Abstract
The high level of abscission of developing reproductive organs in soybean (Glycine max L. Merr.) causes severe yield loss. The role of ethylene in hastening organ abscission has been well documented. However, little is known about the regulatory mechanism by which ethylene influences abscission in soybean. Ethylene synthesis inhibitor [0.5 mM silver thiosulfate (STS)], ethylene-releasing compound [400 mg/L ethephon (ETH)], and water were applied twice with a 5-day interval between the applications. The STS-treated plants produced more flowers and pods, resulting in 55.6 % greater seed yield. ETH treatment significantly increased (P < 0.05) the abscission rate of flowers and pods. Three digital gene expression libraries (STS-, ETH- and control library) were constructed. Illumina sequencing of the three libraries was used to identify 9.7 to 10.5 million unigenes. Strong correlations were observed among different gene expression profiles and specific metabolite groups (such as plant hormone biosynthesis and signal transduction; starch and sucrose metabolism; and secondary metabolism) suggesting the importance of these metabolic pathways during ethylene regulation. Potential ethylene target genes such as 1-aminocyclopropane-1-carboxylate (ACC)-oxidase, 2C protein phosphatases (PP2Cs), MAT1, acetyltransferase, bidirectional sugar transporter SWEET1-like, and so on were identified. These results suggest that ethylene regulates flower and pod abscission in soybean by direct transcriptional regulation of genes that are involved in the metabolic and regulatory processes related to both floral organ development and abscission. Thus, these findings further elucidate the critical role of ethylene in transcriptional regulation of reproductive organ development and abscission.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Soybean (Glycine max L. Merr.) plants develop numerous flowers and pods. However, under field conditions, most of the flowers and young pods abscise rather than develop into mature pods. The abscission of flowers and pods is the primary cause of yield loss in soybean (Heindl and Brun 1984; Rylott and Smith 1990). The abscission percentage ranges from 67 % to 82 % across different soybean cultivars (Wiebold et al. 1981). Biotic (insect and microbial infection) and abiotic (temperature changes, drought, salinity) stresses can influence abscission of soybean flowers and young pods. Ethylene is a hormone involved in early senescence and abscission of vegetative and reproductive organs under stress conditions (Xue et al. 2008). However, the metabolic network triggered by ethylene during organ abscission is not well understood (Singh et al. 2011). Genome-wide gene expression profiles of insertional mutants can help elucidate the relationship between gene function and plant development (Hirayama and Shinozaki 2010). However, no organ abscission mutants have been identified in soybean. In addition, a major challenge faced by scientists in the field of functional genomics is the prevalence of gene families (Alonso and Ecker 2006; Stangeland et al. 2005) with family members that encode proteins with redundant functions (Gu et al. 2003; Pasek et al. 2006). Due to the presence of gene-families with a functional redundancy it is often the case that mutations of a gene will not display any detectable phenotype (Kubis et al. 2004). To overcome this problem, the physiological effect of ethylene was previously characterized by employing precursors/inhibitors involved in ethylene biosynthesis or signaling (Nambeesan et al. 2012; Trivellini et al. 2011; Lin et al. 2009; Schaller 2012).
Ethylene biosynthesis starts with the conversion of S-adenosyl-l-methione (SAM) into 1-aminocyclopropane-1-carboxylic acid (ACC) catalyzed by the enzyme 1-aminocyclopropane-1-carboxylate synthase (ACS). ACC can then be converted to either 1-(malonylamino) cyclopropane-1-carboxylic acid (MACC) by ACC N-malonyl transferase, or to the end product, ethylene, by 1-aminocyclopropane-1-carboxylate oxidase (ACO; Adams and Yang 1979). The success in molecular cloning of ACS and ACO (Sato and Theologis 1989; Hamilton et al. 1991) genes suggests that these enzymes belong to multigene families and are regulated by developmental and environmental stimuli. Ethylene targets a family of membrane-localized receptors, which actively repress ethylene responses. The key elements in ethylene signaling pathway include Raf-like kinase constitutive triple response-1 (CTR1), transmembrane protein EIN2 (ethylene insensitive 2), and the EIN3 (ethylene insensitive 3)-like family of transcription factors (Schaller 2012). Ethylene binding to its receptors relieves the repression on downstream signaling components that results in the activation of EIN3-like transcription factors, which initiate the transcriptional response to ethylene. Ethylene induces genes encoding several other families of transcription factors, such as the ethylene response factor (ERF) and ethylene response DNA-binding factor (EDF) families, indicating that a transcriptional cascade acts downstream of the EIN3-like proteins (Schaller 2012; Dong et al. 2010). On the basis of this knowledge, some inhibitors and inducers of ethylene synthesis and action have been used to study the function of ethylene on plant growth and development. The pharmacological agent aminoethoxyvinylglycine (AVG) and silver can be used to inhibit ethylene responses through their ability to target ethylene biosynthesis or ethylene receptors, respectively (Schaller 2012). Ethephon (ETH) is an ethylene-releasing compound that can release ethylene in the presence of water at a pH above 3.5 (Lorbiecke and Sauter 1999). Typically, either ETH or the ethylene precursor ACC is employed to induce ethylene production (Zhang and Wen 2010).
