Abstract
Between 1991 and 2010, 45 scientists were honored with Nobel prizes in Physiology or Medicine. It is shown that these 45 Nobel laureates are separated, on average, by at most 2.8 co-authorship steps from at least one cross-disciplinary broker, defined as a researcher who has published co-authored papers both in some biomedical discipline and in some non-biomedical discipline. If Nobel laureates in Physiology or Medicine and their immediate collaborators can be regarded as forming the intuitive “center” of the biomedical sciences, then at least for this 20-year sample of Nobel laureates, the center of the biomedical sciences within the co-authorship graph of all of the sciences is closer to the edges of multiple non-biomedical disciplines than typical biomedical researchers are to each other.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
It is intuitively tempting to visualize scientific disciplines as spheres, with highly productive, well-funded intellectual and political leaders such as Nobel laureates occupying their centers and less productive, less well-funded researchers being increasingly peripheral. As preferential attachment mechanisms as well as the economics of employment tend to give the well-known and well-funded more collaborators than the less well-known and less well-funded (e.g. Barabási and Albert 1999; Barabási et al. 2002), one can expect the average degree of vertices in the co-authorship graph of a spherical discipline to decrease as one moves from the center to the periphery. On this intuitive view, one can expect typical pairs of the most peripheral researchers to be separated by roughly the graph diameter from each other, and by roughly half of the graph diameter from the center (for formal definitions of graph-theoretic concepts, see Diestel 2010, or for a briefer, application-specific summary, Börner et al. 2007). In this case, most co-authorship paths between peripheral researchers would traverse or at least pass near the center, so degree, distance and betweenness centrality would all at least roughly coincide [see e.g. Freeman (1978/1979), Borgatti and Everett (2006) or Landherr et al. (2010) for definitions and comparisons of these centrality measures]. Cross-disciplinary brokers would be peripheral researchers in one discipline who interact occasionally with peripheral researchers in another discipline, and who have little influence on the discipline’s overall evolution. As co-authorship connections between researchers are influenced by factors such as geography, academic lineage, personalities, institutional structure and inter-institutional relations, and even international politics as well as by attachment preferences, this intuitive picture of disciplines as spherical is clearly an over-simplification of the fine structure of disciplinary co-authorship graphs. However, it appears to guide much of the institutional management of science, and seems accurately to describe disciplinary organizations and boundaries that appear surprisingly refractory to managerial initiatives toward interdisciplinarity (e.g. Jacobs and Frickel 2009).
Even a cursory examination of the documented Erdős numbers—the minimal co-authorship distances from the late mathematician Paul Erdős—of Nobel laureates, however, challenges this intuitive picture of disciplines as spheres. The Erdős numbers of Nobel laureates in Physics, Chemistry, Economics, and Physiology or Medicine, where known (De Castro and Grossman 1999; see http://www.oakland.edu/enp/erdpaths/ for more current data), tend to be closer to the average co-authorship distances between researchers in their respective disciplines than to half of the relevant graph diameters. Physics, for example, had a graph diameter of 20 and an average co-authorship distance between researchers of 5.9 in the latter half of the 1990s; the corresponding numbers for biomedical science are 24 and 4.6 (Newman 2001, Table 1). The average Erdős numbers during the somewhat larger period 1991–2005 for the incomplete sample of Nobel laureates documented by the Erdős Number Project are 5.4 for physicists and 3.8 for biomedical scientists.Footnote 1 These Nobel laureates are, therefore, considerably closer to the boundaries separating their respective disciplines from mathematics than half of the relevant graph diameter and hence are considerably closer than the spherical model would predict; if these Nobel laureates indeed occupy the centers of their respective disciplines, their disciplines cannot be co-authorship spheres.
That mathematics is not a special case is suggested by recent work that examined co-authorship paths that begin in one discipline, traverse another discipline and end in a third discipline. Co-authorship paths with lengths less than three can be found that traverse subdisciplines as diverse as discrete mathematics, nuclear physics, macroeconomics and theoretical computer science (Fields 2014a). While such short subdiscipline-crossing paths are exceptional, their existence indicates that cross-disciplinary brokers can at least sometimes be found in close proximity. These cross-disciplinary brokers are, moreover, typically highly-collaborative, highly-cited researchers and are in some cases Nobel laureates, including Francis Crick (Physiology or Medicine, 1962), Richard Feynman (Physics, 1965), Max Delbrück (Physiology or Medicine, 1969), Murray Gell-Mann (Physics, 1969) and Herbert Simon (Economics, 1978). This result suggests an alternative picture in which the co-authorship graphs of disciplines are highly non-spherical, with their “centers” in relatively close mutual proximity and their most peripheral researchers located not just half but possibly approaching a full graph diameter away from their respective centers.
The present paper tests the validity of this alternative picture of disciplinary co-authorship graphs by asking how close Nobel laureates in Physiology or Medicine are to cross-disciplinary brokers and hence to researchers in some other discipline. By analyzing citations in the biomedical literature by source discipline, Chen et al. (2014) have recently demonstrated both the successive emergence, between 1910 and 2010, of the core disciplines of current biomedical science and the influence of other disciplines on this process. The work presented here complements this previous study by examining direct co-authorship connections between biomedical researchers and scientists with different disciplinary backgrounds. It shows, in particular, that such direct cross-disciplinary connections are at least sometimes made in close proximity to Nobel laureates.
For the present purposes, a “cross-disciplinary broker” is defined as a researcher who has published co-authored papers meeting the selection criteria outlined below both in biomedicine, the broad domain of scientific work honored by Nobel Prizes in Physiology or Medicine, and in some non-biomedical discipline. As will be seen below, the general discipline of biomedicine includes the five Klavans and Boyack (2009) “consensus” disciplines of Biology, Biochemistry (though biochemists are also sometimes awarded Nobel Prizes in Chemistry), Infectious Disease, Medical Specialties and Neuroscience (capitalization is used throughout to indicate Klavans and Boyack consensus disciplines). The other 11 Klavans and Boyack consensus disciplines are considered to be “distinct disciplines” from biomedicine for the purposes of identifying cross-disciplinary brokers. The traditional biological subdisciplines of taxonomy, phylogeny and systematics are also considered to compose a “distinct discipline” here termed “evolutionary biology.” Nobel Prizes in Physiology or Medicine are not awarded for research in this discipline. Administrative divisions between academic departments emphasizing laboratory studies using the tools of molecular biology and biochemistry and those emphasizing field and museum studies using observational methods, which began to appear as early as the mid-1970s, moreover assure that many biomedically-oriented biologists have little exposure to these traditional, evolutionarily-oriented parts of biology. Hence for the purposes of this study, a cross-disciplinary broker is someone who has published co-authored papers in at least one of the five “biomedical” Klavans and Boyack consensus disciplines, and has also published co-authored papers either in at least one of the other 11 Klavans and Boyack consensus disciplines or in evolutionary biology. No a priori restriction is placed on the relative timing of these papers, so individuals who qualify as brokers due to field mobility are not distinguished a priori from those who publish in multiple fields in parallel; this question of mobility versus parallelism will be considered further below.
To minimize ascertainment bias, co-authorship connections of all Physiology or Medicine Nobel laureates between 1991 and 2010 are examined. The specializations of these 45 laureates range from genetics and molecular biology through cell and developmental biology, virology, microbiology and neuroscience to reproductive physiology. Both the extent to which these Nobel laureates collaborate among themselves and hence form a coherent, highly connected “center” of the biomedical sciences and the co-authorship distances from these Nobel laureates to cross-disciplinary brokers as defined above are examined. Co-authorship data are also presented for 12 additional Nobel laureates, nine in Physiology or Medicine and three in Chemistry, who are closely connected to the 1991–2010 cohort. As the graph search methods used are heuristic as described below, all co-authorship distances reported are upper limits. These upper-limit measurements show, first, that the Klavans and Boyack disciplines of Biology, Biochemistry, Infectious Disease, Medical Specialties and Neuroscience are essentially indistinguishable at the level of co-authorship connections between Nobel laureates; specialists in these disciplines cannot even be identified as forming exclusive disciplinary cliques. Second, they show that Nobel laureates in these disciplines are closely connected, via cross-disciplinary brokers, to at least ten other disciplines ranging from mathematics to philosophy; indeed they are closer, on average, to researchers in at least one other discipline than they are, on average, to other biomedical researchers. Hence if these biomedical Nobel laureates can be regarded as “central” to biomedicine—as surely they can be on any socially or politically meaningful notion of centrality—then at least in terms of co-authorship, the “center” of biomedicine is surprisingly close to the “edge” of biomedicine. Even when they are considered together, therefore, the biomedical disciplines do not form a co-authorship sphere.
Data and methods
Names and specializations of Nobel laureates in Physiology or Medicine were obtained from Nobelprize.org.Footnote 2 Co-authorship data for Nobel laureates and their co-authors were obtained by manual searches of Google Scholar™ between January and June, 2014. The use of Google Scholar™ for bibliometric analysis has been controversial; recent large-scale studies indicate high coverage of the literature (e.g. Harzing 2013) but with low quality control compared to commercial indices (e.g. Aguillo 2012). Only primary and secondary research papers, review articles, research-based science-policy papers and scholarly books were included in the present analysis; otherwise-unpublished technical reports, textbooks, joint editing of collections, and editorial or opinion pieces were not included. Excluding these “grey literature” sources may lead to over-estimation of co-authorship distances, but cannot lead to under-estimation of such distances. All publications employed to establish co-authorship connections are listed in “Appendix 1” so that their titles, co-authors and sources may be examined.
Nobel laureates tend to have many—often hundreds—of co-authors, who may themselves have hundreds co-authors. To make searches for co-authorship paths from laureates to either other laureates or brokers reasonably efficient in the face of this complexity, co-authorship paths from laureates that traverse other authors known to be near either cross-disciplinary brokers or other Nobel laureates were followed preferentially. The present author is himself a cross-disciplinary broker who specialized for several years in bioinformatics; the search process employed here may, therefore, be biased toward identifying other cross-disciplinary brokers associated with bioinformatics over brokers with other backgrounds or specialities. Searches were generally terminated when some cross-disciplinary broker with co-authored publications in at least one non-biomedical discipline as defined above was encountered; where relevant to the main objective of establishing upper limits on laureate-broker distances, co-authorship connections between Nobel laureates and between identified brokers were also considered. This search procedure effectively implements a greedy heuristic and cannot be regarded as globally optimal; it is possible, in particular, that more exhaustive search techniques might reveal additional cross-disciplinary brokers at distances equal to or even smaller than those reported here. All reported co-authorship distances that are greater than one must, therefore, be viewed as upper limits only.
Upper limits on co-authorship distances were measured by counting co-authored publications along the shortest paths found connecting individuals of interest in the co-authorship graph; distances were not weighted by citation counts, numbers of joint publications between pairs of authors, or other specialized metrics. Where necessary, authors with similar names were disambiguated by tracing their histories of institutional appointments. Citations counts are reported where particularly significant; these counts were obtained from Google Scholar™ in early June, 2014. It should be noted that the method used here systematically underestimates interdisciplinarity by discounting all single-author publications. As single-author publications are increasingly rare in the sciences (Porter and Rafols 2009), any effect of this bias is expected to be small.
