Abstract
Flowering, transition from vegetative to reproductive phase in plants, is regulated by both endogenous and environmental signals. FLOWERING LOCUS C (FLC) in Arabidopsis encodes a dosage-dependent repressor of flowering. We have isolated a FLC-related sequence from non-heading Chinese cabbage (Brassica campestris ssp. chinensis Makino) and named it as BcFLC1. In this study, we found that fertility was affected by the overexpression of BcFLC1 in Arabidopsis. The floral morphology of shortening anther filaments and the phenotype of reduced or small siliques were observed in BcFLC1 overexpression plants. RT-PCR showed that the reduced fertility in Arabidopsis by the overexpression of FLC was related with the enhanced expression of anther filaments suppressor, such as RGA and RGL genes, and declined expression of SEPALLATA3 (SEP3) gene, encoding a MADS-box transcription factor involved in flower and ovule development.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In flowering plants, the switch from vegetative to reproductive growth in plants is controlled by a number of developmental and environmental signals. Extensive genetic and physiological analyses on Arabidopsis thaliana have revealed that floral induction is regulated by at least four major genetic pathways, namely, long-day, autonomous, vernalization, and gibberellin-dependent pathways (Onouchi et al. 2000; Samach et al. 2000; Wang et al. 2011). All of these four pathways commonly regulate the so-called flowering pathway integrators including FT, SUPPRESSOR OF OVEREXPRESSION OF CO1 (SOC1), and LFY, and the exact flowering time is determined by the expression level of these integrators (Blázquez and Weigel 2000; Lee et al. 2000; Moon et al. 2005; Ma et al. 2011).
FLC, which encodes a MADS-box transcription factor, functions as a repressor of flowering (Sheldon et al. 1999, 2000; He and Amasino 2005; Barreto et al. 2011). Two flowering-time genes, FT and SOC1, have been reported to be targeted by FLC (Searle et al. 2006). FLC binds to the promoter of SOC1 as well as to the first intron of FT. This binding prevents the transcriptional activation of these genes (Helliwell et al. 2006). The floral meristem identity gene LFY is regulated both by CO, a component of the photoperiod pathway, and GA (Blázquez and Weigel 2000). These act through different motifs within the LFY promoter (Samach et al. 2000).
Constitutive expression of FLC in many plants could delay flowering (Kim et al. 2007; Michaels and Amasino 1999; Sheldon et al. 1999). Meanwhile, overexpression of FLC also caused defects in floral morphology and reproduction. The phenotypes of short stamens, reduced pollen production, and larger carpel were detected, and fertility was slightly reduced in the transgenic lines (Tadege et al. 2001; Hepworth et al. 2002). However, there are no data to show the mechanism by which FLC affects floral morphology and reproduction.
Non-heading Chinese cabbage, which originated in China, is one of the most widely cultivated vegetables in China, especially in the south of the country. Non-heading Chinese cabbage plants remain in the vegetative growth phase until they have experienced prolonged exposure to cold temperature, known as vernalization. This inhibition of flowering is caused by the high levels of FLC expression. To increase the product value of non-heading Chinese cabbage by inhibiting the floral transition, BcFLC1 (not logged), homologous to the AtFLC1 gene, which encodes a floral repressor, was isolated from the non-heading Chinese cabbage “NJ074” (Rong et al. 2010).
In this study, we performed quantitative real-time PCR (qRT-PCR) experiments using both a 35S::BcFLC1 overexpression line and wild-type A. thaliana, and we report that the expression of filament suppressors, RGA and RGL genes (Tyler et al. 2004; Achard et al. 2006), was remarkably enhanced in 35S::BcFLC1 line, while SEP3 gene, encoding a MADS-box transcription factor involved in flower and ovule development (Kaufmann et al. 2009; Dornelas et al. 2010), was slightly downregulated.
Materials and Methods
Plant Materials and Growth Conditions
Surface-sterilized seeds of A. thaliana cv. Columbia (Col) were placed on sterile horizontal agar plates at 4°C for 5 days and then incubated in environmentally controlled sterile growth chambers at 23/18°C day/night under 60 % relative humidity. Cool white fluorescent lights supplied photons at 120 μmol m−2 s−1 with a 16-h light and 8-h dark photoperiod.