Genome-wide gene profiling techniques are evolving rapidly. A combination of ethylene inhibitor/inducer and genome-wide gene expression profiling may help to reveal the complicated physiological networks triggered by ethylene in organ abscission in soybean. RNA-Seq is a recently developed approach for transcriptome profiling and is enabled by deep-sequencing technologies to provide a far more precise measurement of the levels of transcripts and isoforms than those provided by other methods (Wang et al. 2009). The holistic view of the transcriptome and its organization provided by the RNA-Seq method reveals many novel transcribed regions, splice isoforms, single nucleotide polymorphisms (SNPs), and the precise location of transcription boundaries (Cloonan and Grimmond 2008; Li et al. 2008; Lister et al. 2008; Wilhelm et al. 2008). RNA-Seq generates absolute, rather than relative, gene expression measurements, providing greater insight and accuracy than microarrays (Hoen et al. 2008; Marioni et al. 2008; Mortazavi et al. 2008). For example, Illumina sequencing technology offers millions of sequence reads from a single instrument run and thereby permits gene expression profiling experiments with an improved dynamic range and considerable cost savings (Xue et al. 2010).
To better understand the role of ethylene in regulating organ abscission in soybean, the RNA-Seq technique was used to elucidate the regulatory network triggered by ethylene. Genome-wide comparative transcriptome analysis was conducted using RNA from three treatments: (1) distilled water (control), (2) STS treatment (inhibitor of ethylene action), and (3) ETH treatment (ethylene releaser). The digital gene expression (DGE) libraries for each of the three treatments were constructed by the RNA-seq method and compared to identify the regulatory network involved in ethylene-mediated abscission of developing reproductive organs in soybean.
Materials and Methods
Plant Materials and Experimental Design
In 2011, soybean (cv. Tiefeng-31) was planted in a greenhouse under irrigated conditions in Jilin Normal University, Siping, Jilin province, China. On 2 May (21–29 °C), four seeds were planted per pot (7.6 L) and two plants were selected after 14 days. Twenty-four replicate pots per treatment were arranged in a completely randomized block design. At the beginning bloom-R1 stage (Fehr et al. 1971), plants (in their entirety) were sprayed with 0.5 mM STS or 400 mg/L ETH dissolved in distilled water twice with a 5-day interval between applications. The control plants were sprayed with distilled water. It is known that plants often increase ethylene production in response to environmental stress (Djanaguiraman et al. 2011). Additionally, it has been reported previously that when stress occurs at the flowering (R1 and R2), or early pod development (R3 and R4) stage, the pod number is reduced because of high abscission rate of flower and pod (Heindl and Brun 1984; Hirayama and Shinozaki 2010). To obtain complete gene-expression data of flower- and pod-development stages, a total of nine plants (three from each treatment: STS, ETH, and distilled water) were randomly selected, following which the auxiliary buds and surrounding node tissue (third node from the top) were dissected separately on days 0, 7, 14, 21, 28, and 35 (from R1 to R4 stage) after the second spraying. The plant samples were quickly frozen in liquid nitrogen and stored at −80 °C and later used for RNA extraction. The three remaining replicate pots were used to estimate the pod number by recording the abscission rate of flowers and pods during the reproductive stage and to measure the dry matter weight of the seeds and the straw at harvest. The numbers of flowers and pods were determined by artificial counting. Just before flower initiation, four plants from each treatment set were selected randomly and tagged for subsequent flower and pod counting. All emerging flower buds were marked and counted with an oil-based pen. The sepals were marked to avoid a repeat count in subsequent visits, which occurred at 2-day intervals. Flowers and pods on the stem were also counted. Pod location was counted every 2 days by touching the tips of all the young pods, which were not less than 2 cm long, with an oil-based pen. Percent flower and pod abscission were counted every 7 days. Percent of flower and pod abscission at different stages = number of flowers and pods with abscission × 100 %/(number of flowers and pods with abscission + the number of flowers and pods retained on the plant).
RNA Extraction
Total RNA of every sample was extracted from the dissected tissue using the RNA Easyspin Isolation System (Aidlab Biotech, Beijing, China) and treated with RNase-free DNase to eliminate the residual genomic DNA. All steps were followed according to the manufacturer’s instructions (Promega, Beijing, China). Total RNA was then quantified using a spectrophotometer and loaded on a denaturing agarose gel to check RNA concentration and integrity.
DGE Library Preparation and Sequencing
Total RNA was isolated from samples collected at different developmental stages. Equal amounts of RNA from each developmental stage were mixed to make a total RNA pool for library preparation and sequencing. Three pools of mRNA samples, one representing each treatment, were used to build libraries. Total mRNA was enriched using the oligo (dT) magnetic beads. Fragmentation buffer, which fragments the mRNA into short fragments (about 200 bp), was added. First strand cDNA was synthesized using random hexamer-primers, and the mRNA fragments were used as templates. Buffer, dNTPs, RNase H, and DNA polymerase I were added to synthesize the second strand. Double stranded cDNA was purified with QiaQuick PCR extraction kit and washed with EB buffer for end repair and single nucleotide A (adenine) addition. Finally, sequencing adaptors were ligated to the fragments. The required fragments were purified by agarose gel electrophoresis and enriched by PCR amplification. The library products were then ready for sequencing analysis using Illumina HiSeq™ 2000 (http://www.illumina.com).
