatment and RNA extraction in sweetpotato. Sweetpotato plantlets bearing a single leaf and petiole (single-leaf plantlets) grown in distilled water for 3 weeks were treated in a solution containing various concentrations (0, 100, 200, and 500 μM) of BR for 3 h. Total RNAs were then extracted from the fibrous roots of the BR-treated sweetpotato plants and subjected to analysis by real-time RT-PCR using IbEXP1-specific primers. We found that the transcript level of IbEXP1 increased in response to BR treatment (Fig. 5a). The IbEXP1 transcript level increased following treatment at 100, 200, and 500 μM BR, although there was a slight dip in the relative increase at 200 μM BR. These results indicate that IbEXP1 expression is BR-inducible.
Transcript levels of BR-responsive genes in IbEXP1-ox plants. a Effect of BR on the transcript level of IbEXP1. Total RNA was extracted from the sweetpotato fibrous roots treated with various concentrations of BR for 3 h. Real-time RT-PCR data were normalized to those for the endogenous β-tubulin gene. b Transcript level of KCS1 in IbEXP1-ox plants. c Transcript level of δ-TIP in IbEXP1-ox plants. b, c Total RNA was extracted from the seedlings of IbEXP1-ox or Col-0 plants. d–f Transcript levels of seed size-related genes in IbEXP1-ox plants. Total RNA was extracted from siliques at 4- to 5-days after pollination. b–f Real-time RT-PCR data were normalized to those for the elongation factor 1α (EF1α) expression. a–f Error bars indicate the standard deviation between three technical replicates measured on sweetpotato fibrous roots, Arabidopsis seedlings or siliques collected from at least three different plants and subsequently pooled for analysis
Fatty acid elongase 3-ketoacyl-CoA synthase 1 (KCSI) and tonoplast integral protein (δ-TIP) have been reported to be BR-responsive genes whose transcript levels are upregulated by BR treatment (Müssig et al. 2002; Coll-Garcia et al. 2004). We therefore investigated any alteration in the transcript levels of KCSI and δ-TIP in the seedlings of IbEXP1-ox Arabidopsis lines #1 and #22 by real-time RT-PCR analysis using KCSI- or δ-TIP-specific primers. The transcript level of KCSI increased in IbEXP1-ox lines #1 and #22, but that of δ-TIP markedly decreased (Fig. 5b, c), indicating that the transcript level of these BR-responsive genes was altered in the IbEXP1-ox plants and further suggesting the possibility that overexpression of BR-inducible IbEXP1 regulates the transcript levels of BR-responsive genes.
Jiang et al. (2013) recently demonstrated that BR regulates seed size and seed shape by transcriptionally modulating the seed size-related genes, including SHORT HYPOCOTYL UNDER BLUE1 (SHB1), MINISEED3 (MINI3), HAIKU1 (IKU1), HAIKU2 (IKU2), APETALA2 (AP2), and AUXIN RESPONSIVE FACTOR2 (ARF2). However, the direction of the regulation varies, with BR activating expression of SHB1, MINI3, IKU1 and IKU2, which are known positive regulators of seed size, but repressing the expression of AP2 and ARF2, which are negative regulators of seed size (Jiang et al. 2013). Since seed size increased in our IbEXP1-ox plants, we examined the transcript levels of the positive regulators (SHB1, MINI3, IKU1) in the developing siliques from IbEXP1-ox lines #1 and #22. The transcript levels of MINI3 and IKU1 were elevated in the siliques from the IbEXP1-ox plants compared to those from Col-0 plants, but SHB1 expression was unaltered (Fig. 5d–f), indicating that certain BR-activating positive regulators of seed size are transcriptionally up-regulated in the IbEXP1-ox plants. These results suggest the possibility that overexpression of IbEXP1 activates at least one BR-activating pathway related to seed development and leads to an increase in seed size.
