Abstract
The dehydration-responsive element-binding (DREB) protein specifically binds the dehydration-responsive element and is part of a plant-specific family of transcription factors that play important roles in regulating the expression of genes in response to a variety of abiotic stresses, including drought, high salt, and low temperature. In this study, a DREB-like gene containing a conserved APETLA2 (AP2)/ethylene responsive element binding factor (ERF) domain, termed DRE binding factor (CkDBF), was isolated from Caragana korshinskii. The K, R-rich motif in the N-terminal region was shown to play an essential role in nuclear localization of CkDBF. DNA-binding specificity and transactivation activity of CkDBF were verified by yeast one-hybrid experiments. Expression of CkDBF was shown to be upregulated by high salt, dehydration, low temperature, and the phytohormone abscisic acid (ABA). Overexpression of CkDBF in transgenic tobacco plants resulted in higher tolerance to high salinity and osmotic stresses and induction of a downstream target gene under normal conditions. These results suggest that CkDBF may play an essential role as a DREB transcription factor in regulation of stress-responsive signaling in C. korshinskii.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Various abiotic stresses, including low temperature, water deficit, and soil salinization, heavily influence plant growth and crop productivity (Boyer 1982). During long-term evolution, plants have developed complex molecular mechanisms to survive harsh environments. Transcription factor (trans-factor) genes play important roles in stress survival by serving as master regulators of sets of downstream stress-responsive genes. Transcription factors regulate downstream gene expression via binding to specific elements (cis-elements) in target genes, and consequently, enhance stress tolerance in plants (Chen and Zhu 2004; Yamaguchi-Shinozaki and Shinozaki 2006).
Several hundred types of transcription factors have been isolated from higher plants. Important families of stress-responsive transcription factors include the ethylene-responsive element binding factors (ERF), basic-domain leucine-zipper (bZIP) MYB (from avian myeloblastosis virus), and WRKY transcription factors (Agarwal et al. 2006; Liu et al. 1999; Riechmann et al. 2000). The dehydration-responsive element-binding (DREB) proteins make up a subfamily of the AP2/ERF transcription factor family (Magnani et al. 2004; Riechmann and Meyerowitz 1998). The first isolated DREB family member was CRT/DRE-binding factor 1 (CBF1) (Stockinger et al. 1997). Subsequently, two DREB genes regulating drought and low-temperature-responsive gene expression were isolated from Arabidopsis (Liu et al. 1998). At present, many DREB complementary deoxyribonucleic acid (cDNAs) have been cloned and characterized in different plants, including rice (Oryza sativa) (Dubouzet et al. 2003; Tian et al. 2005), soybean (Glycine max) (Li et al. 2005), maize (Zea mays) (Kizis and Pages 2002; Qin et al. 2004), cotton (Gossypium hirsutum) (Huang and Liu 2006a, b), (Hordeum vulgare) (Choi et al. 2002; Xu et al. 2009), wheat (Triticum aestivum) (Xu et al. 2008), and Populus euphratica (Chen et al. 2009). DREB transcription factors specifically bind to C-repeat (CRT)/dehydration responsive element (DRE) elements found in the promoters of stress-responsive genes, such as rd29A, kin1, and erd10, and regulate expression of these genes (Yamaguchi-Shinozaki et al. 2002; Dubouzet et al. 2003). The dehydration-responsive element (DRE) was identified by analyzing the rd29A promoter in both transgenic Arabidopsis and tobacco (Yamaguchi-Shinozaki and Shinozaki 1994). The core sequence of the CRT/DRE element (CCGAC) is essential for specific interaction with DRE-binding factors (DBFs/DREBs) in response to cold, drought, and/or high-salt treatments (Liu et al. 2000; Yamaguchi-Shinozaki and Shinozaki 2005; Sakuma et al. 2002).
DREBs contain a conserved AP2/ERF domain of approximately 60 amino acids, which has been proposed to be essential for deoxyribonucleic acid (DNA) binding (Stockinger et al. 1997; Riechmann and Meyerowitz 1998). Some studies have demonstrated that Val14 (V14) and Glu19 (E19), found in the AP2/ERF domain, play central roles in consensus recognition and binding to the DRE cis-element (Liu et al. 1999; Sakuma et al. 2002). However, recent studies have indicated that Glu19 is not conserved in DREB1 from rice and barley, and is instead replaced by valine (Dubouzet et al. 2003). In most OsDREB1-type proteins, valine is found at both the 14th and 19th positions, with the exception of OsDREB1C. The other DREB1-type proteins in monocots (barley, wheat, and rye) also have a valine in the 19th position (Agarwal et al. 2006). In addition to these two residues, a conserved alanine at position 37 in the AP2/ERF domain has also been indicated to be essential in binding to DRE elements (Liu et al. 2006).
DREB genes form a large multigene family and can be classified into six small groups named as A-1 to A-6 (Sakuma et al. 2002). DREB members of different subgroups play multiple roles in plants. Expressions of the A-1 group genes are induced by low temperature, but not by drought or high-salt stress (Liu et al. 1998; Shinwari et al. 1998), while A-2 group genes are regulated by salt and drought, but not by cold (Dubouzet et al. 2003; Liu et al. 1998; Nakashima et al. 2000). However, recent reports have shown some conflicts with respect to these trends. Some A-2 group genes are also induced by ABA or cold (Xu et al. 2008; Chen et al. 2009), indicating crosstalk between different groups. Until now, most reports about DREB/CBFs focused on DREBA1 and A2, while investigation into other groups is limited. TINY2 (A-4), PpDBP1(A-5), GhDBP1(A-5), and ZmDBP1(A-6) were identified as stress-response regulation genes (Kizis and Pages 2002; Wei et al. 2005; Huang and Liu 2006b; Liu et al. 2007).
Although the expression of DREB genes has been extensively investigated, little information is available regarding tissue-specific expression of DREBs. Expression of AtDREB2A and AhDREB1 was observed in roots, stems, and leaves under normal growth conditions. Under salt-stress conditions, AhDREB1 was highly expressed in roots, but expressed at lower levels in stems and leaves (Shen et al. 2003). Transcription of GmDREBa/b was induced by cold, salt, and drought in soybean leaves. In roots, the expression of GmDREBc was upregulated by salt, drought, and ABA treatment, but was not affected by cold (Li et al. 2005). DREB/CBF genes have been overexpressed in rice, wheat, Paspalum notatum, and tobacco, resulting in improved stress responses under various stress conditions and increased expression of downstream DREB target genes under normal conditions (Gilmour et al. 2000; Haake et al. 2002; Kasuga et al. 2004; Oh et al. 2005; James et al. 2007; Xu et al. 2008; Agarwal et al. 2009; Gao et al. 2009).