Digital gene expression (DGE) tags were mapped by sequencing raw reads and filtering them to remove: (1) reads with adaptors, (2) reads in which unknown bases were more than 10 %, and (3) low quality reads. [50 % of the read had low quality bases (quality value, ≤5).] For tag annotation, the clean reads were mapped to reference sequences using SOAPaligner/soap2 (Li et al. 2009). Mismatches of no more than two bases were allowed in the alignment. The clean tags were designated as unambiguous clean tags. The gene expression was calculated by the number of reads mapped to the reference sequence and every gene.
Screening of Differentially Expressed Genes
To compare the differences in gene expression, the tag frequency in each differentially expressed gene (DGE) library was analyzed statistically according to the method described by Audic and Claverie (1997). A strict algorithm was used to identify DEGs between two samples. If x denotes the number of unambiguous clean tags from gene A, then, given that every gene’s expression occupies only a small part of the library, p(x) will closely follow the Poisson distribution.
If the total number of clean tags in sample 1 is N1, and the total number of clean tags in sample 2 is N2; gene A holds x tags in sample 1 and y tags in sample 2, then the probability of gene A expressed equally between two samples can be calculated by the following equation:
The P-value corresponds to the differential gene expression test. False discovery rate (FDR) is a method used to determine the threshold of P-value in multiple tests. Assume that we identified R DEGs in which S genes really show differential expression and the other V genes are false positive. If we decide that the error ratio “Q = V/R” must stay below a cutoff (e.g., 1 %), we should preset the FDR to a number no larger than 0.01 (Benjamini and Yekutieli 2001). We use “FDR ≤ 0.001 and the absolute value of log2Ratio ≥ 1” as the threshold to judge the significance of gene expression difference. More stringent criteria with smaller FDR and bigger fold-change value can be used to identify DEGs. The DEGs were used for gene ontology (GO) enrichment analyses.
GO Analysis of DEGs
Gene ontology enrichment analysis provides all GO terms that were significantly enriched in the DEGs as compared to those in the genome background, and filters the DEGs that correspond to biological functions. This method can be used to: (1) map all the DEGs with the GO terms in the database (http://www.geneontology.org/), (2) calculate gene numbers for every term, (3) find significantly enriched GO terms in DEGs as compared to those in the genome background by using hypergeometric test.
The calculation formula is:
In the equation, N is the number of all genes with GO annotation; n is the number of DEGs in N; M is the number of all genes that are annotated to the certain GO terms; m is the number of DEGs in M. The calculated P-value goes through Bonferroni correction, taking corrected P-value ≤ 0.05 as a threshold. GO terms fulfilling this condition are defined as significantly enriched GO terms in DEGs. Using this analysis, we were able to recognize the main biological functions of DEGs. The GO functional enrichment analysis enabled us to integrate the cluster analysis of expression patterns.
Pathway Enrichment Analysis of DEGs
Genes commonly interact with each other and these interactions can play crucial roles in certain biological functions (Amit et al. 2009). Pathway-based analysis helps to further understand biological functions of genes. KEGG is the major public pathway-related database (Kanehisa et al. 2008). Pathway enrichment analysis identifies significantly enriched metabolic pathways or signal transduction pathways in DEGs compared to those in the whole genome background. The calculating formula is the same as that in GO analysis. Here N is the number of all genes with KEGG annotation, n is the number of DEGs in N, M is the number of all genes annotated to specific pathways, and m is number of DEGs in M.
Validations of RNA-Seq Data by RT-qPCR
Real-time quantitative PCR (RT-qPCR) was performed to validate the RNA-Seq results for ethylene-responsive gene transcripts, whose change in tissues treated by ETH treatment was in the opposite direction to that in STS-treated tissues. Total RNA was extracted and pooled as described for DGE library preparation. At least two independent biological replicates of each treatment and three technical replicates of each biological replicate were used for the RT-PCR analysis. We used 100 ng total RNA for reverse transcription to cDNA using the PrimeScript RT reagent Kit (TaKaRa, Tokyo, Japan) in a 10-μL reaction mixture containing 1× PrimeScript buffer, 50 pmol random 6-mer primer, and 0.5 μL PrimeScript RT Enzyme Mix I. RT-qPCR was performed with 1 μL cDNA template, 1× reaction buffer, 0.2 mM dNTP mixture, 0.4 μM each primer, 0.6× SYBR Green I dye (Generay Biotech), and 0.3 units of rTaq DNA polymerase (TaKaRa) in a Rotor-Gene 2000 thermocycler (Corbett Research, Sydney, Australia) and analyzed using Rotor-Gene Real-Time Analysis Software 6.0 Build 14 (Corbett Research, Mortlake, Australia). Melting curves for each PCR were monitored carefully to avoid nonspecific amplifications. The results were normalized to the expression level of the constitutive β-tubulin gene. A relative quantitative method was used to evaluate the quantitative variation. The Primers used in RT-qPCR study are shown in Table 1.
