Abstract
Although transgenic crops expressing either Cry1Ab or Cry1Ac, both derived from Bacillus thuringiensis (Bt), have been used commercially, the evolution of insects resistance to these CRY proteins has become a challenge. Thus, it has been proposed that co-expression of two Bt proteins with different modes of action may delay the development of resistance to Bt. However, few Bt proteins have been identified as having different modes of action from those of Cry1Ab or Cry1Ac. In this study, transgenic lines of maize over-expressing either Cry1Ie or Cry1Ac gene have been developed. Several independent transgenic lines with one copy of the foreign gene were identified by Southern blot analysis. Bioassays in the laboratory showed that the transgenic plants over-expressing Cry1Ie were highly toxic against the wild-type cotton bollworm (Heliothis armigera), producing mortality levels of 50 % after 6 days of exposure. However, the mortality caused by these plants was lower than that caused by the Cry1Ac transgenic plants (80 %) and MON810 plants expressing Cry1Ab (100 %), which both exhibited low toxicity toward the Cry1Ac-resistant cotton bollworm. In contrast, three transgenic maize lines expressing Cry1Ie induced higher mortality against this pest and were also highly toxic to the Asian corn borer (Ostrinia furnacalis) in the field. These results indicate that the Cry1Ie protein has a different mode of action than the Cry1Ab and Cry1Ac proteins. Therefore, the use of transgenic plants expressing Cry1Ie might delay the development of Bt-resistant insects in the field.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Transgenic insect-resistant crops producing Bacillus thuringiensis (Bt) insecticidal proteins have been used commercially since 1996. Although the wide use of transgenic Bt crops has led to the reduction of pesticide usage and production costs, the evolution of insects resistant to these crops presents a challenge. Many insects which are resistant to Bt insecticidal proteins have been selected in the laboratory, and the occurrence of insect resistance in the field has also been reported (Tabashnik et al. 2008, 2011).
Several models of the action of Bt toxins have been proposed. The pore formation model proposes that the α-loop of domain I of Bt toxins binds to larval mid-gut membrane-associated receptors, including cadherin, aminopeptidase and alkaline phosphatase (Jurat-Fuentes et al. 2011; Tabashnik et al. 2011), and then creates pores in the midgut membrane, leading to cell death. Another possibility is the signal transduction model, which proposes that the binding of Bt toxins to cadherin triggers an adenylyl cyclase/ protein kinase A (PKA) signal pathway, activating PKA, which leads to oncotic cell death (Zhang et al. 2006). Insects have developed resistance to Bt toxins as a result of the mutation of midgut receptors, leading to the disruption of Bt toxin binding to receptors, which is the most common mechanism of insect resistance (Ferre and Van Rie 2002).
Several strategies have been used to delay insect resistance. According to the refuge strategy, nontransgenic crops are planted near Bt crops to promote the survival of susceptible insects, which will potentially mate with the resistant insects that survive on the Bt crops (Bates et al. 2005; Tabashnik et al. 2008). The pyramid strategy refers to the expression of two or more Bt toxins with different modes of action in the same plants. Cotton plants expressing Cry1Ac and Cry2Ab were found to be more toxic to bollworms than the transgenic plants expressing one gene (Chitkowski et al. 2003). Indeed, Monsanto Company has developed transgenic maize pyramiding eight different genes in one plant (Gatehouse 2008; Soberon et al. 2007; Tabashnik et al. 2011). Recently, it was found that modified Cry1Ab and Cry1Ac toxins lacking the helix α-1 domain could counter insect resistance in laboratory bioassays (Soberon et al. 2007; Tabashnik et al. 2011), providing another strategy for delaying insect resistance.