Caragana korshinskii is a deciduous perennial shrub found in sandy grassland and desert ecosystems. The species is distributed across half-fixed and fixed sandy regions in northwest China and Mongolia (Fu 1989). Although the literature states that the species is highly stress tolerant (Zhang 1994; Wang et al. 2007), most studies of C. korshinskii have focused only on the introduction and domestication of the species. Therefore, the stress-tolerance signal-regulating mechanisms in this species are poorly understood. In the present study, we cloned an AP2/ERF-like gene from C. korshinskii and characterized its subcellular localization, gene expression patterns, and DNA-binding and transactivation activities. Overexpression of the novel gene in transgenic tobacco plants led to enhanced tolerance of high salinity and osmotic stress. Molecular analysis of stress tolerance in C. korshinskii may be useful in improving stress tolerance in pasture or other crops using transgenic plant techniques.
Materials and Methods
Plant Material and Stress Treatments
The seeds of C. korshinskii were sown in 10-cm diameter plastic pots containing a mixture of vermiculite and perlite (3:1, v/v). Seedlings were grown under natural light in a greenhouse at temperatures ranging from 25°C to 30°C and irrigated appropriately. Plants were harvested for experiments when they were 4 weeks old. For the dehydration treatment, plants were carefully removed from the soil to avoid injury and subjected to dehydration on filter papers at room temperature at a relative humidity of approximately 30% for 0.5, 1, 2, 4, 8, 10, and 12 h. For salt and ABA treatments, plants were carefully removed from the soil and hydroponically grown in a solution containing 200 mM NaCl and 100 µM ABA for 0, 0.5, 1, 2, 4, 8, 12, and 24 h. For the cold-stress treatment, potted plants were transferred to a 4°C incubator and were subjected to the stress treatment for 0, 0.5, 1, 2, 4, 8, 12, and 24 h.
Isolation of CkDBF Fragment and Sequence Analysis
Total ribonucleic acid (RNA) was prepared from dehydrated C. korshinskii leaves using TRIzol reagent (Invitrogen, California, USA). First-strand cDNA was synthesized from total RNA using a commercial cDNA synthesis kit (Promega, California, USA) in the presence of oligo(dT)18. Degenerate primers (DP1, 5′-GGNAAGTGGGTBGSTGAAAT-3′; DP2, 5′-CATCATANGCDAGNGCAGC-3′) for DREB were synthesized based on sequences from homologous genes from Arabidopsis (Liu et al. 1998), rice (Dubouzet et al. 2003), and soybean (Li et al. 2005). The first-strand cDNA was used as a template for amplification of CkDBF using primers DP1 and DP2. The polymerase chain reaction (PCR) fragment was purified using the Axygen Gel Extraction Kit (Axygen, California, USA). The purified PCR fragment was then ligated into the PMD-18T vector, and single clones were sequenced. The 5′ and 3′ends of the cDNA sequence were amplified using the smart RACE cDNA Amplification Kit (Roche, Mannheim, Germany) according to the manufacturer's protocol. The gene-specific primers (GSP) for 5′RACE (Rapid Amplification of cDNA Ends) was GSP5′ (5′-TGCGTTTGGGGAGTATAAGG-3′), and for 3′RACE was GSP3′ (5′-TCAGAGGTGACTTTGCGAGG-3′). The amplified PCR fragment was then ligated into the PMD-18T vector and sequenced.
Sequence analysis was performed using LASERGENE software (DNAstar, Madison, WI, USA). Multiple alignment and phylogenetic analyses were performed using the bio-software programs DNAMAN6.0 and MEGA4.0.
Subcellular Localization of CkDBF
The full-length cDNA coding region and a cDNA fragment lacking the N-terminal region of CkDBF were fused to the 5′ end of the green fluorescent protein (GFP) coding sequence, and the fused gene was subcloned into the vector pCAMBIA 1302 under the control of the 35S promoter. Approximately 2 μg of the plasmid construct was used to coat gold particles, and a plasmid containing the GFP open reading frame (ORF) alone was used as a negative control. Inner epidermal peels of onion were cultured in hypertonic medium (one half Murashige and Skoog (MS) medium supplemented with 0.4 mol L−1 sorbitol) for 4 h and were then transformed with the plasmid-coated gold particles using the PDS-1000 bombardment system (Bio-Rad, Canada) at 1,100 psi. Subsequently, the plates containing the bombarded peels were incubated in hypertonic medium at 25°C overnight. GFP fluorescence in the onion epidermal cells was visualized using a laser scanning confocal microscope (Leica, Germany) at a wavelength of 488 nm.
DNA Gel-Blot Analysis
Genomic DNA was isolated from young leaves of C. korshinskii plants using the cetyl-trimethyl ammonium bromide method (Murray and Thompson 1980). The genomic DNA was then digested with restriction endonucleases HindIII, EcoRI, and EcoRV and separated on a 0.8% agarose gel. After electrophoresis, the digested DNA fragments were transferred to a Hybond-N+ nylon membrane (Pharmacia, New Jersey, USA) and hybridized with a digoxigenin (DIG)-11-dUTP-labeled CkDBF cDNA probe at 42°C. The blot was washed 2× for 5 min in 2× standard saline sitrate (SSC), 0.1% sodium dodecylsulfonate (SDS) at 25°C with constant agitation, and then washed 2 × for 15 min in 0.5× SSC, 0.1% SDS (pre-warmed to wash temperature) at 68°C with constant agitation.
Quantitative Real-Time PCR Analysis
For quantitative PCR analysis, total RNA was prepared from treated or untreated leaves or roots of 4-week-old plants grown on soil using TRIzol reagent (Invitrogen USA). First-strand cDNA was synthesized from total RNA using a commercial cDNA synthesis kit (Promega, California, USA) and stored at −20°C before use. The primers used in real-time PCR analyses were as follows: CkDBF forward 5′-GGGGAAAATGGGTTGCTGAGATAAG-3′, CkDBF reverse 5′-CCGTTTTGGTGGTTCTTCAGGTTCG-3′, ACTIN forward 5′-GTGGTCGTACAACTGGTATTGTG-3′, and ACTIN reverse 5′-GAACCTCCAATCCAGACACTG-3′.
