Abstract
Henneguya tunisiensis n. sp., a new myxosporean, is described from the gill-arches of the East Atlantic peacock wrasse Symphodus tinca (L.) collected from off the Kerkennah Islands, Tunisia. It is characterised by the presence of elongate white plasmodia of 1–1.5 × 1.5–2 mm in size. The mature spores are rounded in frontal view and have two identical polar capsules and two caudal appendages which taper considerably at the end. Both light and electron microscopical data show that this species differs in several morphological features from all previously described Henneguya spp. A molecular analysis, based on 18S rDNA sequence data, indicates that H. tunisiensis n. sp. is readily distinguishable from other myxozoan DNA sequences in GenBank. Phylogenetically, the new species is placed in the marine Henneguya clade, which is a sister group of marine Myxobolus spp. from perciform fishes in Tunisian waters.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
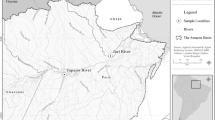
The East Atlantic Peacock wrasse Symphodus [Crenilabrus] tinca (L.) is the most common fish of the family Labridae in Tunisian waters (Bradai, 2000). The spawning period of this species is from April to June (Ouannes-Ghorbel et al., 2002). Off the Kerkennah Islands (34°37′57″N, 11°02′44″E), S. tinca is mainly caught during spring by nets or fixed fisheries called ‘Charfia’ or ‘Chrafi’, and is much appreciated as a food-fish.
During a survey of the parasites of S. tinca caught off the Kerkennah Islands between 2007 and 2008, large white plasmodia were observed in the gill-arches. On dissection, they were found to be plasmodia of a myxosporean species belonging to Henneguya Thélohan, 1892. This parasite has previously been reported by Bahri & Marques (2008) from the same host and locality, with description of its pathology.
In this paper, we describe Henneguya tunisiensis n. sp., a new myxozoan parasite infecting the gill-arches of Symphodus tinca off the Kerkennah Islands, Tunisia. The spore morphology was examined using light (LM) and scanning electron microscopy (SEM), and the cyst location was studied in histological sections. The species description is supplemented by its molecular characterisation, with a phylogenetic analysis based on 18S rDNA sequences.
Materials and methods
Collection of the host fish
Between 2007 and 2008, 250 specimens of Symphodus tinca ranging from 40–100 g in weight were caught at two locations of the Kerkennah Islands (Location 1: Attaya; Location 2: Kraten) using fixed fisheries (‘Chrafi’), which is a traditional fishing system built with lines of palm sheets and composed of a ‘foot’, a ‘large house’, a ‘small house’, a ‘lamp’, two bow nets and a ‘room of capture’. The system is supposed to guide fish towards enclosures matted and encircled by the bow nets, genuine traps with fish, which the fishermen raise from their boats. The fish were transported to the laboratory alive and examined for the presence of myxozoans.
Morphological methods
The cysts located on the gill-arches were removed from the infected fish. Fresh isolated spores obtained directly from the mature plasmodia were photographed with a Nikon E600 microscope using DIC optics. Measurements were taken of 30 spores according to the criteria described by Lom & Arthur (1989). Some of the spores were collected in Eppendorf tubes and stored at −20°C for subsequent molecular examination.
For histological examination, fragments of gill containing young and mature plasmodia were fixed in 4% neutral buffered formalin for 24 h and embedded in paraffin. The 5 μm thin sections were stained with H & E and examined under a microscope.
For SEM, free spores were placed on a slide coated with poly-L-lysine then fixed in 2.0% glutaraldehyde and washed with 1 M sodium-cacodylate buffer before dehydration in a graded ethanol series. After critical point drying, the samples were coated with gold and examined with a JEOL JSM scanning electron microscope.
