Abstract
Alternaria brassicicola is a necrotrophic pathogenic fungus that severely affects both the yield and quality of Gladiolus cut flowers and cormels. The jasmonate signaling pathway plays an important role in plant disease resistance. The protein CORONATINE INSENSITIVE 1 (COI1) is a key regulator of this pathway, and the COI1 gene can modulate jasmonate signaling-induced defenses against necrotrophic fungi. Here, we characterize a COI1 gene from Gladiolus, GhCOI1. GhCOI1 transcript levels were greatest in the leaves, followed by the roots. A low MeJA concentration promoted GhCOI1 expression, which remained high after 6 h. Silencing GhCOI1 via VIGS reduced GhCOI1 transcript levels. The length of fungal hyphae and amount of fungal DNA were significantly greater in silenced plants than in control plants. Moreover, GhCOI1 was necessary for the induction of SOD, POD, PPO, and CAT in the Gladiolus defense response to A. brassicicola infestation. GhCOI1 silencing reduced the accumulation of H2O2, allowing the fungus to enter leaf tissue through stomata and cause necrosis directly. Taken together, these results suggest that GhCOI1 is an essential signaling component that controls the JA-regulated defense response against A. brassicicola in Gladiolus plants.
Key message
The results of this study reveal that silencing GhCOI1 reduced the disease resistance of Gladiolus
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Gladiolus is vulnerable to many diseases that severely affect both the yield and quality of its cut flowers and corms, causing extensive economic losses (Gao and Wu 2008; Li et al. 2007; Liu 2008; Pieterse et al. 2009; Pratibha et al. 2014). Pathogens that cause disease in Gladiolus include fungi, viruses, bacteria, and red spiders (Jiao and Guo 1999). The fungi that damage Gladiolus are mainly necrotrophic pathogenic fungi, among which Alternaria brassicicola is a typical pathogen that infects the leaves of plants, leading to leaf spot disease (Si and Si 2007). A. brassicicola can infect hosts directly or can enter through wounds or stomata (Sharma et al. 2014).
Numerous studies have shown that jasmonic acid (JA) is involved in plant growth and development (Cheng et al. 2009; Schommer et al. 2008), with important regulatory roles in the response to biotic and abiotic stresses (Browse and Howe 2008; Reymond and Farmer 1998; Farmer et al. 1992; Lorenzo et al. 2003; Stintzi et al. 2001). In addition, JA signaling is involved in response to diseases caused by necrotrophic pathogens. The JA signaling pathway begins with the recognition of hormone signals by the protein CORONATINE INSENSITIVE 1 (COI1), a JA receptor and key regulator of the JA signal transduction pathway (Katsir et al. 2008). The JA pathway is blocked in the Arabidopsis mutant coi1, resulting in enhanced susceptibility to A. brassicicola (Thomma et al. 1998). Furthermore, silencing of the COI1 gene in rice plants by virus-induced gene silencing (VIGS) has been shown to reduce the activity of peroxidase (POD) and polyphenol oxidase (PPO) by 48.5% and 27.2%, respectively; the resistance of rice plants to rice leafroller also significantly decreased when COI1 was silenced, suggesting that the COI1 gene is vital for regulating JA-mediated resistance (Ye et al. 2012).
Differently, the resistance of Gladiolus to A. brassicicola has not been reported, and the overall role of the COI gene in disease resistance also remains unclear. In this study, we cloned a COI gene from Gladiolus, GhCOI1, and measured its expression in different organs. We also analyzed the changes in GhCOI1 expression after methyl jasmonate (MeJA) treatment, verified the role of GhCOI1 in the JA signaling pathway and explored the ability of A. brassicicola to infect Gladiolus. We further silenced the GhCOI1 gene with VIGS and analyzed the Gladiolus defense response. Finally, we clarified the effect and function of GhCOI1 in the susceptibility of Gladiolus to leaf spot disease. Our results provide a theoretical basis for improving the quality of both cut flowers and cormels.
Materials and methods
Plant materials and growth conditions
Gladiolus hybridus cv. ‘Rose Supreme’ was planted in the Science Research Garden of China Agricultural University. Samples were collected at 26 weeks after planting to determine the expression profile of the GhCOI1 gene. Cormels (Fig. S1) with diameters of 1.0 cm were used as materials for VIGS assays. The cormels were cultivated in a greenhouse (22 °C temperature, 16 h/8 h light/dark photoperiod, 54 μmol m−2 s−1 illumination intensity). Four-week-old plantlets grown in the greenhouse (one plant per pot) were individually sprayed with 1 mL of MeJA that consisted of 0.01% Tween 20.
Isolation and sequence analysis of GhCOI1
Total RNA was extracted from Gladiolus cormels via an RNA extraction reagent kit (Tiangen, Beijing, China), and cDNA was synthesized using Moloney murine leukemia virus (M-MLV) reverse transcriptase (TaKaRa, Shiga, Japan) and the oligonucleotide adapter primer AP1 (Table 1). Rapid amplification of cDNA ends (RACE) was used for GhCOI1 gene cloning, and nested PCR was performed according to the manufacturer’s protocol (Clontech, CA). The protein characteristics were assessed via ExPASy proteomics tools (http://au.expasy.org/tools/). Amino acid multiple alignments were performed by ClustalX 1.8 and BioEdit 7.0, and phylogenetic trees were constructed via MEGA 7.0 software.
