Abstract
Winter survival develops efficient tolerance mechanisms in plants by regulating cold-responsive and cold-regulated genes at the transcriptional level. Hence, an insight into the expression would provide the molecular function of these cold responsive genes. In this study, an uncharacterized gene encoding a cold-regulated (Cor413) protein identified from Saccharum spontaneum (wild relative species of sugarcane) low-temperature transcriptome with environmental adaptability is isolated and characterized. The full-length coding region possesses an open reading frame of 642 bp, which encodes a putative polypeptide of 213 amino acids of molecular weight 25.6 kDa and an isoelectric point (Pi) of 9.69. The SsCor413 sequence showed a high similarity to monocot Cor413 proteins comprising a WCOR413 domain. Bioinformatics analysis revealed that Cor413 protein has multispanning transmembrane helices along with highly conserved phosphorylation sites. String analysis suggested that SsCor413 is grouped with LEA and Rab proteins that are involved in freezing tolerance. Gene ontology analysis assigned the protein to terms such as “plasma membrane,” “cold acclimation,” and “response to cold.” Sub-cellular localization experiments of sugarcane callus and onion epidermal cells indicated the nuclear localized expression. Quantitative gene expression analysis indicated that the SsCor413 gene is up-regulated in leaf and root tissues of S. spontaneum under low temperature, salinity, and water deficit stress conditions. These results highlight the potential role of SsCor413 in abiotic stress tolerance, and this gene could be a new candidate for combating multiple stresses in sugarcane.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plants exhibit a maximum growth rate and development at an optimum temperature (Fitter and Hay 1981) and when this temperature gets altered, physiological and molecular changes occur. Low temperature (LT) is one of the major abiotic stress factors that disturb plant growth, regulation, and yield (Chinnusamy et al. 2007). LT stress is known to induce several abnormalities at various organizational levels of the cell and its symptoms include changes in the membrane system, ion homeostasis disturbance, inhibition of photosynthesis, reduction in water and mineral uptake, and induction of oxidative stress.
Tolerance mechanisms such as chilling tolerance and cold acclimation process will be in use during low-temperature stress as a defense response. The chilling tolerance is the ability of a plant to tolerate LT (0–15 °C) without injury or damage (Somerville 1995). Cold acclimation is an enhanced tolerance process (Guy 1990). Cold stress-induced genes are known as cold-regulated genes. The transcription of gene Cor is differentially regulated under LT conditions (Baker et al. 1994; Chinnusamy et al. 2006). The regulation of Cor genes results in cell wall modifications, cold signaling transduction, transcriptional regulation, and molecular changes under low-temperature stress conditions (Thomashow 1998).
There are different types of Cor genes, viz., Cor6.6, Cor15A, Cor47, Cor78, and Cor413 that have been identified in plants. Among the Cor genes, Cor413/Cor413-like gene has been well studied from several plant species, including Triticum aestivum (Breton et al. 2003), Gossypium barbadense (Wang et al. 2007), Phlox subulata (Qu et al. 2015), and Solanum lycopersicum (Ma et al. 2017). The Cor413 is a plant-specific protein family that is involved in the cold acclimation process (Breton et al. 2003). Amino acids of GbCor413 protein exhibits hydrophobic nature (Wang et al. 2007) and is predicted to have many phosphorylation sites (Blom et al. 1999). The Cor413 protein was also predicted to consist of a minimum of four transmembrane domains (Krogh et al. 2001) which are required for targeting the protein toward the plasma membrane and thylakoid membrane (Breton et al. 2003). The sequence of Cor413 has not been predicted for the presence of signal sequence. However, its mechanism and sub-cellular localization studies are far from being fully understood.
Saccharum spontaneum (wild sugarcane) belongs to the Poaceae family and is a tall perennial grass with a deep root system (Amalraj et al. 2008). The genetic background of sugarcane is complex as Saccharum hybrids are highly polyploid and derived from interspecific hybridization between S. officinarum and S. spontaneum, suggesting that each gene has 8–10 copies (Premachandran et al. 2017). S. spontanuem is known to have developed tolerance to low-temperature stress conditions (Friesen et al. 2014; Dharshini et al. 2016, 2018). Studying the genetic nature of abiotic tolerance mechanisms would help to understand the gene networks that are employed for cold tolerance. Though several cold-responsive genes have been identified, their functions are still unknown. Hence, studying SsCor413 is relatively necessary to understand its role in different abiotic stress response mechanisms and its likeliness to be selected as a candidate gene for the improvement of sugarcane (Sun et al. 2018).
So far, Cor413 gene has not been studied in Saccharum complex. Hence, the present study was undertaken to isolate and characterize the S. spontaneum homolog of the Cor gene. Our earlier studies on transcriptome analysis of S. spontaneum revealed the Cor413 gene family has an important role in the regulation of cold stress (Dharshini et al. 2016, 2018). Utilizing the Cor413 gene sequence information from the transcriptome data, a coding region of SsCor413 was isolated from S. spontaneum. In silico analyses, such as domain prediction, conserved blocks, protein structure prediction, string analysis, and gene ontology annotation were performed. The temporal and spatial expression of SsCor413 was studied during different abiotic stress, such as low-temperature stress, water deficit, and salinity stress, and the transient expression of SsCor413 protein was studied to identify the sub-cellular localization in sugarcane and onion epidermal cells.
Materials and Methods
Plant Material and Stress Treatments
Saccharum spontaneum IND 00–1037 clone from the high altitude regions of Arunachal Pradesh, North eastern India was previously used for the experiment to develop low-temperature transcriptome profiling (Dharshini et al. 2016). The same cultivar was raised under glasshouse conditions at Indian Council of Agricultural Research—Sugarcane Breeding Institute (ICAR-SBI), Coimbatore, Tamil Nadu, India. Ninety days grown S. spontaneum IND 00-1037 seedlings were selected and shifted to aerated hydroponics setup supplied with Hoagland solution and maintained at 26 ± 2 °C in a glass chamber. Leaf samples were collected from plants exposed to 10 °C for 24 h. As described above, the experiment was repeated and root samples excised at different time intervals (3 h, 6 h, 12 h, 24 h, and 48 h). Leaf and root samples collected from non-treated served as control. All samples collected were immediately frozen in liquid nitrogen and stored at − 80 °C until RNA extraction.