Results
The primary results of this analysis are presented as co-authorship subgraphs demonstrating laureate-to-broker connections (Figs. 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13) to facilitate a visual grasp of laureate-to-broker distances; a summary is presented in tabular form in “Appendix 2”. Nobel laureates are indicated by a “*” and a two-digit award date. Researchers other than Nobel laureates are included in these subgraphs only to indicate minimal identified co-authorship paths between laureates or between laureates and cross-disciplinary brokers, or when they serve as brokers. Cross-disciplinary brokers are indicated by an edge connecting to an italicized discipline name, e.g. Physics; in these cases a representative publication in the indicated discipline co-authored by the broker is provided. Subgraphs were constructed for each year’s Nobel laureates in Physiology or Medicine, ordered by award year, unless that year’s laureates have already been included in a previous subgraph. Like the search procedure employed, this method of subgraph construction effectively implements a greedy heuristic that may overestimate, but cannot underestimate, the upper limits on laureate-to-broker co-authorship distances that are of interest.
Many if not most of the researchers shown in these subgraphs have well over 100 collaborators and some of the papers shown as edges have well over 100 co-authors; hence these subgraphs are far less complex than the region of the complete co-authorship graph from which they are abstracted. As these subgraphs are drawn for the specific purpose of displaying identified inter-laureate and laureate-to-broker connections, they cannot be regarded as representative of the structure of the co-authorship graph as a whole, and formal measures of centrality or of other features of the full co-authorship graph cannot be considered meaningful when applied only to these subgraphs. The subgraphs shown may all be joined along shared vertices to construct a single connected subgraph linking laureates to brokers; join vertices are indicated explicitly. Some of the co-author pairs shown have co-authored multiple papers together (e.g. at least 50 in the case of Hamilton Smith and J. Craig Venter); in such cases, a prominent paper also co-authored by other authors included as vertices in one or more subgraphs is chosen for display. Papers are employed as edges in multiple subgraphs where possible, allowing the subgraphs to be joined along shared edges as well as shared vertices as discussed in specific cases below. Inferred upper limits on the Erdős numbers of all laureates are included in the tabular results provided in “Appendix 2”. Notable citation counts and qualitative data relevant to centrality are provided in the accompanying text.
1991: Erwin Neher and Bert Sakmann. Labels are: a Hamill et al. (1991), b Markram et al. (1997), c Schuster and Just (2006), d Niebur et al. (1991), e Anastassiou et al. (2011), f Li and Itti (2011), g Itti et al. (1998), h Crick and Koch (1990), i Itti and Baldi (2006), j Baldi and Rinott (1989), k Watson and Crick (1953)
The 1991 Nobel Prize in Physiology or Medicine was awarded to Bert Sakmann and Erwin Neher for their development of the patch-clamp technique, a novel and significant application of cell biological methods to neuroscience. The paper of Hamill et al. (1991) introduced this technique and has received 17,154 citations. Markram et al. (1997) is one of many papers applying these methods to characterize interneuronal signalling; it links the Klavans and Boyack (2009) consensus disciplines of Biology—here, cell biology—and Neuroscience. As is well-known, James Watson and Francis Crick shared the 1962 Nobel Prize in Physiology or Medicine, with Maurice Wilkins, for their characterization of the double-helix structure of DNA; Watson and Crick (1953) has 9,703 citations. Some years later, Crick turned his attention to neuroscience; Crick and Koch (1990) introduced an influential neurobiological account of conscious awareness and has 1,631 citations. Crick is, therefore, another connection between Biology—in this case, molecular biology—and Neuroscience.
The co-authorship links shown in Fig. 1 place upper limits of two, three, three and four, respectively, on the maximum co-authorship distances between Nobel laureates Francis Crick, James Watson, Bert Sakmann and Erwin Neher and the borders between the biomedical sciences and the Klavans and Boyack consensus disciplines of Physics and Computer Science. It also shows, incidently, that the Physics—Computer Science distance is only two co-authorship steps, and that Computer Science is separated from a third Klavans and Boyack consensus discipline, Mathematics, by just one co-authorship step. As Pierre Baldi has an Erdős number of two (http://www.oakland.edu/enp/thedata/), the Erdős numbers of Sakmann and Neher are at most six and seven, respectively. Both Watson and Crick have Erdős numbers of at most four (http://www.oakland.edu/enp/erdpaths/).
Christof Koch is Chief Scientific Officer of the Allen Institute for Brain Science and is well-known in neural modelling circles; Itti et al. (1998) describes a computational model of visual attention and has 5,227 citations. Henry Markram currently directs the European Human Brain Project, a multi-national effort to fully characterize human cerebral cell types and connectivity. Both can be considered central figures in contemporary neuroscience; their proximity to the borders between the biomedical sciences and Physics, Computer Science and Mathematics is, therefore, significant in the present context.
The 1992 Nobel Prize in Physiology or Medicine honors the characterization, by Edmond Fischer and Edwin Krebs, of protein phosphorylation as a ubiquitous regulator of biochemical activity; Fischer and Krebs (1958) reports some of this work in the Journal of Biological Chemistry. Figure 2 thus links the Klavans and Boyack (2009) consensus discipline of Biochemistry to neuroscience, cell biology and molecular biology, the specialties of Nobel laureates Eric Kandel, Tim Hunt, and Hamilton Smith respectively. It also shows that Hamilton Smith is only one co-authorship step away from four boundaries between biomedicine and other disciplines: those of evolutionary biology, physics, electrical engineering and cognitive science, and is only two co-authorship steps from the boundaries between biomedicine and philosophy or computer science. Edwin Krebs, Edmond Fischer and Eric Kandel are, therefore, separated from the first four borders by no more than three, four and four co-authorship steps, respectively, and are separated from the latter two borders by no more than four, five and five co-authorship steps, respectively. As Hamilton Smith and Eric Kandel have Erdős numbers of at most three (Fields 2014b) and four (see Fig. 8) respectively, Krebs and Fischer have Erdős numbers of at most six and seven, respectively.
1992: Edmond H. Fischer and Edwin G. Krebs. Labels are: a Malumbres et al. (2009), b Mollinari et al. (2002), c Job et al. (1981), d Huang et al. (1995), e Fischer and Krebs (1958), f Scott et al. (1987), g Zoller et al. (1979), h Fleischmann et al. (1995), i Farris et al. (1994), j Natale et al. (2011), k Kitching et al. (1978), l DeYong et al. (1992), m Dietrich and Fields (1996), n Grenon and Smith (2004), o Mulligan et al. (1984)
As discussed earlier, evolutionary biology is a component of the Klavans and Boyack consensus discipline of Biology, but is not part of biomedicine. Klavans and Boyack include electrical engineering in their consensus discipline of Computer Science; they are named separately here to distinguish the hardware-oriented work of DeYong et al. (1992), which describes the design and testing of novel integrated circuits, from the algorithm-oriented work of Grenon and Smith (2004) or Li and Itti (2011) from Fig. 1. Cognitive science is an amalgam of components from the Klavans and Boyack disciplines of Psychology (mainly cognitive psychology), Computer Science (artificial intelligence), Social Sciences (anthropology and linguistics) and Humanities (philosophy of mind). Philosophy—in Barry Smith’s case, ontology—is a part of the Klavans and Boyack discipline of Humanities. Figures 1 and 2 together, therefore, already demonstrate links between biomedicine and six of the other 11 Klavans and Boyack consensus disciplines.
The subgraph of 14 Nobel laureates, including Chemistry laureate Tom Cech, and five of their collaborators shown in Fig. 3 spans 50 years of Nobel prizes and forms a natural “center” of the biomedical sciences to which the other Nobel laureates between 1991 and 2010 may be referred. It joins directly with 11 of the other 12 subgraphs presented here. The three-laureate clique defined by Akil et al. (2010) links neuroscientist Eric Kandel with geneticist Sydney Brenner and molecular biologist James Watson. The four-clique defined by Berg et al. (1974) links molecular biologists Watson and Dan Nathans to David Baltimore, who received his 1975 Nobel Prize for work in virology. Roberts and Szostak (1997) links molecular biologist Richard Roberts to cell biologist Jack Szostak; Burke et al. (1987) links Szostak to biochemist Tom Cech. The Klavans and Boyack (2009) consensus disciplines of Biology, Biochemistry, Infectious Disease and Neuroscience are, therefore, all represented by Nobel laureates in this subgraph.
Figure 3 also shows that Dan Nathans, Phillip Sharp and Richard Roberts are within two co-authorship steps of the disciplinary borders crossed by the present author, and that Nathans and Roberts are within two co-authorship steps of the disciplinary border crossed by Carol Bult (cf Fig. 2). It places upper limits of four on the Erdős numbers of Nathans and Roberts, five on those of Jack Szostak and Phillip Sharp, and six on those of Elizabeth Blackburn, Carol Greider and Tom Cech. The Erdős numbers of Sydney Brenner and Andrew Fire are at most four, while that of David Baltimore is at most five (Fields 2014b). As Hamilton Smith and James Watson have Erdős numbers of at most three and four as noted earlier, that of Joshua Lederberg is at most six (Lederberg’s Erdős number is in fact at most 5; see http://www.oakland.edu/enp/erdpaths/).
1993: Richard J. Roberts and Phillip A. Sharp; 2009: Elizabeth H. Blackburn, Carol W. Greider and Jack W. Szostak. Labels are: a Akil et al. (2010), b Kandel and Schwartz (1982), c Berg et al. (1974), d Nathans et al. (1962), e Zinder and Lederberg (1952), f Singh et al. (1986), g Smith and Nathans (1973), h Adams et al. (2000); i Padgett et al. (1983), j Manley et al. (1980), k Venter et al. (2001), l Fleischmann et al. (1995), m Mount et al. (1992), n Roberts and Szostak (1997), o Blackburn et al. (2006), p Burke et al. (1987)
Figure 3 includes three major papers of the Human Genome Project. Venter et al. (2001) is one of two initial reports of the complete sequence of the human genome and has garnered 12,061 citations. Adams et al. (2000) reports the complete sequence of the Drosophila melanogaster (fruit fly) genome and has 5,074 citations. Fleischmann et al. (1995), also shown as edge “h” in Fig. 2, reports the first complete sequence of a microbial genome and has 5,214 citations. Fleischmann et al. (1995) has 40 co-authors while Adams et al. (2000) and Venter et al. (2001) both have well over 100, providing a glimpse of the complexity of the full co-authorship graph from which the subgraphs shown are abstracted. J. Craig Venter, whose connections with additional Nobel laureates are shown in Fig. 12, is a pioneer in high-throughput, highly-automated DNA sequencing for whole-genome characterization, environmental sequencing to discover new organisms, and synthetic biology. Currently President of the J. Craig Venter Institute, he is a central figure in both biomedical research and the biotechnology industry.