Expression analysis of GhCOI1 by qRT-PCR
Via an RNA extraction kit (Tiangen, Beijing, China), total RNA was isolated from samples in liquid nitrogen. The RNA samples were reverse-transcribed using a reverse transcriptase kit (BioTeke, Beijing, China). Four hundred nanograms of cDNA was used as a template for qRT-PCR via a KAPA™ SYBR® FAST qPCR kit (Kapa Biosystems, Woburn, MA, USA) and an Applied Biosystems StepOnePlus™ real-time PCR system. The Gladiolus actin gene (GenBank Accession No. JF831193) was used as a control in Gladiolus. The PCR procedure was performed according to the manufacturer’s instructions. Each qRT-PCR assay was run in three biological replicates. The specific primers used for qRT-PCR are shown in Table 1, and the qRT-PCR procedure was performed as described previously (Seng et al. 2016). The relative transcription levels were calculated according to the 2−ΔΔCT method (Livak and Schmittgen 2001).
Inoculation of A. brassicicola onto Gladiolus leaves
The spore/mycelium mixture (obtained from the Jianqiang Li Laboratory) was shaken at 100 rpm for 30–45 min at 26 °C, which facilitated the complete dispersion of the spores. Sterile gauze was folded into 5–6 layers, and the mixture was then filtered into a new triangular bottle. The new mixture was subsequently centrifuged at 10,000×g for 5 min at room temperature. A. brassicicola spores were collected in sterile water by suspension. The A. brassicicola spores were then diluted to a concentration of 1 × 106 cells/mL in sterile water, after which 0.2% Tween 20 was added as an emulsifier for adhesion to the leaf surface. The solution was sprayed evenly onto the upper and lower surfaces of the leaves of Gladiolus plants. The plants were completely covered with transparent plastic cloth and placed in an incubator (22 °C, 16 h/8 h light/dark photoperiod, 54 μmol m−2 s−1 illumination intensity).
Trypan blue staining
Gladiolus leaves inoculated with fungi were placed in 0.25 mg/mL trypan blue staining solution (lactic acid:glycerol:phenol:sterile water = 1:1:1:1) and infiltrated under vacuum for 20 min. The leaves were then heated for 1–3 min in decolorizing solution (95% ethanol:lactic phenol = 2:1). After the mixture cooled, the leaves were rinsed in 50% ethanol solution for thorough decolorization. The samples were subsequently observed under a microscope, and ImageJ software was used to measure the lengths of hyphae and the necrotic areas of cells.
3,3-Diaminobenzidine (DAB) staining
Gladiolus leaves inoculated with fungi were placed in 1 mg/mL DAB staining solution (pH 5.5) for 20 min, after which the leaves were heated in 95% ethanol solution for 2 min. After the mixture cooled, the leaves were rinsed in 50% ethanol solution for thorough decolorization (Liao et al. 2019). The samples were then observed under a microscope, and ImageJ software was used to measure the stained area.
Superoxide dismutase (SOD), catalase (CAT), POD and PPO activity assays
A 0.5 g leaf sample from tobacco rattle virus (TRV) control and silenced plants was harvested and extracted in liquid nitrogen, followed by homogenization with 10 mL of extraction buffer consisting of 100 mM potassium phosphate buffer (pH 7.0) containing 1 mM EDTA. The mixture was filtered through 4 layers of cheesecloth and centrifuged at 12,000×g for 20 min at 2–4 °C. The supernatant was then used as the enzyme extract. SOD activity was assayed by the photochemical nitro blue tetrazolium (NBT) method, in which 3 mL of reaction mixture contained 100 mM potassium phosphate buffer (pH 7.0), 13 mM methionine, 0.1 mM EDTA, 50 mM sodium carbonate and 25 mM NBT and 0.1 mL of enzyme. SOD activity was determined according to the method of Beyer and Fridovich (1987): one unit of SOD activity was defined as the amount of enzyme that inhibited the photoreduction of NBT by 50% at 560 nm of optical density measured with a spectrophotometer (Hitachi U-2000). CAT activity was assayed by tracking the consumption of hydrogen peroxide (H2O2) at 240 nm for 3 min (Aeby 1984). The 3 mL reaction mixture consisted of 100 mM phosphate buffer (pH 7.0), 5 mM H2O2, and 50 μL of enzyme extract, and the final volume was brought to 3 mL with water. One unit of CAT consumes 1 nmol of H2O2/min under the assay conditions. POD activity was measured by following the change in absorbance at 470 nm due to guaiacol oxidation (Polle et al. 1994). The activity was assayed for 1 min in a 3 mL reaction solution comprising 100 mM potassium phosphate buffer (pH 7.0), 20 mM guaiacol, 10 mM H2O2 and 50 μL of enzyme extract. PPO activity was assayed in a 4 mL reaction solution composed of 100 mM potassium phosphate buffer (pH 7.0), 0.2 M catechol and 50 μL of enzyme extract at 420 nm.