Generation of Transgenic Plants
To express the BcFLC1 genes in Arabidopsis, BcFLC1 gene was inserted into the pEarleyGate103 binary vector along with an herbicide-resistance gene (bar; Interuniversity Institute for Biotechnology; http://www.psb.rug.ac.be/gateway) as a selectable maker using Gateway Technology (Invitrogen; https://www.invitrogen.com) to construct pGate-BcFLC1. The final binary vectors, pGate-BcFLC1, were introduced into Agrobacterium tumefaciens GV3101.
A. thaliana Columbia was transformed with the BcFLC1 construct. Flowers were sprayed with A. tumefaciens GV3101 suspended in 5 % sucrose, and then the plants were incubated in a growth chamber at 25°C and 100 % humidity for 1 day and finally allowed to grow in a growth chamber under a 16-h photoperiod at 23°C. Transgenic plants were confirmed by semiquantitative PCR.
Analysis of Gene Expression
Total RNA was isolated from plant leaves by the RNeasy plant mini kit (Qiagen) according to the manufacturer’s instructions. For cDNA production, 4 μg of total RNA was reverse transcribed with oligo(dT)18 primer (Fermentas) in a 20 μL reaction mixture using RevertAid M-MLV reverse transcriptase (Fermentas). After heat inactivation, total volume of the reaction mixture was diluted in 580 μL of sterilized water, and 4 μL was used for the real-time quantitative reverse-transcription (RT)-PCR. All quantitative RT-PCR analyses were performed by iQ5 multicolor real-time PCR detection system (Bio-Rad) using 2× SYBR Green SuperMix (Bio-Rad 170-8882). We adopted the guidelines for the experimental design and statistical analysis of quantitative RT-PCR data (Rieu and Powers 2009). The PCR condition was as follows: 40 cycles of PCR (95°C for 30 s, 60°C for 30 s, and 72°C for 20 s) after the initial denaturation step of 2 min at 95°C. Data were collected at 72°C in each cycle, and the expression levels of genes were calculated by iQ5 optical system software version 2.0 using TUB2 as the reference gene. The quantitative RT-PCR analysis was biologically repeated three times, and each time consisted of three technical replicates. Real-time RT-PCR primer sequences are shown in Table 1.
Results
Constitutive Expression of BcFLC1 Genes Delays Flowering in Transgenic Arabidopsis
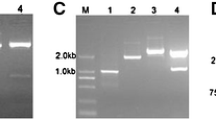
Arabidopsis thaliana Columbia was transformed with A. tumefaciens GV3101 containing the pGate-BcFLC1 vector (Fig. 1). Twelve independent lines of transgenic plants with BcFLC1 (T1 generation) were obtained by using two applications of herbicide (Basta, 0.3 % solution), and transgenic plants of the T2 generation were confirmed by semiquantitative PCR (Fig. 2). Among them, one line (named OE5) having high expression of BcFLC1 was chosen for further experiments.
T2 plants derived from parental lines that showed late flowering produced many rosette leaves (up to 52 leaves) and uniformly showed a late-flowering phenotype (Fig. 3a). Transgenic Arabidopsis plants overexpressing the BcFLC1 gene flowered approximately 15–30 days later than nontransgenic plants (data not shown). Some transgenic plants failed to flower during the experiment. This result indicated that BcFLC1 affected flowering time in the next generation.
Comparison of Col and 35S::BcFLC1 lines. a Effects of BcFLC1 overexpression on the flowering time of transgenic A. thaliana Columbia-ecotype lines. b Effects of BcFLC1 overexpression on the fertility of transgenic A. thaliana Columbia-ecotype lines. c The number of mature seeds in the silique of Col. d The number of mature seeds in the silique of 35S::BcFLC1. e The length of anther filaments in Col. f The length of anther filaments in 35S::BcFLC1. g Comparison of the number of seeds between Col and 35S::BcFLC1 line
Fertility Associated with Overexpression of BcFLC1
Overexpression of BcFLC1 affected floral morphology and reproduction. As shown in Fig. 4, the phenotype of the reduced number of siliques and small siliques were detected in 35S::BcFLC1 overexpression line (Fig. 3b). Meanwhile, in 35S::BcFLC1, the number of mature seeds in a silique is less (about ten seeds) than that in Col (Fig. 3c, d, and g). This result indicated that fertility is reduced in the transgenic lines. Next, we detected the morphology of flowers in Col and 35S::BcFLC1 overexpression line. Comparing to Col, shortening of anther filaments were found in transgenic lines (Fig. 3e and f). Meanwhile, some flowers of transgenic Arabidopsis plants produced anthers with little pollen. So we suggest BcFLC1 affects not only flowering time, but also floral morphology and reproduction in the next generation.