Figure 4 links cell biologists Alfred Gilman and Martin Rodbell, honored in 1994 for their work in cellular signal transduction, to Drosophila geneticists Edward Lewis, Christiane Nüsslein-Volhard and Eric Wieschaus, who shared the 1995 Nobel Prize for work in developmental genetics. It also illustrates the effect of genome-level biology, here represented by Adams et al. (2000), Lander et al. (2001) and Drosophila 12 Genomes Consortium (2007), on the historically somewhat insular Drosophila genetics community. Figure 4 also shows that Nobel laureate Alfred Gilman is at most three co-authorship steps of mathematician László Székely, a direct collaborator of Paul Erdős (http://www.oakland.edu/enp/thedata/), giving Gilman an Erdős number of at most four. Via Eugene Koonin’s co-authorship of Lander et al. (2001), Gilman is three co-authorship steps from the three disciplinary boundaries crossed by Eric Lander. Like Koonin’s, Lander’s Erdős number is two (http://www.oakland.edu/enp/thedata/), confirming Gilman’s Erdős number of at most four by a second route. Nobel laureates Christiane Nüsslein-Volhard, Eric Wieschaus, Edward Lewis, and Martin Rodbell receive Erdős numbers of at most four, five, five and seven, respectively, via Lander, Koonin or both. Besides these connections with mathematics, Nüsslein-Volhard, Wieschaus and Lewis are within two, three and three co-authorship steps, respectively, of physicist James Yorke, a well-known chaos theorist, as well as three, four and three co-authorship steps, respectively, of the present author. This subgraph also shows that biomedical science can be traversed, from physics to mathematics, in four co-authorship steps (Yorke to Székely), three co-authorship steps (Fields to either Székely or Lander), two co-authorship steps (Fields to Ehrenfeucht), or even one co-authorship step (Yorke to Lander), traversal widths comparable to those demonstrated in Fields (2014a) using different co-authorship paths.
1994: Alfred G. Gilman and Martin Rodbell; 1995: Edward B. Lewis, Christiane Nüsslein-Volhard and Eric F. Wieschaus. Labels are: a Erdős et al. (1999), b Pohl et al. (1971), (c) Rogozin et al. (2002), d Montell et al. (2002), e Daigle et al. (2002), f Gardner et al. (1998), g Adams et al. (2000), h Montell and Rubin (1989), i Coleman et al. (1994), j Rubin and Lewis (2000), k Freissmuth et al. (1989), l Karch et al. (1985), m Bahmanyar et al. (2008), n Fleischmann et al. (1995), o Peifer and Wieschaus (1990), p O’Toole et al. (2003), q Nüsslein-Volhard and Wieschaus (1980), r Li et al. (2004), s Ray et al. (1991), t Schneider et al. (1986), u Mount et al. (1992), v Drosophila 12 Genomes Consortium (2007), w Ehrenfeucht and Rozenberg (1990), x Ott et al. (1990); y Farrell and Lander (1989), z Arratia and Lander (1990), aa Lander et al. (2001), bb Lander et al. (1989)
Cross-disciplinary broker Eric Lander earned his D.Phil. in mathematics, was one of the founders, in the late 1980s, of the new subdiscipline of bioinformatics, and served as Director of the Whitehead Institute during the initial stage of the Human Genome Project. He is the first author of Lander et al. (2001), the other of the two initial reports of the complete sequence of the human genome, which appeared in the same week as Venter et al. (2001) and has garnered 16,575 citations. Currently Director of the Broad Institute, he is central to biomedicine on any reasonable definition of centrality. Here he provides additional links from biomedicine to the Klavans and Boyack (2009) disciplines of Mathematics, Computer Science and Social Sciences.
The 1996 Nobel Prize in Physiology or Medicine honored Peter Doherty and Rolf Zinkernagel for their work in immunology; Susumu Tonegawa’s 1987 Nobel Prize similarly honors work in immunology. Figure 5 thus connects immunology, a subdiscipline of Klavans and Boyacks’s (2009) consensus discipline of Medical Specialties, with the Klavans and Boyack disciplines of Neuroscience (via Eric Kandel) and Infectious Disease (via David Baltimore). Given Eric Kandel’s Erdős number of at most four, this subgraph gives Erdős numbers of at most five to Susumu Tonegawa and at most six to both Rolf Zinkernagel and Peter Doherty. It also connects these Nobel laureates to both physics and computer science via Christof Koch’s connections (see Fig. 1).
Figure 6 illustrates the central position in biomedicine of Francis Collins, currently Director of the US National Institutes of Health. It connects Collins to seven Nobel laureates, including biochemist Kary Mullis, a laureate in Chemistry. Günter Blobel’s specialty is cell biology, Mario Capecchi’s is molecular biology and John Sulston’s is genetics. Stanley Prusiner’s Nobel Prize honors his discovery that prions are infectious agents, while Harold Varmus’ honors work in oncology. Figure 6 thus connects Nobel laureates in the Klavans and Boyack disciplines of Biology, Biochemistry, Infectious Disease and Medical Specialties. As Francis Collins is only one co-authorship step from the disciplines of both Eric Lander and the present author, all of these Nobel laureates are close to multiple cross-disciplinary boundaries. Nobel Laureate Günter Blobel, two steps from Collins, is only one co-authorship step from the boundary between biomedical science and evolutionary biology, having co-authored a paper with well-known evolutionary biologist Stephen J. Gould, a cross-disciplinary broker who also did significant work in Blobel’s field of cell biology. Eric Lander’s Erdős number of two confers low Erdős numbers on all of the other scientists in this subgraph. The largest clique in this subgraph is that defined by Lander et al. (2001), which as noted earlier is one of the two initial reports of the complete sequence of the human genome. All co-authors of this paper are also co-authors of Eugene Koonin, and hence link to Fig. 4 through Koonin as well as Lander.
1997: Stanley B. Prusiner; 1999: Günter Blobel; 2007: Mario R. Capecchi. Labels are: a Gould and Lewontin (1979), b Distel et al. (1996), c Saiki et al. (1985), d Oesch et al. (1985), e Boehnke et al. (1989), f Manolio et al. (2009), g Lander et al. (2001), h Austin et al. (2004), i McCombie et al. (1992), j Collins and Watson (2003)
The 1998 Nobel Prize in Physiology and Medicine honored the discovery that nitric oxide (NO) serves as a signalling molecule in the cardiovascular system (Fig. 7). The research leading to this discovery employed biochemical methods far removed from those of molecular biology and genetics; the Nobel laureates of 1998 are correspondingly far, in terms of co-authorship, from the “center” shown in Fig. 3. Like Hamilton Smith in Fig. 2 or Günter Blobel in Fig. 6, Nobel laureate Richard Furchgott is only one co-authorship step from the boundary of biomedical science, having co-authored several papers with physicist-turned-physiologist William Sleator, Jr. Laureates Ferid Murad and Louis Ignarro, however, appear to be at least six and eight co-authorship steps, respectively, from the edges of biomedicine. The Erdős numbers of the 1998 laureates are also among the highest in the 1991–2010 time period. As the present author has an Erdős number of at most three,Footnote 3 Richard Furchgott’s is at most nine, although it may be smaller due to paths to Erdős within physics. John Sulston’s Erdős number is at most three (Fields 2014b); Ferid Murad’s Erdős number is, therefore, at most eight and Louis Ignarro’s is at most ten.
1998: Robert F. Furchgott, Louis J. Ignarro and Ferid Murad. Labels are: a Blair et al. (1948), b Furchgott et al. (1960), c Olney et al. (1979), d Olney et al. (1999), e Newcomer et al. (1999), f Lee et al. (2012), g McCombie et al. (1992), h Schein et al. (1993), i Grishok et al. (2001) j Ruvkun et al. (1989), k Ogg et al. (1997) l Tolias et al. (2005), m Burette et al. (2002), n Förstermann et al. (1991), o Schmidt et al. (1996), p Ignarro et al. (1993), q Wink et al. (1998)
Nobel laureate Peter Mansfield, one of the developers of biomedical magnetic resonance imaging (MRI), is a cross-disciplinary broker in Fig. 8, as is his collaborator Penny Gowland, currently a Professor of Physics at the University of Nottingham. Even more strikingly, cognitive psychologist Michael Posner is traversed by what appears to be the shortest co-authorship path between biomedical Nobel laureates Arvid Carlsson and Peter Mansfield on the one hand and Eric Kandel and his collaborators—and hence the “center” of the biomedical sciences shown in Fig. 3—on the other. This subgraph shows that Eric Kandel is only one co-authorship step from the boundary between the biomedical sciences and the Klavans and Boyack discipline of Psychology and only three steps from the boundary with Mathematics. As Charles Cantor is a co-author, with Francis Collins, of Smith et al. (1987), this subgraph is also linked to Fig. 6 and hence to the cross-disciplinary connections of both Eric Lander and the present author. Noga Alon’s Erdős number of one (http://www.oakland.edu/enp/thedata/) gives Richard Axel an Erdős number of three and Kandel an Erdős number of four as noted earlier. Linda Buck, therefore, also has Erdős number four, while Paul Greengard and Oliver Smithies have Erdős numbers of at most five. This subgraph gives both Arvid Carlsson and Peter Mansfield Erdős numbers of at most nine, although Mansfield’s may be lower via co-authors in the experimental physics community.
2000: Arvid Carlsson, Paul Greengard and Eric R. Kandel; 2003: Peter Mansfield; 2004: Richard Axel and Linda B. Buck; 2007: Oliver Smithies. Labels are: a Kegeles et al. (2000), b Baker et al. (1994), c Mansfield and Grannell (1973), d Petridou et al. (2009), e Innis et al. (2007) f Hervé et al. (2011), g Fox et al. (1988), h Mazziotta et al. (2001), i Petersen et al. (1988), j Posner et al. (1980), k Albright et al. (2000), l Castellucci et al. (1980), m Scheller et al. (1982), n Buck and Axel (1991), o Argarana et al. (1986), p Wigler et al. (1979), q Ressler et al. (1994), r Alon et al. (2006), s Efstratiadis et al. (1980), t Myers et al. (2006), u Alon and Spencer (2000)
Figure 9 shows that Nobel laureates Leland Hartwell, Tim Hunt and Paul Nurse, honored in 2001 for their work on the genetics of the cell-division cycle, are within three, four and two co-authorship links, respectively, of the disciplinary boundaries crossed by both Eric Lander and the present author. They have Erdős numbers of five, four and four, respectively, via Eric Lander, Eugene Koonin or both. D. Carleton Gajdusek, a specialist in tropical medicine, received the 1976 Nobel Prize in Physiology and Medicine for his work on what would later be recognized as prion diseases; Fig. 9 shows that he is only two co-authorship links from multiple disciplinary boundaries, and gives him an Erdős number of at most four.