Silencing of the GhCOI1 gene in Gladiolus
Silencing of GhCOI1 in Gladiolus by VIGS was performed as described previously (Zhong et al. 2014). A specific fragment of GhCOI1 was inserted into a pTRV2 vector to construct a pTRV2-GhCOI1 plasmid. The resulting pTRV1, pTRV2 and pTRV2-GhCOI1 plasmids were transformed separately into Agrobacterium tumefaciens GV3101, which was cultured at 28 °C overnight in Luria–Bertani medium supplemented with 50 μg/mL kanamycin and 50 μg/mL rifampicin. The cultures were collected by centrifugation at 5000 rpm for 15 min, after which the cells were resuspended in infiltration buffer (10 mM MgCl2, 10 mM 2-[N-morpholino]ethanesulfonic acid [MES], and 200 mM acetosyringone) at a final optical density at 600 nm (OD600) of approximately 2.0. Holes were poked into the top and bottom of cormels whose diameter was 1.0 cm. After infection, the cormels were grown in pots in a greenhouse (22 °C, 16 h/8 h light/dark photoperiod).
Results
Isolation and sequence analysis of the GhCOI1 gene
The full-length GhCOI1 gene was 2603 bp and contained an open reading frame (ORF) that was 1842 bp in length; this gene encoded a polypeptide of 613 amino acids with a predicted molecular mass of 69.46 kD, and the isoelectric point is predicted to be 5.89. The molecular formula of the GhCOI1 protein is C3054H4875N855O913S40. This protein is soluble, with a stability coefficient of 53.83. There is no transmembrane domain, and the protein does not belong to a family of transmembrane proteins.
Analysis of the functional domain structure of GhCOI1 showed that this protein is a member of a class of F-box proteins (Fig. 1). GhCOI1 contains two important types of characteristic features: one F-box domain located in the N-terminal region and 9 leucine-rich repeat (LRR) motifs. The protein contains 6 JA-binding sites (Arg102, Arg367, Glu369, Val406, Arg428, and Arg515), which is consistent with AtCOI1 of Arabidopsis. In addition, GhCOI1 has 6 phosphorylation sites, including 4 serine phosphorylation sites (Ser14, Ser16, Ser21, and Ser25), 1 threonine phosphorylation site (Thr359) and 1 tyrosine phosphorylation site (Tyr476).
Comparison of the predicted amino acid sequences of GhCOI1 and other plant COIs. Multiple sequence alignments of COI1 were performed using ClustalX 1.8 and BioEdit 7.0 software. Sequences for the following species were obtained from the NCBI database: 1, Gladiolus hybrid; 2, Pisum sativum (EF486643.1); 3, Hevea brasiliensis (EU136026.1); 4, Lycopersicon esculentum (AY423550.1); 5, Nicotiana attenuata (EF025087.1); 6, Arabidopsis thaliana (NM_129552.3); 7, Brassica rapa subsp. chinensis (GU263836.1); 8, Zea mays (NM_001156957.1); 9, Triticum aestivum (HM447645.1); and 10, Oryza sativa (AY168645.1). Black box: F-box domain; black lines: leucine-rich repeats (LRRs); black triangles: jasmonate-binding sites
Phylogenetic analysis revealed that GhCOI1 is closely related to SiCOI1 from Sesamum indicum (Fig. 2). Phyre2 (http://www.sbg.bio.ic.ac.uk/phyre2/htmL) was used to analyze the tertiary structure of the COI proteins. The results showed that the tertiary structures of GhCOI1 and AtCOI1 are highly similar (66%). In addition, the Arabidopsis auxin receptor AtTIR1 is closely related to AtCOI1, and phylogenetic analysis grouped these proteins onto the same branch. AtTIR1 also contains LRR and F-box domains. The tertiary structures of GhCOI1 and AtTIR1 are 33% similar (Fig. 3).
Expression profiles of GhCOI1
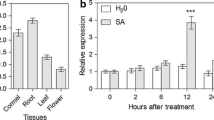
The relative expression levels of GhCOI1 were determined by qRT-PCR assays. GhCOI1 was expressed constitutively in different organs at distinct levels. The highest levels of GhCOI1 mRNA were detected in the leaves, followed by the roots; relatively low levels were detected in the flowers, and the lowest levels were detected in the cormels (Fig. 4a).
GhCOI1 transcript levels in Gladiolus. a Relative GhCOI1 expression levels in various organs of Gladiolus. b Expression of GhCOI1 in the leaves of plants treated with different concentrations of MeJA for 12 h. c Expression of GhCOI1 in the leaves of plants treated with 0.1 mM MeJA after different incubation times. The error bars represent the SEs of three biological replicates (n = 3). The different letters indicate significant differences (p < 0.05; Duncan’s multiple range test); the same letters indicate no significant differences
GhCOI1 expression in the leaves was evaluated at 12 h after the plants were sprayed with different concentrations of MeJA. The highest GhCOI1 expression level, which was approximately 1.5-fold greater than the control level, was detected after treatment with 0.1 mM MeJA. However, compared with those in response to the control treatment, the changes in GhCOI1 expression in response to 0.05 mM and 0.5 mM MeJA treatment were not significantly different (Fig. 4b). An appropriate concentration (0.1 mM) of MeJA promoted GhCOI1 expression, while high concentrations of MeJA (0.75 and 1.00 mM) repressed GhCOI1 expression.
