Abstract
The PsbL protein is one of three low-molecular-weight subunits identified at the monomer–monomer interface of photosystem II (PSII) [Ferreira et al. (2004) Science 303:1831–1838; Loll et al. (2005) Nature 438:1040–1044]. We have employed site-directed mutagenesis to investigate the role of PsbL in Synechocystis sp. PCC 6803 cells. Truncation of the C-terminus by deleting the last four residues (Tyr-Phe-Phe-Asn) prevented association of PsbL with the CP43-less monomeric sub-complex and therefore blocked PSII assembly resulting in an obligate photoheterotrophic strain. Replacement of these residues with Ala created four photoautotrophic mutants. Compared to wild type, the F37A, F38A, and N39A strains had reduced levels of assembled PSII centers and F37A and F38A cells were readily photodamaged. In contrast, Y36A and Y36F mutants were similar to wild type. However, each of these strains had elevated levels of the CP43-less inactive monomeric complex. Mutations targeting a putative hydrogen bond between Arg-16 and sulfoquinovosyldiacylglycerol resulted in mutants that were also highly susceptible to photodamage. Similarly mutations targeting a conserved Tyr residue (Tyr-20) also destabilized PSII under high light and suggest that Tyr-20–lipid interactions or interactions of Tyr-20 with PsbT influence the ability of PSII to recover from photodamage.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Photosystem II (PSII) is embedded in the thylakoid membrane of cyanobacteria, algae and plants where it catalyses the light-driven oxidation of water and reduction of plastoquinone (Wydrzynski and Satoh 2005). Recent X-ray crystal structures from Thermosynechococcus elongatus have shown PSII to contain at least 20 subunits (Ferreira et al. 2004; Loll et al. 2005). Of these polypeptides, only the D1 and D2 subunits together with cytochrome b 559 (Cyt b 559 ) and PsbI, form the photochemically active PSII core complex (Namba and Satoh 1987; Nixon et al. 2005). However, the minimum protein assembly able to support water splitting also contains the chlorophyll a-binding proteins CP47 and CP43 together with additional low-molecular-weight polypeptides (Eaton-Rye and Putnam-Evans 2005; Bricker and Burnap 2005; Thornton et al. 2005).
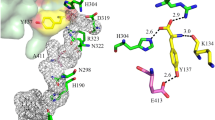
The PsbL protein, encoded in the psbEFLJ operon, is one of ~12 low-molecular-weight proteins identified in the X-ray crystal structures of PSII (Ferreira et al. 2004; Loll et al. 2005). This transmembrane subunit has a molecular weight of ~4.5 kDa with its N-terminus exposed to the cytosol (Fig. 1). Deletion of psbL in Synechocystis sp. PCC 6803 prevented the formation of active PSII complexes producing an obligate photoheterotrophic mutant, and disruption of psbL in Nicotiana tabacum also resulted in plants that were not photoautotrophic (Anbudurai and Pakrasi 1993; Swiatek et al. 2003).
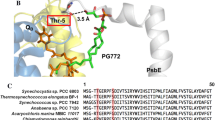
Comparison of PsbL protein sequences. The alignment was performed using Clustal_X default parameters (Thompson et al. 1997). a Alignment of cyanobacterial PsbL sequences obtained from (http://www.ncbi.nlm.nih.gov/sites/entrez). Fully conserved residues (*), highly conserved residues with conservative substitutions (:), and highly conserved residues with non-conservative substitutions (.). The thick solid black line above the alignment represents the α-helical membrane-spanning domain and the thin line represents an extended N-terminal region on the cytosolic side of the membrane. The C-terminus of PsbL does not protrude into the lumen. b Comparison between the cyanobacterial and Viridiplantae consensus sequences. The Viridiplantae consensus sequence is derived form 56 PsbL sequences obtained from http://www.ncbi.nlm.nih.gov/sites/entrez. The consensus sequences for the cyanobacterial alignment in (a) and the Viridiplantae alignment (not shown) indicate residues that are conserved in all of the sequences (upper case) and those conserved in at least 50% of the sequences (lower case)
In vitro studies using isolated PSII from spinach thylakoid membranes found PsbL was required to support electron transfer from the electron donor, diphenyl carbazide to the electron acceptor, 2,6-dichlorophenol-indophenol (Kitamura et al. 1994). Reconstitution studies were undertaken with recombinant PsbL expressed in Escherchia coli (Ozawa et al. 1997). Using this approach to evaluate mutations introduced into PsbL, Toyoshima and co-workers found the C-terminus was crucial for recovering the ability to reduce the primary quinone acceptor, QA and this was supported by electron paramagnetic resonance spectroscopy studies which showed a slowing down of electron transfer from the secondary electron donor, Tyr-Z on the D1 protein, to the oxidized primary donor, P680+ in the PSII reaction center (Ozawa et al. 1997; Hoshida et al. 1997).