2001: Leland H. Hartwell, Tim Hunt and Paul M. Nurse. Labels are: a Lander et al. (2001), b Uhlmann et al. (2000), c Nasmyth and Hunt (1993), d Breeden and Nasmyth (1985), e McCombie et al. (1992), f Wood et al. (2002), g Nurse et al. (1976), h Yarus et al. (1986), i Goldfarb et al. (1991), j Nurse et al. (1998), k Schneider et al. (1984), l Dutcher and Hartwell (1982), m Li et al. (2004)
2002: Sydney Brenner, H. Robert Horvitz and John E. Sulston; 2006: Andrew Z. Fire and Craig C. Mello. Labels are: a Chalfie et al. (1985), b Xue et al. (1992), c Ruvkun et al. (1989), d Fleming et al. (1997), e Herr et al. (1988), f Grishok et al. (2001), g Dupuy et al. (2007), h Schein et al. (1993), i Buldyrev et al. (1992), j Barabási and Albert (1999)
The subgraph of seven Nobel laureates, including Chemistry laureate Martin Chalfie, shown in Fig. 10 demonstrates the important role of the nematode worm Caenorhabditis elegans, the subject of all of the papers shown here except those of Albert-László Barabási that are not in biomedical science, in the late twentieth century development of molecular genetics and genomics. It also shows that these Nobel laureates are all close to the borders between biomedicine and both physics and the theory of networks, an emerging discipline with components from the Klavans and Boyack (2009) disciplines of Social Sciences, Mathematics and Physics. As both John Sulston and the present author have Erdős numbers of three, all of the scientists in this subgraph also have low Erdős numbers. Grishok et al. (2001) is one of the first papers describing the gene-regulating function of small RNAs and has 1,546 citations. The most highly-cited paper in this subgraph, however, is the pioneering work of Barabási and Albert (1999) on scale-free networks; with 20,121 citations, this paper is the most highly-cited work included in the present study.
Paul Lauterbur’s Nobel Prize (Fig. 11) honors his contribution to the development of biomedical MRI; his proximity to the boundary between biomedicine and materials science, a amalgam of the Klavans and Boyack (2009) consensus disciplines of Physics, Chemistry and Engineering, is not surprising. His co-authorship connection to the present author gives him an Erdős number of at most eight. Note the reappearance in this graph of McCombie et al. (1992), which also serves as an edge in Figs. 6, 7 and 9; this paper links both J. Craig Venter and Francis Collins, as well as Richard McCombie, to Fig. 11.
2005: Barry J. Marshall and J. Robin Warren; 2008: Harald zur Hausen, Françoise Barré-Sinoussi and Luc Montagnier. Labels are: a Marshall and Warren (1984), b Yamada et al. (1994), c Gallo et al. (1983), d Broder et al. (1985), e Klausner et al. (2003), f Venter et al. (2001), g Adams et al. (1995), h Gallo et al. (1988), i Haseltine et al. (1976), j Schuler et al. (1996), k Chinwalla et al. (2002), l Barré-Sinoussi et al. (1983)
Figure 12 shows that Nobel laureates David Baltimore and Luc Montagnier are only two co-authorship steps from the cross-disciplinary boundaries crossed by Carol Bult, Eric Lander and the present author, while Françoise Barré-Sinoussi, Harald zur Hausen, Barry Marshall and Robin Warren are three, three, four and five steps from these boundaries, respectively. The Erdős numbers of Montagnier, zur Hausen, Barré-Sinoussi, Marshall and Warren are at most five, five, six, six and seven, respectively, via Eric Lander. Anthony Fauci is Director of the U.S. National Institute of Allergies and Infectious Disease, while Samuel Broder is a former Director of the US National Cancer Institute; these scientists are, therefore, also central figures in the biomedical sciences.
Martin Evans (Fig. 13) is the third of the 2007 Nobel laureates in Physiology or Medicine, all honored for their research on stem cells. He is only two co-authorship steps from evolutionary biology via Carol Bult, and is three steps from the disciplines of Eric Lander. Evans’ Erdős number is at most five, via Lander. The 2010 Nobel Prize in Physiology or Medicine honored Robert Edward’s development of human in vitro fertilization. His closest connection to the “center” defined by Fig. 3 is via Eric Lander, also giving him an Erdős number of at most five.
Discussion
The co-authorship data shown in Figs. 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12 and 13 can be summarized as follows. The 45 Nobel laureates in Physiology or Medicine between 1991 and 2010 are separated, on average, from at least one other Nobel laureate in this set by at most \(d_{l} = 2.0\) co-authorship steps and from a cross-disciplinary broker by at most \(d_{b} = 2.8\) co-authorship steps (see “Appendix 2”, Tables 1 and 2 for complete data). For the 1991–2000 decade only, the average upper limits are \(d_{l} = 2.2\) and \(d_{b} = 3.2\); for the 2001–2010 decade only, the average upper limits are \(d_{l} = 1.7\) and \(d_{b} = 2.5\). Hence these Nobel laureates are, on average, less than one co-authorship step more distant from a boundary of their discipline of biomedical science than they are from at least one other laureate, and this distance relation is roughly constant in time. For comparison, the other 12 Nobel laureates appearing in Figs. 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, and 13, together with Max Delbrück, a physicist who was one of the founders of molecular biology (Fields 2014a), have an average \(d_{l} = 1.3\) and \(d_{b} = 2.2\) (“Appendix 2”, Table 3).
It should be emphasized that the co-authorship distances obtained here are upper limits as discussed above. It should also be noted that the definition of “cross-disciplinary broker” employed here is very stringent. If a “cross-disciplinary broker” was considered to be someone who has collaborated with a researcher in another discipline, as opposed to someone who has published co-authored papers in multiple disciplines, then the difference of one co-authorship step between the average \(d_{l}\) and the average \(d_{b}\) reported here would vanish.
The average Erdős number of the 45 Nobel laureates in Physiology or Medicine between 1991 and 2010 is 5.5; it is 6.0 for the 1991–2000 cohort and 5.1 for the 2001–2010 cohort. For comparison, the average Erdős number of the 13 Nobel laureates in Physics between 1991 and 2010 listed by the Erdős Number Project (http://www.oakland.edu/enp/erdpaths/; accessed June 29, 2014) is 5.5, while the average Erdős number of mathematicians as of Grossman (2005) was 4.7. The average Erdős number of the 13 other Nobel laureates considered here is significantly lower than that of the 1991–2010 laureates, at 4.5.
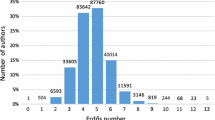
That the 1991–2010 Nobel laureates in Physiology or Medicine form a closely-connected cluster is already suggested by their average \(d_{l}\) of 2.0. The structure of this cluster becomes evident in Fig. 14, which shows the average upper-limit distances from Nobel laureates in Physiology or Medicine, by award year, to some researcher in the informal “center” of biomedicine defined by Fig. 3. The 20-year mean upper-limit distance of Nobel laureates from this subgraph is only 2.7 co-authorship steps, considerably smaller that the average co-authorship distance of 4.6 between authors of papers listed in Medline, an index representing biomedicine broadly, between 1995 and 1999 (Newman 2001). The large-distance outliers in Fig. 14 are of interest: the Nobel prizes of 1991, 1998, 2003 and 2009 were all awarded for work in areas relatively distant from the core area of molecular genetics represented by Fig. 3.
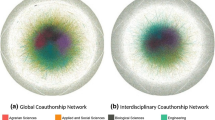
The specializations of the Nobel laureates considered here vary widely within the broad domain of biomedicine. Direct co-authorship connections between Nobel laureates in different Klavans and Boyack (2009) consensus disciplines are shown in Fig. 15a; at most two-step (i.e. average \(d_{l}\)) connections between laureates in different Klavans and Boyack disciplines are shown in Fig. 15b. These graphs show that, at the level of Nobel laureates, the Klavans and Boyack consensus disciplines of Biology, Biochemistry, Infectious Disease, Medical Specialties and Neuroscience are connected even at a co-authorship distance of one, and that all but Biochemistry form a complete graph at length two. The maximum co-authorship distances from Nobel laureates specialized in Biochemistry to laureates in Medical Specialties or Infectious Disease are three and four, respectively, in this sample. These numbers can again be compared with the average distance of 4.6 between authors of papers listed in Medline between 1995 and 1999 (Newman 2001). Newman (2001) reported lower within-discipline clustering in biomedicine than in physics or computer science; the present results are consistent with this observation.
The 19 cross-disciplinary brokers identified here represent 11 distinct disciplines or subdisciplines and eight Klavans and Boyack consensus disciplines: Biology (evolutionary), Computer Science (including electrical engineering), Engineering, Humanities, Mathematics, Physics, Psychology and Social Sciences. Seven (37 %) of the identified brokers are physicists or have published in physics; five (26 %) are mathematicians or have published in mathematics. Twenty-nine (64 %) of the 1991–2010 Nobel laureates are either closest to physics, or are as close to physics as they are to any other non-bioscience discipline, while 26 (58 %) are either closest to mathematics, or as close to mathematics as to any other non-bioscience discipline. The distributions of upper-limit distances from physics and mathematics for all 45 Nobel laureates in Physiology or Medicine between 1991 and 2010 are shown in Fig. 16; only two (4 %), Louis Ignarro and Ferid Murad, are more than five co-authorship steps from the boundaries of both of these disciplines. As noted earlier, the search procedure employed here may be biased toward brokers who have published research in bioinformatics, an inter-disciplinary specialty that attracted both physicists and mathematicians to biology and that played a key enabling role in the human genome project (e.g. Fields 2014b). The upper-limit distance results shown in Fig. 16 cannot, however, be considered artifacts of such a bias.
The co-authorship diameter of physics between 1995 and 1999 was 20 while the average co-authorship distance between physics researchers was 5.9 (Newman 2001); hence Nobel laureates in Physiology or Medicine were, on average, closer to physics during this period than physicists were, again on average, to each other. The average co-authorship distance between mathematicians between 1940 and 1999 was 7.8 (Grossman 2005); assuming that this number did not decrease by half for the partially overlapping interval 1991–2010, Nobel laureates in Physiology or Medicine were, on average, also closer to mathematics during this period than mathematicians were to each other. It is also interesting that a consideration of co-authorship reveals the significant impact of mathematics on late twentieth century biomedicine well before it is evident from citation analysis; Chen et al. (2014) show citations from the biomedical literature to the mathematics literature beginning in 1993 (their Fig. 8), well after the landmark mathematically-oriented papers of, e.g. Schneider et al. (1986) or Lander and Waterman (1988).
The boundaries separating the biomedical sciences from physics and mathematics described here have an interesting asymmetry: while the mathematicians described all have low Erdős numbers and hence are closely connected within mathematics, the physicists are in some cases far closer to each other via collaborations with biologists than they are within physics. For example, James Yorke and the present author share two co-authors who are biologists (Stephen Mount and Ewan Kirkness), but their apparent closest connection within physics requires eight co-authorship steps (data not shown). Albert-László Barabási is similarly separated from the present author by only two steps in biology as shown in Fig. 10, but by ten steps in physics and network theory. For comparison, Yorke and Barabási are separated by four steps, via the present author, within biology and also by four steps within physics and network theory. The co-authorship distance from H. G. Schuster to the present author is five within biology, but six within physics. On the other hand, William Sleator, Jr. is separated from the present author by six co-authorship steps in biology and five within physics; Sleator is separated from H. G. Schuster by at most eight steps in biology and at most seven steps in physics. Such shortcuts across biology are not restricted to physicists. Evolutionary biologist Stephen J. Gould is only four co-authorship steps from the present author in Fig. 6 and hence five steps from Carol Bult. He is six steps from Bult on a path within evolutionary biology, and five steps from the present author on a path traversing evolutionary biology, philosophy and cognitive science. Philosopher Barry Smith is only two steps from the present author in Fig. 2, but eight steps away on a path traversing computer science and artificial intelligence.