GA Pathway Was Affected in 35S::BcFLC1 Overexpression Line
The flc-11 overexpression mutant in Arabidopsis requires more GA3 to flower than the parental C24 (Sheldon et al. 1999). This suggested that FLC may have a quantitative effect on GA action or biosynthesis. In this study, to address the relationship between the overexpression of BcFLC1 and reproduction in transgenic lines, we used qRT-PCR assays detecting the expression of RGA and RGL, anther filaments suppressor involving in GA pathway (Tyler et al. 2004; Achard et al. 2006), and GAMYB and SEP3, involving in anthers and ovule development (Kaufmann et al. 2009; Dornelas et al. 2010). In qRT-PCR assays, the expression level of RGA and RGL mRNA were upregulated (Fig. 4). Meanwhile, the expression of SEP3 was slightly downregulated (Fig. 4). However, the GAMYB expression level was not obviously changed (Fig. 4). This result indicated that BcFLC1 affected GA action and SEP3 transcription in 35S::BcFLC1 overexpression line.
Discussion
Constitutive expression of BcFLC1 causes late flowering in Arabidopsis (Fig. 3). Some transgenic plants failed to flower during the experiment. These results indicated that BcFLC1 have effects on flowering time similar to AtFLC (Tadege et al. 2001). Semiquantitative analysis was performed to confirm the BcFLC1 expression levels in the transgenic Arabidopsis plants. Transgenic plants with the highest BcFLC1 expression showed a severe late-flowering phenotype, whereas transgenic plants that flowered similarly to the wild-type control plants showed much lower BcFLC1 transcript levels (Fig. 2). These results indicated that the BcFLC1 transcript level was very closely correlated with the timing of flowering in these transgenic plants.
Morphological changes in floral organs with the overexpression of FLC have been reported in Arabidopsis and Brassica napus (Tadege et al. 2001; Hepworth et al. 2002). Tadege et al. (2001) reported the stamens of the sterile transgenic plants were shorter than the carpel, and the anthers remained below the stigma making self-pollination difficult. This is consistent with our result that RGA and RGL, inhibitors of anther filaments elongation, were upregulated (Fig. 4). Some of the flowers of transgenic Arabidopsis plants produced anthers with little or no pollen and produced petioles at the bases of the mature siliques and inflorescences inside some siliques (Hepworth et al. 2002). Shore and Sharrocks (1995) reported that these morphological changes could be explained by ectopic expression of MADS-box gene which causes abnormal plant developments, especially in floral organs, possibly because of disruption of MADS-box protein complexes. Deng et al. (2011) suggested that FLC is involved in floral pattern regulation. FLC, which binds to a CArG box 2.7 kb upstream of the translation start of SEP3, involving in flower development, affected floral morphology and reproduction. SEP3, involving in anthers and ovule development, was slightly downregulated in 35S::BcFLC1 overexpression line (Fig. 4). The reduced SEP3 and shortening of anther filaments may explain the cause of defects in fertility (Fig. 3).
Meanwhile, some papers showed the flc-11 overexpression mutant in Arabidopsis requires more GA3 to flower than the parental C24 (Sheldon et al. 1999). This suggested that FLC may have a quantitative effect on GA action or biosynthesis. The abnormality of GA action or biosynthesis affects flower development (Fig. 5). Currently, it is not clear how FLC regulates GA; thus, it would be worthwhile to determine whether FLC affects GA action or biosynthesis.