Discussion
In one of our earlier studies, we observed that the elongation growth of various organs was limited in IbEXP1-antisense transgenic sweetpotato plants, resulting in shorter fibrous roots, stems, petioles, and internodes, and smaller leaves (Noh et al. 2013). These changes indicate that the biological function of IbEXP1 has an effect on the elongation growth of various sweetpotato organs. To examine this effect in more detail, in the present study we overexpressed IbEXP1 in heterologous Arabidopsis plants (i.e., the IbEXP1-ox transgenic Arabidopsis lines) and found that ectopic expression of IbEXP1 led to an enhanced growth rate during the vegetative growth stage (Fig. 1b). This result implies that the growth-promoting activity of IbEXP1 was maintained in the heterologous plant and suggests that IbEXP1 has the potential to improve the growth rate of heterologous plants. Expansins in heterologous plants have also been observed to have universal growth-promoting activities when a soybean expansin gene (GmEXP1), Chinese fir expansin genes (CIEXPA1 and CIEXPA2), and a wheat expansin gene (TaEXPB23), respectively, were overexpressed in tobacco (Lee et al. 2003; Li et al. 2011; Wang et al. 2011). However, in another study, no morphological changes resulted with overexpression of a rose dehydration-inducible expansin gene (RhEXPA4) in RhEXPA4-ox Arabidopsis plants, although plant height was reduced (Dai et al. 2012). Also, ectopic expression of a soybean root-specific β-expansin gene (GmEXPB2) led to no observed alterations in the growth rate of the aerial part of Arabidopsis plants, and only root growth was accelerated (Guo et al. 2011). Therefore, it is most likely that not all of the expansin genes can accelerate overall growth rate in the expansin-ox heterologous plants.
In our study, seed yield was 2.1- to 2.5-fold higher in IbEXP1-ox Arabidopsis plants than in the wild-type Col-0 plants (Table 1; Fig. 4). Seed yield has two main components, i.e., seed size and number, and both the increased number and the increased size of the seeds of IbEXP1-ox plants resulted in an increased seed yield of 2.1- to 2.5-fold, which is relatively higher than those reported in earlier studies (Giroux et al. 1996; Choe et al. 2001; Ashikari et al. 2005; van Daele et al. 2012; Wang et al. 2012). Extensive molecular studies of seed development have led to the identification and characterization of many genes involved in seed development (Garcia et al. 2003; Luo et al. 2005; Zhou et al. 2009; Wang et al. 2010). These include IKU1, IKU2, MINI3, and SHB1, which have been found to positively regulate seed size and whose expressions are activated by BR treatment (Garcia et al. 2003; Luo et al. 2005; Zhou et al. 2009; Wang et al. 2010). We found that the transcript levels of IKU1 and MINI3 were increased in the siliques of IbEXP1-ox plants, but that the expression of SHB1 remained unchanged (Fig. 5d–f), suggesting that expansin probably functions upstream of IKU1 and MINI3. The results of earlier studies demonstrated that IKU1 and MINI3 act in the same seed development-related pathway, while SHB1 in an independent pathway (Luo et al. 2005; Zhou et al. 2009; Wang et al. 2010; Jiang et al. 2013). Thus, it is likely that overexpression of IbEXP1 activates at least one BR-activating seed development-related pathway in which IKU1 and MINI3 are involved, leading to an increase in seed size, but that the SHB1-dependent pathway remains unaffected. The phenotype of larger seeds in the IbEXP1-ox plants, however, was dependent on pot size: seed size differed when IbEXP1-ox plants were grown in a relatively larger pot (pot volume: 330 cm3 for each plant), but the seed-size phenotype disappeared when each plant was grown in a smaller pot (pot volume: 152 cm3 for each plant). This difference is possibly due to the increased number of rosette leaves in the IbEXP1-ox plants grown in the larger pots (Figs. 1c, S3). The number of rosette leaves were, however, almost identical between IbEXP1-ox and Col-0 plants grown in the smaller pots. It is likely that the higher number of rosette leaves provide more of the photosynthate needed for seed development, thereby leading to an increase in seed size.