The criteria for co-authorship used here are time-independent; hence the present analysis is insensitive to the relative timing of the identified brokers’ publications in different disciplines. It does not, in particular, distinguish brokers who exhibit field mobility from those who work in multiple fields in parallel over an extended period, or those who cross disciplinary boundaries only briefly. Of the brokers considered here, William Sleator, Jr. is perhaps the clearest example of field mobility, having moved from nuclear physics to muscle physiology in 1948. Eric Lander’s publications in computer science, economics and mathematics were brief, early-career excursions into other disciplines; his focus has remained on molecular genetics and genomics since then. James Yorke, on the other hand, pursued biological questions extensively in the 1970s and again in the 2000s, all while continuing work in physics. H. G. Schuster worked on biological problems throughout the 1990s, again in parallel with work in physics. Albert-László Barabási’s work in biology similarly parallels work in both the fundamentals of network theory and other areas to which the theory may be applied. The career of the present author started in physics, moved to cognitive science and bioinformatics in parallel, and has returned to physics more recently. Field mobility is not, therefore, a sufficient explanation of the cross-disciplinary interactions described here, and so is not a sufficient explanation for the exceptional closeness of biomedical Nobel laureates to the boundaries of their discipline.
Conclusion
Nobel laureates provide an elite and tractable sample with which to investigate interdisciplinarity at high resolution. It has been shown here that Nobel laureates in Physiology or Medicine between 1991 and 2010 are close, as measured by co-authorship distance, not only to the disciplinary boundaries within the broad area of biomedicine, but also to the boundaries between biomedicine and a wide range of non-biomedical disciplines. On average, these 45 Nobel laureates are less than three co-authorship steps from some non-biomedical discipline; 96 % are less than five steps from at least one physicist or mathematician. As biomedical scientists were separated from each other by an average of 4.6 co-authorship steps during the first half of this period (Newman 2001), the 1991–2010 Nobel laureates in biomedicine appear to be closer to the edge of their discipline than they are to most of their biomedical colleagues. To the extent that these Nobel laureates, together with the major institute directors and other prominent scientists with whom they collaborate, form the “center” of biomedicine between 1991 and 2010, this center is close to the edge of biomedicine in co-authorship terms. Biomedicine cannot, therefore, be regarded as a co-authorship sphere.
This finding for Nobel laureates in biomedicine is consistent with the observation of short co-authorship paths, all of which include either Nobel laureates or other prominent scientists, traversing many distinct Klavans and Boyack (2009) consensus disciplines (Fields 2014a). It thus supports the suggestion of Fields (2014a) that the co-authorship centers of many if not most scientific disciplines, as least as defined by the presence of Nobel laureates and other scientific and political leaders, may be close to multiple disciplinary boundaries. If this is the case, scientific disciplines in general are not co-authorship spheres.
One can, clearly, ask whether the present results are not due to ascertainment bias. The 1991–2010 timeframe considered here encompasses all but the earliest planning stages of the Human Genome Project as well as the subsequent rise of “systems biology.” The first decade of this period was the “Decade of the Brain” in the United States, an initial major effort in the neurosciences, while the second decade saw major progress in the neurosciences internationally. As genomics, systems biology and neuroscience all tend to generate large data sets that require significant computational analysis, it is perhaps not surprising that physicists, mathematicians, computer scientists and other researchers from outside biomedicine have been heavily involved in these areas. Indeed, the development of modern, biochemically-oriented medicine from the 1920s to the 1950s and the emergence of molecular biology in the 1960s and 1970s may indicate that biomedicine has been in such a transitional state since soon after Nobel prizes and the modern research university were introduced; the citation analysis of Chen et al. (2014) supports this view. Nobel laureates in Physiology and Medicine may, therefore, be an intrinsically biased sample. While the low Erdős numbers of Nobel laureates in other disciplines, the fact that some Nobel laureates in other disciplines are also cross-disciplinary brokers, and preliminary results suggesting that Nobel laureates in Physics are also “close to the edge” during the relevant timeframe (Fields, in prep.) all support biomedicine being typical instead of exceptional, only further high-resolution studies can settle this question.
Notes
Data from http://www.oakland.edu/enp/erdpaths/. Accessed June 28, 2014.
See http://chrisfieldsresearch.com/erdos.htm for supporting data.
References
Aguillo, I. F. (2012). Is Google Scholar useful for bibliometrics? A webometric analysis. Scientometrics, 91, 343–351.
Barabási, A.-L., & Albert, R. (1999). Emergence of scaling in random networks. Science, 286, 509–512.
Barabási, A. L., Jeong, H., Néda, Z., Ravasz, E., Schubert, A., & Vicsek, T. (2002). Evolution of the social network of scientific collaborations. Physica A, 311, 590–614.
Borgatti, S. P., & Everett, M. G. (2006). A graph-theoretic perspective on centrality. Social Networks, 28, 466–484.
Börner, K., Sanyal, S., & Vespignani, A. (2007). Network science. Annual Review of Information Science and Technology, 41, 537–607.
Chen, S., Arsenault, C., Gingras, Y., & Lariviève, V. (2014). Exploring the interdisciplinary evolution of a discipline: The case of Biochemistry and Molecular Biology. Scientometrics (in press). doi:10.1007/s11192-014-1457-6
De Castro, R., & Grossman, J. W. (1999). Famous trails to Paul Erdős. Mathematical Intelligencer, 21(3), 51–53.
Diestel, R. (2010). Graph theory (4th ed.). Berlin: Springer.
Fields, C. (2014a). How small is the center of science? Short cross-disciplinary cycles in co-authorship graphs. Scientometrics (in press). doi:10.1007/s11192-014-1468-3
Fields, C. (2014b). Some effects of the Human Genome Project on the Erdős collaboration graph. Journal of Humanistic Mathematics, 4(2), 3–24.
Freeman, L. C. (1978/79). Centrality in social networks: Conceptual clarification. Social Networks, 1, 215–239.
Grossman, J. W. (2005). Patterns of research in mathematics. Notices of the AMS, 52(1), 35–41.
Harzing, A.-W. (2013). A preliminary test of Google Scholar as a source for citation data: A longitudinal study of Nobel prize winners. Scientometrics, 94, 1057–1075.
Jacobs, J. A., & Frickel, S. (2009). Interdisciplinarity: A critical assessment. Annual Review of Sociology, 35, 43–65.
Klavans, R., & Boyack, K. W. (2009). Toward a consensus map of science. Journal of the American Society for Information Science and Technology, 60, 455–476.
Landherr, A., Friedl, B., & Heidemann, J. (2010). A critical review of centrality measures in complex networks. Business Information Systems Engineering, 6, 371–385.
Newman, M. E. J. (2001). The structure of scientific collaboration networks. Proceedings of the National Academy of Sciences USA, 98, 404–409.
Porter, A. L., & Rafols, I. (2009). Is science becoming more interdisciplinary? Measuring and mapping six research fields over time. Scientometrics, 81, 719–745.
Conflict of interest
The author declares that he has no conflicts of interest relevant to the present work.
Author information
Authors and Affiliations
Corresponding author
Appendices
Appendix 1: References cited in co-authorship graphs or otherwise employed as examples
Adams, M. D., Kerlavage, A. R., Fleischmann, R. D. et al. (85 co-authors) (1995). Initial assessent of human gene diversity and expression patterns based upon 83 million nucleotides of cDNA sequence. Nature 377, Suppl. 3–174.
Adams, M. D., Celniker, S. E., Holt, R. A. et al. (175 co-authors) (2000). The genome sequence of Drosophila melanogaster. Science 287, 2185–2195.
Akil, H., Brenner, S., Kandel, E., Kendler, K. S., King, M.-C., Scolnick, E., Watson, J. D. and Zoghbi, H. Y. (2010). The future of psychiatric research: Genomes and neural circuits. Science 327, 1580–1581.
Albright, T. D., Jessell, T. M., Kandel, E. R. and Posner, M. I. (2000). Neural science: A century of progress and the mysteries that remain. Neuron 25, Suppl. S1–S55.
Alon, N., Asodi, V., Cantor, C., Kasif, S. and Rachlin, J. (2006). Multi-node graphs: A framework for multiplexed biological assays. Journal of Computational Biology 13, 1659–1672.
Alon, N. and Spencer, J. H. (2000). The Probabilistic Method. New York: Wiley Interscience.
Anastassiou, C. A., Perin, R., Markram, H. and Koch, C. (2011). Ephaptic coupling of cortical neurons. Nature Neuroscience 14, 217–223.
Argarana, C. E., Kuntz, I. D., Birken, S., Axel, R. and Cantor, C. R. (1986). Molecular cloning and nucleotide sequence of the streptavidin gene. Nucleic Acids Research 14, 1871–1882.
Arratia, R. and Lander, E. S. (1990). The distribution of clusters in random graphs. Advances in Applied Mathematics 11, 36–48.
Ashton-Rickardt, P. G., Bandeira, A., Delaney, J. R., Van Kaert, L., Pircher, H.-P., Zinkernagel, R. M. and Tonegawa, S. (1994). Evidence for a differential avidity model of T cell selection in the thymus. Cell 76, 651–663.
Austin, C. P., Battey, J. F., Bradley, A. et al. (41 co-authors) (2004). The knockout mouse project. Nature Genetics 36, 921–924.
Bahmanyar, S., Kaplan, D. D., DeLuca, J. G., Giddings, T. H. Jr., O’Toole, E. T., Winey, W., Salmon, E. D., Casey, P. J., Nelson, W. J. and Barth, A. I. M. (2008). \(\beta \)-Catenin is a Nek2 substrate involved in centrosome separation. Genes and Development 22, 91–105.
Baker, P. N., Johnson, I. R., Harvey, P. R., Gowland, P. A. and Mansfield, P. (1994). A three-year follow-up of children imaged in utero with echo-planar magnetic resonance. American Journal of Obstetrics and Gynecology 170, 32–33.
Baldi, P. and Rinott, Y. (1989). On normal approximations of distributions in terms of dependency graphs. Annals of Probability 17, 1646–1650.
Barré-Sinoussi, F., Chermann, J. C., Ray, F. et al. (12 co-authors) (1983). Isolation of a T-lymphotropic retrovirus from a patient at risk for acquired immune deficiency syndrome (AIDS). Science 220, 868–871.
Bavister, B. D., Edwards, R. G. and Steptoe, P. C. (1969). Identification of the midpiece and tail of the spermatazoon during fertilization of human eggs in vivo. Journal of Reproduction and Fertility 20, 159–160.