In this study, we employed a site-directed mutagenesis strategy to investigate the function of PsbL in Synechocystis sp. PCC 6803. Mutations were introduced into PsbL at the C-terminus and at Arg-16. This latter residue was selected since the corresponding Arg in T. elongatus appears to be hydrogen-bonded to a sulfoquinovosyldiacylglycerol (SQDG) lipid molecule (Loll et al. 2005). In addition, Tyr-20 in the Synechocystis sp. PCC 6803 sequence may also be associated with this lipid or with a monogalactosyldiacylglycerol (MGDG) molecule. Our results demonstrate that mutations at Arg-16, Tyr-20 or at the C-terminus increase susceptibility to photodamage and, in addition, we show the C-terminus is required for the association of PsbL with the CP43-less PSII assembly intermediate and is thus required for the biogenesis of active PSII complexes.
Materials and methods
Growth of Synechocystis sp. PCC 6803
The glucose-tolerant strain of Synechocystis sp. PCC 6803 (Williams 1988) was grown in BG-11 media under constant illumination at 25 μE m−2 s−1 at 30°C. All cultures were maintained on solid BG-11 plates in the presence of 5 mM glucose, 20 μM atrazine, 10 mM TES–NaOH (pH 8.2), 0.3% sodium thiosulfate, and appropriate antibiotics (Eaton-Rye 2004). The liquid cultures were grown mixotrophically in the presence of 5 mM glucose and appropriate antibiotics. The antibiotics used were chloramphenicol at 15 μg/ml and kanamycin at 25 μg/ml. Photoautotrophic growth experiments were carried out as described in (Eaton-Rye 2004).
Construction of PsbL mutagenesis system
A 2.1-kb DNA fragment containing the psbEFLJ operon was amplified by PCR using 5′-TGCTCGATCTGTCTTTCCGT-3′ and 5′-ATCCCCGCCCTAATTCTTGA-3′ as the forward and reverse primers, respectively. The product was cloned into pGEM-T Easy (Promega, Madison, WI, USA), verified by sequencing and used to assemble the psbL deletion and mutagenesis constructs.
The psbL deletion strain (ΔpsbL) and mutants carrying mutations in PsbL were created using oligonucleotide-directed mutagenesis. The deletion was made by replacing psbL with a 1.9-kb kanamycin-resistance cassette derived from transposon Tn903 (Oka et al. 1981). The mutagenesis construct was created by inserting a 1.0-kb chloramphenicol-resistance cassette, derived from pBR325, at an NheI site downstream of the psbEFLJ operon (Fig. 2). All mutagenic oligonucleotides for the creation of strains are listed in Table 1. The oligonucleotide-directed mutagenesis was based on the methods described by (Kunkel 1985; Bebenek and Kunkel 1989) with slight modifications (Juntarajumnong et al. 2007). Escherichia coli CJ236 cells were used as the dut ung strain and M13K07 as the helper phage. Phages were collected by centrifugation at 17,000g for 15 min at 4°C. Synechocystis sp. PCC 6803 mutant strains were obtained by transformation as described in Eaton-Rye (2004). The expression level of psbE, psbF, psbL, and psbJ was verified in each mutant by RT-PCR with the primers used in (Bentley et al. 2008).
Deletion mutant system for the introduction of amino acid substitutions and deletions in PsbL. a Restriction map of the psbL region showing: (i) the psbEFLJ operon (shaded) in wild type; (ii) the psbL gene replaced by a ~1.9-kb kanamycin-resistance cassette (KanR) inserted in the psbEFLJ operon of the ∆psbL strain at an introduced HindIII site that is not shown, and (iii) the psbEFLJ operon from the control strain showing the position of a ~1.0-kb chloramphenicol-resistance cassette (CamR) downstream of the operon. b Southern blot of wild type (lane 1), ΔpsbL (lane 2), and control (lane 3). The Synechocystis sp. PCC 6803 genomic DNA was cut with HindII and probed with a PCR-generated 173 bp intragenic psbE fragment. The probe is shown as a black line under the psbEFLJ operon in (i). c Photoautotrophic growth of wild type (closed circles), ΔpsbL cells (closed triangles), and control cells (open squares) in BG-11 media as measured by the optical density at 730 nm
Production of PsbL antibody
For the production of an antibody against the PsbL protein, the expression vector pET-21a(+) (Novagen, Madison, WI, USA) was used to over-express psbL in E. coli cells. The coding region of psbL was amplified by PCR using oligonucleotides 5′-TTCATATGGCAAGAAAT TCAAACCCAAACCGC-3′ and 5′-CTAGAATTCCGTATTAAAAAGTTTAGTTAAAG-3′. Using these primers, an NdeI site and an EcoRI site were introduced (shown underlined) upstream and downstream of the psbL ORF. An additional TAA stop codon was also introduced six base pairs downstream of the TAA stop codon (also underlined). The expression host, BL21 Star™(DE3)pLysS (Invitrogen, Carlbad, CA, USA), was induced with 1 mM isopropylthio-β-d-galactoside at 37°C. Induced cells were lysed at room temperature using 50 mM Tris–HCl (pH 8.0) and 5 mM EDTA. Inclusion bodies were collected at 12,000g for 30 min at room temperature followed by 25,000g for 10 min. The pellet was solubilized with Laemmli loading buffer and fractionated by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) (Laemmli 1970; Schägger 1994). Over-expressed protein was then eluted by passive diffusion at 37°C by soaking in 1% SDS for 1.5 h with constant motion. Purified proteins were injected into New Zealand white rabbits in the presence of Freud’s adjuvant.