GhCOI1 expression was further analyzed in the leaves of plants treated with 0.1 mM MeJA after 0, 3, 6, 9, 12, 24, 48 and 72 h. As shown in Fig. 4c, MeJA gradually induced the expression of GhCOI1 during the first 6 h, after which time negative feedback on the expression of GhCOI1 was detected, with a relatively high level of transcript maintained. Similar results have been detected in rice and grapes treated with MeJA (Ye et al. 2012; Wang et al. 2012), indicating that GhCOI1, OsCOI1 and VvCOI1 may have similar roles in the JA signaling pathway.
Silencing the GhCOI1 gene reduced its expression in Gladiolus
Leaves of Gladiolus inoculated with A. brassicicola were observed via stereomicroscopy. We found that the pathogen successfully infected leaves and caused leaf spot disease. On the 8th day after inoculation, yellow-green and irregularly shaped lesions began to appear on the leaves. Twelve days later, the central color of the lesions had deepened to brown, and there were light green to yellow-green halos surrounding these lesions. By the 20th day, the yellow-green halos had gradually expanded over the surface of the leaves, and the dark-brown lesions had further deepened to a black-brown color. Additionally, adjacent lesions had fused into irregular round lesions (Fig. 6a).
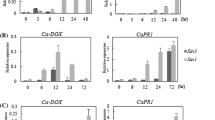
To determine whether GhCOI1 plays a role in disease resistance, we used VIGS to reduce its expression and observed the phenotypes of control and GhCOI1-silenced plants inoculated with A. brassicicola (Fig. 5a). The results showed that, when the control plants were treated with A. brassicicola, GhCOI1 increased sharply in leaves during the first 6 h (0–6 h) but then decreased during the following 6 h (6–12 h). However, the expression of GhCOI1 moderately increased again in the finally 60 h (12–72 h) (Fig. 5b). In the GhCOI1-silenced plants, GhCOI1 barely responded to A. brassicicola (Fig. 5b). Moreover, GhCOI1 expression levels in the leaves of silenced plants were 4.3-fold lower than those in the leaves of the control plants at 6 h. The expression pattern of GhCOI1 in the leaves of control plants (Fig. 5b) was similar to that in the leaves of plants treated with 0.1 mM MeJA (Fig. 4c), suggesting that MeJA and A. brassicicola have similar roles in regulating GhCOI1 expression.
Expression of GhCOI1 in six GhCOI1-silenced plants inoculated with Alternaria brassicicola. a Expression of GhCOI1 in the leaves of six GhCOI1-silenced independent lines. The error bar represents the SEs of three technical replicates. bGhCOI1 is not induced in GhCOI1-silenced plants inoculated with Alternaria brassicicola. The error bar represents the SEs of three biological replicates of three independent plants. The different letters indicate significant differences (p < 0.05; Duncan’s multiple range test); the same letters indicate no significant differences
Silencing the GhCOI1 gene reduced plant disease resistance
Mycelial length is an important index of fungal growth. To determine whether fungal growth is affected by GhCOI1, the mycelial lengths of A. brassicicola inoculated onto Gladiolus leaves were analyzed (Fig. 6). The results showed that the length of mycelia on the leaves of both silenced and TRV control plants was less than 100 μm at 6 h after inoculation; however, the length of mycelia on the leaves of the silenced plants was slightly longer than those on the leaves of the TRV control plants. At 12, 24 and 36 h after inoculation, the lengths of the mycelia on the leaves of the silenced plants were 1.7-, 1.8- and 1.5-fold greater than those of the TRV control plants, respectively, suggesting that GhCOI1 silencing promoted the growth of A. brassicicola on Gladiolus leaves.
Analysis of the length of hyphae on leaves after inoculation. a Leaf appearance at 20 days after inoculation. b Electron microscopy images showing hyphal length after leaf inoculation. The scale bars indicate 5 μm or 20 μm, as indicated. c Statistical analysis of hyphal length after leaf inoculation. The values represent the means, and the error bars represent the SEs of three biological replicates. The different letters indicate significant differences (p < 0.05; Duncan’s multiple range test); the same letters indicate no significant differences
To understand the effect of GhCOI1 on disease, the amount of A. brassicicola DNA within the Gladiolus leaves was quantified by qRT-PCR. At 24 h after inoculation, the amount of fungal DNA within the silenced plants was 2.6-fold greater than that within the TRV control plants, and the difference was significant. After 36 h, the growth rates of the fungi within the control and silenced plants were 58.8% and 33.3%, respectively. Furthermore, at that time, the amount of fungal DNA within the silenced plants was still significantly (2.2-fold) greater than that within control plants (Fig. 7a). Overall, silencing GhCOI1 increased fungal growth efficiency.
Statistical analysis of fungal biomass and the leaf necrotic cell area. a Fungal biomass at 24 h and 36 h after leaf inoculation. DNA was extracted from the leaves of silenced and control plants (n = 3). b Statistical analysis of the leaf necrotic cell area of three different silenced plants. c Necrotic cells showing trypan blue staining of inoculated leaves. The values represent the means, and the error bars represent the SEs of three biological replicates. The different letters indicate significant differences (p < 0.05; Duncan’s multiple range test); the same letters indicate no significant differences. (Color figure online)
To further determine whether GhCOI1 participates in Gladiolus resistance to A. brassicicola, the death of leaf cells infected by pathogens was analyzed by trypan blue staining. As shown in Fig. 7c, a small number of dead cells were observed in the leaves of both silenced and TRV control plants at 24 h after inoculation, and there was no significant difference in the sizes of the necrotic areas. After 36 h, the number of dead cells increased significantly, and the necrotic area was 1.8-fold greater in the silenced plants than in the TRV control plants, which was significantly different (Fig. 7b). These results show that the leaf necrotic area in the silenced and TRV control plants increased gradually with time after inoculation but that leaf necrosis was significantly greater in the silenced plants than in the TRV control plants. Therefore, A. brassicicola was better able to infect GhCOI1-silenced plants, and GhCOI1 gene silencing rendered plants more susceptible to disease.