Bavister, B. D. and Yanagimachi, R. (1997). The effects of sperm extracts and energy sources on the motility and acrosome reaction of hamster spermatozoa in vitro. Biology of Reproduction 16, 228–237.
Berg, P., Baltimore, D., Boyer, H. W., Cohen, S. N., Davis, R. W., Hogness, D. S., Nathans, D., Roblin, R., Watson, J. D., Weissman, S. and Zinder, N. D. (1974). Potential biohazards of recombinant DNA molecules. Science 185, 303.
Blackburn, E., Greider, C. and Szostak, J. (2006). Telomeres and telomerase: The path from maize, Tetrahymena and yeast to human cancer and aging. Nature Medicine 12, 1133–1138.
Blair, J. M., Freier, G., Lampi, E., Sleator, W. Jr. and Williams, J. H. (1948). The angular distributions of the products of the d - d reaction: 1 to 3.5 Mev. Physical Review 74, 1599–1603.
Boehnke, M., Arnheim, N., Li, H. and Collins, F. S. (1989). Fine-structure genetic mapping of human chromosomes using the polymerase chain reaction on single sperm: Experimental design considerations. American Journal of Human Genetics 45, 21–32.
Breeden, L. and Nasmyth, K. (1985). Regulation of the yeast HO gene. Cold Spring Harbor Symposia on Quantitative Biology 50, 643–650.
Broder, S., Collins, J. M., Markham, P. D. et al. (15 co-authors) (1985). Effects of suramin on HTLV-III/LAV infection presenting as Karposi’s sarcoma or AIDS-related complex: Clinical pharmacology and suppression of virus replication in vivo. The Lancet 326, 627–630.
Buck, L. and Axel, R. (1991). A novel multigene family may encode odorant receptors: A molecular basis for odor recognition. Cell 65, 175–187.
Buldyrev, S. V., Barabási, A.-L., Caserta, F., Havlin, S., Stanley, H. E., and Vicsek, T. (1992). Anomalous interface roughening in porous media: Experiment and model. Physical Review A 45, R8313–R8316.
Burette, A., Zabel, U., Weinberg, R. J., Schmidt, H. H. H. W. and Valtschanoff, J. G. (2002). Synaptic localization of nitric oxide synthase and soluble guanylyl cyclase in the hippocampus. Journal of Neuroscience 22, 8961–8970.
Burke, J. M., Belfort, M., Cech, T. R., Davies, R. W., Schweyen, R. J., Shub, D. A., Szostak, J. W. and Tabak, H. F. (1987). Structural conventions for group I introns. Nucleic Acids Research 15, 7217–7221.
Castellucci, V. F., Kandel, E. R., Schwartz, J. H., Wilson, F. D., Nairn, A. C. and Greengard, P. (1980). Intracellular injection of t he catalytic subunit of cyclic AMP-dependent protein kinase simulates facilitation of transmitter release underlying behavioral sensitization in Aplysia. Proceedings of the National Academy of Sciences USA 77, 7492–7496.
Chinwalla, A. T., Cook, L. L., Delehaunty, K. D. et al. (229 co-authors) (2002). Initial sequencing and comparative analysis of the mouse genome. Nature 420, 520–562.
Chalfie, M., Sulston, J. E., White, J. G., Southgate, E., Thomson, J. N. and Brenner, S. (1985). The neural circuit for touch sensitivity in Caenorhabditis elegans. Journal of Neuroscience 5, 956–964.
Coleman, D. E., Berghuis, A. M., Lee, E., Linder, M. E., Gilman, A. G. and Sprang, S. R. (1994). Structures of active conformations of Gi alpha 1 and the mechanism of GTP hydrolysis. Science 265, 1405–1412.
Collins, F. S. and Watson, J. W. (2003). Genetic discrimination: Time to act. Science 302, 745.
Crick, F. R. C. and Koch, C. (1990). Towards a neurobiological theory of consciousness. Seminars in the neurosciences 2, 263–275.
Daigle, D. M., Rossi, L., Berghuis, A. M., Aravind, L., Koonin, E. V. and Brown, E. D. (2002). YjeQ, an essential, conserved, uncharacterized protein from Escherichia coli, is an unusual GTPase with circularly permuted G-Motifs and marked burst kinetics. Biochemistry 41, 11109–11117.
DeYong, M., Findley, R. and Fields, C. (1992). The design, fabrication, and testing of a new VLSI hybrid analog-digital neural processing element. IEEE Transactions on Neural Networks 3, 363–374.
Dietrich, E. and Fields, C. (1996). The role of the frame problem in Fodor’s modularity thesis: A case study of rationalist cognitive science. In: K. M. Ford and Z. W. Pylyshyn (Eds.) The Robot’s Dilemma Revisited. Norwood, NJ: Ablex, 9–24.
Distel, B., Erdmann, R., Gould, S. J. et al. (25 co-authors)(1996). A unified nomenclature for peroxisome biogenesis factors. Journal of Cell Biology 135, 1–3.
Drosophila 12 Genomes Consortium (419 co-authors) (2007). Evolution of genes and genomes on the Drosophila phylogeny. Nature 450, 203–218.
Dupuy, D., Bertin, N., Hidalgo, C. A. et al. (26 co-authors) (2007). Genome-scale analysis of in vivo spatiotemporal promoter activity in Caenorhabditis elegans. Nature Biotechnology 25, 663–668.
Dutcher, S. K. and Hartwell, L. H. (1982). The role of S. cerevisiae cell division cycle genes in nuclear fusion. Genetics 100, 175–184.
Efstratiadis, A., Posakony, J. W., Maniatis, T. et al. (15 co-authors) (1980). The structure and evolution of the human \(\beta \)-globin gene family. Cell 21, 653–668.
Ehrenfeucht, A. and Rozenberg, G. (1990). Partial (set) 2-structures. Acta Informatica 27, 343–368.
Erdős, P. L., Steel, M. A., Szekély, L. and Warnow, T. J. (1999). A few logs suffice to build (almost) all trees (I). Random Structures and Algorithms 14, 153–184.
Evsikov, A. V.,de Vries, W. N., Peaston, A. E. et al. (11 co-authors) (2004). Systems biology of the 2-cell mouse embryo. Cytogenetic and Genome Research 105, 240–250.
Farrell, J. and Lander, E. S. (1989). Competition between and within teams: The Lifeboat Principle. Economics Letters 29, 205–208.
Farris, J. S., Källersjö, M., Kluge, A. G. and Bult, C. (1994). Testing significance of incongruence. Cladistics 10, 315–319.
Fischer, E. H. and Krebs, E. G. (1958). The isolation and crystallization of rabbit skeletal muscle phosphorylase \(b\). Journal of Biological Chemistry 231, 65–72.
Fleischmann, R. D., Adams, M., White, O. et al. (40 co-authors) (1995). Whole genome random shotgun sequencing and assembly of Haemophilus influenzae Rd. Science 269, 496–508.
Fleming, J. T., Squire, M. D., Barnes, T. M. et al. (11 co-authors) (1997). Caenorhabditis elegans levamisole resistance genes lev-1, unc-29, and unc-38 encode functional nicotinic acetylcholine receptor subunits. Journal of Neuroscience 17, 5843–5857.
Förstermann, U., Schmidt, H. H. H. W., Pollock, J. S., Sheng, H., Mitchell, J. A., Warner, T. D., Nakane, M. and Murad, F. (1991). Isoforms of nitric oxide synthase: Characterization and purification from different cell types. Biochemical Pharmacology 42, 1849–1857.
Fox, P. T., Raichle, M. E., Mintun, M. A., and Dence, C. (1988). Nonoxidative glucose consumption during focal physiologic neural activity. Science 241, 462–464.
Freissmuth, M., Casey, P. J. and Gilman, A. G. (1989). G proteins control diverse pathways of transmembrane signaling. FASEB Journal 3, 2125–2131.
Furchgott, R. F., Sleator, W. Jr. and T. de Gubareff, T. (1960). Effects of acetylcholine and epinepherine on the contractile strength and action potential of electrically-driven guinea pig atria. Journal of Pharmacology and Experimental Therapeutics 129, 405–416.
Gallo, R. C., Kalyanaraman, V. S., Samgadharan, M. G. et al. (28 co-authors) (1983). Association of the human type C retrovirus with a subset of adult T-cell cancers. Cancer Research 43, 3892–3899.
Gallo, R. C., Wong-Staal, F., Montagnier, L., Haseltine. A. and Yoshida, M. (1988). HIV/HTLV gene nomenclature. Nature 333, 504.
Gardner, M. J., Tettelin, H., Carucci, D. J. et al. (27 co-authors) (1998). Chromosome 2 sequence of the human malaria parasite Plasmodium falciparum. Science 282, 1126–1132.
Goldfarb, L. G., Brown, P., McCombie, W. R., Goldgaber, D., Swergold, G. D., Wills, P. R., Cervenakova, L., Baron, H., Gibbs C. J. Jr. and Gajdusek, D. C. (1991). Transmissible familial Creutzfeldt-Jakob disease associated with five, seven, and eight extra octapeptide coding repeats in the PRNP gene. Proceedings of the National Academy of Sciences USA 88, 10926–10930.
Gooi, H. C., Feiza, T., Kapadia, A., Knowles, B. B., Solter, D. and Evans, M. J. (1981). Stage-specific embryonic antigen involves \(\alpha 1 \rightarrow 3\) fucosylated type 2 blood group chains. Nature 292, 156–158.
Gould, S. J. and Lewontin, R. C. (1979). The spandrels of San Marco and the Panglossian paradigm: A critique of the adaptationist programme. Proceedings of the Royal Society B 21, 581–598.
Grenon, P. and Smith, B. (2004). SNAP and SPAN: Towards dynamic spatial ontology. Spatial Cognition and Computation 4, 69–104.
Grishok, A., Pasquinelli, A. E., Conte, D., Li, N., Parrish, S., Ha, I., Baillie, D. L., Fire, A., Ruvkun, G. and Mello, C. C. (2001). Genes and mechanisms related to RNA interference regulate expression of the small temporal RNAs that control C. elegans developmental timing. Cell 106, 23–34.
Hamill, O. P., Marty, A., Neher, E., Sakmann, B. and Sigworth, F. J. (1991). Improved patch-clamp techniques for high-resolution current recording from cells and cell-free membrane patches. Pflügers Archiv 391(2), 85–100.
Han, C. J., O’Tuathaigh, C. M., van Trigt, L., Quinn, J. J., Fanselow, M. S., Mongeau, R., Koch, C. and Anderson, D. J. (2003). Trace but not delay fear conditioning requires attention and the anterior cingulate cortex. Proceedings of the National Academy of Sciences USA 100, 13087–13092.
Haseltine, W. A, Kleid, D. G., Panet, A., Rothenberg, E. and Baltimore, D. (1976). Ordered transcription of RNA tumor virus genomes. Journal of Molecular Biology 106, 109–131.
Herr, W., Strum, R. A., Clerc, R. G. et al. (11 co-authors) (1988). The POU domain: A large conserved region in the mammalian pit-1, oct-1, oct-2 and Caenorhabditis elegans unc-86 gene products. Genes and Development 2, 1513–1516.