Results
Effects of STS and ETH on Growth and Development in Soybean
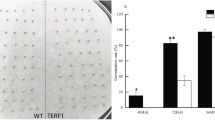
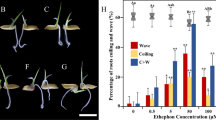
There were significant differences in vegetable growth and reproductive development of soybean among the three treatments (Table 2, Fig. 1). Dry weight of seed and straw at harvest was promoted by STS and reduced by ETH treatment. Plants treated with STS produced more flowers and pods and their seed yield increased by 55.6 % compared with the control. ETH treatment promoted the abscission ratio of flowers and pods significantly (P < 0.05), and the data indicated that ETH might have long-lasting negative effects on flower and pod development. STS treatment did not affect the abscission of flowers and pods significantly (P < 0.05). More lateral branches were observed at the base of fruit branch after ETH treatment (Fig. 1), and STS distinctly suppressed branching growth as compared to that of control and ETH treatment.
Illumina Sequencing and Sequence Mapping
Three cDNA libraries were generated from a mixture of RNA isolated from buds and surrounding node tissue from the third nodes from the apex of the plants, for each of the three treatments. After filtering the low quality tags, the total number of clean tags in each library ranged from 11 to 12 million (Table 3), and the percentage of clean tags among the raw tags in each library ranged from 99.54 % to 99.56 % (Supplement file 1). Reads that failed to meet these criteria or mapped to multiple locations were excluded. Among the clean tags, the number of sequences that could be mapped to unigenes ranged from 9.7 to 10.5 million, and the percentage of these clean tags ranged from 82.8 to 83.7 % in the three libraries (Table 3).
Comparative analysis of the STS and ETH treatment libraries revealed significant expression changes in 1,524 genes. Among these genes, 1,002 up-regulated and 522 down-regulated genes were identified in the STS library as compared to those in the ETH library (Fig. 2). Compared with the control treatment, only 522 DEGs were regulated by STS treatment, but 1,043 DEGs were regulated by ETH treatment. In order to gain further insight into ethylene-regulated target genes during soybean reproductive development, the genes whose expression changes in opposite directions in control-VS-STS and control-VS- ETH DEGs were chosen as the direct ethylene-responsive genes (Table 4).
GO Classification
GO assignments were used to classify the functions of the predicted ethylene regulative genes. Based on DEGs, 2,571 differentially expressed sequences from all the three treatments can be categorized into 28 functional groups (Fig. 3). In each of the three main categories (biological process, cellular component and molecular function) of GO classification, ‘response to stimulus’, ‘external encapsulating structure’ and ‘catalytic activity’ terms are dominant in the DEGs, respectively. In the control-vs-STS treated samples, a high-percentage of genes that showed change in expression belonged to the category of ‘molecular function’, and a smaller number of genes that showed expression changes belonged to the ‘biological process’ category (Fig. 3). DEGs did not show a difference in expression in the ‘cellular component’ category of the control-vs-ETH library. The GO analysis showed that the functions of the 17 ethylene-responsive genes were involved mainly in the various stimulus responses and catalytic activities, which may in turn be involved in the ethylene-mediated response.
Pathway Enrichment Analysis of DEGs
Ethylene-affected biological pathways were evaluated by enrichment analysis of DEGs. Significantly enriched metabolic pathways and signal transduction pathways were identified. Pathways with a Q value < 0.05 were significantly enriched. DEGs with pathway annotation after ETH and STS treatment were listed according to enrichment priority (Tables 5, 6). Secondary metabolism (flavone and flavonol biosynthesis, phenylpropanoid biosynthesis, glyoxylate and dicarboxylate metabolism, caffeine metabolism, some hormone metabolism) and primary metabolism (limonene and pinene degradation, ABC-transporter-dependent pathways and circadian rhythm–plant–reference pathway) were affected by up-regulated DEGs after ETH treatment. After STS treatment, the up-regulated DEGs affected flavonoid biosynthesis, pentose and glucuronate interconversions, starch and sucrose metabolism and some hormone metabolism pathway. Plant–pathogen interaction, hormone biosynthesis and signal transduction pathways were identified to be enriched and were affected by down-regulated DEGs.
Further analysis of pathways showed that the expression of cysteine synthase and ACC oxidase gene expression was enhanced by ETH treatment, and that ACC syntheses was inhibited by STS treatment in ethylene biosynthesis pathway. Similarly, the other regulated plant-hormone biosynthesis pathways included phenylalanine metabolism, carotenoid biosynthesis, linoleic acid metabolism along with the plant hormone signal transduction pathway (see electronic supplementary material, Supplementary files 2, 3). The up-regulated DEGs of control-vs-STS related to plant hormone biosynthesis and signal transduction pathway were plant hormone auxin, cytokine, brassinosteroid and salicylic acid, while the down-regulated DEGs included genes involved in the biosynthesis process of plant hormone gibberellin, abscisic acid, ethylene and jasmonic acid. The ETH treatment had the contrary effects on the biosynthesis and signal transduction pathway of plant hormone auxin, abscisic acid, ethylene, salicylic acid and jasmonic acid. These findings provided clues to unravel the complicated signal networks of phytohormones.