Isolation of thylakoid membranes
Cells were collected at 5000g for 10 min and resuspended in 50 mM HEPES–NaOH (pH 7.2), 10 mM MgCl2, 5 mM CaCl2, and 1 M sucrose in the presence of protease inhibitors (1 mM ε-caproic acid, 1 mM phenylmethylsulfonylfluoride, and 2 mM benzamidine). Cells were broken with 0.1 mm glass beads using a mini bead-beater (Biospec, Bartlesville, OK, USA) for four breaking cycles, where each cycle consisted of 20 s beating at 5000 rpm, followed by 5 min of cooling on ice. Unbroken cells and glass beads were removed by centrifuging twice at 900g at 4°C while thylakoid membranes were collected at 75,000 rpm for 30 min at 4°C (Beckman TLA-100.2 rotor). Pellets were suspended in the suspension buffer without protease inhibitors and stored in liquid nitrogen.
Blue-native PAGE and immunodetection
Blue-native PAGE (BN-PAGE) was performed according to Schägger (1994) and Rokka et al. (2005). Five micrograms of chlorophyll were loaded and native complexes separated using a 5–12% acrylamide gradient gel. Electrophoresis was performed in a Protean II cell (Bio-Rad Laboratories, Hercules, CA, USA) at 4°C and 10 mA for 15 h. On completion, proteins were subjected to immunodetection by electro-blotting to polyvinylidene fluoride membrane and probed with protein-specific antibodies. Binding of antibodies was visualized by chemiluminescence (Whitehead et al. 1979). The secondary antibody was anti-rabbit IgG peroxidase conjugate (Sigma, St Louis, MO, USA).
Herbicide-binding and oxygen evolution assays and western blotting
The relative level of assembled PSII reactions centers was estimated on a chlorophyll basis by employing a [14C]atrazine herbicide-binding assay (Summerfield et al. 2005; Vermaas et al. 1990). Oxygen evolution measurements were carried out using a Clark-type electrode (Hansatech, King’s Lynn, UK) using 1 mM K3Fe(CN)6 and 0.2 mM 2,5-dimethyl-p-benzoquinone (DMBQ) as described in Summerfield et al. (2005). For western blots, two micrograms of chlorophyll were loaded and protein separated according to (Laemmli 1970) or (Schägger 1994) using SDS-PAGE (12% acrylamide). Proteins were then immunodetected by protein-specific antibodies and visualized by chemiluminscence.
Photoinactivation and recovery assays
Cells were incubated at 30°C at a chlorophyll concentration of 10 μg ml−1 in buffered BG-11 media containing 25 mM HEPES–NaOH (pH 7.5). Samples were first subjected to 2 mE m−2 s−1 white light illumination from a Kodak Ektalite 1000 slide projector for 45 min followed by recovery under low light at 30 μE m−2 s−1 for 120 min provided by metal halide bulbs as described in Morgan et al. (1998). Oxygen evolution was measured as described above at 15 min intervals.
Results
PsbL alignment
In the X-ray crystal structure of PSII from Thermosynechococcus elongatus, PsbL consists of a cytosol-exposed hydrophilic region (13 amino acids) and a transmembrane helix (24 amino acids). As shown in Fig. 1, PsbL is highly conserved, especially in the transmembrane helix region, with, for example, E13LNRTSL(Y/F)LG(L/V)L24 (Synechocystis sp. PCC 6803 numbering) present in all cyanobacterial sequences. Similarly, the C-terminal FSSYFFN sequence is highly conserved. To investigate the role of individual residues within these regions, we constructed Synechocystis sp. PCC 6803 strains allowing the introduction of oligonucleotide-directed mutations into PsbL.
Verification of PsbL mutagenesis system
A ∆psbL strain was constructed by deleting the corresponding ORF in the psbEFLJ operon and inserting a 1.9-kb kanamycin-resistance cassette into an introduced HindIII site (Fig. 2a and Table 1). At the same time, a control strain was assembled with a reintroduced copy of psbL and a 1.0-kb chloramphenicol-resistance cassette immediately downstream of the psbEFLJ operon (Fig. 2a).
Figure 2b presents the Southern analysis to verify these strains. Genomic DNA was isolated from wild-type, ΔpsbL and control cells, and digested with HindII. The digested DNA was probed with a 173 bp intragenic psbE fragment and the expected 1.6 kb, 3.3 kb and 2.5 kb hybridization bands were detected (Fig. 2b). As shown in Fig. 2c and Table 2, the ΔpsbL strain was unable to grow photoautotrophically, evolve oxygen, or assemble active PSII reaction centers. However, the control strain was similar to wild type except for the chloramphenicol-resistance phenotype and therefore confirmed that our mutagenesis system could be used for introducing mutated copies of psbL into our ∆psbL mutant.