Hervé, P.-Y., Cox, E. F., Lotfipour, A. K., Mougin, O. E., Bowtell, R. W., Gowland, P. A. and Paus, T. (2011). Structural properties of the corticospinal tract in the human brain: A magnetic resonance imaging study at 7 Tesla. Brain Structure and Function 216, 255–262.
Huang, Y.-Y., Kandel, E. R., Varshavsky, L., Brandont, E. P., Qi, M., Idzerda, R. L., McKnight, G. S. and Bourtchouladz, R. (1995). A genetic test of the effects of mutations in PKA on mossy fiber LTP and its relation to spatial and contextual learning. Cell 83, 1211–1222.
Humpherys, D., Eggan, K., Akutsu, H., Friedman, A., Hochedlinger, K., Yanagimachi, R., Lander, E. S., Golub, T. R. and Jaenisch, R. (2002). Abnormal gene expression in cloned mice derived from embryonic stem cell and cumulus cell nuclei. Proceedings of the National Academy of Sciences USA 99, 12889–12894.
Ignarro, L. J., Fukato, J. M., Griscavage, J. M., Rogers, N. E. and Byrns, R. E. (1993). Oxidation of nitric oxide in aqueous solution to nitrite but not nitrate: Comparison with enzymatically formed nitric oxide from L-arginine. Proceedings of the National Academy of Sciences USA 90, 8103–8107.
Innis, R. B., Cunningham, V. J., Delforge, J. et al. (30 co-authors) (2007). Consensus nomenclature for in vivo imaging of reversibly binding radioligands. Journal of Cerebral Blood Flow & Metabolism 27, 1533–1539.
Irizarry, M. C., Kim, T.-W., McNamara, M., Tanzi, R. E., George, J. M., Clayton, D. F. and Hyman, B. T. (1996). Characterization of the precursor protein of the non-A\(\beta \) component of senile plaques (NACP) in the human central nervous system. Journal of Neuropathology & Experimental Neurology 55, 889–895.
Itti, L., Koch, C. and Niebur, E. (1998). A model of saliency-based visual attention for rapid scene analysis. IEEE Transactions on Pattern Analysis and Machine Intelligence 20, 1254–1259.
Itti, L. and Baldi. P. (2006). Bayesian surprise attracts human attention. Advances in Neural Information Processing Systems 18, 547–554.
Job, D., Fischer, E. H. and Margolis, R. L. (1981). Rapid disassembly of cold-stable microtubules by calmodulin. Proceedings of the National Academy of Sciences USA 78, 4679–4682.
Kandel, E. R. and Schwartz, J. H. (1982). Molecular biology of learning: modulation of transmitter release. Science 218, 433–443.
Karch, F., Weiffenbach, B., Peifer,M., Bender, W., Duncan, I., Celniker, S., Crosby, M. and Lewis, E. B. (1985). The abdominal region of the bithorax complex. Cell 43, 81–96.
Kegeles, L. S., Abi-Darghama, A., Zea-Ponce, Y., Rodenhiser-Hill, J., Mann, J. J., Van Heertum, R. L., Cooper, T. B., Carlsson, A. and Laruelle, M. (2000). Modulation of amphetamine-induced striatal dopamine release by ketamine in humans: Implications for schizophrenia. Biological Psychiatry 48, 627–640.
Kitching, J. E., Batay-Csorba, P. A., Fields, C. A., Ristinen, R. A. and Smith, B. L. (1978). High-spin states in \(^{88,87,86}\)Zr. Nuclear Physics A 302, 159–172.
Klausner, R. D., Fauci, A. S., Corey, L. et al. (24 co-authors) (2003). Enhanced: The need for a global HIV vaccine enterprise. Science 300, 2036–2039.
Lander, E. S., Linton, L. M., Birren, B. et al. (250 coauthors) (2001). Initial sequencing and analysis of the human genome. Nature 409, 860–921.
Lander, E. S., Mesirov, J. P. and Taylor IV, W. (1989). Study of protein sequence comparison metrics on the Connection Machine CM-2. Journal of Supercomputing 3(4), 255–269.
Lander, E. S. and Waterman, M. S. (1988). Genomic mapping by fingerprinting random clones: A mathematical analysis. Genomics 2, 231–239.
Lee, J.-M., Ramos, E. M., Lee, J.-H. et al. (150 coauthors) (2012). CAG repeat expansion in Huntington disease determines age at onset in a fully dominant fashion. Neurology 78, 690–695.
Li, J. B., Gerdes, J. M., Haycraft, C. J. et al. (21 co-authors) (2004). Comparative genomics identifies a flagellar and basal body proteome that includes the BBS5 human disease gene. Cell 117, 541–552.
Li, Z. and Itti, L. (2011). Saliency and gist features for target detection in satellite images. IEEE Transactions on Image Processing 20, 2017–2029.
Malumbres, M., Harlow, E., Hunt, T., Hunter, T., Lahti, J. M., Manning, G., Morgan, D. O., Tsai, L.-H., and Wolgemuth, D. J. (2009). Cyclin-dependent kinases: a family portrait. Nature Cell Biology 11, 1275–1276.
Manley, J. L., Fire, A., Cano, A., Sharp, P. A. and Gefter, M. L. (1980). DNA-dependent transcription of adenovirus genes in a soluble whole-cell extract. Proceedings of the National Academy of Sciences USA 77, 3855–3859.
Manolio, T. A., Collins, F. S., Cox, N. J. et al. (27 co-authors) (2009). Finding the missing heritability of complex diseases. Nature 461, 747–753.
Mansfield, P. and Grannell, P. K. (1973). NMR ‘diffraction’ in solids? Journal of Physics C: Solid State Physics 6, L422–L426.
Markram, H., Lübke, J., Frotscher, M. and Sakmann, B. (1997). Regulation of synaptic efficacy by coincidence of postsynaptic APs and EPSPs. Science 275, 213–215.
Marshall, B. J. and Warren, J. R. (1984). Unidentified curved Bacilli in the stomach of patients with gastritis and peptic ulceration. The Lancet 323, 1311–1315.
Mazziota, J., Toga, A., Evans, A. et al. (27 co-authors) (2001). A probabilistic atlas and reference system for the human brain: International Consortium for Brain Mapping (ICBM). Philosophical Transactions of the Royal Society B 356, 1293–1322.
McCombie, W. R., Martin-Gallardo, A., Gocayne, J. D. et al. (21 co-authors) (1992). Expressed genes, Alu repeats and polymorphisms in cosmids sequenced from chromosome 4p16.3. Nature Genetics 1, 348–353.
McHugh, T. J., Jones, M. W., Quinn, J. J., Balthasar, N., Coppari, R., Elmquist, J. K., Lowell, B. B., Fanselow, M. S., Wilson, M. A. and Tonegawa, S. (2007) Science 317, 94–99.
Meffert, M. K., Chang, L. M., Wiltgen, B. J., Fanselow, M. S. and Baltimore, D. (2003). NF-\(\kappa \)B functions in synaptic signaling and behavior. Nature Neuroscience 6, 1072–1078.
Meral, F. C., Royston, T. J. and Magin, R. (2010). Fractional calculus in viscoelasticity: An experimental study. Communications in Nonlinear Science and Numerical Simulation 15, 939–945.
Mollinari, C., Kleman, J.-P., Jiang, W., Schoehn, G., Hunter, T. and Margolis, R. L. (2002). PRC1 is a microtubule binding and bundling protein essential to maintain the mitotic spindle midzone. Journal of Cell Biology 157, 1175–1186.
Montell, C. and Rubin, G. (1989). Molecular characterization of the Drosophila trp locus: A putative integral membrane protein required for phototransduction. Neuron 2, 1313–1323.
Montell, C., Birnbaumer, L. and Flockerzi, V. (2002). The TRP Channels, A remarkably functional family. Cell 108, 595–598.
Mount, S. M., Burks, C., Herts, G., Stormo, G. D., White, O. and Fields, C. (1992). Splicing signals in Drosophila: Intron size, information content, and consensus sequences. Nucleic Acids Research 20, 4255–4262.
Mulligan, K., Simons, P. and Smith, B. (1984). Truth-makers. Philosophy and Phenomenological Research 44, 287–321.
Myers, K. M., Ressler, K. J. and Davis, M. (2006). Different mechanisms of fear extinction dependent on length of time since fear acquisition. Learning & Memory 13, 216–223.
Nasmyth, K. and Hunt, T. (1993). Cell cycle: Dams and sluices. Nature 366, 634–635.
Natale, D. A., Arighi, C. N., Barker, W. C., Blake, J. A., Bult, C. J., Caudy, M., Drabkin, H. J., D’Eustachio, P., Evsikov, A. V., Huang, H., Nchoutmboube, J., Roberts, N. V., Smith, B., Zhang, J. and Wu, C. H. (2011). The Protein Ontology: A structured representation of protein forms and complexes. Nucleic Acids Research 39 (Suppl. 1), D539–D545.
Newcomer, J. W., Selke, G., Melson, A. K., Hershey, T., Craft, S., Richards, K. and Alderson, A. L. (1999). Decreased memory performance in healthy humans induced by stress-level cortisol treatment. Archives of General Psychiatry 56, 527–533.
Niebur, E., Schuster, H. G., Kammen, D. M. and Koch, C. (1991). Oscillator-phase coupling for different two-dimensional network connectivities. Physical Review A 44, 6895–6905.
Nurse, P., Masui, Y. and Hartwell, L. (1998). Understanding the cell cycle. Nature Medicine 4, 1103–1106.
Nurse, P., Thuriaux, P. and Nasmyth, K. (1976). Genetic control of the cell division cycle in the fission yeast Schizosaccharomyces pombe. Molecular and General Genetics 146, 167–178.
Nüsslein-Volhard, C. and Wieschaus, E. (1980). Mutations affecting segment number and polarity in Drosophila. Nature 287, 795–801.
Oesch, B., Westaway, D., Wälchli, M., McKinley, M. P., Kent, S. B. H., Aebersold, R., Barry, R. A., Tempst, P., Teplow, D. B., Hood, L. E., Prusiner, S. B. and Weissmann, C. (1985). A cellular gene encodes scrapie PrP 27-30 protein. Cell 40, 735–746.
Ogg, S., Paradis, S., Gottlieb, S., Patterson, G. I., Lee, L., Tissenbaum, H. A. and Ruvkun, G. (1997). The Fork head transcription factor DAF-16 transduces insulin-like metabolic and longevity signals in C. elegans. Nature 389, 994–999.
Olney, J. W., Fuller, T. and de Gubareff, T. (1979). Acute dendrotoxic changes in the hippocampus of kainate treated rats. Brain Research 176, 91–100.
Olney, J. W., Newcomer, J. W. and Farber, N. B. (1999). NMDA receptor hypofunction model of schizophrenia. Journal of Psychiatric Research 33, 523–533.