To characterize the expression of stress-induced target genes in transgenic tobacco plants, primers used in the real-time PCR reactions were as follows: NtERD10A (AB049335) forward 5′-GGGGAGCATCATCAT-3′, NtERD10A reverse 5′-TCTCCTTAATCTTGTCCA-3′, NtERD10B (AB049336) forward 5′-CGGCAATCCCATTCGTCAAACC-3′, NtERD10B reverse 5′-ATGATGCTCCCCTTCCATACCATAAC-3′, NtERD10C (AB049337) forward 5′-AGGAGGAAGAAATAGGAGAGGACGG-3′, NtERD10C reverse 5′-TCTTCTCTTTTCCCTCAGCCTCGTG-3′, NtERD10D (AB049338) forward 5′-AGGACACGGCTGTACCAGTG-3′, NtERD10D reverse 5′-TTTCCCTCAGCCTCGTGCTC3′, NtZfp (AF053077) forward 5′-TGCCCCACCGACTGAAGAAGAGTATT-3′, and NtZfp reverse 5′-GAGGCGGCGGTAGCAGTAGTAGT-3′. Quantitative real-time PCR was performed with an ABI 7500 sequence detection system according to the manufacturer's protocol (Applied Biosystems, Foster, CA). Each experiment was repeated three times. Triplex quantitative assays were performed on each cDNA sample. The relative quantification method was used to evaluate quantitative variation between replicates examined. To account for differences in total RNA present in each sample, the amount of cDNA calculated was normalized using the amount of amplified actin cDNA detected in the same sample. Data were then analyzed with ABI Prism 7500 SDS software version 1.1 (Applied Biosystems, Foster, CA).
Generation of 35S::CkDBF Transgenic Tobacco Plants
The overexpression construct was made by inserting the full-length CkDBF cDNA into the binary plant vector pCAMBIA1300 (35S). The full-length CkDBF cDNA was amplified by reverse transcription polymerase chain reaction (RT-PCR) using gene-specific primers (DBFF: 5′-ATGGCAGGTAGAATGGATTTC-3′, CkDBFR: 5′-TCACAGAGAATCCCAATCAATCT-3′). The PCR product was subcloned into the pMD-18T vector. The plasmid containing the cDNA was digested using KpnI and SalI, and the cDNA was inserted into the pCAMBIA1300 vector under the control of CaMV 35S promoter by directional cloning using the KpnI and SalI sites. The resulting vector was introduced into the Agrobacterium tumefaciens strain LBA4404. Transformation was carried out using the leaf-disc method (Horsch et al. 1985). Primary transgenic plants (T0) were transplanted to soil, and seeds (T1) were collected. T1 seeds were selected on hygromycin (100 mg/L) medium. Positive seedlings were transferred to soil and grown in a greenhouse. T2 seeds were used for further analysis.
Stresses Tolerance Analysis of Transgenic Tobacco Plants
T2 seeds were surface-sterilized in 10% (v/v) commercial bleach (2.6% NaOCl) for 15 min, stratified at 4°C for 2–4 days, and sown on MS medium supplemented with 200 mM NaCl or 250 mM mannitol. To determine the germination rate, seeds were considered germinated when radicles completely penetrated the seed coat. Germination seeds were counted when there are no seeds to germinate any more. Each experiment was repeated three times, 100 seeds were used in each replication.
For primary root growth tests, individual seedlings of wildtype (WT) tobacco and transgenic tobacco seedlings grown on MS medium were selected for uniformity. After 14 days, root length was quantified by measuring with a ruler. Each experiment was repeated three times, and 20 seedlings were used in each replication. For mature plants' tolerance test, the WT and transgenic plants were grown on MS basal medium and MS medium with 200 mM NaCl and 250 mM mannitol for 8 weeks. The photos representing test result were taken using Canon EOS50D digital camera. In each experiment, 15–20 plants were used. For survival rate from dehydration, the WT and transgenic plants grown in pots were not watered for 2 weeks, after which, watering resumed. To examine the tolerance of the transgenic plants to cold stress, plants grown in pots were exposed to temperature of 2°C for 2 days, then returned to 25°C, and grown for 7 days. Survival plants were counted. The survival plants for every treatment were counted. Each experiment was repeated three times, 20 plants were used in each replication. Statistical analysis was performed using the SPSS program (SPSS Inc.).
Results
Identification of the CkDBF Gene from C. korshinskii
A cDNA fragment of 280 base pair (bp), with homology to the AP2/ERF gene, was identified from C. korshinskii. Subsequently, RACE experiments were performed to clone the 3′ and 5′ ends of the sequence. A 2,088-bp full-length cDNA, including an ORF of 975 bp encoding a peptide of 324 amino acids with a 745-bp 5′ Un-Translation Region (UTR) and a 368-bp 3′ UTR, was obtained (Fig. 1a). Analysis of the deduced amino acid sequence indicated that an AP2/ERF domain was found in the sequence, and the corresponding gene was named CkDBF (Genbank accession GU573848). Amplification of the genomic sequence of CkDBF was performed using primers for the full-length cDNA. The alignment of the genomic sequence and cDNA sequence indicated that no introns were present in CkDBF.