Influence of the PsbL C-terminus
Previous studies had indicated that the C-terminal residues of PsbL, and in particular Tyr-34 (spinach numbering), were important for Tyr-Z oxidation in isolated PSII complexes from spinach (e.g., Toyoshima et al. 1998). To evaluate the role of the C-terminus of PsbL in Synechocystis sp. PCC 6803, the Y36F and ∆(Y36-N39) mutants were constructed. In Fig. 3a, the photoautotrophic growth of Y36F cells was similar to wild type (cf. Fig. 2c), and oxygen evolution and PSII assembly were essentially unaltered (Table 3); however, the ∆(Y36-N39) strain, which introduced a 4 amino acid truncation of PsbL, was unable to support photosynthesis. To investigate this further BN-PAGE was performed and the PSII complexes probed with antibodies to the D1 and PsbL proteins (Fig. 3b). Replacing Tyr-36 with Phe had little effect on the relative levels of PSII monomers and dimers. In contrast, only the inactive CP43-less monomeric complex was assembled in the ∆(Y36-N39) strain which is similar to the result observed with a psbL disruption mutant (Bentley et al. 2008). In Fig. 3c, trace levels of PsbL could be detected in thylakoid membranes from the ∆(Y36-N39) mutant when a western blot was performed comparing levels of PsbL and the constitutively expressed PsbO protein in isolated thylakoids. However, it is also possible that our PsbL-specific antibody only reacted weakly with the truncated form of the PsbL protein.
Characterization of Δ(Y36-N39) and Y36F strains. a Photoautotrophic growth as measured by the optical density at 730 nm. Symbols are: Δ(Y36-N39) (closed squares) and Y36F (open triangles). b Immunodetection of PSII complexes from thylakoid membranes using antibodies against D1 and PsbL following separation by BN-PAGE. c Immunodetection of PsbL and PsbO from thylakoid membranes following separation by SDS-PAGE
The deletion from Tyr-36 to Asn-39 was created by a point mutation introducing a stop codon (Table 1), and this did not change the expression level of the psbEFLJ operon (data not shown). This suggests that PsbL in the ∆(Y36-N39) mutant cannot be incorporated into PSII complexes. To evaluate if this was a result of the deletion or due to the absence of one or more specific residues, the deleted residues in the ∆(Y36-N39) strain were individually substituted with Ala.
The Y36A, F37A, F38A, and N39A mutants grew photoautotrophically with doubling times of 15–16 h; however, wild type had a doubling time of 14 h (Table 3). Oxygen evolution rates obtained with the Y36A and N39A cells were also similar to the wild-type rate; however, the F37A and F38A strains exhibited rates of 85% and 73%, respectively, when compared with wild-type cells and PSII assembly was reduced (Table 3). We next evaluated the susceptibility of these mutants to high-light-induced stress (Fig. 4a). In wild type and Y36F cells, oxygen evolution initially fell upon exposure to 2 mE m−2 s−1 but then these cells acclimated to the high light conditions. In contrast, the F37A and F38A mutants exhibited pronounced photodamage with oxygen evolution depressed to ~40% and ~30% of the initial rates observed for each strain, respectively. In addition, Y36A and N39A cells showed a less severe response with oxygen evolution reduced to ~70% of the original activity (Fig. 4a). However, all mutants were able to recover under low light at 30 μE m−2 s−1 and exhibited rates of oxygen evolution that exceeded their initial rates before high light exposure. The recovery was observed to occur within 15 min of returning the cells to the low light conditions, and a similar stimulation of oxygen-evolving activity upon transition from high to low light was also seen in the wild type.
Characterization of C-terminal mutants carrying Ala or Phe substitutions. a Susceptibility of different mutants to photodamage and their subsequent recovery under low light. Symbols are: wild type (closed circles); Y36A (close diamonds); Y36F (open triangles); F37A (open diamonds); F38A (closed inverted triangles); N39A (open circles). High light (HL) intensity was 2.0 mE m−2 s−1 and low light (LL) was 0.03 mE m−2 s−1. Oxygen evolution data are the average ± the standard error from three independent experiments. b Immunodetection of PSII complexes from thylakoid membranes using antibodies against D1 and PsbL following separation by BN-PAGE. c Immunodetection of CP43, D1, and PsbL from thylakoid membranes following separation by SDS-PAGE
The composition of PSII native complexes in the Y36A, F37A, F38A, and N39A mutants was also examined by BN-PAGE followed by probing with D1-specific and PsbL-specific antibodies. All mutants displayed an increased level of the inactive CP43-less monomer compared to wild-type cells when probed with the D1 antibody and the level of PsbL in monomeric forms of PSII was reduced (Fig. 4b). However, western analysis showed all mutants acquired levels of D1, CP43 and PsbL in their thylakoid membranes that were comparable to those detected in wild type (Fig. 4c). In the case of D1, processed and partially processed polypeptides were detected in all strains (Satoh and Yamamoto 2007).
Mutations of Arg-16
The X-ray crystal structure from Thermosynechococcus elongatus identified a putative hydrogen-bonding interaction between Arg-14 of PsbL and an SQDG lipid molecule (Loll et al. 2005). The corresponding Arg in Synechocystis sp. PCC 6803 is Arg-16 (Fig. 1). To investigate the role of this residue in Synechocystis sp. PCC 6803, three mutants were created: R16A, R16E, and R16K. Photoautotrophic growth in R16A and R16E cells exhibited a doubling time of 24 h while the R16K strain had a doubling of time of 14 h, which was similar to wild type (Fig. 5a, Table 3). Table 3 also shows that whereas rates of oxygen evolution were reduced to ~88% in the R16A and R16E strains, the rate in the R16K mutant remained similar to wild type. Moreover, PSII assembly remained comparable in wild type and mutants.