O’Toole, E. T., Giddings, T. H., McIntosh, J. R. and Dutcher, S. K. (2003). Three-dimensional organization of basal bodies from wild-type and \(\delta \)-tubulin deletion strains of Chlamydomonas reinhardtii. Molecular Biology of the Cell 14, 2999–3012.
Ott, E., C. Grebogi and Yorke, J. A. (1990). Controlling chaos. Physical Review Letters 64, 1996–1199.
Padgett, R. A., Mount, S. M., Steitz, J. A. and Sharp, P. A. (1983). Splicing of messenger RNA precursors is inhibited by antisera to small nuclear ribonucleoprotein. Cell 35, 101–107.
Peck, T. L., Magin, R. L. and Lauterbur, P. C. (1995). Design and analysis of microcoils for NMR microscopy. Journal of Magnetic Resonance, Series B 108, 114–124.
Peifer, M. and Wieschaus, E. (1990). The segment polarity gene armadillo encodes a functionally modular protein that is the Drosophila homolog of human plakoglobin. Cell 63, 1167–1178.
Petersen, S. E., Fox, P. T., Posner, M. I., Mintun, M. and Raichle, M. E. (1988). Positron emission tomographic studies of the cortical anatomy of single-word processing. Nature 331, 585–589.
Petridou, N., Schäfer, A., Gowland, P. and Bowtell, R. (2009). Phase vs. magnitude information in functional magnetic resonance imaging time series: Toward understanding the noise. Magnetic Resonance Imaging 27, 1046–1057.
Pohl, S. L., Birnbaumer, L. and Rodbell, M. (1971). The Glucagon-sensitive Adenyl Cyclase System in Plasma Membranes of Rat Liver I: Properties. Journal of Biological Chemistry 246, 1849–1856.
Posner, M. I., Snyder, C. R. and Davidson, B. J. (1980). Attention and the detection of signals. Journal of Experimental Psychology: General 109, 160–174.
Ray, R. P., Arora, K., Nüsslein-Volhard, C. and Gelbart, W. M. (1991). The control of cell fate along the dorsal-ventral axis of the Drosophila embryo. Development 113, 35–54.
Ressler, K. J., Sullivan, S. L. and Buck, L. R. (1994). Information coding in the olfactory system: Evidence for a stereotyped and highly organized epitope map in the olfactory bulb. Cell 79, 1245–1255.
Roberts, R. and Szostak, J. (1997). RNA-peptide fusions for the in vitro selection of peptides and proteins. Proceeding of the National Academy of Sciences USA 94, 12297–12302.
Rogozin, I. B., Makarova, K. S., Murvai, J., Czabarka, E., Wolf, Y. I., Tatusov, R. L., Székely, L. A. and Koonin, E. V. (2002). Connected gene neighborhoods in prokaryotic genomes. Nucleic Acids Research 30, 2212–2223.
Rosen, D. R., Siddique, T., Patterson, D. et al. (33 co-authors) (1993). Mutations in Cu/Zn superoxide dismutase gene are associated with familial amyotrophic lateral sclerosis. Nature 362, 59–62.
Rubin, G. M. and Lewis, E. B. (2000). A brief history of Drosophila’s contributions to genome research. Science 287, 2216–2218.
Ruvkun, G., Ambros, V., Coulson, A., Waterston, R., Sulston, J. E. and Horvitz, H. R. (1989). Molecular genetics of the Caenorhabditis elegans heterochronic gene lin-14. Genetics 121, 501–516.
Saiki, R. K., Scharf, S., Faloona, F., Mullis, K. B., Horn, G. T., Erlich, H. A. and Arnheim, N. (1985). Enzymatic amplification of \(\beta \)-globin genomic sequences and restriction site analysis for diagnosis of sickle-cell anemia. Science 230, 1350–1354.
Scheller, R. H., Jackson, J. F., MacAllister, L. B., Schwartz, J. H., Kandel, E. R. and Axel, R. (1982). A family of genes that codes for ELH, a neuropeptide eliciting a stereotyped pattern of behavior in Aplysia. Cell 28, 707–719.
Schein, J. E., Marra, M. A., Benian, G. M., Fields, C. and Baille, D. L. (1993). The use of deficiencies to determine essential gene content in the let-56 - unc-22 region of Caenorhabditis elegans. Genome 36, 1148–1156.
Schmidt, H. H. H. W., Hofmann, H., Schindler, U., Shutenko, Z. S., Cunningham, D. D. and Feelisch, M. (1996). No \(\cdot \)NO from NO synthase. Proceeding of the National Academy of Sciences USA 93, 14492–14497.
Schneider, T. D., Stormo, G. D., Gold, L. and Ehrenfeucht, A. (1986). Information content of binding sites on nucleotide sequences. Journal of Molecular Biology 188, 415–431.
Schneider, T. D., Stormo, G. D., Yarus, M. and Gold, L. (1984). Delila system tools. Nucleic Acids Research 12, 129–140.
Schuler, G. D., Boguski, M. S., Stewart, E. A. et al. (104 co-authors) (1996). A gene map of the human genome. Science 274, 540–546.
Schuster, H. G. and Just, W. (2006). Deterministic Chaos: An Introduction. Weinheim: Wiley-VCH.
Scott, J. D., Glaccum, M. B., Zoller, M. J., Uhler, M. D., Helfman, D. M., McKnight, G. S. and Krebs, E. G. (1987). The molecular cloning of a type II regulatory subunit of the cAMP-dependent protein kinase from rat skeletal muscle and mouse brain. Proceeding of the National Academy of Sciences USA 84, 5192–5196.
Shows, T. B., McAlpine, P. J., Boucheix, C. et al. (25 co-authors) (1987). Guidelines for human gene nomenclature. An international system for human gene nomenclature (ISGN, 1987). Cytogenetics and Cell Genetics 46, 11–28.
Singh, H., Sen, R., Baltimore, D. and Sharp, P. A. (1986). A nuclear factor that binds to a conserved sequence motif in transcriptional control elements of immunoglobulin genes. Nature 319, 154–158.
Smith, C. L., Lawrance, S. K., Gillespie, G. A., Cantor, C. R., Weissman, S. M. and Collins, F. S. (1987). Strategies for mapping and cloning macroregions of mammalian genomes. Methods in Enzymology 151, 461–489.
Smith, H. O. and Nathans, D. (1973). A suggested nomenclature for bacterial host modification and restriction systems and their enzymes. Journal of Molecular Biology 81, 419–423.
Swain, R. A., Harris, A. B., Wiener, E. C., Dutka, M. V., Morris, H. D., Theien, E., Konda, S., Engberg, K., Lauterbur, P. C. and Greenough, W. T. (2003). Prolonged exercise induces angiogenesis and increases cerebral blood volume in primary motor cortex of the rat. Neuroscience 117, 1037–1046.
Tolias, K. F., Bikoff, J. B., Burette, A., Paradis, S., Harrar, D., Tavazoie, S., Weinberg, R. J. and Greenberg, M. E. (2005). The Rac1-GEF Tiam1 couples the NMDA receptor to the activity-dependent development of dendritic arbors and spines. Neuron 45, 525–538.
Tsien, J. Z. Chen, D. F., Gerber, D., Tom, C., Mercer, E. H., Anderson, D. J., Mayford, M., Kandel, E. R. and Tonegawa, S. (1996). Subregion- and cell type restricted gene knockout in mouse brain. Cell 87, 1317–1326.
Uhlmann, F., Wernic, D., Poupart, M.-A., Koonin, E. V. and K. Nasmyth, K. (2000). Cleavage of cohesin by the CD clan protease separin triggers anaphase in Yeast. Cell 103, 375–386.
Van Kaert, L., Ashton-Rickardt, P. G., Eichelberger, M., Gaczynska, M., Nagashima, K., Rock, K. L., Goldberg, A. L., Doherty, P. C. and S. Tonegawa, S. (1994). Altered peptidase and viral-specific T cell response in LMP2 mutant mice. Immunity 1, 533–541.
Venter, J. C., Adams, M. D., Myers, E. W. et al. (274 co-authors) (2001). The sequence of the human genome. Science 291, 1304–1351.
Wallace, C. S., Withers, G. S., Weiler, I. J., George, J. M., Clayton, D. F. and Greenough, W. T. (1995). Correspondence between sites of NGFI-A induction and sites of morphological plasticity following exposure to environmental complexity. Molecular Brain Research 32, 211–220.
Watson, J. D. and Crick, F. R. C. (1953). Molecular structure of nucleic acids. Nature 171, 737–738.
Wigler, M., Sweet, R., Sim, G. K., Wold, B., Pellicer, A., Lacy, E., Maniatis, T., Silverstein, S. and Axel, R. (1979). Transformation of mammalian cells with genes from procaryotes and eucaryotes. Cell 16, 777–785.
Wink, D. A., Feelisch, M., Fukuto, J., Chistodoulou, D., Jourd’heuil, D., Grisham, M. B., Vodovotz, Y., Cook, J. A., Krishna, M., DeGraff, W. G. Kim, S.-M., Gamson, J. and Mitchell, J. B. (1998). The cytotoxicity of nitroxyl: Possible implications for the pathophysiological role of NO. Archives of Biochemistry and Biophysics 351, 66–74.
Wood, V., Gwilliam, R., Rajandream, M.-A. et al. (133 co-authors) (2002). The genome sequence of Schizosaccharomyces pombe. Nature 415, 871–880.
Xue, D., Finney, M., Ruvkun, G. and Chalfie, M. (1992). Regulation of the mec-3 gene by the C. elegans homeoproteins UNC-86 and MEC-3. EMBO Journal 11, 4969–4979.
Yamada, T., Searle, J. G., Ahnen, D. et al. (51 co-authors) (1994). Helicobacter pylori in peptic ulcer disease. Journal of the American Medical Association 272, 65–69.
Yarus, M., Cline, S. W, Wier, P., Breeden, L. and Thompson, R. C. (1986). Actions of the anticodon arm in translation on the phenotypes of RNA mutants. Journal of Molecular Biology 192, 235–255.
Zinder, N. and Lederberg, J. (1952). Genetic exchange in Salmonella. Journal of Bacteriology 64, 679–699.
Zinkernagel, R. M. and Doherty, P. C. (1979). MHC-restricted cytotoxic T cells: studies on the biological role of polymorphic major transplantation antigens determining T-cell restriction-specificity, function, and responsiveness. Advances in Immunology 27, 51–177.
Zoller, M. J., Kerlavage, A. R. and Taylor, S. S. (1979). Structural comparisons of cAMP-dependent protein kinases I and II from porcine skeletal muscle. Journal of Biological Chemistry 254, 2408–2412.
Appendix 2: Tabular results
The following tables summarize the results depicted in Figs. 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12 and 13 and described in the text. Note that the distances reported are in some cases along paths traversing multiple subgraphs (Tables 1, 2, 3).
Rights and permissions
About this article
Cite this article
Fields, C. Close to the edge: co-authorship proximity of Nobel laureates in Physiology or Medicine, 1991–2010, to cross-disciplinary brokers. Scientometrics 103, 267–299 (2015). https://doi.org/10.1007/s11192-015-1526-5
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11192-015-1526-5