The development of Bt plants expressing novel Bt toxins is an effective strategy to delay insect resistance. Although more than 700 different Bt proteins have been classified into 72 Cry groups and 3 Cyt groups (http://www.lifesci.sussex.ac.uk/home/Neil_Crickmore/Bt/toxins2.html), only approximately a dozen (Cry1Aa, Cry1Ab, Cry1Ac, Cry1C, Cry1D, Cry1E, Cry1F, Cry2Aa, Cry2Ab, Cry3A, Cry3B and Cry34/Cry35) are used commercially as sprays or in Bt crops (Bravo and Soberon 2008; Tohidfar et al. 2013). Cry1Ie is a novel Cry1 gene that was obtained by the PCR-restriction fragment length polymorphism (RFLP) method (Song et al. 2003). Our previous work showed that the Cry1Ie proteins expressed in Escherichia coli and transgenic tobacco plants were toxic to the corn borer (Liu et al. 2004a) and that transgenic tobacco plants expressing synthetic Cry1Ac and Cry1Ie genes were more toxic to cotton bollworm (Heliothis armigera) than those containing one gene (Lian et al. 2008). It was recently shown that one Cry1Ab-resistant Ostrinia furnacalis strain that was selected by the inclusion of the Cry1Ab protein in an artificial diet for 34 generations was susceptible to the Cry1Ie protein that was purified from E. coli, indicating that there is no cross-resistance to Cry1Ab and Cry1Ie in insects (Xu et al. 2010).
Bt cotton expressing Cry1Ac has been used commercially since 1997. Corn, soybean, peanut and other crops in the mixed-planting system in China have been used as natural refuge crops for Bt cotton (Wu and Guo 2005). For corn, it is preferable to use a Bt gene with a different mode of action than Cry1Ac to delay the development of insects resistant to Bt crops. In this study, we demonstrate that over-expression of the Cry1Ie gene confers transgenic lines of maize plants with a high tolerance to both wild-type and Cry1Ac-resistant insects. This suggests that Cry1Ie might be a good candidate for the development of Bt corn in China.
Materials and methods
Maize transformation
The A. tumefaciens strain EHA105 containing the binary vectors p3301UbiAc and p3301UbiIe (Liu et al. 2004a, b) was used for transformation. The donor plant for immature embryos was the maize inbred line Z31. The ears were sterilized in 70 % ethanol for 2 min, and the kernel crowns of the ear were cut off using a sharp blade. Immature embryos of 1.0–2.0 mm in length were isolated and suspended in liquid infection medium [Murashige & Skoog (MS) basal medium, 68.5 g L−1 sucrose, 36.0 g L−1 glucose, 0.115 g L−1 proline and 100 μM acetosyringone, pH 5.2] and washed twice with this medium. The final wash was transferred to an A. tumefaciens suspension and incubated for 5 min. Following inoculation, the embryos were transferred to solid co-cultivation medium [MS basal medium, 20 g L−1 sucrose, 10 g L−1 glucose, 0.85 mg L−1 silver nitrate, 0.115 g L−1 proline, 1.5 mg L−1 2,4-dichlorophenoxyacetic acid (2,4-D), 100 μM acetosyringone and 8 g L−1 agar, pH 5.8] and incubated in the dark at 23 °C. After 3 days, the embryos were transferred to 28 °C in resting medium [CHU (N6) basal medium, 20 g L−1 sucrose, 10 g L−1 glucose, 0.85 mg L−1 silver nitrate, 0.115 g L−1 proline, 1.5 mg L−1 2,4-D, 250 mg L−1 cefotaxime and 8 g L−1 agar, pH 5.8] for 7 days. The embryos were moved to a selection medium, which was identical to the resting medium with the addition of 1.5 mg L−1 bialaphos, and maintained for 2 weeks in the dark at 28 °C. The strength of bialaphos in the selection medium was then increased to 3 mg L−1 for 2 rounds of 2-week selection. The resistant calli were placed in regeneration medium [MS basal medium, 0.115 g L−1 proline, 50 g L−1 sucrose, 5 mg L−1 6-benzylaminopurine, 250 mg L−1 cefotaxime, 1 mg L−1 bialaphos and 8 g L−1 agar, pH 5.8], and 2–3-cm shoots were transferred to 1/2 MS rooting medium. The calli for regeneration and regenerated plantlets were grown in a growth chamber at 28 °C under fluorescent white light and a 16/8-h light:dark cycle.
PCR and southern blot analysis
Cry1Ie is a novel Cry1 gene that was obtained by the PCR-RFLP method (Song et al. 2003). The PCR primers for Cry1Ie were designed as follows: forward primer, 5′-AACAGCCAGATCAGCACCTT-3′, and reverse primer, 5′-CTGTACACCAGGGCCTTCAC-3′. An 830-bp PCR product was obtained using this pair of primers.