Characterization of mutants with substitutions at Arg-16. a Photoautotrophic growth of Synechocystis sp. PCC 6803 strains as measured by the optical density at 730 nm. Symbols are: R16A (open inverted triangles); R16E (closed squares); R16K (closed circles). b Susceptibility of different mutants to photodamage and their subsequent recovery under low light. Symbols are the same as per panel (a). High light (HL) intensity was 2.0 mE m−2 s−1 and low light (LL) was 0.03 mE m−2 s−1. Oxygen evolution data are the average ± the standard error from three independent experiments. c Immunodetection of PsbL from thylakoid membranes following separation by SDS-PAGE. d Immunodetection of PSII complexes from thylakoid membranes using antibodies against D1 and PsbL following separation by BN-PAGE
We again evaluated the sensitivity of cells to high-light-induced stress. The R16K mutant acclimated to the high light as previously observed for wild type and Y36F cells. In contrast, the R16A and R16E strains were highly susceptible to photodamage, and oxygen evolution declined to ~50% and ~30% of their initial rates, respectively, during the high light exposure. However, on returning cells to 30 μE m−2 s−1, the R16A and R16E strains were observed to fully recover with apparently biphasic kinetics.
In Fig. 5c western blotting found similar levels of PsbL in wild type and mutants. However, BN-PAGE in Fig. 5d revealed an increased amount of the CP43-less intermediate in both R16A ands R16E cells when probed with both D1 and PsbL antibodies.
Tyr-20, Leu-24 and Leu25, and Phe-33
Figure 1a shows Tyr-20 as well as Leu-24 and Leu-25 to be highly conserved with Leu-25 also found in all green plant sequences. In addition, Phe-33 is found in the majority of cyanobacterial sequences and all green plant sequences. Hence these residues were elected for further study leading to the creation of three further mutants: Y20A, L24A:L25A, and F33A. As summarized in Table 3, all mutants grew photoautotrophically, assembled similar levels of PSII centers and the Y20A and L24A:L25A strains exhibited oxygen-evolution rates that were comparable to those of the wild type. However, the F33A cells showed impaired oxygen evolution. When tested for their susceptibility to photodamage, Y20A cells showed a dramatic decrease to 10–16% of their initial activity after 45 min under high light. Similarly, the L24A:L25A and F33A strains were susceptible to photodamage but exhibited less severe responses with rates declining by ~50%. All strains were able to recover when returned to 30 μE m−2 s−1. However, the Y20A mutant did not exhibit rates of oxygen evolution beyond the initial rate measured before high light exposure (cf. Fig. 6a with Figs. 4a and 5b).
Characterization of mutants targeting additional conserved residues in the transmembrane domain. a Susceptibility of different mutants to photodamage and their subsequent recovery under low light. Symbols are: Y20A (closed diamonds), L24A:L25A (open squares), and F33A (closed triangles). High light (HL) intensity was 2.0 mE m−2 s−1 and low light (LL) was 0.03 mE m−2 s−1. Oxygen evolution data are the average ± the standard error from three independent experiments. b Immunodetection of PSII complexes from thylakoid membranes using antibodies against D1 and PsbL following separation by BN-PAGE. c Immunodetection of CP43, D1, and PsbL from thylakoid membranes following separation by SDS-PAGE
In addition, BN-PAGE was performed and the separated PSII complexes probed with antibodies to D1 and PsbL. The interaction between PsbL and the inactive CP43-less monomer was destabilized in the L24A:L25A and F33A strains although the levels of active monomers and dimers were apparently not affected (Fig. 6b). However, in the Y20A strain, PsbL was found to be associated with the CP43-less monomer where this complex was elevated relative to the level observed in the wild type. Moreover, western blotting indicated that CP43 levels were reduced in the L24A:L25A and F33A strains whereas the level of D1 and PsbL remained essentially unchanged in the thylakoid membranes of these mutants (Fig. 6c). At present, the explanation for the lower levels of CP43 in L24A:L25A and F33A cells is unknown.
Discussion
The role of the C-terminus of PsbL
Experiments employing isolated CP47/D1/D2/Cyt b 559 /PsbI/PsbW PSII subcomplexes from spinach revealed electron transfer between Tyr-Z and P680+ required addition of PsbL (Hoshida et al. 1997). In these studies, Toyoshima and colleagues demonstrated reconstitution of the CP47/D1/D2/Cyt b 559 /PsbI/PsbW complex with recombinant protein carrying three Leu substitutions at Tyr-34, Phe-35, and Phe-36 (spinach numbering) prevented formation of Tyr-Z+.P680.Pheo−, (where Pheo is the pheophytin that transfers an electron to QA (Kern and Renger 2007)). In particular, an essential role for Tyr-34 was indicated (Toyoshima et al. 1998). These results suggest the C-terminus of PsbL facilitates donor side electron transfer in spinach PSII. However, in the X-ray structure for PSII from Thermosynechococcus elongatus, PsbL has been assigned to the monomer–monomer interface where it is found together with PsbM and PsbT and therefore the C-terminus of PsbL is not located in the vicinity of Tyr-Z or P680 (Ferreira et al. 2004; Loll et al. 2005). We have therefore investigated the role of the C-terminus of PsbL, in intact photosystems in Synechocystis sp. PCC 6803.