Molecular methods
Samples were thawed and then homogenised in 1.5 ml microtubes with a sterile plastic Eppendorf pestle. Then microtubes containing the homogenates were filled with dH2O, mixed by vortexing and centrifuged at 10,000 rpm for 5 min. Pellets were dissolved in 500 μl lysis buffer (100 mM NaCl, 10 mM Tris, 10 mM EDTA, 0.2% SDS and 0.4 mg ml−1 Proteinase K) and incubated at 55°C for 5 h. DNA was then purified using the MiniPrep Express Matrix (BIO101, Qbiogene) as per Eszterbauer (2004). Genomic DNA was amplified with the primer pair 18e (5′-CTG GTT GAT TCT GCC AGT-3′) (Hillis & Dixon, 1991) and 18r (5′-CTA CGG AAA CCT TGT TAC-3′) (Whipps et al., 2003). The total volume of the PCR reactions was 50 μl, which contained approx. 10–50 ng DNA, 1× Taq PCR reaction buffer (MBI Fermentas), 1.5 mM MgCl2, 0.2 mM dNTP mix (Sigma), 25 μM of each primer and 2 units of Taq DNA Polymerase (MBI Fermentas). Amplification conditions were: 95°C for 50 s, 58°C for 50 s and 72°C for 80 s for 35 cycles, with a terminal extension at 72°C for 7 min. PCR products were purified with QIAquick Gel Extraction Kit (Qiagen).
The purified PCR product was cloned with the CloneJET PCR Cloning Kit (Fermentas) following the manufacturer’s manual. Four positive clones were sequenced with sequencing primers supplied with the cloning kit, using the ABI BigDye Terminator v3.1 Cycle Sequencing Kit with an ABI 3100 Genetic Analyzer automated DNA sequencer (Applied Biosystems). Furthermore, the purified PCR products of two Henneguya samples were sequenced directly in both strands. The following primers were used for direct sequencing: amplification primers 18e and 18r; Myx4r and Act1f as in Hallett & Diamant (2001); MB5 described by Eszterbauer (2004); SphR by Eszterbauer & Székely (2004) and the reverse complements of Act1f and Myx4r. For sequence assembling, the STADEN Sequence Analysis Package version 2001.0 (Staden, 1996) was used. DNA sequence similarities were calculated with the Sequence Identity Matrix of the software BioEdit (Hall, 1999).
Phylogenetic analysis
18S rRNA gene sequences of several myxozoan species (both actinospores and myxospores) were selected from GenBank for phylogenetic analysis. They consist of: Henneguya pagri (AB183748), H. lateolabracis (AB183747), H. sp. (DQ377706), H. akule (EU016076), H. ictaluri (AF195510), H. doori (U37549), H. sp. (U13826), H. weishanensis (AY165182), H. sp. (AB447996), H. sp. (AB447995), H. sp. (AB447994), Tetraspora discoidea (AF306793), Myxobolus exiguus (AY129317), M. bizerti (AY129318), M. spinacurvatura (AF378341), M. ichkeulensis (AY129315), M. episquamalis (AY129312), M. osburni (AF378338), M. neurophilus (FJ468489), Sphaeractinomyxon ersei (AF306790), Endocapsa (DQ473516), Triactinomyxon (DQ473515), Aurantiactinomyxon mississippiensis (AF021878), Aurantiactinomyxon (AF378356), and Ceratomyxa shasta (AF001579) (the outgroup).
Nucleotide sequences were aligned with the software Multalin (Corpet, 1988) available online. The alignment was adjusted manually using the GeneDoc sequence alignment editor program. The dataset for the alignment was chosen on the basis of the results of BLAST searches and morphological findings. Phylogenetic analysis using the maximum likelihood algorithm was conducted in PAUP* Version 4.0b10 (Swofford, 2001). An optimal evolutionary model (GTR+I+G) for the alignment was determined with AIC in Modeltest 3.06 (Posada & Crandall, 1998). Maximum likelihood (ML) analysis employed a heuristic search algorithm with random sequence addition (10 replicates) and TBR branch swapping. Bootstrap confidence values were calculated with 100 repetitions. Bayesian Inference analysis was performed using MrBayes v3.1.2 (Ronquist & Huelsenbeck, 2003). A general time reversible model (GTR) with gamma-shaped rate variations across sites (Invgamma) was chosen for the analysis. Two independent runs were conducted with four chains for 1 million generations. Trees were sampled every 100 generations. The first 25% of the samples were discarded from the cold chain (burninfrac = 0.25), and a 50% majority-rule consensus tree was created, which was visualised by TreeView.