Sequences information, comparison, and phylogenetic analysis of the CkDBF gene. a Nucleotide and deduced amino acid sequences of the CkDBF gene. The AP2/ERF domain is underlined. Bold italic letters show potential nuclear location signals. The predicted transcriptional activation domain is double-underlined. b Amino acid multiple alignment of CkDBF with homologs from 9 other plants: GhDBP2 (AAT39542), GmDREBb (AAQ57226), RAP2.4 (NP_177931), PgDREB2A (AAV90624), AtDREB2A (BAA33794), OsDREB2A (AAN02487), AtDREB1A/CBF3(BAA33434), AtDREB1B/CBF1(BAA33435), BnCBF (AAL38243). c Phylogenetic relationship of AP2-type transcription factors based on amino acid sequence comparison of the AP2 regions. The neighbor-joining method was used to generate the phylogenetic tree. ABI4 (AF085279), ZmABI4 (AY125490), AhDREB1 (AF274033), AsCBF1 (AM071406), AtERF1 (NM_113225), AtERF2 (NM_124093), AtERF4 (NM_112384), AtERF5 (NM_124094), CaDREBLP1 (AY496155), MtDREB2A (DQ908959), AtDREB1C (AB007789), AtDREB2B (AB007791), OsDREB1A (AF300970), OsDREB1B (AF300972), OsDREB1E (AY829439), GhDBP1 (AY174160), ZmDBP2 (FJ805750), GmDREB1 (AF514908), GmDREB3 (DQ208969), BnCBF5 (AF499031), HvCBF1 (AF298230), HvCBF3 (AF239616), HvCBF6 (AY785860), LeCBF1 (AY034473), TaCBF1 (AF376136), TaCBF6 (AY785903), ScCBF (AF370728), ScCBF2 (AF370730), TaAIDFb (DQ022953), TaAIDFc (DQ022952), TaAIDFa (AY781361), TINY (AC005405), ZmDBF1 (AF493800), ZmDREB1A (AF450481), RAP2.10 (NM_119854), GmDREBc (AY244760), GmDREBa (AY542886), PpDREB2 (AY553331), TINY2 (NM_121197), RcTINY (XM_002516456), RAP2.1 (AY086838), ZmDBP2 (FJ805750), CdDREB1 (AY462117), CbCBF (AY391121)
The AP2/ERF domain of CkDBF was made up of 58 amino acid residues, which were predicted to compose a secondary structure made up of three β-sheets and one α-helix using Anthepro software (Fig. 1b). This structure may be responsible for the DNA-binding activity (Allen et al. 1998; Sakuma et al. 2002). A potential nuclear location signal (NLS) was found near the N-terminus, suggesting that CkDBF may localize to the nucleus. An acidic amino acid region (ESEEGSDDSSPLSDLTFGDVTEPQWEAGSEHLLNLQKFPSYEID) was identified in the C-terminal region of CkDBF that may act as a transcriptional regulation domain (Fig. 1a). As shown in Fig. 1b, a conserved valine (V) residue was found at the 14th amino acid position in the second β-sheet of the AP2/ERF domain. Furthermore, the 19th amino acid was identified as leucine (L), similar to the DREBA-6 group homologs.
The alignment analysis with other homologous sequences of higher plants indicated that CkDBF shared high homology with other proteins in the AP2/ERF domain (Fig. 1b). By using the MEGA4.0 program, phylogenetic analysis was performed according to the Sakuma et al. (2002). The result suggested that CkDBF can be classified into DREBA-6 group of DREB protein (Fig. 1c).
For the gene copy analysis, C. korshinskii genomic DNA was digested with HindIII, EcoRI, and EcoRV. DNA blot results showed that CkDBF hybridized with only one genomic fragment, indicating that CkDBF is a single-copy gene in the legume forage genome (Fig. 2).
Subcellular Localization of CkDBF Within the Nucleus
Two fragments of CkDBF were fused to the GFP gene and then transferred into onion epidermal cells by particle bombardment for analysis of its subcellular localization. The control GFP protein was distributed throughout the cell, as expected. However, fluorescence from CkDBF::GFP was found primarily in the nucleus. The CkDBF construct lacking the N-terminus, including the KKRKTRRK motif (CkDBF-N::GFP) exhibited a similar localization pattern to the 35S::GFP control (Fig. 3). These results suggest that the N-terminal region, containing the predicted NLS, plays central role in nuclear localization of CkDBF.
CkDBF Specifically Binds to DRE Elements
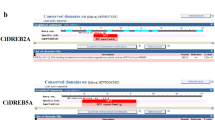
The DNA-binding and transactivation activities of the CkDBF protein were investigated in a yeast one-hybrid system. Two YM4271 yeast strains carrying integrated copies of the HIS3 and lacZ reporter genes and three-time tandem repeats of a DNA fragment of the rd29A promoter containing either a DRE motif (wDRE) or a mutated DRE motif (mDRE) were used. The full-length cDNAs of CkDBF and AtDREB2A from Arabidopsis were inserted into the yeast expression vector pGAL4-AD and transformed into the two yeast strains (Fig. 4a). The pGAL4-AD construct containing AtDREB2A was used as positive control. Growth of the yeast strains on selective media was then analyzed. As shown in Fig. 4b, the control and wildtype CRT/DRE yeast transformants grew on synthetic defined (SD) media lacking histidine in the presence of 10 mmol L-1 3-AT, whereas the mutant yeast transformants did not. The positive response of the WT-recombination strain in β-galactosidase analysis further confirmed that CkDBF possesses the DNA-binding function required for transcription factors and can specifically bind to DRE elements (Fig. 4b).
Analysis of DNA-binding and transactivation activities of CkDREB2 protein by the yeast one-hybrid assay. a Map of the pAD-CkDREB2 vector. The plasmid expressed the fusion protein in the yeast and activated the expression of two reporter genes, HIS3 and lacZ. b The growth of transformed yeast strains on SD/−His + 10 mM 3-AT (left); β-galactosidase activity analysis (middle); location of different transformed yeast strains on plate (right). c Map of the pBridge-CkDREB2 vector. d The growth of transformed yeast strains on SD/−His–Trp medium (left); β-galactosidase activity analysis (middle); location of different transformed yeast strains on plate (right)
CkDBF Transactivation Activity Assay
The transactivation activity of CkDBF was tested through a second yeast one-hybrid assay. The full-length cDNAs of CkDBF and GAL4AD were inserted into the yeast pBridge expression vector, and the vectors were transformed into the yeast strain YH109 (Fig. 4c). Growth of the transformed strain on selective media was analyzed. As shown in Fig. 4d, the pBridge-CkDBF and pBridge-GAL4AD strains grew on SD media lacking histidine (His) and tryptophan (Trp), while the negative control strain containing the empty pBridge vector did not. Moreover, the pBridge-CkDBF and pBridge-GAL4AD strains also turned blue when subjected to β-galactosidase analysis. These results indicated that the CkDBF protein possesses transactivation activity.
Various Stresses and the Exogenous Phytohomone ABA Upregulate the Expression of CkDBF
First, we analyzed the tissue-specific expression of CkDBF in C. korshinskii. C. korshinskii plants were treated with or without dehydration. The CkDBF transcript level was detected slightly in the leaves, stems, and roots of plants grown under normal conditions, but after dehydration for 4 h, levels of the CkDBF transcript increased dramatically in the leaves and stems, and there was a moderate increase in the level of transcription in the roots (Fig. 5a).