Silencing the GhCOI1 gene reduced the activities of SOD, CAT, POD and PPO
The activities of the SOD, CAT, POD and PPO enzymes in TRV-GhCOI1 plants were significantly lower than those in the TRV control plants (35.1%, 25%, 50.5% and 60% lower, respectively) (Fig. 8). These findings suggest that the GhCOI1 gene is necessary for the induction of SOD, POD, PPO, and CAT enzyme activity in Gladiolus inoculated with A. brassicicola and that this gene may thus play a positive regulatory role in regulating enzyme activity.
Effects of Alternaria brassicicola inoculation on the activity of defense-related enzyme in the leaves of TRV control and silenced plants. a Analysis of SOD activity in the leaves of TRV control and silenced plants. b Analysis of CAT activity in the leaves of TRV control and silenced plants. c Analysis of POD activity in the leaves of TRV control and silenced plants. d Analysis of PPO activity in the leaves of TRV control and silenced plants. The values represent the means, and error bars represent the SEs of three biological replicates of three independent lines. The different letters indicate significant differences (p < 0.05; Duncan’s multiple range test); the same letters indicate no significant differences
Silencing the GhCOI1 gene reduced H2O2 accumulation in plants
To investigate the relationship between A. brassicicola infection and H2O2 accumulation in leaves, the accumulation of H2O2 in the leaves was assessed by DAB staining, and the results were analyzed with ImageJ software. As illustrated in Fig. 9a, H2O2 accumulated in the leaves of both silenced plants and TRV control plants at 24 h after inoculation. However, the area of H2O2 accumulation was 1.5-fold greater in the leaves of the TRV control plants than in the leaves of the silenced plants. At 36 h, the area of H2O2 accumulation in leaves of the TRV control plants was 1.3-fold greater than that in the leaves of silenced plants, indicating that silencing of GhCOI1 reduced the accumulation of H2O2.
Effects of Alternaria brassicicola inoculation on H2O2 accumulation in the leaves of TRV control and silenced plants. a The area of H2O2 staining was analyzed by DAB staining at 24 h and 36 h after inoculation of the leaves of TRV control and silenced plants. b The accumulation of H2O2 was measured via DAB staining at 24 h and 36 h after inoculation of the leaves of TRV control and silenced plants. The scale bars indicate 100 μm. c Hyphae in leaves inoculated in different ways. In the TRV control plants, the hyphae passed near the stomata and penetrated the leaves directly. In the TRV-GhCOI1 plants, the hyphae entered the leaves through the stomata. The scale bars indicate 5 μm. The values represent the means, and the error bars represent the SEs of three biological replicates of three independent lines. The different letters indicate significant differences (p < 0.05; Duncan’s multiple range test); the same letters indicate no significant differences
The images in Fig. 9b show that the color of the tissue with H2O2 accumulation occurring around the stomata was lighter in the leaves of the silenced plants than in leaves of the control plants. We speculate that the poor resistance of the silenced plants to A. brassicicola was related to the method of invasion. More H2O2 accumulated in the subsidiary cells of the leaves of the TRV control plants than in those of the silenced plants, preventing A. brassicicola invasion through stomata and increasing the resistance of the control plants. In contrast, only a small amount of H2O2 accumulated in the leaves of the silenced plants. Thus, the fungus could directly enter the tissue through the stomata and cause cell necrosis (Fig. 9c).
On the basis of the results in which GhCOI1 responded to A. brassicicola and in which silencing of COI1 in Gladiolus increased the susceptibility to A. brassicicola and impaired the induction of SOD, CAT, PPO and POD, we conclude that the JA signal transduction pathway plays a key role in Gladiolus defense against A. brassicicola and that GhCOI1 is specifically required for the regulation of JA-mediated defense in response to A. brassicicola. Moreover, SOD, CAT, PPO and POD are JA-induced defenses that respond to A. brassicicola. SOD, CAT, PPO and POD are mediated by GhCOI1 (Fig. 10). The GhCOI1 gene product serves as a receptor of the JA signal and activates the JA signaling pathway, thus increasing the downstream activities of the SOD, CAT, PPO and POD enzymes, which then leads to increased Gladiolus resistance against A. brassicicola (Fig. 10).