Henneguya tunisiensis n. sp.
Type-host: East Atlantic peacock wrasse Symphodus (Crenilabrus) tinca (L.) (Perciformes: Labridae).
Localities: Location 1: Off Attaya (34°44′34″N, 11°17′31″) (type-locality); Location 2: Off Kraten (34°49′39″N, 11°15′17″E), the Kerkennah Islands, Tunisia
Site: Gill-arches.
Prevalence of infection: 33 of 250 (13.2%), i.e. 11 of 150 (7.33%) at Location 1, and 22 of 100 (22%) at Location 2.
Type-material: The syntype spores and histological sections are deposited in the parasitological collection of the National Museum of Natural History, Paris, Coll. No. ZS53.
The 18S rDNA sequence was deposited in GenBank under the accession number GQ340975.
Description (Figs. 1–9)
Vegetative stages
Plasmodia opaque white, ellipsoidal or spherical, 1.0–1.5 × 1.5–2.0 mm, located in gill-arches (Fig. 1). Development asynchronous. Plasmodia in advanced stages contain mature and immature spores (Fig. 2).
Spores
Spores round-oval in frontal view (Figs. 3, 5a), ellipsoidal in sutural view (Figs. 4, 5b). Valves thin, symmetrical, smooth (Fig. 6). Sutural rim around spore present 4–6 distinct marginal markings (Fig. 7). Mature spores have total length of 41.8 ± 3.6 (38–50) μm; length of spore body 13.1 ± 0.5 (13–14) μm, width 9.1 ± 0.2 (9–10) μm and thickness 8 ± 0.1 (7.5–8.5) μm. Two polar capsules pyriform, equal in size, 4 ± 0.2 (3.5–4.0) μm long, 2 ± 0.1 (1.8–2.0) μm wide. Polar filaments coiled, with 4–5 turns, situated perpendicularly to longitudinal axis of polar capsule. Caudal processes tapered and flexible distally, with total length (including flexible portion) of 28.4 (25–32) μm.
Henneguya tunisiensis n. sp. 6–7. Scanning electron micrographs: 6. Frontal view of spore; note the flexible the distal region of the caudal processes (arrows). 7. Spore body showing the pore (white arrow) for the discharge of the polar filament situated at the anterior extremity of the spore and sutural markings over surface of spore (arrows). 8–9. Histological sections showing the localisation of the plasmodia: 8. Young plasmodium (P) situated under the mucosal epithelium (E) of the gill-arch, H&E. 9. Mature plasmodium (P) compressing the neighbouring tissue of the gill-arch, H&E. Abbreviations: ca, cartilage; mu, muscle Scale-bars: 6, 7.5 μm; 7, 1.5 μm; 8, 50 μm; 9, 100 μm
SEM
Spores with smooth valves (Figs. 6, 7); valves with prominent sutural line and internal sutural markings (Fig. 7). Two caudal processes appear to unite close to caudal end of spore body; they are filamentous and bent distally (Fig. 6).
Histology
Large, oval plasmodia are located in the connective tissue of the gill-arches (1–2 plasmodia per arch). Young plasmodia are situated under the mucosal epithelium of the gill-arch (Fig. 8). When plasmodia grow, they compress the neighbouring tissues, inducing atrophy of the distal region of the infected area (Fig. 9).
Molecular results
The primer pair 18e-18r successfully amplified a 2,067 nt long fragment of the 18S rDNA. The PCR products originating from the amplifications of two cysts collected separately were sequenced from both localities, yielding 100% identical sequences to the cloned PCR fragments. BLASTn search revealed that H. tunisiensis n. sp. has the greatest genetic similarity (91.5%) to H. pagri Yokoyama, Ito & Tanaka, 2005 (AB183748) described by Yokoyama et al. (2005) from the bulbus arteriosus of Red Sea bream Pagrus major in Japanese waters.