Expression pattern of CkDBF a The tissue-specific expression of CkDBF. Expression of CkDBF in response to high-salinity (200 µM NaCl) stress in leaves (b) and roots (c). The expression of CkDBF in response to dehydration (d), low-temperature (4°C) (e) and exogenous ABA (100 µM) (f) for the indicated times. The ACTIN gene was amplified as a control to normalize samples. Experiments were repeated three times
Next, the expression pattern of the CkDBF gene in plants under various abiotic stresses was investigated using quantitative real-time PCR. The messenger ribonucleic acid (mRNA) levels were very low in the leaves and roots of control plants. After treatment with 200 mM NaCl for 0.5 h, levels of the CkDBF transcript began to increase in the leaves and roots. The transcription level peaked at 4 h in roots, and then began to decline at 12 h, while in the leaves, expression peaked later than in roots and at 12 h. The levels of the transcript were lower in roots than in leaves on average (Fig. 5b, c). Dehydration treatment also sharply induced the transcription of CkDBF, and the transcript level is much higher than in other treatments (Fig. 5d). Expression of CkDBF was induced after 0.5 h of dehydration, peaked at 4 h, then slightly decreased and remained at a high level for an extended time period (Fig. 5d). Similarly, expression of CkDBF was also induced by cold-stress treatment in a time-dependent manner and peaked at 2 h (Fig. 5e). These observations were similar to the results of Xu et al. (2008), who showed that transcription of TaAIDFa was also activated by cold treatment, but the highest levels of transcription occurred after 12 h.
Since abscisic acid (ABA) plays an important role in regulation of gene expression in response to various abiotic stresses (Leung and Giraudat 1998), the response of the CkDBF transcript to exogenous application of the phytohormone ABA was assessed. As is shown in Fig. 5f, CkDBF transcript expression was significantly induced by exogenous ABA. Similar to other stress treatments, the expression followed a time-dependent pattern, peaking at 2 h, indicating that CkDBF may function in an ABA-mediated signaling pathway.
Overexpression of CkDBF Enhances the Tolerance for High-Salinity and Mannitol Stresses in Transgenic Plants
Three independent lines of T2 progenies of transgenic tobacco plants were used in tolerance tests. The germination rate and primary root length was determined in MS media supplemented with 200 mM NaCl or 250 mM mannitol. No differences were observed in germination rate between wildtype (WT) and transgenic seeds on MS basal medium. On medium containing NaCl or mannitol, the germination rates of both WT and transgenic lines were reduced. However, the germination rate of WT plants was more dramatically reduced than that of transgenic lines, which showed only a moderate reduction in generation rate (Fig. 6a).
Enhanced stress tolerance of 35S::CkDBF transgenic tobacco. a Germination rates of WT (control) and transgenic lines grown on MS medium (1), or MS medium supplemented with 200 mM NaCl (2) or 250 mM mannitol (3) (n = 100, each experiment was repeated three times). b Primary root growth of WT and transgenic lines tobacco seedlings under normal condition (1), treated with 150 mM mannitol (2), 250 mM mannitol (3), 100 mM NaCl (4), or 200 mM NaCl (5) (n = 20). c Growth of the WT and transgenic lines plants under salt and osmotic stresses. d Expression analysis of downstream genes NtERD10A, NtERD10B, NtERD10C, NtERD10D, and NtZfp in transgenic tobacco by using real-time PCR. Genes were amplified with specific primers. The ACTIN gene was used to normalize samples. Experiments were repeated three times
To further investigate the enhanced stress tolerance of the transgenic lines, the primary root growth of seedlings from WT and transgenic lines was tested in the presence of 200 mM NaCl or 250 mM mannitol. The primary root lengths of the WT and transgenic lines were relatively consistent on MS basal medium. However, when plants were grown on NaCl or mannitol plates, the primary root growth of the WT lines was dramatically retarded compared to transgenic lines (Fig. 6b). For the mature plants, the growth of WT and transgenic lines was similar under normal condition. Under salt and mannitol stress, the growth of WT plants was dramatically retarded, while in transgenic lines, plant growth was only slightly restrained (Fig. 6c). These results suggest that overexpression of CkDBF can enhance tolerance for salt and osmotic stress in transgenic plants.
Drought tolerance analysis indicated that after drought treatment, 3.3% WT tobacco plants survived, whereas the survival rate of 35S::CkDBF transgenic line1, line2, and line3 is 30%, 41.7%, and 48.3%, respectively. After cold stress, the survival rate of line1, line2, and line3 was 18.3%, 26.7%, and 31.7%, respectively, while only 8.3% in WT (Table 1).
Overexpression of CkDBF Induces the Expression of Downstream Stress-Responsive Genes
Many past studies have shown that overexpression of DREB family genes regulates the expression of downstream stress-induced genes containing CRT/DRE element in their promoter in transgenic plants (Jaglo-Ottosen et al. 1998; Gilmour et al. 2000; Xu et al. 2008). To test whether this is the case for CkDBF, the expression of four NtERD10 genes and one NtZfp gene in the WT and transgenic lines were analyzed. As shown in Fig. 6d, all these genes were induced even under normal growth conditions compared with WT plants. The transcription levels of these genes were significantly induced after dehydration stress, and the expression level in transgenic lines were much higher than that in WT (Fig. 6d). These results imply that overexpression of CkDBF induces abiotic stress-response genes containing a DRE element in their promoters and that CkDBF is involved in regulation of stress-responsive signaling.
Discussion
In this study, we isolated a novel DREB homolog from C. korshinskii, termed CkDBF. Sequence analysis identified an AP2/ERF domain in CkDBF. The conserved AP2/ERF domain contains 58 amino-acid residues and is predicted to fold into a structure containing three anti-parallel β-sheets and one α-helix. This structure is thought to play a key role in recognizing and binding to specific cis-elements (Sakuma et al. 2002; Allen et al. 1998). The second β-sheet and the α-helix contained conserved residues at the 14th position (V) and the 37th position (A). Alignment analysis and domain comparison suggested that CkDBF is homologous to DREBA-6 proteins. DNA-binding and transactivation assays further suggested that CkDBF encodes a DREB transcription factor. Various abiotic stresses, specifically dehydration, high salinity, low temperature, and exogenous ABA, upregulated the expression of CkDBF in a time-dependent fashion. Furthermore, overexpression of CkDBF was found to activate the expression of target stress-responsive genes containing DRE element in those promoter and to enhance stress tolerance in the resulting transgenic tobacco. Therefore, we propose that CkDBF is involved in the abiotic stress-signaling pathway as a DREB transcription factor.