DEG Functional Annotation and RT-qPCR Confirmation
The putative function of the 17 ethylene-responsive genes was determined on the basis of sequence similarity with the genomes of soybean, Medicago truncatula, and Arabidopsis thaliana using the BLASTN program. In the similarity range, 16 transcripts were classified as “known protein,” and only 1 unigene had unknown annotations. The seven up-regulated expression unigenes in ETH treatment included two ACC-oxidase genes (gnl|UG|Gma#S53085092, gnl|UG|Gma#S53086117), one central component PP2Cs gene of abscisic acid (ABA) signal transduction (gnl|UG|Gma#S48312502), two genes responding to stimulus [maturation protein gene (MAT1), and an uncharacterized acetyltransferase gene (At3g50280)], cinnamyl alcohol dehydrogenase gene (gnl|UG|Gma#S48316018) involved in cell-wall degradation and Glycine max glucan endo-1, 3-beta-glucosidase 14-like gene involved in cellulose degradation. The remaining nine up-regulated unigenes in STS treatment and down-regulated genes in ETH treatment included two anthocyanidin synthase genes (547615, gnl|UG|Gma#S39303376) and two non-specific lipid-transfer protein genes (100305883, 100306032), one aconitate hydratas gene (gnl|UG|Gma#S47723140), one putative cell wall protein gene (gnl|UG|Gma#S4992524), one bidirectional sugar transporter SWEET1-like gene (gnl|UG| Gma#S53088986), and one glutathione S-transferase F11-like gene (plays a role in plant growth and development in vivo and shoot regeneration in vitro). On the basis of the GO functional classification, the nine up-regulated genes in control-VS-STS treatment were thought to be involved in the biosynthesis of carbohydrate metabolism, translocation, plant growth, and development, and the seven up-regulated genes in control-vs-ETH were involved in ABA and ethylene metabolism, stress stimulus, abscission, ripeness, and senescence processes.
We further analyzed the expression of these 17 responsive genes by RT-qPCR to confirm their gene expression pattern after different treatments. Transcript levels for each of the 17 genes were quantified using SYBR Green technology (Fig. 4), and the results showed good correlation with those obtained by RNA-seq.
Gene Expression Variations During Different Developmental Stages
We also focused on gene expression variations related to ethylene biosynthesis at the reproductive developmental stage. ACS and ACO gene expression variations were assessed by RT-qPCR (Figs. 5, 6). The results showed that gene expression levels of ACO and ACS were both low on day 0 and reached their peak value on days 14 and 21 in the control treatment, respectively. The ACO gene expression in ETH treatment was significantly higher compared with that during the corresponding period in the control treatment (FDR ≤ 0.001), but their change dynamics were similar to each other. ACS gene expression was inhibited by STS and ETH treatment on days 7 and 14. On days 21–35, ACS gene expression was inhibited by STS treatment and promoted by ETH treatment.
Discussion
Ethylene is a hormone involved in the senescence and abscission of vegetative and reproductive organs under stress conditions. Our results showed that application of STS at the R1 stage promoted seed yield significantly (P < 0.05), as indicated by the increased flower and pod number (Table 2); a similar phenomenon was observed previously by Chen et al. (2011). Aminoethoxyvinylglycine and 1-methylcyclopropene—two ethylene perception inhibitors—were both beneficial for flower-bud and pollen formation due to the enhanced photosynthetic rate of source leaves (Cheng et al. 2008, 2010; Djanaguiraman et al. 2011). In contrast, ETH treatment promoted the abscission of flowers and pods and lateral branch differentiation (Table 2, Fig. 1), suggesting that ethylene plays an important role in early abscission of flowers and pods and in the transition process between the vegetative and reproductive development (Cheng et al. 2010; Chen et al. 2011; Djanaguiraman et al. 2011; Gniazdowska et al. 2010).
Recent studies have highlighted the significance of high-throughput expression data, particularly with the integration of large, diverse data sets, in constructing biochemical and regulatory networks (Hoen et al. 2008; Marioni et al. 2008; Mortazavi et al. 2008; Eveland et al. 2010; Amit et al. 2009). In this study, we analyzed the short-read, sequence-based expression profiles related to ethylene-mediated organ abscission using Illumina’s DGE technology. Our results suggest that deep sequencing of 20- to 21-nucleotide DGE tags can be used to successfully establish genome-wide expression profiles in soybean and detect differences in transcript abundance over a broad dynamic range. We identified 11–12 million clean tags from three DGE libraries. Of these, 82.8 % to 83.7 % clean tags could be mapped to unigenes. Results from these expression analyses provided testable hypotheses for resolving which regulatory and biochemical processes contribute to ethylene-regulated soybean reproductive development and abscission. To investigate the functions of the identified ethylene responsive genes, we explored their annotations and functional category in the soybean Genome Annotation Project database (Table 4). Several members of the ethylene receptors gene families, which are known to be induced rapidly by exogenous ethylene (Zhang and Wen 2010; Chervin and Deluc 2010), are listed in Supplement file 4. These results confirmed the reliability of the RNA-seq. Our homogenous expression data sets allowed us to perform counterpart comparisons with ethylene inducers and ethylene inhibitors.