Seo et al. (2009) suggested the soc1-2 mutants, which have relatively higher CBF1, CBF2, CBF3, and COR15a expression, showed enhanced resistance to cold resistance. However, the soc1-101D mutants, which have a higher SOC1 expression, showed much weaker induction of COR15a expression compared with wild type. This result suggests that SOC1 attenuates the induction of COR15a gene in response to cold. The same as Seo et al. (2009), in 35S::BcFLC1 overexpression line, the expression of SOC1 was repressed, and then the CBFs expression was increased (data not shown). Achard et al. (2006) concludes that CBF1 enhances RGA and RGL accumulation and, thereby, inhibits GA bioactive. RGA and RGL, belonging to DELLAs (Silverstone et al. 2001; Lee et al. 2002; Wen and Chang 2002), are key components of the GA-signaling pathway. GA promotes growth by overcoming DELLA-mediated growth restraint (Dill and Sun 2001; King et al. 2001; Silverstone et al. 2001; Harberd 2003). Accordingly, we speculated 35S::BcFLC1 transgenic plants that enhanced CBFs accumulate less bioactive GA and, as a consequence, exhibit shortening of anther filaments and reduced fertility (model 1, Fig. 5).
Abbreviations
- Bc:
-
Brassica campestris ssp. chinensis Makino
- GA:
-
Gibberellic acid
- RT-PCR:
-
Real-time polymerase chain reaction
References
Achard P, Cheng H, De Grauwe L, Decat J, Schoutteten H, Moritz T, Van Der Straeten D, Peng J, Harberd NP (2006) Integration of plant responses to environmentally activated phytohormonal signals. Science 331:91–94
Barreto HG, Lazzari F, Ságio SA, Chalfun-Junior A, Paiva LV, Benedito VA (2011) In silico and quantitative analyses of the putative FLC-like homologue in coffee (Coffea arabica L.). Plant Mol Biol Report 30:29–35
Blázquez MA, Weigel D (2000) Integration of floral inductive signals in Arabidopsis. Nature 404:889–892
Deng W, Ying H, Helliwell CA, Taylor JM, Peacock WJ, Dennis ES (2011) FLOWERING LOCUS C (FLC) regulates development pathways throughout the life cycle of Arabidopsis. Proc Natl Acad Sci USA 108:6680–6685
Dill A, Sun TP (2001) Synergistic derepression of gibberellin signaling by removing RGA and GAI function in Arabidopsis thaliana. Genetics 159:777–785
Dornelas MC, Patreze CM, Angenent GC, Immink RG (2010) MADS: the missing link between identity and growth? Trends Plant Sci 16:89–97
Harberd NP (2003) Relieving DELLA restraint. Science 299:1853–1854
He Y, Amasino RM (2005) Role of chromatin modification in flowering-time control. Trends Plant Sci 10:30–35
Helliwell CA, Wood CC, Robertson M, James Peacock W, Dennis ES (2006) The Arabidopsis FLC protein interacts directly in vivo with SOC1 and FT chromatin and is part of a high molecular weight protein complex. Plant J 46:183–192
Hepworth SR, Valverde F, Ravenscroft D, Mouradov A, Coupland G (2002) Antagonistic regulation of flowering-time gene SOC1 by CONSTANS and FLC via separate promoter motifs. EMBO J 21:4327–4337
Kaufmann K, Muino JM, Jauregui R, Airoldi CA, Smaczniak C, Krajewski P, Angenent GC (2009) Target genes of the MADS transcription factor SEPALLATA3: integration of developmental and hormonal pathways in the Arabidopsis flower. PLoS Biol 7:854–875
Kim SY, Park BS, Kwon SJ, Kim J, Lim MH, Park YD, Kim DY, Suh SC, Jin YM, Ahn JH, Lee YH (2007) Delayed flowering time in Arabidopsis and Brassica rapa by the overexpression of FLOWERING LOCUS C (FLC) homologs isolated from Chinese cabbage (Brassica rapa L. ssp. pekinensis). Plant Cell Rep 26:327–336
King K, Moritz T, Harberd NP (2001) Gibberellins are not required for normal stem growth in Arabidopsis thaliana in the absence of GAI and RGA. Genetics 159:767–776
Lee H, Suh SS, Park E, Cho E, Ahn JH, Kim SG, Lee JS, Kwon YM, Lee I (2000) The AGAMOUS-LIKE 20 MADS domain protein integrates floral inductive pathways in Arabidopsis. Genes Dev 14:2366–2376