DREB proteins are important members of the AP2/ERF transcription factor family. The DNA-binding region of these proteins specifically recognizes the DRE element in the promoter region of stress-responsive genes, and the transactivation region activates the expression of these genes, activating signaling pathways in the plant stress response (Liu et al. 1998; Dubouzet et al. 2003). CkDBF was found to specifically bind to the DRE element, but not the mutant mDRE, through yeast one-hybrid analysis. Previous results have suggested that Val14 and Glu19 in the AP2/ERF domain are essential for specific binding to DRE (Liu et al. 1998; Cao et al. 2001; Sakuma et al. 2002). However, in CkDBF, Glu19 is replaced by leucine (L). Similar amino acid changes have also been observed in other plants, including soybean, rice, and maize. Furthermore, in the monocotyledons rice, wheat, and barley, the 19th amino acid in the AP2/ERF domain of DREB1-type factors is valine (Dubouzet et al. 2003). These observations suggest that the function of the 14th amino acid is likely more important than the 19th for specific DNA-binding activity.
In many previous studies, nuclear localization signals (NLS), which are required for nuclear localization, have been identified in the N-terminus of AP2/ERF transcription factor genes. In this present study, a basic amino acid-rich region (KKRKTRRK) was identified at the N-terminus of CkDBF. In order to test the function of this short sequence, we fused a fragment lacking the N-terminal region, which includes the short sequence and the full cDNA sequence with GFP. Through the aid of particle biolistic technology, the subcellular localization of these constructs was tested. As expected, without the KR-rich region, CkDBF could not enter the nucleus. Therefore, we propose that this short basic amino acid-rich region acts as an NLS in CkDBF.
The yeast one-hybrid assay showed that CkDBF possesses transactivation activity and could activate expression of the reporter gene. Sequence analysis of CkDBF identified a Gln (Q)-rich region N-terminal to the AP2/ERF domain. In general, Q-rich regions in the C-terminus can act as structural motifs required for transcriptional activation activity (Courey and Tjian 1988; Bruhn et al. 1992). However, the glutamine-rich region of CkDBF is located at the N-terminus. We propose that this glutamine-rich region maybe responsible for enhancing the DRE-binding ability of CkDBF.
Another member of DREB A-6, ZmDBF1, was reported to be induced by drought, salt, and ABA, but not by cold (Kizis and Pages 2002). In the present study, expression of CkDBF, which belongs to DREBA-6 type, was upregulated by multiple abiotic stresses, including low temperature (Fig. 5e), and the 35S::CkDBF transgenic plant exhibited enhanced tolerance to cold (Table 1). A more recent study also showed that a DREB2/CBF-type gene from wheat was not only induced by NaCl and drought, but also by low temperature (Xu et al. 2008); although in general, the DREB2 gene only responded to NaCl and drought stress. So it could be proposed that DREB factor genes from different organisms maybe display different expression patterns.
Expression analysis of DREB genes has been primarily focused on the leaves, and the transcription level of DREB genes from other nutritional organs remains to be investigated. In limited reports, the expression levels of DREBs were higher in roots than in leaves. However, in the present study, the transcription level of CkDBF in leaves and stems is higher than in roots (Fig. 4b), indicating that DREB genes from different organisms show different tissue-specific expression patterns. The expression of other DREB type genes from C. korshinskii needs to be investigated in the future.
The phytohormone ABA plays a crucial role in several physiological processes and in the adaptation of plants to different environmental stresses (Zeevaart and Creelman 1988). In some studies regarding DREB transcription factors, ABA was found to regulate the expression of DREB genes (Egawa et al. 2006; Xu et al. 2008). In this study, exogenous ABA induced expression of CkDBF in a time-dependent manner. So there might exist an ABA-dependent pathway for the regulation of CkDBF.
A number of studies have explored engineering stress tolerance by overexpression of DREBs. The overexpression of AtDREB1A and OsDREB1A led to enhanced freezing and dehydration tolerance in transgenic Arabidopsis; however, these plants showed growth retardation when compared to WT plants (Liu et al. 1998; Kasuga et al. 1999; Dubouzet et al. 2003). However, in 35S::AtDREB2A and 35S::OsDREB2A transgenic lines, the downstream genes were not shown to be activated (Liu et al. 1998; Dubouzet et al. 2003). Some studies found that the Ser/Thr-rich motif, a putative phosphorylation site may be a negative regulation motif (Xu et al. 2008; Agarwal et al. 2006; Sakuma et al. 2006). In the present study, overexpression of CkDBF enhanced high salinity and osmotic tolerance in transgenic tobacco plants and induced the expression of downstream target genes under normal conditions. These observations suggest that CkDBF may be expressed in a constitutively active form and is involved in the regulation of multiple stress-responsive signaling transduction pathways. Furthermore, expression of CkDBF could maintain a high level for a longer period under a variety of abiotic stresses, especially drought and salt stress (Fig. 4). Under high-salt treatment, the CkDBF mRNA could accumulate up to 24 h. In some plants, the transcription level of DREB significantly decreased after stress for 1–2 h (Choi et al. 2002; Xu et al. 2008). CBF1 from Arabidopsis was induced by drought stress, the expression of gene peaked at 0.5 h, then sharply decreased (Gilmour et al. 1998). In hot pepper, the DRE-binding-factor-like gene CaDREBLP1 was immediately induced by 200 mM NaCl, and peaked at 0.5 h (Hong and Kim 2005). These plants all belong to glycophyte ones, while C. korshinskii is an important pasture legume with high tolerance for drought in sandy grassland and desert environments. We propose that different stress-signaling responses and regulation mechanisms exist between stress-tolerant and sensitive plants on DREB-like transcription factors. Exploration of these resources and investigation of important transcription factor genes from C. korshinskii may be useful for breeding of stress-tolerant pasture or other crops.