GO analysis and the pathway enrichment analysis of DEGs identified two major functional and process-based categories, which were involved in catalytic activity and response to stimulus, respectively, in the DEGs of control-vs-ETH and control-vs-STS. Pathway enrichment analysis revealed the most significantly affected pathways was the “secondary metabolism, ABA signal and ethylene signal “pathway after ETH treatment. STS treatment affected metabolism of starch and sucrose and signal transduction of auxin, zeatin, brassinosteroid, and salicylic acid. Therefore, this finding implies that the role of ethylene was related to photosynthesis, stress response, carbohydrate transportation, signal transduction and hormone crosstalk (Cheng et al. 2010; Chervin and Deluc 2010; Zhu et al. 2011; Iqbal et al. 2011). In the present study, 17 ethylene-responsive genes were identified as potential key components in complicated ethylene signaling networks. ACO—the key enzyme responsible for ethylene biosynthesis—was up-regulated during soybean reproductive development after ETH treatment, and therefore ethylene promotes ethylene biosynthesis during soybean reproductive development by positive feedback regulation of ACO (Schaller 2012). The results of qRT-PCR analysis confirmed that the expression of ACO was up-regulated after 14 days of treatment, and this implied that there was a sustained feedback regulation of ethylene. Expression of PP2C was also up-regulated after ETH treatment. The Arabidopsis PP2C enzymes, such as the ABI1/ABI2 PP2Cs, have been identified as central components in abscisic acid (ABA) signal transduction (Ma et al. 2009). An increase in ethylene due to ETH treatment, stress-induced hormones (ABA and ethylene), and some genes associated with stress responses (MAT1, BAHD acyltransferase, CAD) could probably lead to the abscission of flowers and pods in soybean. Furthermore, glucan endo-1, 3-beta-glucosidase 14-like gene, which is involved in cellulose degradation, was significantly up-regulated in the ETH treatment as compared with that in STS treatment (FDR ≤ 0.001), suggesting that stress-induced ethylene is closely related with organ abscission.
The function of up-regulated genes in the STS treatment included genes related to sugar biosynthesis and transport; cell wall metabolism and structure; and synthesis of flower pigment. The expression of Aconitate hydratase, an iron-containing enzyme gene that catalyzes the dehydration of citrate to cis-aconitate (a reaction of significance in the tricarboxylic acid cycle) and a bidirectional sugar transporter SWEET1-like gene were significantly up-regulated (FDR ≤ 0.001). Changes in cell wall extensibility appears to be the primary factor regulating segmental growth rates in the root elongation zone (Neumann 1995). Among the 17 ethylene-responded genes, we found 2 non-specific lipid-transfer protein gene and 1 putative cell wall protein gene in the STS treatment, which may play a role in cell-wall metabolism and structure and in suppressing the abscission rate of organs (Nieuwland et al. 2005). Briefly, the expression of genes involved in sugar metabolism and sugar transport in both flowers and pods would contribute to efforts to improve the yield of soybean.
In conclusion, the results of the present study show that inhibition of ethylene could inhibit the abscission of flowers and pods in soybean. Potential target genes regulated by ethylene were identified as follows: ACC-oxidase, PP2Cs, maturation protein gene(MAT1), acetyltransferase, BAHD acyltransferase, CAD, bidirectional sugar transporter SWEET1-like, aconitate hydratas, and Anthocyanidin synthase. These identified genes provided new insights into the molecular mechanism of ethylene-mediated networks regulating abscission and development.