Lee S, Cheng H, King KE, Wang W, Husssain A, Lo J, Harberd NP, Peng J (2002) Gibberellin regulates Arabidopsis seed germination via RGL2, a GAI/RGA-like gene whose expression is up-regulated following imbibition. Genes Dev 16:646–658
Ma G, Ning G, Zhang W, Zhan J, Bao M (2011) Overexpression of Petunia SOC1-like gene FBP21 in tobacco promotes flowering without decreasing flower or fruit quantity. Plant Mol Biol Report 29:573–581
Michaels SD, Amasino RM (1999) FLOWERING LOCUS C encodes a novel MADS domain protein that acts as a repressor of flowering. Plant Cell 11:949–956
Moon J, Lee H, Kim M, Lee I (2005) Analysis of flowering pathway integrators in Arabidopsis. Plant Cell Physiol 46:292–299
Onouchi H, Igeño MI, Périlleux C, Graves K, Coupland G (2000) Mutagenesis of plants overexpressing CONSTANS demonstrates novel interactions among Arabidopsis flowering-time genes. Plant Cell 12:885–900
Rieu I, Powers SJ (2009) Real-time quantitative RT-PCR: design, calculations, and statistics. Plant Cell 21:1031–1033
Rong ZL, Hou XL, Shi GJ, Xiao D, Hao HN (2010) Cloning and expression analysis of late bolting BcFLC1 gene from Brassica campestris ssp. chinensis. J Nanjing Agricult Univ 33:23–27
Samach A, Onouchi H, Gold SE, Ditta GS, Schwarz-Sommer Z, Yanofsky MF, Coupland G (2000) Distinct roles of CONSTANS target genes in reproductive development of Arabidopsis. Science 288:1613–1616
Searle I, He Y, Turck F, Vincent C, Fornara F, Krober S, Amasino RA, Coupland G (2006) The transcription factor FLC confers a flowering response to vernalization by repressing meristem competence and systemic signaling in Arabidopsis. Genes Dev 20:898–912
Seo E, Lee H, Jeon J, Park H, Kim J, Noh YS, Lee I (2009) Crosstalk between cold response and flowering in Arabidopsis is mediated through the flowering-time gene SOC1 and its upstream negative regulator FLC. Plant Cell 21:3185–3197
Sheldon CC, Burn JE, Perez PP, Metzger J, Edwards JA, Peacock WJ, Dennis ES (1999) The FLF MADS box gene: a repressor of flowering in Arabidopsis regulated by vernalization and methylation. Plant Cell 11:445–458
Sheldon CC, Rouse DT, Finnegan EJ, Peacock WJ, Dennis ES (2000) The molecular basis of vernalization: the central role of FLOWERING LOCUS C (FLC). Proc Natl Acad Sci USA 97:3753–3758
Shore P, Sharrocks AD (1995) The MADS-box family of transcription factors. Eur J Biochem 229:1–13
Silverstone AL, Jung HS, Dill A, Kawaide H, Kamiya Y, Sun TP (2001) Repressing a repressor: gibberellin-induced rapid reduction of the RGA protein in Arabidopsis. Plant Cell 13:1555–1566
Tadege M, Sheldon CC, Helliwell CA, Stoutjesdijk P, Dennis ES, Peacock WJ (2001) Control of flowering time by FLC orthologues in Brassica napus. Plant J 28:545–553
Tyler L, Thomas SG, Hu J, Dill A, Alonso JM, Ecker JR, Sun TP (2004) DELLA proteins and gibberellin-regulated seed germination and floral development in Arabidopsis. Plant Physiol 135:1008–1019
Wang ZJ, Huang JQ, Huang YJ, Chen FF, Zheng BS (2011) Cloning and characterization of a homologue of the FLORICAULA/LEAFY gene in hickory (Carya cathayensis Sarg.). Plant Mol Biol Report. doi:10.1007/s11105-011-0389-z
Wen CK, Chang C (2002) Arabidopsis RGL1 encodes a negative regulator of gibberellin responses. Plant Cell 14:87–100
Acknowledgments
The research was supported by a project funded by the Priority Academic Program Development of Jiangsu Higher Education Institutions, the Fundamental Research Funds (KYZ200912) for the Central Universities, and the project supported by China Agriculture Research System (CARS-25-A-12).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Liu, T., Li, Y., Zhang, C. et al. Overexpression of FLOWERING LOCUS C, Isolated from Non-Heading Chinese Cabbage (Brassica campestris ssp. chinensis Makino), Influences Fertility in Arabidopsis . Plant Mol Biol Rep 30, 1444–1449 (2012). https://doi.org/10.1007/s11105-012-0469-8
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-012-0469-8