References
Agarwal PK, Agarwal P, Reddy MK, Sopory SK (2006) Role of DREB transcription factors in abiotic and biotic stress tolerance in plants. Plant Cell Rep 25:1263–1274
Agarwal P, Agarwal PK, Joshi AJ, Sopory SK, Reddy MK (2009) Over-expression of PgDREB2A transcription factor enhances abiotic stress tolerance and activates downstream stress-responsive genes. Mol Biol Rep 37:1125–1135
Allen MD, Yamasaki K, Ohme-Takagi M, Tateno M, Suzuki M (1998) A novel mode of DNA recognition by a beta-sheet revealed by the solution structure of the GCC-box binding domain in complex with DNA. EMBO J 17:5484–5496
Boyer JS (1982) Plant productivity and environment. Science 218:443–448
Bruhn L, Hwang-Shum JJ, Sprague GF Jr (1992) The N-terminal 96 residues of MCM1, a regulator of cell type-specific genes in Saccharomyces cerevisiae, are sufficient for DNA binding, transcription activation, and interaction with α1. Mol Cell Biol 12:3563–3572
Cao ZF, Li J, Chen F, Li YQ, Zhou HM, Liu Q (2001) Effect of two conserved amino acid residues on DREB1A function. Biochemistry 66:623–627
Chen WJ, Zhu T (2004) Networks of transcription factors with roles in environmental stress response. Trends Plant Sci 9:591–596
Chen JH, Xia XL, Yin WL (2009) Expression profiling and functional characterization of a DREB2-type gene from Populus euphratica. Biochem Biophys Res Commun 378:483–487
Choi DW, Rodriguez EM, Close TJ (2002) Barley Cbf3 gene identification, expression pattern, and map location. Plant Physiol 129:1781–1787
Courey AJ, Tjian R (1988) Analysis of Spl in vivo reveals multiple transcriptional domains, including a novel glutamine-rich activation motif. Cell 55:887–898
Dubouzet JG, Sakuma Y, Ito Y, Kasuga M, Dubouzet EG, Miura S, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2003) OsDREB genes in rice, Oryza sativa L., encode transcription activators that function in drought-, high-salt- and cold-responsive gene expression. Plant J 33:751–763
Egawa C, Kobayashi F, Ishibashi M, Nakamura T, Nakamura C, Takumi S (2006) Differential regulation of transcript accumulation and alternative splicing of a DREB2 homolog under abiotic stress conditions in common wheat. Genes Genet Syst 81:77–91
Fu HC (1989) Caragana. In: Fu HC (ed) Flora intramongolica, 2nd edn. Inner Mongolia People Press, Huhhot, pp 236–238
Gao SQ, Chen M, Xia LQ, Xiu HJ, Xu ZS, Li LC, Zhan CP, Cheng XG, Ma YZ (2009) A cotton (Gossypium hirsutum) DRE-binding transcription factor gene, GhDREB, confers enhanced tolerance to drought, high salt, and freezing stresses in transgenic wheat. Plant Cell Rep 28:301–311
Gilmour SJ, Zarka DG, Stockinger EJ, Salazar MP, Houghton JM, Thomashow MF (1998) Low temperature regulation of the Arabidopsis CBF family of AP2 transcriptional activators as an early step in cold-induced COR gene expression. Plant J 16:433–442
Gilmour SJ, Sebolt AM, Salazar MP, Everard JD, Thomashow MF (2000) Overexpression of Arabidopsis CBF3 transcriptional activator mimics multiple biochemical changes associated with cold acclimation. Plant Physiol 124:1854–1865
Haake V, Cook D, Riechmann JL, Pineda O, Thomashow MF, Zhang JZ (2002) Transcription factor CBF4 is a regulator of drought adaptation in Arabidopsis. Plant Physiol 130:639–648
Hong JP, Kim WT (2005) Isolation and functional characterization of the Ca-DREBLP1 gene encoding a dehydration-responsive element binding-factor-like protein in hot pepper (Capsicum annuum L. cv. pukang). Planta 220:875–888
Horsch RB, Fry JE, Hoffmann N, Wallroth M, Eichholtz D, Rogers SG, Fraley RT (1985) A simple and general method for transferring genes into plants. Science 227:1229–1231
Huang B, Liu JY (2006a) Cloning and functional analysis of the novel gene GhDBP3 encoding a DRE-binding transcription factor from Gossypium hirsutum. Biochim Biophys Acta 1759:263–269
Huang B, Liu JY (2006b) A cotton dehydration responsive element binding protein functions as a transcriptional repressor of DRE mediated gene expression. Biochem Biophys Res Commun 343:1023–1031
Jaglo-Ottosen KR, Gilmour SJ, Zarka DG, Schabenberger O, Thomashow MF (1998) Arabidopsis CBF1 overexpression induces COR genes and enhances freezing tolerance. Science 280:104–106
James VA, Neibaur I, Altpeter F (2007) Stress inducible expression of the DREB1A transcription factor from xeric, Hordeum spontaneum L. in turf and forage grass (Paspalum notatum Flugge) enhances abiotic stress tolerance. Transgenic Res 17:93–104
Kasuga M, Liu Q, Miura S, Yamaguchi-Shinozaki K, Shinozaki K (1999) Improving plant drought, salt, and freezing tolerance by gene transfer of a single stress-inducible transcription factor. Nat Biotechnol 17:287–291
Kasuga M, Miura S, Shinozaki K, Yamaguchi-Shinozaki K (2004) A combination of the Arabidopsis DREB1A gene and stress-inducible rd29A promoter improved drought- and low-temperature stress tolerance in tobacco by gene transfer. Plant Cell Physiol 45:346–350
Kizis D, Pages M (2002) Maize DRE-binding proteins DBF1 and DBF2 are involved in rab17 regulation through the drought responsive element in an ABA-dependent pathway. Plant J 30:679–689
Leung J, Giraudat J (1998) Abscisic acid signal transduction. Annu Rev Plant Physiol Plant Mol Biol 49:199–222
Li XP, Tian AG, Luo GZ, Gong ZZ, Zhang JS, Chen SY (2005) Soybean DRE-binding transcription factors that are responsive to abiotic stresses. Theor Appl Genet 110:1355–1362
Liu Q, Kasuga M, Sakuma Y, Abe H, Miura S, Yamaguchi-Shinozaki K, Shinozaki K (1998) Two transcription factors, DREB1 and DREB2, with an EREBP/AP2 DNA binding domain separate two cellular signal transduction pathways in drought- and low-temperature-responsive gene expression, respectively, in Arabidopsis. Plant Cell 10:1391–1406
Liu LS, Michael J, White MJ, MacRae TH (1999) Transcription factors and their genes in higher plants. Functional domains, evolution and regulation. Eur J Biochem 262:247–257