Diagram showing the involvement of GhCOI1 in response to Alternaria brassicicola. a When normal Gladiolus is invaded by A. brassicicola, endogenous JA is induced, which further activates the JA signaling response, including the key regulator GhCOI1. GhCOI1 is necessary for the induction of SOD, POD, PPO, and CAT in the Gladiolus defense response to A. brassicicola infestation. GhCOI1 also induces H2O2 accumulation, blocking the ability of the fungus to directly enter leaf tissues through the stomata. b When A. brassicicola invades GhCOI1-silenced Gladiolus, the JA signal transduction is weak and cannot induce a sufficient amount of SOD, POD, PPO and CAT for a defense response. The level of H2O2 is relatively low, contributing to the infestation through the stomata. The yellow dots indicate symptoms of A. brassicicola. (Color figure online)
Discussion
In Arabidopsis, the F-box motif at the N-terminus of AtCOI1 mediates the interaction between COI1 and ASK1, and the 18 LRR motifs at the C-terminus have specificity for the recognition of target proteins (Jin et al. 2004). The integrity of the AtCOI1 protein contributes to its stability in vivo (Yan et al. 2013). In wheat, the predicted TaCOI1 protein has an F-box domain, several LRRs and a conserved AMN1 domain (Liu et al. 2017). In the present study, we found the F-box motif to be present at the N-terminus of the GhCOI1 protein. The GhCOI1 protein also contains 9 LRR motifs and 6 JA-binding sites, consistent with AtCOI1 and TaCOI1 (Fig. 1).
AtCOI1 expression has been detected in the roots, stems, leaves, petals and stamens of Arabidopsis (Adie et al. 2007; Benedetti et al. 1998; Melotto et al. 2006). In Capsicum, CaCOI1 is expressed in the roots, stems, leaves and flowers, with the greatest expression occurring in the flowers (Hu et al. 2013). However, the expression of COI1 in Gladiolus differs from that in Capsicum and Arabidopsis. In our study, the greatest levels of GhCOI1 mRNA expression were detected in the leaves, followed by the roots; relatively low levels were detected in the flowers, and the lowest levels were detected in the cormels (Fig. 4). In recent years, homologous COI1 genes have been found in plant species such as rice and wheat. For example, there are three homologous COI genes in the rice genome: OsCOI1a, OsCOI1b, and OsCOI2 (Lee et al. 2013). Notably, inoculation of Pseudomonas fluorescens onto wheat roots induces expression of COI2 (Okubara et al. 2010). It remains to be explored whether homologous COI genes are expressed in other Gladiolus tissues or organs.
It has been demonstrated that the COI protein is a key regulator within the JA pathway (Farmer and Ryan 1990). As a signaling component of plant stress resistance, MeJA participates in the host response to pathogenic microbial stress and induces plant disease resistance. The stress resistance induced by JA is closely related to its concentration (Mehari et al. 2015; Cohen and Flescher 2009). A certain concentration of MeJA has a positive effect on the defense response of host plants. When the concentration of MeJA is not suitable, its induction effect decreases rapidly and can even become toxic (Feng et al. 2009). In the present study, we also found that GhCOI1 is induced by low concentrations of MeJA and repressed by high concentrations of MeJA in Gladiolus (Fig. 4); a suitable concentration of MeJA is 0.1 mM.
In this study, GhCOI1 expression increased sharply during the first 6 h in response to 0.1 mmol/L MeJA treatment but then decreased slightly, after which a high level was maintained (9–72 h) (Fig. 4). A similar expression pattern was also observed in rice and grape (Ye et al. 2012; Wang et al. 2012). We hypothesize that there may be negative feedback or counteraction by other signals to repress the expression of GhCOI1 after 6 h. It seems that GhCOI1 is tightly regulated in plants. Thus, it will be interesting to explore the regulatory mechanism. In JA signaling, an alternative splice variant of JAZ10 was shown to exert negative feedback on its own signal transduction (Moreno et al. 2013).
Silencing GhCOI1 in Gladiolus reduced GhCOI1 transcript levels. Moreover, the fungal biomass within TRV-GhCOI1 plants was significantly greater than that within control plants, suggesting that the GhCOI1 gene is involved in the regulation of Gladiolus disease resistance. To verify this effect of GhCOI1, we used the trypan blue staining method to observe cell necrosis and fungal infection. The results showed that the necrotic area of leaf cells in the TRV-GhCOI1 plants was significantly greater than that of leaf cells in the TRV control plants at 36 h after inoculation (Fig. 7c). Such increased necrosis was beneficial to the growth of the fungus and improved infectivity. Additionally, the hyphae on the TRV-GhCOI1 plants were longer than those on the TRV control plants at 12 h after inoculation (Fig. 6c), demonstrating that silencing GhCOI1 promotes the hyphal growth.
Previous studies have shown that A. brassicicola can infect plants directly or through stomata (Sharma et al. 2014), and reactive oxygen species can induce stomatal closure, thereby preventing hyphae from entering plant tissue through stomata. Figure 9b shows that H2O2 accumulated around some of the stomata of the TRV control plants, thereby inhibiting fungal invasion. Indeed, trypan blue staining confirmed that in the TRV-GhCOI1 plants, fungal hyphae were able to enter stomata but that hyphae did not enter the stomata of the TRV control plants; rather the hyphae expanded around these structures (Fig. 9c).