The topology of the two consensus trees generated by Bayesian inference and maximum likelihood analyses was almost identical. Slight differences were observable in the confidence values, but the main clusters and, most of all, the position of H. tunisiensis was the same in both consensus trees. H. tunisiensis grouped with the marine Henneguya spp. (H. pagri, H. akule Work, Tanaka, Whipps & Kent, 2008, H. lateolabracis Yokoyama, Kawakami, Yasuda & Tanaka, 2003) infecting the bulbus arteriosus of the heart, and they formed a sister group with the freshwater Henneguya spp. (H. doori Guilford, 1963 and H. ictaluri Pote, Hanson & Shivaji, 2000) and aurantiactinomyxon type actinospores (Fig. 10).
Remarks
The shape and dimensions of the spores of Henneguya tunisiensis n. sp. were compared with the Henneguya spp. reported by Lom & Dyková (1992, 2006) and Eiras (2002). Of these, none is identical to the present parasite. Furthermore, we compared the spore measurements of H. symphodae Lubat, Radujkovic, Marques & Bouix, 1989, Henneguya sp. 1 and Henneguya sp. 2, all described from fishes of the family Labridae on the Montenegran coast by Lubat et al. (1989), with our samples, and it appeared that H. symphodae from the gall-bladder of Symphodus tinca, S. cinereus, S. rostratus and S. mediterraneus has a smaller spore body (9.5–11 × 6–7.5 μm) and shorter polar capsules (3 μm in length). However, Henneguya sp. 1 from the ovary of S. ocellatus differs from our material in the total length of the spore, which is much shorter (20–23 μm) and possesses a smaller spore body (6–8 × 5.5–7 μm). Nevertheless, Henneguya sp. 2 from the gills of S. rostratus is of a similar size (40–48 μm), but differs in the length (10.5–11.75 μm) and the width (7.5–8 μm) of the spore body, which is smaller. Hence, these Henneguya spp., previously described from labrid fishes in a different geographical area, are morphologically different from the present species. Unfortunately, no DNA sequences are available in GenBank for a genetic comparison.
During the course of our survey, we found no Henneguya cysts in the heart of S. tinca. Comparing the morphology of H. tunisiensis with those of its phylogenetic sister species from the heart of fish, it appears that H. pagri, H. lateolabracis and H. akule have a similar spore shape to our material. Moreover, all of them have long caudal appendages, which taper away distally. H. pagri, from Pagrus major off Japan (Yokoyama et al., 2005), is similar in spore shape to the present species but possesses a smaller spore body (10.5 × 7.5 μm). However, spores of H. lateolabracis from Lateolabrax sp. off Japan (Yokoyama et al., 2003) were shorter in length than those of H. tunisiensis. Nevertheless, H. akule from the big-eyed scad Selar crumenophthalmus in Hawaiian waters (Work et al., 2008) have a relatively similar size to our material but with smaller polar capsules (3.4 × 1.4 μm). On the basis of a 2,062 bp long alignment of 18S rDNA sequences, the genetic similarity of H. tunisiensis was 91.5% with H. pagri, 85.7% with H. lateolabracis and 85.9% with H. akule. Despite the slight morphological differences, 18S rDNA data indicated distinct difference between H. tunisiensis and the abovementioned species. Therefore, the parasite observed in Symphodus tinca is considered to be a new species of Henneguya, for which we propose the name H. tunisiensis n. sp.
Discussion
Henneguya spp. are rarely found in marine fishes of the Mediterranean Sea. Hitherto, only Henneguya sp. infecting the gill-lamellae or the heart of the gilthead sea bream Sparus aurata off the Tunisian coast has been reported (Bahri et al., 1996), and the same species has been observed by Caffara et al. (2003) in an Italian fish farm.
The phylogenetic relationships between myxosporean taxa are studied mainly based on 18S rDNA sequences. Kent et al. (2001) placed freshwater and marine Henneguya spp. within the same clade, suggesting a single marine invasion from freshwater. Further analyses with additional taxa supported this (Fiala, 2006). Molecular data have also revealed that species of Henneguya probably arose from ancestral Myxobolus Bütschli, 1882 several times during myxosporean evolution, and both marine and freshwater species of these genera form a polyphyletic clade (Kent et al., 2001; Eszterbauer et al., 2005; Fiala, 2006). Moreover, previous phylogenetic studies have suggested that the caudal appendages might not represent a valid character for distinguishing Myxobolus and Henneguya (see Kent et al., 2001; Bahri et al., 2003; Fiala, 2006).