On the contrary, the phenotype of a higher number of seeds was constantly observed in the IbEXP1-ox plants regardless of pot size. IbEXP1-ox plants stably produced a higher number of lateral inflorescence stems, which subsequently gave rise to more siliques (Table 1; Figs. 4, S5). Choe et al. (2001) also observed a similar phenotype in DWARF4 (BR-biosynthetic gene)-overexpressing Arabidopsis plant. These observations suggest a possible regulation of the IbEXP1 gene by BR, which is supported by our finding that the transcript level of IbEXP1 was upregulated in response to BR treatment (Fig. 5a). This result is in agreement with those of studies showing the upregulation of AtEXP5 by BR treatment (Müssig et al. 2002; Coll-Garcia et al. 2004; Park et al. 2010). In two of these earlier studies (Müssig et al. 2002; Coll-Garcia et al. 2004), the authors found that the transcript levels of KCSI and δ-TIP were also up-regulated in response to BR treatment. In our study, we also found the transcript levels of these two genes to be altered in IbEXP1-ox plants (Fig. 5b, c). The KCS1 transcript level was increased in the IbEXP1-ox plants relative to Col-0, suggesting that expansin and KCS1 genes are probably in the same BR signaling pathway and that KCS1 is most likely to function downstream of expansin. The transcript level of δ-TIP was, however, relatively lower in the IbEXP1-ox plants compared to the Col-0 plants. This result suggests two possible mechanisms. First, it is possible that the increased transcript level of δ-TIP found in earlier studies (Müssig et al. 2002; Coll-Garcia et al. 2004) is a transient effect induced by the BR treatment and that ultimately the transcript level of δ-TIP is downregulated by prolonged BR treatment. The fact that these earlier studies investigated the transcript level of δ-TIP in BR-treated samples harvested at one specific time point (1 or 7 h after BR treatment) supports this possibility. The second possibility is that BR treatment may affect diverse BR-signaling pathways and that expansin belongs to one of the pathways; in this case, overexpression of expansin alone is not sufficient to induce the expression of δ-TIP as the latter functions in an independent BR-signaling pathway. This hypothesis is supported by the above-mentioned finding that, in the siliques of IbEXP1-ox plants, overexpression of IbEXP1 activated the expression of IKU1 and MINI3, which are known to act in the same seed development-related pathway (Luo et al. 2005; Wang et al. 2010; Jiang et al. 2013), but unaffected that of SHB1, which is known to act in an independent pathway (Zhou et al. 2009)—although both of the pathways are known to be BR-inducible (Jiang et al. 2013). These results therefore imply that overexpression of IbEXP1 increases the transcript levels of certain BR-activating genes, thereby leading to the similar morphological alterations observed in the BR-treated plants.
It has been proposed that modulation of cell-wall extensibility could play a key role in plant morphogenesis (Green 1997). Expansins are unique cell-wall proteins that function in vivo to modulate cell-wall extensibility and thereby regulate organ growth and morphogenesis (Fleming et al. 1997; Lee et al. 2001; Pien et al. 2001; Cosgrove et al. 2002; Li et al. 2003). The role of expansins in leaf formation has been demonstrated in earlier studies (Fleming et al. 1997; Pien et al. 2001). Fleming et al. (1997) elicited the development of leaf-like structures at the shoot apical meristem of tomato plants through the local application of expansin protein, and Pien et al. (2001) induced local expression of an expansin gene at the shoot apical meristem which led to the formation of an entire leaf. In addition, Zenoni et al. (2011) demonstrated the role of expansin in axillary meristem formation by showing that overexpression of PhEXPA1 promoted axillary meristem development in Petunia hybrid. In the present study, we observed an increased number of rosette leaves and of primary and secondary inflorescence stems at the reproductive stage in IbEXP1-ox plants compared to Col-0 plants (Fig. 1c; Table 1). Thus, it is most likely that the higher expansin levels in IbEXP1-ox plants promote the development of rosette leaves and of primary and secondary inflorescence stems, resulting in the higher number of these organs and an increase in seed yield. To date, a phenotype for seed yield has not been reported in expansin gene-overexpressing plants. Thus, IbEXP1 is the first expansin gene to be identified which is able to enhance seed yield as well as overall vegetative growth in heterologous plants.