For the Southern blot analysis of the transgenic plants containing Cry1Ie, a 1,403-bp probe for Cry1Ie was amplified using the primers 5′-ACCCACACCTGCTGGACTT-3′ and 5′-TTGCGCGCTATATTTTGTTTT-3′ (Fig. 1). A 953-bp probe for Cry1Ac was amplified using the primers 5′-CATTCAACATCGGCATCAAC-3′ and 5′-GCGCGCTATATTTTGTTTTCT-3′ (Supplemental Fig. 2). Approximately 100 μg genomic DNA from the transgenic plants and the non-transgenic controls was digested with EcoRI, HindIII or KpnI, separated by electrophoresis on 0.8 % agarose gels and transferred to a nylon membrane (Amersham). The membrane was hybridized with the DIG-labeled probes using the DIG High Primer DNA Labeling and Detection Starter Kit III (Roche). The hybridization was performed according to the kit manual.
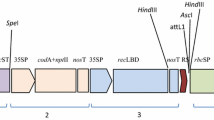
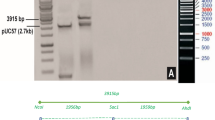
Southern blot analysis of IE034 T4 generation plants. a T-DNA cassette of plasmid p3301UbiIe. LB and RB, left and right borders of the T-DNA; 35S polyA, 35S terminator; Bar, phosphinothricin acetyltransferase gene; 35S, CaMV 35S promoter; Ubiquitin, maize ubiquitin promoter; Cry1Ie, Cry1Ie gene; Nos, Nos terminator. The probe fragment used for the Southern blotting is denoted with a line. b Southern blot analysis. M, Fermentas 1-kb DNA marker; 1, plasmid p3301UbiIe digested with EcoRI; 2, non-transgenic maize Z31 genomic DNA digested with EcoRI; 3–4, individual IE034 genomic DNA digested with EcoRI; 5, non-transgenic maize Z31 genomic DNA digested with HindIII; 6–7, individual IE034 genomic DNA digested with HindIII; 8, plasmid p3301UbiIe digested with HindIII
Insect bioassay
For the bioassay in the field, 50–60 Asian corn borer (O. furnacalis) larvae were placed into the whorl of each plant at the 8–9-leaf-stage, and the infestation was repeated 7 days later. The leaf damage was recorded according to a described method (He et al. 2000).
For the bioassay in the laboratory, the resistance of the transgenic maize plants against the Asian corn borer or cotton bollworm was assayed by exposing fresh leaf-disks from 7–8-leaf-stage plants to 1st-instar larvae. The resistant cotton bollworms were selected using solubilized Cry1Ac protoxin, according to our previously described method (Liang et al. 2008). The Cry1Ac-R1 and Cry1Ac-R2 cotton bollworms exhibited 456.47- and 1,530.93-fold resistance to Cry1Ac, respectively. The bioassays were performed in 24-well Petri dishes: each well was filled with leaf samples and infested with one 1st-instar cotton bollworm larva. The bioassay was performed in an environmental chamber at 70–80 % RH, 26–28 °C and a photoperiod consisting of 14/10-h light:dark cycle. The assays were evaluated every 24 h by counting the number of surviving larvae. The experiment was replicated three times.
Western blot analysis
Approximately 0.1 g sample was ground in liquid nitrogen, and the total proteins were extracted in 1 mL of 1 × SDS-PAGE sample buffer. Approximately 20 μg total protein was resolved by one-dimensional 10 % SDS-PAGE, and the proteins were wet-blotted onto polyvinylidene difluoride membranes in transfer buffer. Cry1Ie was detected with a monoclonal antibody specific for the Cry1Ie protein developed by Abmart Company (Shanghai, China). As a positive control, actin was detected with monoclonal antibodies purchased from Abmart Company. Antibody recognition was visualized using horseradish peroxidase-conjugated secondary antibodies.
Statistical analysis
Comparisons of the values for significant differences were performed using Student’s t-test in Excel (Microsoft) at P < 0.05.