The importance of site selection in myxosporean evolution has been revealed in numerous phylogenetic studies (Andree et al., 1999; Eszterbauer, 2004; Holzer et al., 2004; Whipps et al., 2004; Fiala, 2006; Work et al., 2008). According to the results of the 18S rDNA analysis in this study, H. tunisiensis n. sp. was genetically most similar to H. pagri, H. akule and H. lateolabracis, which parasitise the bulbus arterious of marine fishes (Fig. 10). Furthermore, H. tunisiensis, which develops plasmodia in the gill-arches clustered with the heart-infecting species of Henneguya in the phylogenetic trees. This phylogenetic position of H. tunisiensis is rather interesting, because these species differ in host and organ specificity. However, histological examination demonstrated that the plasmodia of all four species develop in connective tissue. Furthermore, the similar spore morphology of the heart-infecting Henneguya spp. and H. tunisiensis is also remarkable. Despite slight differences in measurements, the spore-shape of these species is very similar. The relatively long tails, which taper towards the distal end of the tail, and the flexible nature of the distal third of the tail both indicate great morphological similarity.
The majority of Henneguya spp. infect the gills of freshwater and marine fish. In his synopsis of Henneguya, Eiras (2002) listed 146 species, 50% of which infect the gills. The pathology of Henneguya spp. is often related to the site of infection in the gill (lamellae, filaments or arch). With regard to gill-arch infections, data from the literature suggest that different types of development occur in the epithelium, blood vessels or connective tissue (Sakiti et al., 1991; Molnár, 2002). In the case of H. tunisiensis, the parasite develops plasmodia in the connective tissue elements of the gill-arch, under the mucosal epithelium. Large plasmodia are usually situated at the ends of the gill-arch and induce the compression of the capillaries and retraction of the neighbouring tissue. In a similar histopathological study of the gills of Astyanax altiparanae parasitised by H. chydadea Barassa, Cordeiro & Arana, 2003, which develops an intralamellar-type of plasmodia, Barassa et al. (2003) also reported compression of the capillaries and retraction of the neighbouring lamellae, with a resulting reduction in the surface available for gas-exchange. In another work, Molnár (1998) studied the gills of Stizostedion lucioperca parasitised by H. creplini (Gurley, 1894). The plasmodia located in gill lamellae caused epithelial hyperplasia, and the formation of a thick layer of granular tissue was also reported.
Although we have not found H. tunisiensis in the heart of Symphodus tinca, the close genetic relationship with its sister species and the same tissue tropism would suggest that this parasite may occasionally appear in the heart. The pathological effects of the heart-infecting species of Henneguya are more severe in cultured fish. In fact, H. pagri and H. lateolabracis have induced internal haemorrhage in the pericardial cavity and sometimes degenerative cardiopathy (Yokoyama et al., 2003, 2005). However, in the case of H. akule in wild Selar crumenophthalmus off Hawaii, Work et al. (2008) reported only a host immune response and did not observe any gross lesions.
Although massive infections by H. tunisiensis were not detected in Symphodus tinca during our survey, the histopathological changes indicated that this parasite is potentially pathogenic and that a high parasite load may compromise heart and gill functions.
References
Andree, K. B., Székely, Cs., Molnár, K., Gresoviac, S. J., & Hedrick, R. P. (1999). Relationships among members of the genus Myxobolus (Myxozoa; Bivalvulida) based on small subunit ribosomal DNA sequences. Journal of Parasitology, 85, 68–74.
Bahri, S., Andree, K. B., & Hedrick, R. P. (2003). Morphological and phylogenetic studies of marine Myxobolus spp. from mullet in Ichkeul Lake, Tunisia. Journal of Eukaryotic Microbiology, 50, 463–470.
Bahri, S., Ben Hassine, O. K., & Marques, A. (1996). Henneguya sp. (Myxosporea, Bivalvulida) infecting the gills of wild gilthead sea bream Sparus aurata L., from the coast of Tunisia. Bulletin of the European Association of Fish Pathologists, 16, 51–53.