PPO catalyzes the formation of lignin and other phenolic oxides, producing a protective barrier against invading pathogens. CAT, SOD and POD are defensive enzymes that scavenge reactive oxygen species in cells, and inoculation of Solanum leaves with Alternaria solani was shown to increase phenylalanine ammonia lyase (PAL), POD and PPO activities (Patykowski 2006). In addition, PPO, CAT and SOD activities increase after inoculation of Brassica with A. brassicicola, and the activities of PPO, SOD and PAL are greater in resistant varieties than in susceptible varieties (Hou and Wang 2008). In the present study, infection by A. brassicicola increased the activities of SOD, POD, PPO and CAT. In addition, the POD, PPO and CAT activities were significantly lower in the GhCOI1-silenced plants than in the TRV control plants (Fig. 8), suggesting that COI1 is a positive regulator of the disease resistance process. Silencing of the GhCOI1 gene thus impairs inducible defenses and increases susceptibility to A. brassicicola.
In summary, the GhCOI1 gene was identified and functionally characterized in this study. The GhCOI1 protein acts as a receptor for JA and is a key regulator of JA signaling. Silencing the GhCOI1 gene hinders the plant response to A. brassicicola and ultimately leads to susceptibility, which can result in leaf cell death. On the basis of these findings, we will apply genetic engineering techniques in the future to improve the disease resistance of Gladiolus and to provide a theoretical and technical foundation for improving the quality of cormels and cut flowers.
References
Adie BAT, Pérez-Pérez J, Pérez-Pérez MM, Godoy M, Sánchez-Serrano JJ, Schmelz EA, Solano R (2007) ABA is an essential signal for plant resistance to pathogens affecting JA biosynthesis and the activation of defenses in Arabidopsis. Plant Cell 19:1665–1681
Aeby H (1984) Catalase in vitro. Methods Enzymol 105:121–126
Benedetti CE, Costa CL, Turcinelli SR, Arruda P (1998) Differential expression of a novel gene in response to coronatine, methyl jasmonate, and wounding in the coi1 mutant of Arabidopsis. Plant Physiol 116:1037–1042
Beyer WF, Fridovich I (1987) Assaying for super oxide dismutase activity: some large consequence of minor changes in conditions. Annal Biochem 161:559–566
Browse J, Howe GA (2008) New weapons and a rapid response against insect attack. Plant Physiol 146:832–838
Cheng H, Song SS, Xiao LT, Soo HM, Cheng ZW, Xie DX, Peng JR (2009) Gibberellin acts through jasmonate to control the expression of MYB21, MYB24, and MYB57 to promote stamen filament growth in Arabidopsis. PLoS Genet 5:e1000440
Cohen S, Flescher E (2009) Methyl jasmonate: a plant stress hormone as an anti-cancer drug. Phytochemistry 70:1600–1609
Farmer EE, Ryan CA (1990) Interplant communication: airborne methyl jasmonate induces synthesis of proteinase inhibitors in plant leaves. PNAS 87:7713–7716
Farmer EE, Johnson RR, Ryan CA (1992) Regulation of expression of proteinase inhibitor genes by methyl jasmonate and jasmonic acid. Plant Physiol 98:995–1002
Feng YJ, Wang JW, Luo SM (2009) Timing and concentration effects on the defense response of Zea mays seedlings after application of jasmonic acid to leaves. Chin J Plant Ecol 33:812–823
Gao F, Wu YH (2008) Progresses in the biocontrol of plant diseases caused by Alternaria. Plant Prot 03:1–6
Hou XL, Wang LY (2008) Induced expression of pathogenesis-related protein genes in interaction between non-heading cabbage and black spot pathogen. Chin Soc Hortic Sci 11:201–204
Hu TZ, Yang YW, Zeng H, Hu ZL, Chen ZG, Chen GP, Wu YM (2013) Transgenic pepper plants carrying RNA interference constructs of CaCOI1 gene show severe abnormality. Mol Breed 31:971–979
Jiao P, Guo T (1999) Control of common diseases and pests of Gladiolus. Spec Econ Anim Plant 3:43
Jin J, Cardozo T, Lovering RC, Elledge SJ, Pagano M, Harper JW (2004) Systematic analysis and nomenclature of mammalian F-box proteins. Genes Dev 18:2573–2580
Katsir L, Schilmiller AL, Staswick PE, He SY, Howe GA (2008) COI1 is a critical component of a receptor for jasmonate and the bacterial virulence factor coronatine. PNAS 1057:100–7105
Lee HY, Seo JS, Cho JH, Jung H, Kim JK, Lee JS, Rhee S, Choi YD (2013) Oryza sativa COI homologues restore jasmonate signal transduction in Arabidopsis coi1-1 Mutants. PLoS ONE 8:e528021
Li QF, Wang LM, Liu Y (2007) Study on the epidemiological law of tobacco red spot disease and its control by chemicals. Mod Agric 07:20–21
Liao JJ, Wang CH, Xing QJ, Li YP, Liu XF, Qi HY (2019) Overexpression and VIGS system for functional gene validation in oriental melon (Cucumis melo var. makuwa Makino). Plant Cell Tissue Ogran Cult 137:275–284
Liu F (2008) Identification and control of common diseases and pests in gladiolus. Spec Econ Anim Plants 2:51–52