Results
Production and molecular identification of transgenic maize
In our previous work, Cry1Ac and Cry1Ie were artificially synthesized using the bias code from maize and inserted in plasmids p3301UbiAc or p3301UbiIe under the control of the maize ubiquitin promoter (Liu et al. 2004a, b). In the present study, both genes were transformed into the maize inbred line Z31, and the independent transgenic lines were confirmed by Southern blot analysis. The genomic DNA of transgenic or non-transgenic plants was digested with EcoRI, HindIII or KpnI, for which there are no recognition sites in the probe region (Fig. 1, Supplemental Figs. 1, 2). Only one band was detected by Southern blot when genomic DNA was digested with each of these enzymes, indicating that only one copy of Cry1Ie or Cry1Ac was integrated into the maize genome. The results showed that the independent transgenic lines IE034, IE1103 and IE012 contain a single copy of the Cry1Ie gene and that the AC400 transgenic line has one Cry1Ac copy (Fig. 1, Supplemental Figs. 1, 2). These lines were self-crossed to obtain homozygous plants, which were then used for further investigation.
Transgenic plants over-expressing Cry1Ie are resistant to Cry1Ac-resistant cotton bollworm
Transgenic insect-resistant cotton expressing Cry1Ac has been commercially grown for many years in China, and several cotton bollworm strains have developed Cry1Ac resistance during laboratory selection (Liang et al. 2008; Zhang et al. 2009). To investigate whether transgenic maize expressing Cry1Ie could tolerate the Cry1Ac-resistant cotton bollworm, leaf pieces of the transgenic plants grown in the field were fed to Cry1Ac-resistant cotton bollworms with different levels of resistance. Both the Monsanto MON810 plants expressing Cry1Ab and AC400 plants expressing Cry1Ac resulted in a high mortality of the susceptible cotton bollworm. The transgenic lines IE034, IE1103 and IE012, which express Cry1Ie, were also toxic to the susceptible cotton bollworm, though the mortality was lower than that induced by the MON810 and AC400 plants (Fig. 2a).
The mortality of the Cry1Ac-resistant cotton bollworm. Fresh leaf-disks from 7–8-leaf-stage plants were infested with the 1st-instar larvae of the susceptible cotton bollworm (a), Cry1Ac-R1 cotton bollworm that had 456.47-fold resistance to Cry1Ac (b) or Cry1Ac-R2 cotton bollworm that had 1,530.93-fold resistance to Cry1Ac (c). The bioassays were performed in 24-well Petri dishes, and four Petri dishes were used for each experiment. The mortality was recorded 1 day later. The data are presented as the mean ± S.E. of 3 independent experiments
When the maize leaves were infested with Cry1Ac-R1 cotton bollworm larvae with a 456.47-fold resistance level, higher mortality was observed for the transgenic plants expressing Cry1Ie than for the MON810 and AC400 plants (Fig. 2b). However, when Cry1Ac-R2 cotton bollworm larvae with 1,530.93-fold resistance were used for the bioassay, the Cry1Ac-R2 larvae feeding on the MON810 and AC400 plants exhibited the same mortality as those feeding on the non-transgenic plants, indicating that the MON810 and AC400 plants are sensitive to the Cry1Ac-R2 cotton bollworm. However, three transgenic lines expressing Cry1Ie were still able to efficiently kill the Cry1Ac-R2 larvae with 1,530.93-fold resistance (Fig. 2c). Although some larvae survived on the leaf samples from transgenic plants expressing Cry1Ie, the growth of these larvae was strongly inhibited, and they exhibited a smaller size than the larvae that survived on the Z31 or MON810 leaves (Fig. 3). Significantly lower larval weights were observed when the MON810 and IE034 samples were fed to the Cry1Ac-R1 larvae than when non-transgenic plants were used. When the IE034 samples were fed to the Cry1Ac-R2 larvae, significantly lower larval weights were observed compared to the larvae that were fed the control and MON810 leaves (Fig. 4). A high Cry1Ie content was observed in the IE034 and IE1103 transgenic lines, whereas a relatively lower Cry1Ie content was observed in the IE012 line (Fig. 5a), which was correlated with the mortality of the Cry1Ac-R2 cotton bollworm larvae (Fig. 2).