Bahri, S., & Marques, A. (2008). Gill infection of Symphodus tinca by Henneguya sp. (Myxozoa, Myxobolidae) in Kerkennah Islands, Tunisia. Bulletin of the European Association of Fish Pathologists, 28, 42–45.
Barassa, B., Cordeiro, N. S., & Arana, S. (2003). A new species of Henneguya, a gill parasite of Astyanax altiparanae (Pisces: Characidae) from Brazil with comments on histopathology and seasonality. Memórias do Instituto Oswaldo Cruz, 98, 761–765.
Bradai, M. N. (2000). Diversité du peuplement ichtyque et contribution à la connaissance des sparidés du golfe de Gabès. Thèse Doctorat d’Etat, Faculté des Sciences de Sfax, 600 pp.
Caffara, M., Marcer, F., Florio, D., Quaglio, F., & Fioravanti, M. L. (2003). Heart infection due to Henneguya sp. (Myxozoa, Myxosporea) in gilthead sea bream (Sparus aurata) cultured in Italy. Bulletin of the European Association of Fish Pathologists, 23, 108–112.
Corpet, F. (1988). Multiple sequence alignment with hierarchical clustering. Nucleic Acids Research, 16, 10881–10890.
Eiras, J. C. (2002). Synopsis of the species of the genus Henneguya Thélohan, 1892 (Myxozoa: Myxosporea: Myxobolidae). Systematic Parasitology, 52, 43–54.
Eszterbauer, E. (2004). Genetic relationship among gill-infecting Myxobolus species (Myxosporea) of cyprinids: molecular evidence of importance of tissue-specificity. Diseases of Aquatic Organisms, 58, 35–40.
Eszterbauer, E., Kallert, D. M., Székely, Cs., & Molnár, K. (2005). The phylogeny of the genus Henneguya (Myxosporea: Bivalvulida) on the basis of 18S rDNA sequences. Abstracts. 12th EAFP International Conference on ‘Diseases of Fish and Shellfish’, 11th–16th September 2005, Copenhagen, p. 111.
Eszterbauer, E., & Székely, Cs. (2004). Molecular phylogeny of the kidney-parasitic Sphaerospora renicola from common carp (Cyprinus carpio) and Sphaerospora sp. from goldfish (Carassius auratus auratus). Acta Veterinaria Hungarica, 52, 469–478.
Fiala, I. (2006). The phylogeny of Myxosporea (Myxozoa) based on small subunit ribosomal RNA gene analysis. International Journal for Parasitology, 36, 1521–1534.
Hall, T. A. (1999). BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symposium Series, 41, 95–98.
Hallett, S. L., & Diamant, A. (2001). Ultrastructure and small-subunit ribosomal DNA sequence of Henneguya lesteri n. sp. (Myxosporea), a parasite of sand whiting Sillago analis (Sillaginidae) from the coast of Queensland, Australia. Diseases of Aquatic Organisms, 46, 197–212.
Hillis, D. M., & Dixon, T. (1991). Ribosomal DNA: molecular evolution and phylogenetic inference. Quarterly Review of Biology, 66, 411–453.
Holzer, A. S., Sommerville, C., & Wootten, R. (2004). Molecular relationships and phylogeny in a community of myxosporeans and actinosporeans based on their 18S rDNA sequences. International Journal of Parasitology, 34, 1099–1111.
Kent, M. L., Andree, K. B., Bartholomew, J. L., El-Matbouli, M., Desser, S. S., Devlin, R. H., et al. (2001). Recent advances in our knowledge of the Myxozoa. Journal of Eukaryotic Microbiology, 48, 395–413.
Lom, J., & Arthur, J. R. (1989). A guideline for the preparation of species description in Myxosporea. Journal of Fish Diseases, 12, 151–157.
Lom, J., & Dyková, I. (1992). Protozoan parasites of fishes. Developments in aquaculture and fisheries science. Vol. 26. Amsterdam: Elsevier, 315 pp.