S. spontaneum plants were raised and planted in 16-inch pots (containing soil, sand, and farmyard manure in 1:1:1 ratio) and maintained with regular irrigation inside a glasshouse at ICAR-SBI. Water deficit stress was imposed on the plants at the tillering phase (90 days after planting) by withholding irrigation for 7 days and was released on the 8th day, and normal irrigation was continued (Augustine et al. 2015). Leaf samples were excised from the plant on the 0th day, 1st day, 2nd day, 3rd day, 5th day, and 7th day of drought induction. The collected samples were ground in liquid nitrogen and stored at − 80 °C for further analysis.
Sixty days old plantlets were treated with 200 mM of sodium chloride (NaCl) for salt treatment. Leaf samples were harvested during 1 h, 3 h, 12 h, and 24 h of salt stress along with the control samples under normal irrigation. The collected samples were frozen in liquid nitrogen and stored at − 80 °C until further use.
RNA Extraction and cDNA Synthesis
Total RNA was extracted from frozen tissues using the RNeasy plant media kit (Qiagen, MD) and DNA contamination was removed using RNase free DNase I (Thermo Scientific, USA). The quality and quantity of total RNA were analyzed by agarose gel and NanoDrop Spectrophotometer (Thermo Scientific, USA), respectively. RNA integrity was checked using Agilent RNA Bioanalyzer chip (Agilent Technologies Inc., Santa Clara, CA). The RNA samples with 260–280 ratios of more than 2.0 and RIN (RNA integrity number) of more than 9.0 were selected and used for the further experiment. First-strand cDNAs were synthesized from 100 ng of total RNA using RevertAid First strand cDNA Synthesis Kit (Thermo Fisher Scientific Company, USA) following the manufacture’s instruction.
Gene Isolation and Cloning
The gene-specific primers were designed for SsCor413 gene using the sequence information available from a previous cold transcriptome study. Cor413FP (5’ATGGGGAAGGGGTTCGCGTCGTACT-3′) and Cor413RP (5′- CTACAGGATTTGCAGCACCCCGGTC-3′) primers were used for PCR amplification. Using leaf cDNA (LT-treated sample) as a template, PCR amplification was carried out as follows: initial denaturation of 4 min at 94 °C, 35 cycles (94 °C for 45 s, 63 °C for 45 s, and 72 °C for 45 s) and final extension of 10 min at 72 °C in a thermocycler (Eppendorf, Hamburg, Germany). The PCR product was analyzed on 1.2% agarose gel and gel purified using the GeneJET Gel Extraction Kit (Thermo Fisher Scientific USA). The eluted DNA fragment was ligated into pTZ57R/T vector using the InsTAclone PCR Cloning Kit (Thermo Fisher Scientific, USA) and the ligated product was transformed into E. coli DH5α cells. Recombinant colonies were selected in media containing ampicillin (AmpR). Positive colonies were confirmed using M13 and Cor413 gene-specific primers. The recombinant plasmid was isolated using a Plasmid Isolation Kit (Qiagen, MD) and was Sanger sequenced using the facilities available at the University of Delhi, South Campus, New Delhi, India.
Bioinformatics Analysis
Database searches to identify SsCor413 homologs were performed using the National Center for Biotechnology Information (http://www.ncbi.nlm.nih.gov/BLAST) Web implementation of BLAST (Altschul et al. 1990) against the GenBank non-redundant sequence database. The ORF of the SsCor413 gene was analyzed using the ORF Finder (https://www.ncbi.nlm.nih.gov/orffinder/). Simple modular architecture research tool (SMART) (http://smart.embl-heidelberg.de/) was used for conserved domain prediction (Schultz et al. 1998; Letunic et al. 2014). The conserved domains were also predicted using batch web, CD-search tool (http://www.ncbi.nlm.nih.gov/Structure/bwrpsb) for ten monocot Cor413 protein sequences. Expasy’s Protparam Server (http://web.expasy.org/protparam/) (Gasteiger et al. 2005) was used for physicochemical characterization including theoretical molecular weight (MW), isoelectric point (pI), instability index (II) (Guruprasad et al. 1990), and aliphatic index (AI) (Ikai 1980). Secondary structure was predicted using a PSIPRED server (McGuffin et al. 2000) (http://bioinf.cs.ucl.ac.uk/psipred/) and HNN (Hierarchical Neural Network) (Guermeur et al. 1999) (https://npsa-prabi.ibcp.fr/cgi-bin/npsa_automat.pl?page=/NPSA/npsa_hnn.html). SignalP was used for detection of signal peptides (Nielsen and Krogh 1998) (http://www.cbs.dtu.dk/services/SignalP/). ProtScale program (http://web.expasy.org/protscale/) was used to generate Kyte and Doolittle hydropathy plot with the Kyte and Doolittle option and a window of nine amino acids (Kyte and Doolottle 1982). ngLOC, an n-gram-based 209Q7 Bayesian classifier that predicts subcellular localization of proteins both in prokaryotes and eukaryotes used to predict sub cellular localization (King and Guda 2007) (http://genome.unmc.edu/ngLOC/index.html). Importin α-dependent nuclear localization signals were predicted using cNLS mapper (http://nls-mapper.iab.keio.ac.jp/cgi-bin/NLS_Mapper_form.cgi). LocSigDB server predicts with protein sorting signals using experimental and literature database (http://genome.unmc.edu/LocSigDB/index.html). TMHMM server v2.0 (http://www.cbs.dtu.dk/services/TMHMM/) was used for transmembrane domains (TMD) prediction (Krogh et al. 2001).
STRING is a database of known and predicted protein networks derived from genomic context, co-expression, high-throughput experiments, and previous knowledge. STRING database (http://string-db.org/) with default parameters was used to search the protein network for protein query sequence and provide protein networks with known functional links as well as from predicted molecular models. The SsCor413 sequence was used as a query and searched against Zea mays database and the network was analyzed with different categories such as “Network,” “Experiments,” “Text mining,” “Databases,” “Co-occurrence,” “Co-expression”, and “Neighborhood,” and Gene ontology components such as Molecular function, Biological process, and Cellular component using AmiGo server (http://amigo.geneontology.org/amigo) with default parameters against Arabidopsis thaliana.