References
Ashikari M, Sakakibara H, Lin S, Yamamoto T, Takashi T, Nishimura A, Angeles ER, Qian Q, Kitano H, Matsuoka M (2005) Cytokinin oxidase regulates rice grain production. Science 309:741–745
Brummell DA, Harpster MH, Civello PM, Palys JM, Bennett AB, Dunsmuir P (1999) Modification of expansin protein abundance in tomato fruit alters softening and cell wall polymer metabolism during ripening. Plant Cell 11:2203–2216
Cho HT, Cosgrove DJ (2000) Altered expression of expansin modulates leaf growth and pedicel abscission in Arabidopsis thaliana. Proc Natl Acad Sci USA 97:9783–9788
Choe S, Fujioka S, Noguchi T, Takatsuto S, Yoshida S, Feldmann KA (2001) Overexpression of DWARF4 in the brassinosteroid biosynthetic pathway results in increased vegetative growth and seed yield in Arabidopsis. Plant J 26:573–582
Choi D, Lee Y, Cho HT, Kende H (2003) Regulation of expansin gene expression affects growth and development in transgenic rice plants. Plant Cell 15:1386–1398
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Coll-Garcia D, Mazuch J, Altmann T, Müssig C (2004) EXORDIUM regulates brassinosteroid-responsive genes. FEBS Lett 563:82–86
Cosgrove DJ, Li LC, Cho HT, Hoffmann-Benning S, Moore RC, Blecker D (2002) The growing world of expansins. Plant Cell Physiol 43:1436–1444
Dai F, Zhang C, Jiang X, Kang M, Yin X, Lü P, Zhang X, Zheng Y, Gao J (2012) RhNAC2 and RhEXPA4 are involved in the regulation of dehydration tolerance during the expansion of rose petals. Plant Physiol 160:2064–2082
Fleming AJ, McQueen-Mason S, Mandel T, Kuhlemeier C (1997) Induction of leaf primordia by the cell wall protein expansin. Science 276:1415–1418
Fu J, Thiemann A, Schrag TA, Melchinger AE, Scholten S, Frisch M (2010) Dissecting grain yield pathways and their interactions with grain dry matter content by a two-step correlation approach with maize seedling transcriptome. BMC Plant Biol 10:63
Garcia D, Saingery V, Chambrier P, Mayer U, Jürgens G, Berger F (2003) Arabidopsis haiku mutants reveal new controls of seed size by endosperm. Plant Physiol 131:1661–1670
Giroux MJ, Shaw J, Barry G, Cobb BG, Greene T, Okita T, Hannah LC (1996) A single mutation that increases maize seed weight. Proc Natl Acad Sci USA 93:5824–5829
Green PB (1997) Expansin and morphology: a role for biophysics. Trends Plant Sci 2:365–366
Guo W, Zhao J, Li X, Qin L, Yan X, Liao H (2011) A soybean β-expansin gene GmEXPB2 intrinsically involved in root system architecture responses to abiotic stresses. Plant J 66:541–552
Han YY, Li AX, Li F, Zhao MR, Wang W (2012) Characterization of a wheat (Triticum aestivum L.) expansin gene, TaEXPB23, involved in the abiotic stress response and phytohormone regulation. Plant Physiol Biochem 54:49–58
Jiang WB, Huang HY, Hu YW, Zhu SW, Wang ZY, Lin WH (2013) Brassinosteroid regulates seed size and shape in Arabidopsis. Plant Physiol 162:1965–1977
Kwon YR, Lee HJ, Kim KH, Hong SW, Lee SJ, Lee H (2008) Ectopic expression of Expansin3 or Expansinβ1 causes enhanced hormone and salt stress sensitivity in Arabidopsis. Biotechnol Lett 30:1281–1288