Lom, J., & Dyková, I. (2006). Myxozoan genera: definition and notes on taxonomy, life-cycle terminology and pathogenic species. Folia Parasitologica, 53, 1–36.
Lubat, V., Radujković, B., Marques, A., & Bouix, G. (1989). Parasites des poissons marins du Montenegro: Myxosporidies. In Radujković, B. & Raibaut, A. (Eds.), Faune des parasites de poissons marins des côtes du Monténegro (Adriatic Sud). Acta Adriatica, 30, 31–50.
Molnár, K. (1998). Taxonomic problems, seasonality and histopathology of Henneguya creplini (Myxosporea) infection of the pikeperch Stizostedion lucioperca in Lake Balaton. Folia Parasitologica, 45, 261–269.
Molnár, K. (2002). Site preference of fish myxosporeans in the gill. Diseases of Aquatic Organisms, 48, 197–2002.
Ouannes-Ghorbel, A., Bradai, M. N., & Bouain, A. (2002). Période de reproduction et maturité sexuelle de Symphodus (Crenilabrus) tinca (Labridae), des côtes de Sfax (Tunisie). Cybium, 26, 89–92.
Posada, D., & Crandall, K. A. (1998). MODELTEST: testing the model of DNA substitution. Bioinformatics, 14, 817–818.
Ronquist, F., & Huelsenbeck, J. P. (2003). MRBAYES 3: Bayesian phylogenetic inference under mixed models. Bioinformatics, 19, 1572–1574.
Sakiti, N., Blanc, E., Marques, A., & Bouix, G. (1991). Myxosporidies (Myxozoa: Myxosporea) du genre Myxobolus Bütschli, 1882 parasites des poissons Cichlidae du lac Nokoué au Bénin (Afrique de l’Ouest). Journal Africain de Zoologie, 1, 173–186.
Staden, R. (1996). The Staden sequence analysis package. Molecular Biotechnology, 5, 233–241.
Swofford, D. L. (2001). PAUP*: Phylogenetic analysis using parsimony (*and other methods). Version 4.0b8. Sunderland, MA: Sinauer Associates.
Whipps, C. M., Adlard, R. D., Bryant, M. S., Lester, R. J., Findlay, V., & Kent, M. L. (2003). First report of three Kudoa species from eastern Australia: Kudoa thyrsites from Mahi mahi (Coryphaena hippurus), Kudoa amamiensis and Kudoa minithyrsites n. sp. from sweeper (Pempheris ypsilychnus). Journal of Eukaryotic Microbiology, 50, 215–219.
Whipps, C. M., Grossel, G., Adlard, R. D., Yokoyama, H., Bryant, M. S., Munday, B. L., et al. (2004). Phylogeny of the Multivalvulidae (Myxozoa: Myxosporea) based on comparative ribosomal DNA sequence analysis. Journal of Parasitology, 90, 618–622.
Work, T. M., Takara, G., Whipps, C. M., & Kent, M. L. (2008). A new species of Henneguya (Myxozoa) in the big-eyed scad (Selar crumenophthalmus) from Hawaii. Journal of Parasitology, 94, 524–529.
Yokoyama, H., Kawakami, H., Yasuda, H., & Tanaka, S. (2003). Henneguya lateolabracis sp. n. (Myxozoa, Myxosporea), the causative agent of cardiac henneguyosis in Chinese sea bass Lateolabrax sp. Fisheries Science, 69, 1116–1120.
Yokoyama, H., Itoh, N., & Tanaka, S. (2005). Henneguya pagri n. sp. (Myxozoa: Myxosporea) causing cardiac henneguyosis in red sea bream, Pagrus major (Temminck & Schlegel). Journal of Fish Diseases, 28, 479–487.
Acknowledgement
The molecular part of the study was supported by the Hungarian Scientific Research Fund (OTKA) grant no. K75873.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bahri, S., Marton, S., Marques, A. et al. Henneguya tunisiensis n. sp. (Myxosporea: Bivalvulida), a new gill parasite of Symphodus tinca (L.) (Teleostei: Labridae) off Tunisia. Syst Parasitol 76, 93–101 (2010). https://doi.org/10.1007/s11230-010-9237-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11230-010-9237-z