Chromosomal Mapping of Cor Gene
Chromosomal location of Cor gene was identified by mapping the Cor sequence to mosaic monoploidy sugarcane sequence (Garsmeur et al. 2018) using CLC workbench version 11 with default parameters.
Phylogenetic Construction
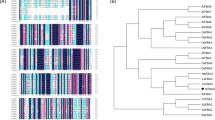
A total of ten Cor413 sequences of different plants were retrieved from the NCBI database, and multiple sequence alignment was performed using ClustalX (Thompson et al. 1997) using default settings. For building up of the evolutionary tree, Maximum Parsimony (MP) analysis was performed with MEGA7 (Kumar et al. 2016) using the tree-bisection-regrafting (TBR) algorithm (Nei and Kumar 2000) where the random addition of sequences (10 replicates) with search level 1 was employed for obtaining the initial tree. Average pathway methods (Nei and Kumar 2000) were used to calculate the branch lengths and represented as units of the number of changes over the whole sequence. The resulting percentages of replicate trees (1000 replicates) where the associated taxa clustered together were indicated next to the branches (Felsenstein 1985).
Development of GFP Constructs
The sub-cellular localization of SsCor413 protein was studied using both N′-terminal and C′-terminal GFP fused constructs. The ZmUbi-GFP vector was used as template. Fusion GFP-SsCor413 (ZmUbi-GFP– SsCor413) was constructed by amplifying the SsCor413 using primers GFPCOR-F (5’-GATCACTAGTATGGGGAAGGGGTTCGCGT-3’) and GFPCOR-R (5’-TAGCGGCCGCCTCAGGACTCG-3’) with restriction sites for SpeI and NotI, respectively, and cloned into ZmUbi-GFP vector digested with same restriction enzymes according to Xue (2002) and Shivalingamurthy et al. (2018). Also, to generate C-terminal SsCor413-GFP fusion (CaMV35S-SsCor413-GFP), SsCor413 without a stop codon was amplified with primers CORGFP-F (5’-ATACCATGGATATGGGGAAGGGGTTCGCG-3’) and CORGFP-R (5’-GCGCCTAGTCAGGACTCGCAGCACCGC-3’) with restriction sites for NcoI and SpeI, respectively, and cloned into a pCAMBIA1302 vector (CaMV35S-GFP) digested with the same set of restriction enzymes.
Particle Bombardment and GFP Localization
Sugarcane calli and young onion epidermal cells were used for sub-cellular localization experiments. The young shoot tips from 4 to 6 months old sugarcane variety Co 86032 was used as a starting material for raising the embryogenic calli. The surface sterilized sugarcane meristematic explants were selected, and young leaves surrounding the sub-apical meristematic portion were cut into pieces, placed on the MS + 2,4-D medium, and incubated in the dark at 25 °C. The proliferated calli were sub-cultured every 15 days in fresh MS + 2,4-D medium for the production of embryogenic calli for particle bombardment.
Sugarcane embryogenic calli and onion epidermal tissue were placed in concentric circles on MS + osmotic medium (MS + 50 g/L mannitol and 50 g/L sorbitol) for 3 h prior to bombardment. A mixture of 1.5–3 μm-sized gold particles were sterilized and used as a microcarrier for particle bombardment. The gold suspension was added to the SsCor413-GFP localization plasmids (6 μg) in a 1.5 ml siliconized microcentrifuge tube. 20 μl of 0.1 M spermidine was added with constant vortexing followed by 50 μl of 2.5 M calcium chloride solution was added drop by drop to the mixture with constant vortexing. The cocktail was set aside for 10 min and centrifuged at 10,000 RPM for 10 s. The pellet containing DNA-coated gold particles was washed with 200 μl of absolute alcohol, and the supernatant was decanted after centrifugation. Finally, the pellet was resuspended with 60 μl of absolute alcohol. A 10 μl of DNA-coated gold suspension was coated onto the sterile macrocarrier for particle bombardment. Bio-Rad PDS 1000/He Biolistic System at a pressure of 1100 Psi of helium was used for bombardment. The explants were bombarded at a distance of 4 and 8 cm from stopping screen. The bombarded explants were incubated in the dark at 25 °C for 24 h. The slides were prepared by staining the bombarded calli with 0.1% propidium iodide (nuclear stain) for 1 h in the dark (Palaniswamy et al. 2016) and examined in both RFP (530 nm -593 nm) and GFP (470 nm – 525 nm) channels of an EVOS FL color fluorescence microscope (Life Technologies, USA).
Quantitative Real-Time PCR (qRT-PCR) Analysis
For qRT-PCR experiments, SsCor413 primers (Forward Primer-5’-AGCTTCCTGGTTCCATCATC-3′, Reverse Primer-5’-CATCCAATCGCAAGGCATATC-3′) were designed using a FastPCR tool (Kalendar et al. 2017). Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) gene (Forward Primer-5’-AAGGGTGGTGCCAAGAAGG-3′, Reverse Primer-5’-CAAGGGGAGCAAGGCAGTT-3′) was used as an endogenous control. To perform qRT-PCR experiments, the cDNA converted using total RNA samples isolated from samples of low temperature (leaf and root), water deficit (leaf), salinity (leaf), and control plants were used as a template. qRT-PCR was performed on a StepOnePlus Real-Time PCR system (Applied Biosystems, Canada) using the SYBR-green dye method. Each reaction was carried out in triplicates. In brief, each qRT-PCR reaction consists of 50 ng cDNA, 2.5 pmol primers, 12.5 μl of 2X MESAGREEN Master Mix (Eurogentec Belgium) and the final volume was made up to 25 μl with sterile water (Dharshini et al. 2018). qRT-PCR reaction conditions used were as follows: denaturation for 10 min at 95 °C followed by annealing and extension at 1 min for 60 °C (40 cycles). The fold change of the target genes was determined using 2-ΔΔCT method (Livak and Schmittgen 2001).