Truncation of Tyr-36, Phe-37, Phe-38, and Asn-39, the four C-terminal residues in Synechocystis sp. PCC 6803, prevented the association of PsbL with the CP43-less monomer subcomplex of PSII thus arresting PSII assembly as observed with psbL deletion or inactivation mutants (Suorsa et al. 2004; Bentley et al. 2008). Moreover, replacing Tyr-36 (equivalent to Tyr-34 in spinach, see Fig. 1) with Phe or Ala resulted in mutants with similar phenotypes to wild type (Figs. 3 and 4) and thus a Tyr at this position does not play an essential role in Synechocystis sp. PCC 6803. However, our F37A and F38A mutants were highly susceptible to photodamage.
Notably the equivalent Phe residues in T. elongatus (Phe-35 and Phe-36) are positioned in close proximity to PsbM and PsbT. For example, Phe-36 is in a pocket formed by Leu-8 from PsbM and Glu-2, Thr-5, and Ty-6 of PsbT. We have recently observed inactivation of psbM and psbT in Synechocystis sp. PCC 6803 creates strains that are highly susceptible to photodamage, and the ∆PsbM:∆PsbT double mutant was unable to assemble PSII dimers as judged by BN-PAGE (Bentley et al. 2008). This may suggest that correct anchoring of the C-terminus of PsbL by PsbM and PsbT contributes to dimer stabilization.
In this study, a feature of wild type and mutants following exposure to high light is an enhanced and/or rapid recovery from photodamage when cells are transferred from high light to low light conditions (e.g., Fig. 4a). We have also observed this phenomenon with ∆PsbT and ∆PsbM cells and found the recovery to be sensitive to protein synthesis inhibitors. A possible underlying mechanism for this response is discussed in (Bentley et al. 2008).
Potential protein–lipid interactions
Loll et al. (2005) located one SQDG and 2 MGDG molecules at the PSII monomer–monomer interface in T. elongatus and identified hydrogen bonds between SQDG and Arg-14 of PsbL and between SQDG and Arg-18 of CP47 from the adjacent monomer. In Synechocystis sp. PCC 6803 the equivalent Arg residue in PsbL is Arg-16, and hence the sensitivity of our R16A and R16E strains to high light may arise from disrupting this putative protein–lipid interaction. We have also constructed mutants at the corresponding Arg residue in CP47 (Luo and Eaton-Rye 2008). The CP47 R18A and R18E strains exhibited a less-pronounced susceptibility to photodamage than the PsbL Arg-16 mutants.
The X-ray-derived structural data for T. elongatus also indicates that Tyr-18 on PsbL (Tyr-20 in Synechocystis sp. PCC 6803) may interact with SQDG and an MGDG molecule as well as being in close proximity to Leu-16, Phe-19, and Phe-23 on PsbT. These additional interactions appear to be important for PSII stability as the Y20A strain in our study exhibited the most extensive sensitivity to photodamage of all mutants examined, and this strain also accumulated CP43-less monomers to a level similar to that seen with the C-terminal mutants. At present, the importance of protein–lipid interactions in PSII function has not been fully established, and it should be noted that species-specific variations have been observed (Güler et al. 1996; Minoda et al. 2003; Sakurai 2007a, b).
Leu-24, Leu 25, and Phe-33
Leu-24 and Leu-25 may contribute to a hydrophobic cavity formed between PsbL and D2 into which the isoprenoid tail of QA may protrude. However, our data suggest that these residues contribute to the ability of PsbL to associate with the CP43-less monomeric complex (Fig. 6b). Interestingly, although located more towards the C-terminus, the conserved Phe-33 was also found to influence the binding of PsbL to PSII. The Phe-33 residue is conserved in all other PsbL sequences except in that of Synechococcus sp. PCC 6301 (Sugita et al. 2007). However, the C-terminus of this strain is unique and potentially contains compensating mutations to preserve the function of this region of the transmembrane helix (Fig. 1).