The surviving Cry1Ac-R2 cotton bollworm with 1,530.93-fold resistance to Cry1Ac. Fresh leaf-disks from 7–8-leaf-stage plants were infested with the 1st-instar larvae of the Cry1Ac-R2 cotton bollworm with 1,530.93-fold resistance to Cry1Ac. The bioassays were performed in 24-well Petri dishes. The pictures represent the surviving cotton bollworms that had been fed leaf samples for 6 days
The weight of each surviving Cry1Ac-resistant cotton bollworm. Fresh leaf-disks from 7–8-leaf-stage plants were infested with the 1st-instar larvae of the Cry1Ac-R1 and Cry1Ac-R2 cotton bollworms that had 456.47-fold and 1,530.93-fold resistance to Cry1Ac, respectively. The bioassays were performed in 24-well Petri dishes, and four Petri dishes were used for each experiment. The weight of the surviving Cry1Ac-resistant cotton bollworms was recorded on the sixth day. The data are presented as the mean weight of each cotton bollworm ± the S.E. of 3 independent experiments. The asterisks indicate a significant difference at the P < 0.05 level
The expression level of Cry1Ie protein. a The expression level of Cry1Ie in the leaves of different transgenic lines. b The expression of Cry1Ie in different tissues of IE034 plants. M, Molecular weight marker; 1, root; 2, stem; 3, leaf; 4, husk leaf; 5, silk; 6, pollen; 7, tassel; 8, seed; 9, leaf of a non-transgenic plant
Transgenic maize plants expressing Cry1Ie are highly resistant to the Asian corn borer
IE034 was chosen to further analyze whether the overexpression of Cry1Ie could provide transgenic maize plants with resistance to the susceptible Asian corn borer. The presence of Cry1Ie was confirmed by PCR analysis (Supplemental Fig. 3), and samples from T3 and T4 generation plants were infested with larvae in the laboratory. The results showed that transgenic plants caused larval mortality rates of 60.83 and 66.66 % at 3 days after infestation, with the rates increasing to 85.42 and 90.62 % 6 days later (Table 1). The Z31 non-transgenic plants resulted in significantly lower larval mortality. The insect bioassays were performed with three generations of transgenic plants in the field over three consecutive years. The non-transgenic plants presented more severe leaf damage after larval infestation, whereas the transgenic maize was almost unharmed (Supplemental Fig. 4). The IE034 transgenic maize plants were highly resistant to the corn borer, with a leaf damage rating below three, whereas the non-transgenic plants were highly sensitive to the corn borer (Table 2). We further investigated the expression level of Cry1Ie in the different tissues of the IE034 plants, and the results showed a higher accumulation of the Cry1Ie protein in the leaf and leaf husk; relatively low expression levels were observed in the root, silk, pollen and tassel. In contrast, no Cry1Ie protein was detected in the stem or seed (Fig. 5b).
Discussion
The development of insect resistance to Bt plants is a major issue that threatens the use of these plants. Therefore, the identification of new Bt proteins that have different modes of action is important for the sustainable use of Bt crops. To date, only a few Bt proteins, i.e., Cry2Ab, Cry2Aa and Cry1C, have been demonstrated to have different modes of action from Cry1Ac/Cry1Ab, and these proteins can be pyramided with Cry1Ac/Cry1Ab to delay the development of resistance (Kota et al. 1999; Zhao et al. 2003; Tabashnik et al. 2009). Here, we showed that a novel Bt protein, Cry1Ie, may be a candidate for pyramiding with Cry1Ac.
Independent transgenic maize lines containing one copy of Cry1Ac or Cry1Ie were identified using Southern blot analysis (Fig. 1, Supplemental Figs. 1, 2). Susceptible and Cry1Ac-resistant cotton bollworms with different resistance levels were fed leaf samples from the transgenic maize, and the AC400 plants expressing Cry1Ac and MON810 plants expressing Cry1Ab caused higher mortality in the susceptible cotton bollworm than the transgenic plants expressing Cry1Ie. It has been reported that Cry1Ie protein purified from E. coli was not toxic to the cotton bollworm or the beet armyworm (Song et al. 2003). Another Cry1I-type protein, Cry1Ia7, has also been shown to be less active than Cry1A (Ruiz de Escudero et al. 2006). Conversely, transgenic maize plants expressing Cry1Ie are highly toxic to the cotton bollworm. Thus, it is possible that the purified Cry1Ie fusion protein did not represent the active form. We also noticed that our AC400 plants caused relatively lower mortality than MON810 plants. The AC400 line used for the bioassay was selected from approximately 20 lines; however, MON810 was obtained from more independent lines. Therefore, by testing additional independent lines, we might be able to identify the best line expressing Cry1Ac.