Lee Y, Choi D, Kende H (2001) Expansins: ever-expanding numbers and functions. Curr Opin Plant Biol 4:527–532
Lee DK, Ahn JH, Song SK, Choi YD, Lee JS (2003) Expression of an expansin gene is correlated with root elongation in soybean. Plant Physiol 131:985–997
Li Y, Jones L, McQueen-Mason S (2003) Expansins and cell growth. Curr Opin Plant Biol 6:603–610
Li F, Asami T, Wu X, Tsang EW, Cutler AJ (2007) A putative hydroxysteroid dehydrogenase involved in regulating plant growth and development. Plant Physiol 145:87–97
Li F, Xing S, Guo Q, Zhao M, Zhang J, Gao Q, Wang G, Wang W (2011) Drought tolerance through over-expression of the expansin gene TaEXPB23 in transgenic tobacco. J Plant Physiol 168:960–966
Lin C, Choi HS, Cho HT (2011) Root hair-specific EXPANSIN A7 is required for root hair elongation in Arabidopsis. Mol Cells 31:393–397
Lizana XC, Riegel R, Gomez LD, Herrera J, Isla A, McQueen-Mason SJ, Calderini DF (2010) Expansins expression is associated with grain size dynamics in wheat (Triticum aestivum L.). J Exp Bot 61:1147–1157
Luo M, Dennis ES, Berger F, Peacock WJ, Chaudhury A (2005) MINISEED3 (MINI3), a WRKY family gene, and HAIKU2 (IKU2), a leucine-rich repeat (LRR) KINASE gene, are regulators of seed size in Arabidopsis. Proc Natl Acad Sci USA 102:17531–17536
McQueen-Mason S, Cosgrove D (1995) Expansin mode of action on cell walls. Analysis of wall hydrolysis, stress relaxation, and binding. Plant Physiol 107:87–100
McQueen-Mason S, Durachko DM, Cosgrove DJ (1992) Two endogenous proteins that induce cell wall extension in plants. Plant Cell 4:1425–1433
Müssig C, Fischer S, Altmann T (2002) Brassinosteroid-regulated gene expression. Plant Physiol 129:1241–1251
Noh SA, Park SH, Huh GH, Paek K-H, Shin JS, Bae JM (2009) Growth retardation and differential regulation of expansin genes in chilling-stressed sweetpotato. Plant Biotechnol Rep 3:75–85
Noh SA, Lee HS, Huh EJ, Huh GH, Paek KH, Shin JS, Bae JM (2010) SRD1 is involved in the auxin-mediated initial thickening growth of storage root by enhancing proliferation of metaxylem and cambium cells in sweetpotato (Ipomoea batatas). J Exp Bot 61:1337–1349
Noh SA, Lee HS, Kim YS, Paek KH, Shin JS, Bae JM (2013) Down-regulation of the IbEXP1 gene enhanced storage root development in sweetpotato. J Exp Bot 64:129–142
Ohto MA, Fischer RL, Goldberg RB, Nakamura K, Harada JJ (2005) Control of seed mass by APETALA2. Proc Natl Acad Sci USA 102:3123–3128
Park CH, Kim TW, Son SH, Hwang JY, Lee SC, Chang SC, Kim SH, Kim SW, Kim SK (2010) Brassinosteroids control AtEXPA5 gene expression in Arabidopsis thaliana. Phytochemistry 71:380–387
Pien S, Wyrzykowska J, McQueen-Mason S, Smart C, Fleming A (2001) Local expression of expansin induces the entire process of leaf development and modifies leaf shape. Proc Natl Acad Sci USA 98:11812–11817
Roxrud I, Lid SE, Fletcher JC, Schmidt ED, Opsahl-Sorteberg HG (2007) GASA4, one of the 14-member Arabidopsis GASA family of small polypeptides, regulates flowering and seed development. Plant Cell Physiol 48:471–483