Results
Isolation and Sequence Analysis of SsCor413 Gene
A low-temperature responsive uncharacterized gene SsCor413 was identified from our previous low-temperature stress transcriptome profiling of Saccharum spontaneum (Dharshini et al. 2016). A full-length coding region (CDS) of Cor413 was isolated from S. spontaneuem collected from a low-temperature region. Sequence analysis of SsCor413 revealed that the length of the ORF is 642 nucleotides which encode a putative protein of 213 amino acids (Fig. 1). The molecular weight of the protein was 25.6 kDa and an isoelectric point (Pi) of 9.69. The instability index (II) is computed to be 23.89 for protein and hence it classifies as stable protein. The basic local alignment search tool for nucleotide (BLASTN) and protein (BLASTP) sequence analysis showed that SsCor413 has 95% and 96% similarity to the ZmCor413 (EU965484.1), respectively.
The full-length cDNA sequence and deduced amino acid sequence of SsCor413. The primers used for amplification were underlined and * represents stop codon. The deduced amino acid sequence was shown beneath the nucleotide sequence and the amino acids were numbered on the right-hand side of the sequence. The cDNA sequence has been deposited in GenBank under accession No. MF680545
The secondary structure analysis of putative SsCor413 showed the dominance of α helices (69.04–76.99%) followed by random coils (21.59–31.60%) and strands (0–5.16%). The prediction of secondary structure using different server is given in supplementary Table S1, and Fig. S1 represents the secondary prediction. The SsCor413 protein was predicted with one WCOR413 domain and its position ranged from 19 to 200 amino acids (supplementary Fig. S2). Further to confirm, ten monocot Cor413 proteins were subjected to Batch CD server, and results indicated that all queries assigned to WCOR413 domain and Pfam 05562 as accession number.
The Kyte and Doolittle hydrophobicity plot of SsCor413 protein showed a pattern of high hydrophobicity with the Grand Average of Hydropathy (GRAVY) value of 0.808 and 35 hydrophobic leu residues (Fig. 2a). TMHMM analysis based on (Krogh et al. 2001) Hidden Markov Model predicted that SsCor413 protein possesses five transmembrane helices at positions “53–75,” “88–105,” “131–150,” “157–175,” and “190–212” which is complemented with high hydrophobic nature of SsCor413 protein (Fig. 2b).
Conserved region analysis revealed the comparison of Cor413 proteins of the Poaceae family showed ten conserved blocks at amino acid position “40’AARKLANHA,” “48’ VLGGGLGF,” “67’ AAVYLL,” “81’NWKTNMLT,” “90’, LLVPYIFFTLP,” “122’LRLFFPRHFPDWLELPGS,” “146’VAP,” “170’LGCYLL,” “177’EHI,” and “207’YPVW” suggesting the highly conserved nature of Cor413 among grass family (Fig. 3). SsCor413 protein showed the presence of serine at positions “7, 59, 103, 147, 178, 192, and 194”, threonine at “13, 58, 82, 86, 96, 142, and 152,” and tyrosine residues at “8, 71, 92, 169, and 203” revealed the possible regulatory mechanism of protein function through post-translation modification. ngLOC protein subcellular localization prediction server reveals that putative SsCor413 protein is likely to be localized in the nucleus. But low prediction score (19.8) in this analysis has given low chance of localization in the nuclear lumen. Hence, SsCor413 protein sequence was further analyzed for NLS using cNLS Mapper which has predicted presence of importin α-dependent few bipartite NLS sequences in SsCor413 amino acid sequence position ranging from 34 to 67 with a minimum score of 2.9 and 60–88 with a score of 2.4. Importin alpha is known to bind to NLS sequence of nucleus-targeted protein which recognize and transports the NLS-containing protein across the nuclear membrane. SignalP server does not predict with any signals and cleavage sites; LocSigDB server displays protein sorting signals of SsCor413 to endoplasmic reticulum at the position 173–176 and lysosomes at positions 7–11, 70–74, 91–95, 202–206.
STRING Network and Gene Ontology of Cor413
Clustered analysis using STRING database indicated that SsCor413 protein grouped with proteins like Cor413-like, Rab28, late embryogenic abundant proteins (LEA)-Lea4 and Lea14-A, oleosin18, and xyloglucan endotransglucosylase, which are reported to be involved in low-temperature tolerance (Fig. 4). SsCor413 protein annotated with gene ontology terms such as integral components of membrane (AT2G15970), plasma membrane (AT2G15970), cold acclimation (AT1G29395), cellular response to cold (AT1G29395), cellular response to water deprivation (AT2G15970), and response to abscisic acid (AT1G29395). These results suggest that SsCor413 protein might respond to cold, drought, and abscisic acid stress. A pictorial representation of SsCor413 protein GO terms are given in supplementary Fig. S3.
Chromosomal Location of Cor Gene
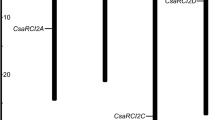
Chromosomal location of Cor gene and its isoforms were identified in this study. The location of Cor gene includes in chromosome 1 at position “7829564–7830587,”; chromosome 3 at positions “30565359–30566967,” “30585588–30586911,” “30594461–30594810,” and “50326281–50326556,”; and chromosome 9 at position “33099810–33103744.” These results indicate that Cor gene located at four positions in chromosome 3 and located at a single position in chromosome 1 and chromosome 9.
Development of SsCor413 Phylogeny
Amino acid sequences of a Cor413 family of monocot species were retrieved from the NCBI database. The phylogenetic analysis of SsCor413 proteins showed its close evolutionary relationship with Sorghum bicolor and Zea mays with high similarity (Fig. 5). The ten Cor413 protein sequences from monocots showed a good similarity level. Also, 154 protein sequences which contain the WCOR413 domain from different species were retrieved and phylogenetic tree analysis also revealed that the WCOR413 domain was highly conserved among monocots.