The transgenic plants expressing Cry1Ie caused higher mortality to the Cry1Ac-R1 cotton bollworm with 456.47-fold resistance than the MON810 and AC400 plants. Similar to the non-transgenic plants, the MON810 and AC400 plants could not kill the 1,530.93-fold resistant Cry1Ac-R2 cotton bollworm. However, transgenic maize expressing Cry1Ie was still highly toxic to the Cry1Ac-R2 cotton bollworm (Fig. 2), and the mortality was related to the level of Cry1Ie protein (Fig. 5). We noticed that both the non-transgenic Z31 plants and IE034 plants resulted in higher mortality in resistant larvae than in susceptible larvae, which might be due to the decreased fitness of resistant larvae. Indeed, it has been demonstrated that the fitness of Helicoverpa armigera decreased with an increase in the resistance level (Liang et al. 2008).
The development of insect resistance to Bt toxin might be due to a mutation in the receptor that disrupts the binding between the Bt toxin and its receptor. Many reports have shown that mutations in cadherin, aminopeptidase and alkaline phosphatase are related to Cry1A resistance in Heliothis virescens, Pectinophora gossypiella and H. armigera (Jurat-Fuentes et al. 2011; Tabashnik et al. 2011). In this study, the transgenic maize plants expressing Cry1Ie were toxic toward the Cry1Ac-resistant cotton bollworm, confirming that Cry1Ie does not compete for the Cry1Ac binding site in the cotton bollworm. It has also been demonstrated that the Cry1Ia7 protein does not compete for the Cry1Ab or Cry1Ac binding site (Ruiz de Escudero et al. 2006). Furthermore, it was recently reported that there is no cross-resistance of insects to the Cry1Ab and Cry1Ie proteins (Xu et al. 2010). Although the successful management of Cry1Ac-resistant cotton bollworms with the Cry1Ie protein in the laboratory is a promising result, long-term tests of transgenic maize expressing Cry1Ie are needed. Regardless, Cry1Ie can be proposed as an appropriate candidate for expression with Cry1Ac or Cry1Ab in second-generation Bt crops.
The Cry1Ie fusion protein purified from E. coli showed no toxicity toward the cotton bollworm but was highly active against the Asian corn borer (Song et al. 2003). Several other Cry1I-type proteins have been identified to have a broad host range (Tailor et al. 1992; Choi et al. 2000; Bergamasco et al. 2011). The transgenic maize plants expressing Cry1Ie were highly resistant to the Asian corn borer (Tables 1, 2). The highest levels of Cry1Ie were detected in the leaf compared to the other organs, and the expression level was approximately 0.41 % of the total soluble protein in the leaf according to an ELISA analysis (data not shown). This result is similar to the reported Cry1Ab level in transgenic maize plants in which the expression of the Cry1Ab gene is driven by the CaMV 35S promoter (Koziel et al. 1993). The high level of resistance of the transgenic plants might be due to the high expression level of Cry1Ie. High doses of Bt protein are necessary for insect control and are also the primary basis for the “refuge” strategy to delay insect resistance (Tabashnik et al. 2008). As the transgenic maize expressing Cry1Ie was associated with reduced insect resistance, our results indicate that it can be commercially grown in China to manage insect damage.
In China, Bt cotton expressing Cry1Ac has been widely grown since 1997, and, currently, approximately 80 % of the cotton grown in the field is transgenic (James 2013). No non-Bt cotton refuges are required in China because the non-transgenic crops, i.e., corn, soybean and others, provide a sufficient natural refuge to delay the development of resistant insects (Wu and Guo 2005). However, scientists in China are concerned about the development of resistance to Cry1Ac in the field (Zhang et al. 2011). For future commercialized Bt maize in China, it would be preferable to use another Bt gene with a mode of action that is different from Cry1Ac to delay the development of pest resistance. This study clearly showed that Cry1Ie might be a good candidate for the development of Bt maize in China.