Liu Q, Zhao NM, Yamaguch-Shinozaki K, Shinozaki K (2000) Regulatory role of DREB transcription factors in plant drought, salt and cold-tolerance. Chin Sci Bull 45:11–16
Liu Y, Zhao TJ, Liu JM, Liu WQ, Liu Q, Yan YB, Zhou HM (2006) The conserved Ala37 in the ERF/AP2 domain is essential for binding with the DRE element and the GCC box. FEBS Lett 580:1303–1308
Liu N, Zhong NQ, Wang GL (2007) Cloning and functional characterization of PpDBF1 gene encoding a DRE-binding transcription factor from Physcomitrella patens. Planta 226:827–838
Magnani E, Sjölander K, Hake S (2004) From endonucleases to transcription factors: evolution of the AP2 DNA binding domain in plants. Plant Cell 16:2265–2277
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8:4321–4325
Nakashima K, Shinwari ZK, Sakuma Y, Seki M, Miura S, Shinozaki K, Yamaguchi-Shinozaki K (2000) Organization and expression of two Arabidopsis DREB2 genes encoding DRE-binding proteins involved in dehydration- and high-salinity-responsive gene expression. Plant Mol Biol 42:657–665
Oh SJ, Song SI, Kim YS, Jang H-J, Kim SY, Kim M, KimYK NBH, Kim JK (2005) Arabidopsis CBF3/DREB1A and ABF3 in transgenic rice increased tolerance to abiotic stress without stunting growth. Plant Physiol 138:341–351
Qin F, Sakuma Y, Liu Q, Li Y-Q, Shinozaki K, Yamaguchi-Shinozaki K (2004) Cloning and functional analysis of a novel DREB1/CBF transcription factor involved in cold-responsive gene expression in Zea mays L. Plant Cell Physiol 45:1042–1052
Riechmann JL, Meyerowitz EM (1998) The AP2/EREBP family of plant transcription factors. Biol Chem 379:633–646
Riechmann JL, Heard J, Martin G, Reuber L, Jiang C, Keddie J, Adam L, Pineda O, RatcliVe OJ, Samaha RR, Creelman R, Pilgrim M, Broun P, Zhang JZ, Ghandehari D, Sherman BK, Yu G (2000) Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes. Science 290:2105–2110
Sakuma Y, Liu Q, Dubouzet JG, Abe H, Shinozaki K, Yamaguchi-Shinozaki K (2002) DNA-binding specificity of the ERF/AP2 domain of Arabidopsis DREBs, transcription factors involved in dehydration- and cold-inducible gene expression. Biochem Biophys Res Commun 290:998–1009
Sakuma Y, Maruyama K, Osakabe Y, Qin F, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2006) Functional analysis of an Arabidopsis transcription factor, DREB2A, involved in drought responsive gene expression. Plant Cell 18:1292–1309
Shen YG, Zhang WK, He SJ, Zhang JS, Liu Q, Chen SY (2003) An EREBP/AP2-type protein in Triticum aestivum was a DRE-binding transcription factor induced by cold, dehydration and ABA stress. Theor Appl Genet 106:923–930
Shinwari ZK, Nakashima K, Miura S, Kasuga M, Seki M, Yamaguchi-Shinozaki K, Shinozaki K (1998) An Arabidopsis gene family encoding DRE/CRT binding proteins involved in low-temperature-responsive gene expression. Biochem Biophys Res Commun 250:161–170
Stockinger EJ, Gilmour SJ, Thomashow MF (1997) Arabidopsis thaliana CBF1 encodes an AP2 domain-containing transcriptional activator that binds to the C-repeat/DRE, a cis-acting DNA regulatory element that stimulates transcription in response to low temperature and water deficit. Proc Natl Acad Sci USA 94:1035–1040
Tian XH, Li XP, Zhou HL, Zhang JS, Gong ZZ, Chen SY (2005) OsDREB4 genes in rice encode AP2-containing proteins that bind specifically to the dehydration- responsive element. J Integr Plant Biol 47:467–476
Wang Z, Gao HW, Wu YQ, Han JG (2007) Genetic diversity and population structure of Caragana korshinskii revealed by AFLP. Crop Sci 47:1737–1743
Wei G, Pan Y, Lei J, Zhu YX (2005) Molecular cloning, phylogenetic analysis, expressional prowling and in vitro studies of TINY2 from Arabidopsis thaliana. J Biochem Mol Biol 38:440–446
Xu ZS, Ni ZY, Liu L, Nie LN, Li LC, Chen M, Ma YZ (2008) Characterization of the TaAIDFa gene encoding a CRT/DRE-binding factor responsive to drought, high-salt, and cold stress in wheat. Mol Genet Genomics 280:497–508
Xu ZS, Ni ZY, Li ZY, Li LC, Chen M, Gao DY, YU XD, Liu P, Ma YZ (2009) Isolation and functional characterization of HvDREB1-a gene encoding a dehydration- responsive element binding protein in Hordeum vulgare. J Plant Res 122:121–130
Yamaguchi-Shinozaki K, Shinozaki K (1994) A novel cis-acting element in an Arabidopsis gene is involved in responsiveness to drought, low temperature, or high-salt stress. The Plant Cell 6:251–264
Yamaguchi-Shinozaki K, Kasugua M, Liu Q (2002) Biological mechanisms of drought stress response. JIRCAS Work Rep 23:1–8
Yamaguchi-Shinozaki K, Shinozaki K (2005) Organization of cis-acting regulatory elements in osmotic- and cold-stress-responsive promoters. Trends Plant Sci 10:88–94
Yamaguchi-Shinozaki K, Shinozaki K (2006) Transcriptional regulatory networks in cellular response and the tolerance to dehydration and cold stresses. Annu Rev Plant Biol 57:781–803
Zeevaart JAD, Creelman R (1988) Metabolism and physiology of abscisic acid. Annu Rev Plant Physiol Plant Mol Biol 39:439–473
Zhang XS (1994) Principles and optimal models for development of Maowusu sandy grassland. Acta Phytoecologica Sin 18:1–16
Acknowledgements
This work was financially supported by the Earmarked Fund for Modern Agro-industry Technology Research System and National Key Technology R&D Program (2007BAD56B01-03, 2008BADB3B01). We thank Professor Shi-Hua Shen for the kind gift of the yeast one-hybrid system, and Dr. Yun Liu for technical support in the yeast one-hybrid analyses.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wang, X., Dong, J., Liu, Y. et al. A Novel Dehydration-Responsive Element-Binding Protein from Caragana korshinskii Is Involved in the Response to Multiple Abiotic Stresses and Enhances Stress Tolerance in Transgenic Tobacco. Plant Mol Biol Rep 28, 664–675 (2010). https://doi.org/10.1007/s11105-010-0196-y
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-010-0196-y