Sub-Cellular Localization of SsCor413
Localization of SsCor413 using C-terminal GFP (SsCor413-GFP) and N-terminal GFP (GFP-SsCor413) was studied using sugarcane calli and onion epidermal cells. The diagrammatic representation of GFP-COR constructs is given in Fig. 6. Control and GFP fused plasmids were bombarded on the sugarcane calli and onion epidermal cells. The microscopic slides were prepared using bombarded explants stained with propidium iodine (Nuclear stain) and observed under a fluorescent microscope. The results revealed that control GFP constructs exhibit fluorescence strongly throughout the cell in sugarcane calli and onion cells. The fused plasmids such as GFP-SsCor413 and SsCor413-GFP showed strong GFP expression in the nucleus compared with plasma membrane of both sugarcane calli (Fig. 7) and onion epidermal cells (Fig. 8).
Subcellular localization of SsCor413-GFP in sugarcane. a Fluorescent microscopic images of untransformed sugarcane calli. b N-terminal control plasmid shows GFP expression throughout the callus. c Fluorescent microscopic images of sugarcane calli showing the transient expression of SsCor413-GFP in nucleus and plasma membrane. d C-terminal control plasmid shows GFP expression throughout the callus. e Fluorescent microscopic images of sugarcane calli showing the transient expression of GFP-SsCor413 in nucleus. Middle vertical lane from A to E represents nuclear staining with the propidium iodide
Subcellular localization study in onion epidermal cells. a Fluorescent microscopic images of untransformed onion epidermal cells. b N-terminal control plasmid shows GFP expression throughout the onion epidermal cells. c Fluorescent microscopic images of onion epidermal cells showing the transient expression of SsCor413-GFP in nucleus and plasma membrane. d C-terminal Control plasmid shows GFP expression throughout the onion epidermal cells. e Fluorescent microscopic images of epidermal cells showing the transient expression of GFP-SsCor413 in nucleus and plasma membrane. Middle vertical lane from A to E represents nuclear staining with the propidium iodide
Expression Pattern of SsCor413 upon LT, Water Deficit, and Salinity Stresses
To understand the regulation of the SsCor413 protein under abiotic stresses such as low temperature, drought, and salinity in sugarcane, the gene expression pattern was studied through quantitative real-time PCR at different points of time in leaf and root samples. The qRT-PCR results indicated that SsCor413 transcripts are strongly up-regulated in leaf and root tissues at 24 h upon LT stress. A gradual up-regulation of SsCor413 was observed in root tissue at 3 h (0.70 fold), 6 h (3.5 fold), 12 h (4.21 fold), and reached a maximum at 24 h (7.26 fold) and then declined at 48 h (0.32 fold) under LT stress. Similarly, SsCor413 transcripts in leaf tissue showed up-regulation at 3 h (1.02 fold), 6 h (3.78 fold), 12 h (3.98 fold), and reached a maximum (5.3 fold) at 24 h and declined at 48 h (4.0 fold) under LT stress (Fig. 9). During water deficit conditions, SsCor413 expression was up-regulated by 0.95-, 1.20-, 3.22-, 3.41-, 3.22-, and 1.47-fold at 1st, 2nd, 3rd, 4th, 5th, and 7th days, respectively (Fig. 10a). Upon 200 mM salinity treatment, SsCor413 showed 3.94-, 2.77-, 1.84-, and 1.48-fold change at 1 h, 3 h, 12 h and 24 h, respectively (Fig. 10b).
Discussion
In this study, a full-length SsCor413 gene, which encodes a polypeptide of 213 amino acids, was cloned from S. spontaneum, a low-temperature tolerant clone (Dharshini et al. 2016). The amino acid sequence of SsCor413 showed 96% homology to ZmCor413 (EU965484.1). The molecular weight of the SsCor413 is 25.6 kDa, which almost equal to ZmCor413 (25.4 kDa) (Breton et al. 2003). Cor413 protein with molecular weight of 22.74 kDa was also cloned from Gossypium barbadense (Wang et al. 2007). Isoelectric point (pI) value indicates the pH at which protein is stable and has no net charge. The computed pI value of SsCor413 is 9.69. Hence, it is classified that the protein is alkali in nature. The Cor413 protein is seen to be stable with instability index (II) of 23.89. Instability index lower than 40 is considered to be stable proteins (Guruprasad et al. 1990). The amino acids of SsCor413 protein are hydrophobic, suggesting that this protein could be a membrane protein (Wang et al. 2007). The amino acid residue ratio is one of the essential criteria that decide the role and the activity of its protein. Multiple sequence alignment was performed to find out the conserved amino acids residue, and this revealed that the leucine (leu) residue is abundant among all cold-regulated proteins. For example, GbCor413 contains 25 leu residues and aids stabilization of the transmembrane structure during cold stress (Wang et al. 2007). In this study, SsCor413 contains 35 leu residues and perhaps plays a vital role in stabilization. The percentage of leu in SsCor413 protein is very close to that of ZmCor413 (Breton et al. 2003) and GbCor413 (Wang et al. 2007) characterized from Z. mays and G. barbadense, respectively. The residues such as serine (ser), tyrosine (tyr), and threonine (thr) are associated with phosphorylation sites of a protein. The Cor413 protein reported to contain phosphorylation sites (Wang et al. 2007). The SsCor413 protein contains 8 ser, 7 thr, and 5 tyr residues. SsCor413 contains eight pro residues. Other amino acids such as pro and cys were also conserved among different Cor413 proteins (Breton et al. 2003), and Cor413 from Arabidopsis and cereals contain five pro residues and is conserved among other Poaceae families. These residues are involved in the possible regulatory mechanism of protein function through post-translational modification. SsCor413 protein possesses five transmembrane helices as that of cold-regulated Cor413 proteins of cereals and Arabidopsis family which are identified as a novel stress-regulated multi-spanning transmembrane protein family (Breton et al. 2003), whereas only four transmembrane helices were found in GbCor413 protein (Wang et al. 2007). SignalP result showed the absence of signal peptide in SsCor413 sequence as reported in Wang et al. (2007). But interestingly, LocSigDB server predicted with protein sorting signals to ER and lysosome. Based on earlier reports and present study, Cor413 protein exhibits different number of transmembranes, different protein sorting signals suggesting us that Cor413 protein could be diverse.