References
Bates SL, Zhao JZ, Roush RT, Shelton AM (2005) Insect resistance management in GM crops: past, present and future. Nat Biotechnol 23(1):57–62
Bergamasco V, Goncalves J, Polanczyk R, Desiderio J, Lemon M (2011) Expression of a new Bacillus thuringiensis cry1Ia gene in Escherichia coli with strong activity against cotton pests. Aust J Bas Appl Sci 5(12):526–533
Bravo A, Soberon M (2008) How to cope with insect resistance to Bt toxins? Trends Biotechnol 26(10):573–579
Chitkowski RL, Turnipseed SG, Sullivan MJ, Bridges WC Jr (2003) Field and laboratory evaluations of transgenic cottons expressing one or two Bacillus thuringiensis var. kurstaki Berliner proteins for management of noctuid (Lepidoptera) pests. J Econ Entomol 96(3):755–762
Choi SK, Shin BS, Kong EM, Rho HM, Park SH (2000) Cloning of a new Bacillus thuringiensis cry1I-type crystal protein gene. Curr Microbiol 41(1):65–69
Ferre J, Van Rie J (2002) Biochemistry and genetics of insect resistance to Bacillus thuringiensis. Annu Rev Entomol 47:501–533
Gatehouse JA (2008) Biotechnological prospects for engineering insect-resistant plants. Plant Physiol 146(3):881–887
He K, Wang Z, Zhou D, Wen L, Song Y (2000) Methodologies and criterions for evaluating maize resistance to Asian maize borer. J Shenyang Agric Univ 31:439–443
James C (2013) Global status of commercialized biotech/GM crops: 2012. ISAAA Brief 44-2012
Jurat-Fuentes JL, Karumbaiah L, Jakka SR, Ning C, Liu C, Wu K, Jackson J, Gould F, Blanco C, Portilla M, Perera O, Adang M (2011) Reduced levels of membrane-bound alkaline phosphatase are common to lepidopteran strains resistant to Cry toxins from Bacillus thuringiensis. PLoS One 6(3):e17606
Kota M, Daniell H, Varma S, Garczynski SF, Gould F, Moar WJ (1999) Overexpression of the Bacillus thuringiensis (Bt) Cry2Aa2 protein in chloroplasts confers resistance to plants against susceptible and Bt-resistant insects. Proc Natl Acad Sci USA 96(5):1840–1845
Koziel MG, Beland GL, Bowman C, Carozzi NB, Crenshaw R, Crossland L, Dawson J, Desai N, Hill M, Kadwell S, Launis K, Lewis K, Maddox D, McPherson K, Meghji MR, Merlin E, Rhodes R, Warren GW, Wright M, Evola SV (1993) Field performance of elite transgenic maize plants expressing an insecticidal protein derived from Bacillus thuringiensis. Nat Biotechnol 11(2):194–200
Lian Y, Jia Z, He K, Liu Y, Song F, Wang B, Wang G (2008) Transgenic tobacco plants expressing synthetic Cry1Ac and Cry1Ie genes are more toxic to cotton bollworm than those containing one gene. Chin Sci Bull 53:1381–1387
Liang GM, Wu KM, Yu HK, Li KK, Feng X, Guo YY (2008) Changes of inheritance mode and fitness in Helicoverpa armigera (Hubner) (Lepidoptera: Noctuidae) along with its resistance evolution to Cry1Ac toxin. J Invertebr Pathol 97(2):142–149
Liu YJ, Song FP, He KL, Yuan Y, Zhang XX, Gao P, Wang JH, Wang GY (2004a) Expression of a modified Cry1Ie gene in E. coli and in transgenic tobacco confers resistance to corn borer. Acta Biochim Biophys Sin (Shanghai) 36(4):309–313
Liu YJ, Yuan Y, Zheng J, Tao YZ, Dong ZG, Wang JH, Wang GY (2004b) Signal peptide of potato PinII enhances the expression of Cry1Ac in transgenic tobacco. Acta Biochim Biophys Sin (Shanghai) 36(8):553–558
Ruiz de Escudero I, Estela A, Porcar M, Martinez C, Oguiza JA, Escriche B, Ferre J, Caballero P (2006) Molecular and insecticidal characterization of a Cry1I protein toxic to insects of the families Noctuidae, Tortricidae, Plutellidae, and Chrysomelidae. Appl Environ Microbiol 72(7):4796–4804