References
Adams DO, Yang SF (1979) Ethylene biosynthesis: identification of 1-aminocyclopropane-l -carboxylic acid as an intermediate in the conversion of methionine to ethylene. Proc Natl Acad Sci USA 76:170–174
Alonso JM, Ecker JR (2006) Moving forward in reverse: genetic technologies to enable genome-wide phenomic screens in Arabidopsis. Nat Rev Genet 7:524–536
Amit I, Garber M, Chevrier N, Leite AP, Donner Y et al (2009) Unbiased reconstruction of a mammalian transcriptional network mediating pathogen responses. Science 326:257–263
Audic S, Claverie JM (1997) The significance of digital gene expression profiles. Genome Res 7:986–995
Benjamini Y, Yekutieli D (2001) The control of the false discovery rate in multiple testing under dependency. Ann Stat 29:1165–1188
Chen MK, Hsu WH, Lee PF, Thiruvengadam M, Chen HI et al (2011) The MADS box gene, FOREVER YOUNG FLOWER, acts as a repressor controlling floral organ senescence and abscission in Arabidopsis. Plant J 68:168–185
Cheng YQ, Yang J, An LJ, Liu JF, Chen ZW (2008) Effects of ethylene on yield and floral organ development in soybean (Glycine max). J Zhejiang Univ (Agric Life Sci) 34:540–545
Cheng YQ, Zhao GL, Liu JF, Chen ZW, Zhang WW (2010) Effects of ethylene inhibitor and promoter on photosynthetic characteristics of soybean (Glycine max) seedling leaves. J Zhejiang Univ (Agric Life Sci) 36:419–426
Chervin C, Deluc L (2010) Ethylene signaling receptors and transcription factors over the grape berry development: gene expression profiling. Vitis 49:129–136
Cloonan N, Grimmond SM (2008) Transcriptome content and dynamics at single-nucleotide resolution. Genome Biol 9:234
Djanaguiraman M, Prasad PVV, Al-Khatib K (2011) Ethylene perception inhibitor 1-MCP decreases oxidative damage of leaves through enhanced antioxidant defense mechanisms in soybean plants grown under high temperature stress. Environ Exp Bot 71:215–223
Dong CH, Jang M, Scharein B, Malach A, Rivarola M et al (2010) Molecular association of the Arabidopsis ETR1 Ethylene receptor and a regulator of ethylene signaling, RTE1. J Biol Chem 285:40706–40713
Eveland A, Satoh-Nagasawa N, Goldshmidt A, Meyer S, Beatty M et al (2010) Digital gene expression signatures for maize development. Plant Physiol 154:1024–1039
Fehr WR, Caviness CF, Burmood DT, Pennington JS (1971) Stage of development descriptions for soybeans, glycine max. (L). Merrill. Fehr & Caviness et al. Crop Sci 11:929–931
Gniazdowska A, Krasuska U, Czajkowska K, Bogatek R (2010) Nitric oxide, hydrogen cyanide and ethylene are required in the control of germination and undisturbed development of young apple seedlings. Plant Growth Regul 61:75–84
Gu Z, Steinmetz LM, Gu X, Scharfe C, Davis RW et al (2003) Role of duplicate genes in genetic robustness against null mutations. Nature 421:63–66
Hamilton AJ, Bouzayen M, Grierson D (1991) Identification of a tomato gene for the ethylene-forming enzyme by expression in yeast. Proc Natl Acad Sci USA 88:7434–7437
Heindl JC, Brun WA (1984) Patterns of reproductive abscission, seed yield, and yield components in soybean. Crop Sci 24:542–545
Hirayama T, Shinozaki K (2010) Research on plant abiotic stress responses in the post-genome era: past, present and future. Plant J 61:1041–1052
Hoen PA, Ariyurek Y, Thygesen HH, Vreugdenhil E, Vossen RH et al (2008) Deep sequencing-based expression analysis shows major advances in robustness, resolution and interlab portability over five microarray platforms. Nucleic Acids Res 36:e141
Iqbal N, Nazar R, Syeed S, Masood A, Khan NA (2011) Exogenously-sourced ethylene increases stomatal conductance, photosynthesis, and growth under optimal and deficient nitrogen fertilization in mustard. J Exp Bot 62:4955–4963
Kanehisa M, Araki M, Goto S, Hattori M, Hirakawa M et al (2008) KEGG for linking genomes to life and the environment. Nucleic Acids Res 36:480–484
Kubis S, Patel R, Combe J, Bedard J, Kovacheva S et al (2004) Functional specialization amongst the Arabidopsis Toc159 family of chloroplast protein import receptors. Plant Cell 16:2059–2077
Li H, Ruan J, Durbin R (2008) Mapping short DNA sequencing reads and calling variants using mapping quality scores. Genome Res 18:1851–1858
Li RQ, Yu C, Li YR, Lam TW, Yiu SM et al (2009) SOAP2: An improved ultrafast tool for short read alignment. Bioinformatics 25:1966–1967
Lin ZF, Zhong SL, Grierson D (2009) Recent advances in ethylene research. J Exp Bot 60:3311–3336
Lister R, O’Malley RC, Tonti-Filippini J, Gregory BD, Berry CC et al (2008) Highly integrated single-base resolution maps of the epigenome in Arabidopsis. Cell 133:523–536
Lorbiecke R, Sauter M (1999) Adventitious root growth and cell cycle induction in deepwater rice (Oryza sativa L.). Plant Physiol 119:21–29
Ma Y, Szostkiewicz I, Korte A, Moes D, Yang Y et al (2009) Regulators of PP2C phosphatase activity function as abscisic acid sensors. Science 324:1064–1068
Marioni JC, Mason CE, Mane SM, Stephens M, Gilad Y (2008) RNA-seq: an assessment of technical reproducibility and comparison with gene expression arrays. Genome Res 18:1509–1517
Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 5:621–628
Nambeesan S, AbuQamar S, Laluk K, Mattoo AK, Mickelbart MV et al (2012) Polyamines attenuate ethylene-mediated defense responses to abrogate resistance to Botrytis cinerea in tomato. Plant Physiol 158:1034–1045
Neumann PM (1995) The role of cell wall adjustment in plant resistance to water deficits (review and interpretation). Crop Sci 35:1258–1266
Nieuwland J, Feron R, Huisman BAH, Fasolino A, Hilbers CW et al (2005) Lipid transfer proteins enhance cell wall extension in tobacco. Plant Cell 17:2009–2019
Pasek S, Risler JL, Brezellec P (2006) The role of domain redundancy in genetic robustness against null mutations. J Mol Biol 362:184–191
Rylott PD, Smith ML (1990) Effects of applied plant growth substances on pod set in broad beans (Vicia faba var. Major). J Agric Sci Camb 114:41–47
Sato T, Theologis A (1989) Cloning the mRNA encoding 1-aminocyclopropane-1- carboxylate synthase, the key enzyme for ethylene biosynthesis in plants. Proc Natl Acad Sci USA 86:6621–6625
Schaller GE (2012) Ethylene and the regulation of plant development. BMC Biol 10:9. doi:10.1186/1741-7007-10-9
Singh AP, Tripathi SK, Nath P, Sane AP (2011) Petal abscission in rose is associated with the differential expression of two ethylene-responsive xyloglucan endotransglucosylase/hydrolase genes, RbXTH1 and RbXTH2. J Exp Bot 62:5091–5103
Stangeland B, Nestestog R, Grini PE, Skrbo N, Berg A et al (2005) Molecular analysis of Arabidopsis endosperm and embryo promoter trap lines: reporter-gene expression can result from T-DNA insertions in antisense orientation, in introns and in intergenic regions, in addition to sense insertion at the 5′ end of genes. J Exp Bot 56:2495–2505
Trivellini A, Ferrante A, Vernieri P, Serra G (2011) Effects of promoters and inhibitors of ethylene and ABA on flower senescence of Hibiscus rosa-sinensis L. J Plant Growth Regul 30:175–184
Wang Z, Gerstein M, Snyder M (2009) RNA–Seq: a revolutionary tool for transcriptomics. Nat Rev Genet 10:57–63
Wiebold W, Ashley D, Boerma HR (1981) Reproductive abscission levels and patterns for eleven determinate soybean cultivars. Agron J 73:43–46
Wilhelm BT, Marguerat S, Watt S, Schubert F, Wood V et al (2008) Dynamic repertoire of a eukaryotic transcriptome surveyed at single-nucleotide resolution. Nature 453:1239–1243
Xue JQ, Li YH, Tan H, Yang F, Ma N et al (2008) Expression of ethylene biosynthetic and receptor genes in rose floral tissues during ethylene-enhanced flower opening. J Exp Bot 59:2161–2169
Xue J, Bao YY, Li BL, Cheng YB, Peng ZY et al (2010) Transcriptome analysis of the brown planthopper Nilaparvata lugens. PLoS One 5:e14233
Zhang W, Wen CK (2010) Preparation of ethylene gas and comparison of ethylene responses induced by ethylene, ACC and ethephon. Plant Physiol Biochem 48:45–53
Zhu H, Dardick CD, Beers EP, Callanhan AM, Xia R et al (2011) Transcriptomics of shading-induced and NAA induced abscission in apple (Malus domestica) reveals a shared pathway involving reduced photosynthesis, alterations in carbohydrate transport and signaling and hormone crosstalk. BMC Plant Biology 11:138
Acknowledgments
We are very grateful to Prof. Guohua Mi for his comments and advice. We also thank anonymous reviewers for critical and constructive suggestions to improve this manuscript. This study was supported by grants from National Natural Science Foundation of China (Grant No.31000691/C130301), Special Foundation for Young Scientists of Jilin Province, China (Grant No.201201084) and China Scholarship Council Project (201208220004).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplement file 1
Classification and percentages of raw reads of STS, Control and ETH treated sample. Only Adaptor Number of reads containing adaptor, Containing N number of reads containing N, Low Quality number of low quality reads, Clean Reads the number of clean reads. (DOC 73 kb)
Supplement file 2
Detailed information of plant hormone signal transduction pathway in KEGG and the DEGs of STS treatment compared to the control treatment in the pathway. Red borders Up-regulated genes green borders down-regulated genes, black borders Non-change genes. (DOC 40 kb)
Supplement file 3
Detailed information of plant hormone signal transduction pathway in KEGG and the DEGs of ETH treatment compared to the control treatment in the pathway. Red borders Up-regulated genes green borders down-regulated genes, black borders Non-change genes. (DOC 44 kb)
Supplement file 4
Several members of the ethylene receptors gene families were induced rapidly after ETH treatment and inhibited by STS treatment in ethylene signaling (DOC 42 kb)
Rights and permissions
About this article
Cite this article
Cheng, YQ., Liu, JF., Yang, X. et al. RNA-seq Analysis Reveals Ethylene-Mediated Reproductive Organ Development and Abscission in Soybean (Glycine max L. Merr.). Plant Mol Biol Rep 31, 607–619 (2013). https://doi.org/10.1007/s11105-012-0533-4
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-012-0533-4