Domains are distinct functional and structural units of a protein containing conserved sequence patterns. They are recognized as building blocks and may rearrange to modulate its protein function. SsCor413 protein was detected with one WCOR413 domain. Domain-based evolutionary studies indicated that WCOR413 domain found in many species such as Lolium temulentum, Brassica rapa, Cucumis sativus, and Gossypium barbadense was grouped within its family and plays a significant role in freezing tolerance mechanism. Early studies reported that there were two conserved blocks in ZmCor413 (Wang et al. 2007). In this study, we were able to identify a few more conserved blocks in Cor413 sequence among monocot families. The phylogenetic analysis revealed that SsCor413 was highly conserved among Poaceae family. The gene ontology (GO) for SsCor413 protein was annotated with cold responsive and cold acclimation terms indicating the SsCor413 might play a significant role upon cold stress. The STRING network analysis revealed that SsCor413 protein is connected to LEA, Rab28, and oleosin18 which was reported to enhance cold tolerance in plants. Rab, an LEA protein and Oleosin help in freezing tolerance in A. thaliana (Puhakainen et al. 2004; Shimada et al. 2008). This level of mining suggests that SsCor413 protein is involved in low-temperature tolerance mechanism, as it connects to LEA proteins. The secondary structure of SsCor413 predicted with alpha-helix structures.
Cellular localization of SsCor413 protein using C′-terminal GFP (SsCor413-GFP) and N′-terminal GFP (GFP-SsCor413) was performed in sugarcane callus and onion epidermal cells. The result indicated that SsCor413 is likely to be localized in nucleus. This localization result correlated with in silico ngLOC prediction and further supported by cNLS Mapper prediction. Prediction of the importin α-dependent nuclear localization signals (NLS) in SsCor413 protein sequence indicated its localization in nucleus. In general, the NLSs are short stretches of amino acids recognized by nucleo-cytoplasmic transporters (karyopherins) that promote active transport of proteins into the nucleus (Xu et al. 2010). Also, our localization study showed low abundance of SsCor413-GFP signals in plasma membrane which might be because of presence of sorting signals to ER and lysosomes. Usually, after ER synthesis, lysosomal transmembrane proteins are transported as glycosylated proteins to the trans Golgi Network (TGN) where they follow the secretory route to the plasma membrane. Thus, the result of localization of SsCor413 was differed from other reports of Cor413 protein which were reported to be localized in thylakoid membrane and plasma membrane. Zhou et al. (2018) reported that GFP construct containing transmembrane regions of Cor413 protein (PsCor413-TM1-TM4-GFP) was localized to the nucleus and the cytoplasm.
SsCor413 transcript up-regulated under low-temperature stress and showed a maximum accumulation in leaf and root tissues at 24 h of stress. TaCor413-pm1 is expressed abundantly in the leaves and root under freezing conditions (Breton et al. 2003). During water deficit stress conditions, SsCor413 expression showed a gradual increase until the 4th day of stress and declines at 5th and 7th day. Zhao et al. (2016) reported that Cor413 was up-regulated during water deficit and low-temperature stress conditions and thus playing a double role and widespread cross talk between the cold and water deficit stress response pathway in sheepgrass. SsCor413 showed an early response to salt stress in sugarcane. Other stresses as well triggered Cor genes such as Cor15A, Cor6.6 in response to cold treatment, ABA, and water deficit stresses (Thomashow 1998). All cereals contain homologous low-temperature responsive genes in their genome, but the gene expression is found only in cold-tolerant cereal variety under low-temperature stress conditions (Sarhan and Danyluk 1998). Accumulation of Cor proteins has a close relationship between freezing tolerance and this protein can be used as a molecular marker to select for freezing tolerance (Houde et al. 1992). In line with previous reports, SsCor413 can be used as molecular markers and would serve as a potential candidate gene for developing abiotic stress tolerance variety in sugarcane.
Conclusion
SsCor413 gene was isolated from S. spontaneum (low-temperature tolerant) and for the first time to gain insight into features of protein sequence, localization, and expression analysis of different stress conditions such as low temperature, water deficit, and salinity stresses were studied. Results of physicochemical characterization showed that SsCor413 protein was stable and hydrophobic in nature. The evolutionary and conserved region analysis suggested ten conserved blocks which showed high conservation among monocots. SsCor413 contains five potential transmembrane domains which are stabilized by the presence of 35 leu amino acids upon adverse conditions. SsCor413 was significantly up-regulated and might actively participate in the cold acclimation process. Also, SsCor413 was up-regulated upon water deficit and salinity stress conditions, suggesting that this protein responds to multiple stresses. Localization study revealed SsCor413 protein presumably to be localized in the nucleus and plasma membrane. This is the first report which studied the transcriptional regulation of SsCor413 under low temperature, salinity, and water deficit conditions in S. spontaneum. SsCor413 has shown potential candidate for developing multiple abiotic stress-tolerant sugarcane varieties either through marker-assisted breeding or transgenic approaches.