Soberon M, Pardo-Lopez L, Lopez I, Gomez I, Tabashnik BE, Bravo A (2007) Engineering modified Bt toxins to counter insect resistance. Science 318(5856):1640–1642
Song F, Zhang J, Gu A, Wu Y, Han L, He K, Chen Z, Yao J, Hu Y, Li G, Huang D (2003) Identification of cry1I-type genes from Bacillus thuringiensis strains and characterization of a novel cry1I-type gene. Appl Environ Microbiol 69(9):5207–5211
Tabashnik BE, Gassmann AJ, Crowder DW, Carriere Y (2008) Insect resistance to Bt crops: evidence versus theory. Nat Biotechnol 26(2):199–202
Tabashnik BE, Unnithan GC, Masson L, Crowder DW, Li X, Carriere Y (2009) Asymmetrical cross-resistance between Bacillus thuringiensis toxins Cry1Ac and Cry2Ab in pink bollworm. Proc Natl Acad Sci USA 106(29):11889–11894
Tabashnik BE, Huang F, Ghimire MN, Leonard BR, Siegfried BD, Rangasamy M, Yang Y, Wu Y, Gahan LJ, Heckel DG, Bravo A, Soberon M (2011) Efficacy of genetically modified Bt toxins against insects with different genetic mechanisms of resistance. Nat Biotechnol 29(12):1128–1131
Tailor R, Tippett J, Gibb G, Pells S, Pike D, Jordan L, Ely S (1992) Identification and characterization of a novel Bacillus thuringiensis delta-endotoxin entomocidal to coleopteran and lepidopteran larvae. Mol Microbiol 6(9):1211–1217
Tohidfar M, Zare N, Jouzani G, Eftekhari S (2013) Agrobacterium-mediated transformation of alfalfa (Medicago sativa) using a synthetic cry3a gene to enhance resistance against alfalfa weevil. Plant Cell Tissue Organ Cult 113:227–235
Wu KM, Guo YY (2005) The evolution of cotton pest management practices in China. Annu Rev Entomol 50:31–52
Xu L, Wang Z, Zhang J, He K, Ferry N, Gatehouse A (2010) Cross-resistance of Cry1Ab-selected Asian corn borer to other Cry toxins. J Appl Entomol 134:429–438
Zhang X, Candas M, Griko NB, Taussig R, Bulla LA Jr (2006) A mechanism of cell death involving an adenylyl cyclase/PKA signaling pathway is induced by the Cry1Ab toxin of Bacillus thuringiensis. Proc Natl Acad Sci USA 103(26):9897–9902
Zhang S, Cheng H, Gao Y, Wang G, Liang G, Wu K (2009) Mutation of an aminopeptidase N gene is associated with Helicoverpa armigera resistance to Bacillus thuringiensis Cry1Ac toxin. Insect Biochem Mol Biol 39(7):421–429
Zhang H, Yin W, Zhao J, Jin L, Yang Y, Wu S, Tabashnik BE, Wu Y (2011) Early warning of cotton bollworm resistance associated with intensive planting of Bt cotton in China. PLoS One 6(8):e22874
Zhao JZ, Cao J, Li Y, Collins HL, Roush RT, Earle ED, Shelton AM (2003) Transgenic plants expressing two Bacillus thuringiensis toxins delay insect resistance evolution. Nat Biotechnol 21(12):1493–1497
Acknowledgments
This work is financially supported by the National Major Project for Transgenic Organism Breeding (2011ZX08003-001). We thank the editors from American Journal Experts for improving the English language in the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Yuwen Zhang and Yunjun Liu have contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhang, Y., Liu, Y., Ren, Y. et al. Overexpression of a novel Cry1Ie gene confers resistance to Cry1Ac-resistant cotton bollworm in transgenic lines of maize. Plant Cell Tiss Organ Cult 115, 151–158 (2013). https://doi.org/10.1007/s11240-013-0348-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-013-0348-5