Liu XY, Wang JY, Fan BL, Shang YT, Sun YF, Dang C, Xie CJ, Wang ZY, Peng YK (2017) A COI1 gene in wheat contributes to the early defence response against wheat powdery mildew. J Phytopathol 166:116–122
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(T)(-Delta Delta C) method. Methods 25:402–408
Lorenzo O, Piqueras R, Sánchez-Serrano JJ, Solano R (2003) ETHYLENE RESPONSE FACTOR1 integrates signals from ethylene and jasmonate pathways in plant defense. Plant Cell 15:165–178
Mehari ZH, Elad Y, Rav-David D, Graber ER, Harel YM (2015) Induced systemic resistance in tomato (Solanum lycopersicum) against Botrytis cinerea by biochar amendment involves jasmonic acid signaling. Plant Soil 395:31–44
Melotto M, Underwood W, Koczan J, Nomura K, He SY (2006) Plant stomata function in innate immunity against bacterial invasion. Cell 126:969–980
Moreno JE, Shyu C, Campos ML, Patel LC, Chung HS, Yao J, He SY, Howe GA (2013) Negative feedback control of jasmonate signaling by an alternative splice variant of JAZ10. Plant Physiol 162:1006–1017
Okubara PA, Call DR, Kwak YS, Skinner DZ (2010) Induction of defense gene homologues in wheat roots during interactions with Pseudomonas fluorescens. Biol Control 55:118–125
Patykowski J (2006) Role of hydrogen peroxide and apoplastic peroxidase in tomato—Botrytis cinerea interaction. Acta Physiol Plant 28:589–598
Pieterse CM, Leon-Reyes A, Van der Ent S, Van Wees SC (2009) Networking by small-molecule hormones in plant immunity. Nat Chem Biol 5:308–316
Polle A, Otteer T, Seeifert F (1994) Apoplastic peroxidases and lignification in needles of norway Spurse (Picea abies L.) Plant Physiol 106:53–56
Pratibha S, Swati D, Bhati DS, Manika S, Chowdappa P (2014) Penetration and infection processes of Alternaria brassicicola on cauliflower leaf and Alternaria brassicae on mustard leaf: a histopathological study. Plant Pathol J (Faisalabad) 13:100–111
Reymond P, Farmer EE (1998) Jasmonate and salicylate as global signals for defense gene expression. Curr Opin Plant Biol 1:404–411
Schommer C, PalatnikJF AggarwalP, Chételat A, Cubas P, Farmer EE, Nath U, Weigel D (2008) Control of jasmonate biosynthesis and senescence by miR319 targets. Plos Bio 16:e230
Seng SS, Wu J, Sui JJ, Wu CY, Zhong XH, Liu C, Liu C, Gong BH, Zhang FQ, He JN, Yi MF (2016) ADP-glucose pyrophosphorylase gene plays a key role in the quality of corm and yield of cormels in gladiolus. Biochem Biophy Res Comm 474:206–212
Sharma P, Deep S, Bhati DS, Sharma M, Chowdappa P (2014) Penetration and infection processes of Alternaria brassicicola on cauliflower leaf and Alternaria brassicae on mustard leaf: a histopath ological study. Plant PathJ 13:100–111
Si F, Si Y (2007) Occurrence and control of black spot disease of cabbage and broccoli. J Changjiang Veg 11:21–22
Stintzi A, Weber H, Reymond P, Browse J, Farmer EE (2001) Plant defense in the absence of jasmonic acid: the role of cyclopentenones. PNAS 98:12837–12842
Thomma BPHJ, Eggermont K, Penninckx IAMA, Mauch-Mani B, Vogelsang R, Cammue BPA, Broekaert WF (1998) Separate jasmonate-dependent and salicylate-dependent defense-response pathways in Arabidopsis are essential for resistance to distinct microbial pathogens. PNAS 95:15107–15111
Wang WY, Yue LX, Zhang YY, Chu JQ, Zhang XY, Fang JG (2012) Cloning of several important genes involved in grapevine SA and JA signaling pathways and their response to exogenous signals. Acta Hortic Sinica 05:817–827
Yan J, Li H, Li S, Yao R, Deng H, Xie Q, Xie D (2013) The Arabidopsis F-box protein CORONATINE INSENSITIVE1 is stabilized by SCFCOI1 and degraded via the 26S proteasome pathway. Plant Cell 25:486–498
Ye M, Luo SM, Xie JF, Li YF, Xu T, Liu Y, Song YY, Zhu-Salzman K, Zeng RS (2012) Silencing COI1 in rice increases susceptibility to chewing insects and impairs inducible defense. PLoS ONE 7:e36214
Zhong XH, Yuan X, Wu Z, Khan MA, Chen J, Li XX, Gong BH, Zhao Y, Wu J, Wu CY (2014) Virus-induced gene silencing for comparative functional studies in Gladiolus hybridus. Plant Cell Rep 33:301–312
Acknowledgements
This work was supported by the National Natural Science Foundation of China (31171991 to MFY and 31701952 to JW) and the Chinese Postdoctoral Science Foundation (2018M641569 to SSS).
Author information
Authors and Affiliations
Contributions
SSS and CYW designed the experiments. SSS, CYW and JW performed the experiments. SSS and XHZ analyzed the data. SSS wrote the manuscript. JW, JNH and MFY revised the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
All authors declare that they have no conflicts of interest.
Additional information
Communicated by Degao Liu.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Seng, S., Wu, C., Wu, J. et al. Silencing GhCOI1 in Gladiolus hybridus increases susceptibility to Alternaria brassicicola and impairs inducible defenses. Plant Cell Tiss Organ Cult 140, 69–81 (2020). https://doi.org/10.1007/s11240-019-01711-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-019-01711-6