Abbreviations
- Cor:
-
Cold regulated
ORF
Open reading frame
GFP
Green Fluorescent Protein
LEA
Late Embryogenic Abundant Protein
GO
Gene Ontology
LT
Low temperature
MS
Murashige and Skoog
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Amalraj VA, Rakkiyappan P, Neelamathi D, Chinnaraj S, Subramanian S (2008) Wild cane as a renewable source for fuel and fiber in paper industry. Curr Sci 95:1599–1602 https://www.jstor.org/stable/24105519
Augustine SM, Cherian AV, Syamaladevi DP, Subramonian N (2015) Erianthus arundinaceus HSP70 (EaHSP70) acts as a key regulator in the formation of anisotropic interdigitation in sugarcane (Saccharum spp. hybrid) in response to drought stress. Plant Cell Physiol 56(12):2368–2380. https://doi.org/10.1093/pcp/pcv142
Baker SS, Wilhelm KS, Thomashow MF (1994) The 5′-region of Arabidopsis thaliana Cor15a has cis-acting elements that confer cold, drought and ABA-regulated gene expression. Plant Mol Biol 24:701–713. https://doi.org/10.1007/BF00029852
Blom N, Gammeltoft S, Brunak S (1999) Sequence and structure-based prediction of eukaryotic protein phosphorylation sites. J Mol Biol 294:1351–1362. https://doi.org/10.1006/jmbi.1999.3310
Breton G, Danyluk J, Charron JB, Sarhan F (2003) Expression profiling and bioinformatic analyses of a novel stress regulated multispanning transmembrane protein family from cereals and Arabidopsis. Plant Physiol 132:64–74. https://doi.org/10.1104/pp.102.015255
Chinnusamy V, Zhu J, Zhu JK (2006) Gene regulation during cold acclimation in plants. Physiol Plant 126:52–61. https://doi.org/10.1111/j.1399-3054.2006.00596.x
Chinnusamy V, Zhu J, Zhu JK (2007) Cold stress regulation of gene expression in plants. Trends Plant Sci 12:444–451. https://doi.org/10.1016/j.tplants.2007.07.002
Dharshini S, Chakravarthi M, Manoj VM, Naveenarani M, Kumar R, Meena M, Appunu C (2016) De novo sequencing and transcriptome analysis of a low temperature tolerant Saccharum spontaneum clone IND 00-1037. J Biotech 231:280–294. https://doi.org/10.1016/j.jbiotec.2016.05.036
Dharshini S, Chakravarthi M, Vignesh D, Nerkar G, Ashwin Narayan J, Manoj VM, Naveenarani M et al (2018) Differential gene expression profiling through transcriptome approach of Saccharum spontaneum L. under low temperature stress reveals genes potentially involved in cold acclimation. 3 Biotech 8(4):195. https://doi.org/10.1007/s13205-018-1194-2
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791. https://doi.org/10.1111/j.1558-5646.1985.tb00420.x
Fitter AH, Hay RKM (1981) Environmental physiology of plants. Academic Press, New York
Friesen PC, Peixoto MM, Busch FA, Johnson DC, Sage RF (2014) Chilling and frost tolerance in Miscanthus and Saccharum genotypes bred for cool temperate climates. J Exp Biol 65(13):3749–3758. https://doi.org/10.1093/jxb/eru105
Garsmeur O, Droc G, Antonise R, Grimwood J, Potier B, Aitken K, Jenkins J, Martin G, Charron C, Hervouet C, Costet L (2018) A mosaic monoploid reference sequence for the highly complex genome of sugarcane. Nat Commun 9(1):2638. https://doi.org/10.1038/s41467-018-05051-5
Gasteiger E, Hoogland C, Gattiker A, Duvaud S, Wilkins MR, Appel RD, Bairoch A (2005) Protein identification and analysis tools on the ExPASy server. In: Walker JM (ed) The proteomics protocols handbook. Humana Press, Totowa, pp 71–607
Guermeur Y, Elisseeff A, Paugam-Moisy H (1999) Estimating the sample complexity of a multi-class discriminant model. In artificial neural networks, ICANN 99. Ninth international conference on (Conf. Publ. No. 470) (Vol. 1, pp. 310-315). IET
Guruprasad K, Reddy BVP, Pandit MW (1990) Correlation between stability of a protein and its dipeptide composition: a novel approach for predicting in vivo stability of a protein from its primary sequence. Protein Eng 4:155–164. https://doi.org/10.1093/protein/4.2.155
Guy CL (1990) Cold acclimation and freezing stress tolerance: role of protein metabolism. Annu Rev Plant Biol 41(1):187–223. https://doi.org/10.1146/annurev.pp.41.060190.001155
Houde M, Dhindsa RS, Sarhan F (1992) A molecular marker to select for freezing tolerance in Gramineae. Mol Gen Genet 243:43–48. https://doi.org/10.1007/BF00272343
Ikai AJ (1980) Thermo stability and aliphatic index of globular proteins. J Biochem 88:1895–1898. https://doi.org/10.1093/oxfordjournals.jbchem.a133168
Kalendar R, Khassenov B, Ramanculov E, Samuilova O, Ivanov KI (2017) FastPCR: an in silico tool for fast primer and probe design and advanced sequence analysis. Genomics. 109:312–319. https://doi.org/10.1016/j.ygeno.2017.05.005
King BR, Guda C (2007) ngLOC: an n-gram-based Bayesian method for estimating the subcellular proteomes of eukaryotes. Genome Biol 8(5):R68. https://doi.org/10.1186/gb-2007-8-5-r68
Krogh A, Larsson B, von Heijne G, Sonnhammer EL (2001) Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J Mol Biol 305:567–580. https://doi.org/10.1006/jmbi.2000.4315
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33(7):1870–1874. https://doi.org/10.1093/molbev/msw054
Kyte J, Doolottle RF (1982) A simple method for displaying the hydropathic character of a protein. J Mol Biol 157:105–132. https://doi.org/10.1016/0022-2836(82)90515-0
Letunic I, Doerks T, Bork P (2014) SMART: recent updates, new developments and status in 2015. Nucleic Acids Res 43(D1):D257–D260. https://doi.org/10.1093/nar/gku949
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25(4):402–408. https://doi.org/10.1006/meth.2001.1262
Ma X, Wang G, Zhao W, Yang M, Ma N, Kong F, Dong X, Meng Q (2017) SlCOR413IM1: a novel cold-regulation gene from tomato, enhances drought stress tolerance in tobacco. J Plant Physiol 216:88–99. https://doi.org/10.1016/j.jplph.2017.03.016
McGuffin LJ, Bryson K, Jones DT (2000) The PSIPRED protein structure prediction server. Bioinformatics 16(4):404–405. https://doi.org/10.1093/bioinformatics/16.4.404
Nei M, Kumar S (2000) Molecular evolution and phylogenetics. Oxford University Press, New York