Schipper O, Schaefer D, Reski R, Flemin A (2002) Expansins in the bryophyte Physcomitrella patens. Plant Mol Biol 50:789–802
Van Daele I, Gonzalez N, Vercauteren I, de Smet L, Inzé D, Roldán-Ruiz I, Vuylsteke M (2012) A comparative study of seed yield parameters in Arabidopsis thaliana mutants and transgenics. Plant Biotechnol J 10:488–500
Wang E, Wang J, Zhu X, Hao W, Wang L, Li Q, Zhang L, He W, Lu B, Lin H, Ma H, Zhang G, He Z (2008) Control of rice grain-filling and yield by a gene with a potential signature of domestication. Nat Genet 40:1370–1374
Wang A, Garcia D, Zhang H, Feng K, Chaudhury A, Berger F, Peacock WJ, Dennis ES, Luo M (2010) The VQ motif protein IKU1 regulates endosperm growth and seed size in Arabidopsis. Plant J 63:670–679
Wang G, Gao Y, Wang J, Yang L, Song R, Li X, Shi J (2011) Overexpression of two cambium-abundant Chinese fir (Cunninghamia lanceolata) α-expansin genes ClEXPA1 and ClEXPA2 affect growth and development in transgenic tobacco and increase the amount of cellulose in stem cell walls. Plant Biotechnol J 9:486–502
Wang S, Wu K, Yuan Q, Liu X, Liu Z, Lin X, Zeng R, Zhu H, Dong G, Qian Q, Zhang G, Fu X (2012) Control of grain size, shape and quality by OsSPL16 in rice. Nat Genet 44:950–954
Werner T, Motyka V, Laucou V, Smets R, Van Onckelen H, Schmülling T (2003) Cytokinin-deficient transgenic Arabidopsis plants show multiple developmental alterations indicating opposite functions of cytokinins in the regulation of shoot and root meristem activity. Plant Cell 15:2532–2550
You MK, Hur CG, Ahn YS, Suh MC, Jeong BC, Shin JS, Bae JM (2003) Identification of genes possibly related to storage root induction in sweetpotato. FEBS Lett 536:101–105
Zenoni S, Reale L, Tornielli GB, Lanfaloni L, Porceddu A, Ferrarini A, Moretti C, Zamboni A, Speghini A, Ferranti F, Pezzotti M (2004) Downregulation of the Petunia hybrida α-expansin gene PhEXP1 reduces the amount of crystalline cellulose in cell walls and leads to phenotypic changes in petal limbs. Plant Cell 16:295–308
Zenoni S, Fasoli M, Tornielli GB, Dal Santo S, Sanson A, de Groot P, Sordo S, Citterio S, Monti F, Pezzotti M (2011) Overexpression of PhEXPA1 increases cell size, modifies cell wall polymer composition and affects the timing of axillary meristem development in Petunia hybrida. New Phytol 191:662–677
Zhou Y, Zhang X, Kang X, Zhao X, Zhang X, Ni M (2009) SHORT HYPOCOTYL UNDER BLUE1 associates with MINISEED3 and HAIKU2 promoters in vivo to regulate Arabidopsis seed development. Plant Cell 21:106–117
Acknowledgments
This work was carried out with the support of “Cooperative Research Program for Agriculture Science & Technology Development” (Next-Generation BioGreen 21 Program No. PJ00810302 and No. PJ00807603), Rural Development Administration, Republic of Korea.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Bae, J.M., Kwak, M.S., Noh, S.A. et al. Overexpression of sweetpotato expansin cDNA (IbEXP1) increases seed yield in Arabidopsis. Transgenic Res 23, 657–667 (2014). https://doi.org/10.1007/s11248-014-9804-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11248-014-9804-1