Abbreviations
- BN-PAGE:
-
Blue-native polyacrylamide gel electrophoresis
- DCMU:
-
3-(3,4-dichlorophenyl)-1,1-dimethylurea
- DM:
-
n-dodecyl β-d-maltoside
- DMBQ:
-
2,5-dimethyl-p-benzoquinone
- EDTA:
-
Ethylenediamine tetra-acetic acid (di-sodium salt)
- HEPES:
-
4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid
- Kb:
-
Kilobases
- kDa:
-
Kilodaltons
- OD:
-
Optical density
- PCC:
-
Pasteur Culture Collection
- PCR:
-
Polymerase chain reaction
- PQ:
-
Plastoquinone
- PSII:
-
Photosystem II
- QA :
-
Primary quinone electron acceptor of photosystem II
- SDS:
-
Sodium dodecyl sulfate
- SDS-PAGE:
-
SDS polyacrylamide gel electrophoresis
- TES:
-
2-[tris(hydroxymethyl)methyl]amino-1-ethanesulfonic acid
- Tris:
-
Tris(hydroxymethyl)aminomethane
References
Anbudurai P, Pakrasi H (1993) Mutational analysis of the PsbL protein of photosystem II in the cyanobacterium Synechocystis sp. PCC 6803. Z Naturforsch 48:267–274
Bebenek K, Kunkel T (1989) The use of native T7 DNA polymerase for site-directed mutagenesis. Nucleic Acids Res 17:5408. doi:10.1093/nar/17.13.5408
Bentley FK, Luo H, Dilbeck P, Burnap RL, Eaton-Rye JJ (2008) Effects of inactivating psbM and psbT on photodamage and assembly of photosystem II in Synechocystis sp. PCC 6803. Biochemistry. doi:10.1021/bi800804h
Bricker TM, Burnap RL (2005) The extrinsic proteins of photosystem II. In: Wydrzynski TJ, Satoh K (eds) Photosystem II: the light-driven water:plastoquinone oxidoreductase, advances in photosynthesis and respiration, vol 22. Springer, Dordrecht, pp 95–120
Eaton-Rye JJ (2004) The construction of gene knockouts in the cyanobacterium Synechocystis sp. PCC 6803. In: Carpentier R (ed) Methods of molecular biology, photosynthesis research protocols. Humana Press, Totowa, pp 309–324
Eaton-Rye JJ, Putnam-Evans C (2005) The CP47 and CP43 core antenna components. In: Wydrzynski TJ, Satoh K (eds) Photosystem II: the light-driven water:plastoquinone oxidoreductase, advances in photosynthesis and respiration, vol 22. Springer, Dordrecht, pp 45–70
Ferreira KN, Iverson TM, Maghlaoui K, Barber J, Iwata S (2004) Architecture of the photosynthetic oxygen-evolving center. Science 303:1831–1838. doi:10.1126/science.1093087
Güler S, Seeliger A, Hartel H, Renger C, Benning C (1996) A null mutant of Synechococcus sp. PCC7942 deficient in the sulfolipid sulfoquinovosyldiacylglycerol. J Biol Chem 271:7501–7507. doi:10.1074/jbc.271.13.7501
Hoshida H, Sugiyama R, Nakano Y, Shiina T, Toyoshima Y (1997) Electron paramagnetic resonance and mutational analysis revealed the involvement of photosystem II-L subunit in the oxidation step of Tyr-Z by P680 + to form the Tyr-Z+P680Pheo− state in photosystem II. Biochemistry 36:12053–12061. doi:10.1021/bi9710885
Juntarajumnong W, Hirani TA, Simpson JM, Incharoensakdi A, Eaton-Rye JJ (2007) Phosphate sensing in Synechocystis sp. PCC 6803: SphU and the SphS-SphR two-component regulatory system. Arch Microbiol 188:389–402. doi:10.1007/s00203-007-0259-0
Kern J, Renger G (2007) Photosystem II: structure and mechanism of the water:plastoquinone oxidoreductase. Photosynth Res 94:183–202. doi:10.1007/s11120-007-9201-1
Kitamura K, Ozawa S, Shiina T, Toyoshima Y (1994) L protein, encoded by psbL restores normal functioning of the primary quinone acceptor, QA, in isolated D1/D2/CP47/Cytb–559/I photosystem II reaction center core complex. FEBS Lett 354:113–116. doi:10.1016/0014-5793(94)01089-7
Kunkel T (1985) Rapid and efficient site-specific mutagenesis without phenotypic selection. Proc Natl Acad Sci USA 82:488–492. doi:10.1073/pnas.82.2.488
Laemmli U (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 277:680–685. doi:10.1038/227680a0
Loll B, Kern J, Saenger W, Zouni A, Biesiadka J (2005) Towards complete cofactor arrangement in the 3.0 Å resolution structure of photosystem II. Nature 438:1040–1044. doi:10.1038/nature04224
Luo H, Eaton-Rye J (2008) Mutations in CP47 that target putative hydrogen bonds with sulfoquinovosyl-diacylglycerol. In: Allen F, Osmond B, Golbeck J, Gant E (eds) Energy from the sun. Springer, Heidelberg, pp 737–739
Minoda A, Sonoike K, Okada K, Sato N, Tsuzuki M (2003) Decrease in the efficiency of the electron donation to tyrosine Z of photosystem II in an SQDG-deficient mutant of Chlamydomonas. FEBS Lett 553:109–112. doi:10.1016/S0014-5793(03)00981-5
Morgan TR, Shand JA, Clarke SM, Eaton-Rye JJ (1998) Specific requirements for cytochrome c-550 and the manganese-stabilizing protein in photoautotrophic strains of Synechocystis sp. PCC 6803 with mutations in the domain Gly-351 to Thr-436 of the chlorophyll-binding protein CP47. Biochemistry 37:14437–14449. doi:10.1021/bi980404s