Nielsen H, Krogh A (1998) Prediction of signal peptides and signal anchors by a hidden Markov model. In: Glasgow J, Littlejohn T, Major F, Lathrop R, Sankoff D, Sensen C (eds) Proceedings of the sixth international conference on intelligent Systems for Molecular Biology. AAAI Press, Menlo Park, pp 122–130
Palaniswamy H, Syamaladevi DP, Mohan C, Philip A, Petchiyappan A, Narayanan S (2016) Vacuolar targeting of r-proteins in sugarcane leads to higher levels of purifiable commercially equivalent recombinant proteins in cane juice. Plant Biotechnol J 14(2):791–807. https://doi.org/10.1111/pbi.12430
Premachandran MN, Sobhakumari VP, Lekshmi M, Viola VR (2017) Genome characterization of in vitro induced amphiploids of an intergeneric hybrid Erianthus arundinaceus × Saccharum spontaneum. Sugar Tech 19(4):386–393. https://doi.org/10.1007/s12355-016-0482-6
Puhakainen T, Hess MW, Makela P, Svensson J, Heino P, Palva ET (2004) Overexpression of multiple dehydrin genes enhances tolerance to freezing stress in Arabidopsis. Plant Mol Biol 54:743–753. https://doi.org/10.1023/B:PLAN.0000040903.66496.a4
Qu Y, Zhou A, Zhang X, Tang H, Liang M, Han H, Zuo Y (2015) De novo transcriptome sequencing of low temperature-treated Phlox subulata and analysis of the genes involved in cold stress. Int J Mol Sci 16:9732–9748. https://doi.org/10.3390/ijms16059732
Sarhan F, Danyluk J (1998) Engineering cold-tolerant crops-throwing the master switch. Trends. Plant Sci 3:289–290. https://doi.org/10.1016/S1360-1385(98)01285-0
Schultz J, Milpetz F, Bork P, Ponting CP (1998) SMART, a simple modular architecture research tool: identification of signaling domains. Proc Natl Acad Sci U S A 95(11):5857–5864. https://doi.org/10.1073/pnas.95.11.5857
Shimada TL, Shimada T, Takahashi H, Fukao Y, Hara-Nishimura I (2008) A novel role for oleosins in freezing tolerance of oilseeds in Arabidopsis thaliana. Plant J 55:798–809. https://doi.org/10.1111/j.1365-313X.2008.03553.x
Shivalingamurthy SG, Anangi R, Kalaipandian S, Glassop D, King GF, Rae AL (2018) Identification and functional characterization of sugarcane invertase inhibitor (ShINH1): a potential candidate for reducing pre-and post-harvest loss of sucrose in sugarcane. Front Plant Sci 9. https://doi.org/10.3389/fpls.2018.00598
Somerville C (1995) Direct tests of the role of membrane lipid composition in low-temperature-induced photoinhibition and chilling sensitivity in plants and cyanobacteria. Proc Natl Acad Sci USA 92(14):6215 PMCID: PMC41488
Sun T, Liu F, Wang W, Wang L, Wang Z, Li J, Que Y, Xu L, Su Y (2018) The role of sugarcane catalase gene ScCAT2 in the defense response to pathogen challenge and adversity stress. Int J Mol Sci 19(9):2686. https://doi.org/10.3390/ijms19092686
Thomashow MF (1998) Role of cold-responsive genes in plant freezing tolerance. Plant Physiol 118:1–8. https://doi.org/10.1104/pp.118.1.1
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882. https://doi.org/10.1093/nar/25.24.4876
Wang J, Zuo KJ, Qin J, Zhang L, Su L, Liu J, Tang KX (2007) Isolation and bioinformatics analyses of a COR413-like gene from Gossypium barbadense. Acta Physiol Plant 29(1):1–9. https://doi.org/10.1007/s11738-006-0001-6
Xu D, Farmer A, Chook YM (2010) Recognition of nuclear targeting signals by Karyopherin-β proteins. Curr Opin Struct Biol 20:782–790. https://doi.org/10.1016/j.sbi.2010.09.008
Xue GP (2002) Characterization of the DNA-binding profile of barley HvCBF1 using an enzymatic method for rapid, quantitative and high throughput analysis of the DNA-binding activity. Nucleic Acids Res 30:e77. https://doi.org/10.1093/nar/gnf076
Zhao P, Liu P, Yuan G, Jia J, Li X, Qi D, Chen S, Ma T, Liu G, Cheng L (2016) New insights on drought stress response by global investigation of gene expression changes in sheepgrass (Leymus chinensis). Front Plant Sci 7:954. https://doi.org/10.3389/fpls.2016.00954
Zhou A, Sun H, Feng S, Zhou M, Gong S, Wang J, Zhang S (2018) A novel cold-regulated gene from Phlox subulata, PsCor413im1, enhances low temperature tolerance in Arabidopsis. Biochem Biophys Res Commun 495(2):1688–1694. https://doi.org/10.1016/j.bbrc.2017.12.042
Acknowledgments
The authors would like to thank ICAR—Sugarcane Breeding Institute, Coimbatore for providing the necessary infrastructure. We thank Dr. Shobakumari, ICAR-SBI for extending microscope facility. We greatly acknowledge Dr. K. Kadirvelu, DRDO-BU, Coimbatore to access the fluorescence microscope. Thanks to Mr. K. Selvamuthu for his technical assistance to carry out the work.
Funding
The authors would like to thank Science and Engineering Research Board (SERB), Department of Science and Technology (DST), New Delhi (Grant No SB/YS/LS-165/2013) for financial support. Authors thank the Department of Science and Technology, India (DST-INSPIRE, IF150891) for financial support to DS.
Author information
Authors and Affiliations
Contributions
DS and AC designed the experiments. DS performed the experiments and wrote the manuscript. SGS provided localization control vectors, assisted in microscopic studies, and manuscript revision. MVM and ANJ supported in cloning. SPTS did the artwork for figs. RK and MRM helped in bioinformatics analysis. MM, BR, and AC revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Key Message
Cor413 is identified as abiotic stress responsive gene in Saccharum spontaneum and acts as potential gene for combating abiotic stresses in sugarcane.
Electronic supplementary material
ESM 1
(DOC 392 kb)
Rights and permissions
About this article
Cite this article
Dharshini, S., Manoj, V.M., Suresha, G.S. et al. Isolation and Characterization of Nuclear Localized Abiotic Stress Responsive Cold Regulated Gene 413 (SsCor413) from Saccharum spontaneum. Plant Mol Biol Rep 38, 628–640 (2020). https://doi.org/10.1007/s11105-020-01224-z
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-020-01224-z