Namba O, Satoh K (1987) Isolation of a photosystem II reaction center consisting of d-1 and d-2 polypeptides and cytochrome b-559. Proc Natl Acad Sci USA 84:109–112. doi:10.1073/pnas.84.1.109
Nixon PJ, Sarcina M, Diner BA (2005) The CP47 and CP43 core antenna components. In: Wydrzynski TJ, Satoh K (eds) Photosystem II: the light-driven water:plastoquinone oxidoreductase, advances in photosynthesis and respiration, vol 22. Springer, Dordrecht, pp 71–94
Oka A, Sugisake H, Takanami M (1981) Nucleotide sequence of the kanamycin resistance transposon Tn903. J Mol Biol 295:1225–1236
Ozawa S, Kobayashi T, Sugiyama R, Hoshida H, Shiina T, Toyoshima Y (1997) Role of PSII-L complex. 1. Over-production of wild-type and mutant versions of PSII-L protein and reconstitution into the PSII core complex. Plant Mol Biol 34:151–161. doi:10.1023/A:1005800909495
Rokka A, Suorsa M, Saleem A, Battchikova N, Aro E-M (2005) Synthesis and assembly of thylakoid protein complexes: multiple assembly steps of photosystem II. Biochem J 388:159–168. doi:10.1042/BJ20042098
Sakurai I, Mizusawa N, Ohashi S, Kobayashi M, Wada H (2007a) Effects of the lack of phosphatidylglycerol on the donor side of photosystem II. Plant Physiol 144:1336–1346. doi:10.1104/pp.107.098731
Sakurai I, Mizusawa N, Wada H, Sato N (2007b) Digalactosyldiacylglycerol is required for stabilization of the oxygen-evolving complex in photosystem II. Plant Physiol 145:1361–1370. doi:10.1104/pp.107.106781
Satoh K, Yamamoto Y (2007) The carboxyl-terminal processing of precursor D1 protein of the photosystem II reaction center. Photosynth Res 94:203–215. doi:10.1007/s11120-007-9191-z
Schägger H (1994) Electrophoretic isolation of membrane-proteins from acrylamide gels. Appl Biochem Biotechnol 48:185–203. doi:10.1007/BF02788741
Sugita C, Ogata K, Shikata M, Jikuya H, Takano J, Furumichi M, Kanehisa M, Omata T, Sugiura M, Sugita M (2007) Complete nucleotide sequence of the freshwater unicellular cyanobacterium Synechococcus elongatus PCC 6301 chromosome: gene content and organization. Photosynth Res 93:55–67. doi:10.1007/s11120-006-9122-4
Summerfield TC, Shand JA, Bentley FK, Eaton-Rye JJ (2005) PsbQ (Sll1638) in Synechocystis sp. PCC 6803 is required for photosystem II activity in specific mutants and in nutrient-limiting conditions. Biochemistry 44:805–815. doi:10.1021/bi048394k
Suorsa M, Regel RE, Paakkarinen V, Battchikova N, Herrmann RG, Aro E-M (2004) Protein assembly of photosystem II and accumulation of subcomplexes in the absence of low molecular mass subunits PsbL and PsbJ. Eur J Biochem 271:96–107. doi:10.1046/j.1432-1033.2003.03906.x
Swiatek M, Regel R, Meurer J, Wanner G, Pakrasi H, Ohad I, Herrmann R (2003) Effects of selective inactivation of individual genes for low-molecular-mass subunits on the assembly of photosystem II, as revealed by chloroplast transformation: the psbEFLJ operon in Nicotiana tabacum. Mol Genet Genomics 268:699–710
Thompson J, Gibson T, Plewniak F, Jeanmougin F, Higgins D (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882. doi:10.1093/nar/25.24.4876
Thornton LE, Roose JL, Pakrasi HB, Ikeuchi M (2005) The low molecular weight proteins of photosystem II. In: Wydrzynski TJ, Satoh K (eds) Photosystem II: the light-driven water:plastoquinone oxidoreductase, advances in photosynthesis and respiration, vol 22. Springer, Dordrecht, pp 121–138
Toyoshima Y, Iwata T, Nakano Y, Hoshida H (1998) Tyr34 in PSII-L protein is essential for oxidation of Tyr-Z in PSII. In: Garab G (ed) Photosynthesis: mechanisms and effects, vol II. Kluwer Academic Publishers, Dordrecht, pp 1383–1386
Vermaas W, Charité J, Shen G (1990) QA binding to D2 contributes to the functional and structural integrity of photosystem-II. Z Naturforsch 45:359–365
Williams J (1988) Construction of specific mutations in photosystem II photosynthetic reaction center by genetic engineering methods in Synechocystis 6803. Methods Enzymol 167:766–778. doi:10.1016/0076-6879(88)67088-1
Whitehead TP, Kricka LJ, Carter TJN, Thorpe GHG (1979) Analytical luminescence: its potential in the clinical laboratory. Clin Chem 25:1531–1546
Wydrzynski TJ, Satoh K (eds) (2005) Photosystem II: the light-driven water:plastoquinone oxidoreductase, advances in photosynthesis and respiration, vol 22. Springer, Dordrecht
Acknowledgment
This work was supported by a grant (UOO309) from the New Zealand Marsden Fund to J.J.E.-R.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Luo, H., Eaton-Rye, J.J. Directed mutagenesis of the transmembrane domain of the PsbL subunit of photosystem II in Synechocystis sp. PCC 6803. Photosynth Res 98, 337–347 (2008). https://doi.org/10.1007/s11120-008-9375-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11120-008-9375-1