Abstract
Forisomes are mechanoproteins that function as valves in the phloem sieve tubes of the Fabaceae. Changes in the Ca2+ level that occur after wounding trigger an instantaneous switch between the condensed and dispersed conformations, allowing forisomes to seal the sieve plate pores of injured sieve elements. Recently, three genes encoding forisome components from Medicago truncatula were identified [Medicago truncatula sieve element occlusion 1 (MtSEO1), MtSEO2 and MtSEO3] and MtSEO1 expression was shown to be restricted to immature sieve elements. Here, we present a detailed molecular analysis of the MtSEO2 and MtSEO3 promoters and show, through reverse transcriptase-polymerase chain reaction and reporter-transgenic experiments in M. truncatula roots and tobacco plants, that their expression profiles are comparable to MtSEO1. The impact of these data on the likelihood of MtSEO gene co-regulation by common transcription factors is discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Forisomes are ATP-independent mechanoproteins found in the phloem sieve tubes of leguminous plants (Fabaceae). Wound-induced increases in the level of intracellular calcium cause forisomes to undergo a rapid conformational change from a condensed to a dispersed state, sealing the conducting tubes and preventing the loss of photo-assimilates (Knoblauch et al. 2001, 2003). Remarkably, this conformational change is fully reversible after wound repair (Knoblauch et al. 2001). The evolutionary importance of forisomes was recently showcased by a comprehensive analysis of the aphid saliva proteome, which identified a number of calcium-binding proteins that probably absorb free Ca2+ ions in order to preserve the contracted forisome state and allow continuous feeding from the phloem (Will et al. 2007).
The forisomes (formerly crystalline P-proteins; Behnke 1991) are grouped according to the presence (group 1) or absence (group 2) of tail-like protrusions at both poles of the protein body (Lawton 1978a, b). Their size differs according to the plant species, e.g. a diameter/length ratio of ∼3/10 µm in Medicago truncatula and ∼6/30 µm in Vicia faba (Knoblauch et al. 2001; Noll et al. 2007). The ultrastructure of forisomes has been described in detail (Palevitz and Newcomb 1971; Wergin and Newcomb 1970; Lawton 1978a, b; Arsanto 1982), and recent investigations describe a fibrillar substructure packed into large, compact bundles (Schwan et al. 2007a, b). These bundles appear to be twisted, as shown by atomic force microscopy using a specially sharpened tip (Jaeger et al. 2008). The complex macromolecular structure of forisomes suggests the presence of multiple proteins, and the three M. truncatula genes designated MtFOR1, MtFOR2, and MtFOR3 were subsequently shown to encode forisome components (Noll 2005; Noll et al. 2007). The genes were renamed Medicago truncatula sieve element occlusion 1 (MtSEO1), MtSEO2, and MtSEO3 (Pélissier et al. 2008) to avoid confusion with the Arabidopsis thaliana FOR gene family. Green fluorescent protein (GFP)-tagged versions of the MtSEO1, MtSEO2, and MtSEO3 proteins were shown to incorporate into wild-type forisomes (Pélissier et al. 2008).
The expression of MtSEO1 is restricted to immature sieve elements (Noll et al. 2007), where forisomes first appear during plant development (Palevitz and Newcomb 1971; Wergin and Newcomb 1970). Here, we show that MtSEO2 and MtSEO3 are also expressed in immature sieve elements with expression profiles very similar to MtSEO1.
Materials and Methods
Plant Material and Cultivation Conditions
Medicago truncatula cv. Jemalong genotype A17 seeds were kindly provided by Dr. Alfred Pühler (University of Bielefeld, Germany). After germination, plants were cultivated in a growth chamber at 22°C with a 16-h photoperiod and a light intensity of 200 μE m−2 s−1. Nicotiana tabacum var. Petit Havana SR1 plants were maintained in a greenhouse at 25°C with a 16-h photoperiod.
Preparation of Genomic DNA and Total RNA
Genomic DNA was prepared according to established protocols (Doyle and Doyle 1990). Total RNA was isolated using the NucleoSpin® RNA Plant Kit (Macherey-Nagel, Düren, Germany) following the manufacturer’s recommendations.
Cloning and Design of Promoter–Reporter Constructs
The MtSEO2 promoter (PMtSEO2) was amplified by polymerase chain reaction (PCR) from M. truncatula cv. Jemalong A17 genomic DNA using sequence information available in public databases (National Center for Biotechnology Information, http://www.ncbi.nlm.nih.gov) and the primer combination P2 forward (5′-CAA ACA ATG CAA ATG AAC-3′) and P2 reverse (5′-TGA TGA TTT GTT TAT AAA TTA A-3′). Due to the lack of sequence information in the above databases, the MtSEO3 promoter (PMtSEO3) was obtained by genome walking using the BD GenomeWalkerTM Universal Kit (BD Biosciences, Clontech, Mountain View, CA, USA) in combination with the walking primer 5′-GCT GAC ACA TTA CTT ACA AGT GAA GAT GG-3′. All fragments were cloned in the pGEM-T vector system (Promega, Heidelberg, Germany) and verified by sequencing (Sanger et al. 1977).
The MtSEO2 and MtSEO3 promoter-β-glucuronidase (GUS) constructs were generated by reamplifying the promoter sequences from the pGEM-T constructs described above, digesting the products with appropriate restriction endonucleases (EcoRI and RcaI for PMtSEO2, EcoRI and NcoI for PMtSEO3) and inserting them into the corresponding restriction sites of pUC103GUS (kindly provided by Dr. Klaus Fritze, Cologne) to obtain PMtSEO2-GUS and PMtSEO3-GUS, respectively. PMtSEO2 was amplified using forward primer 5′-AGA GAA TTC CAA ACA ATG CAA ATG AAC-3′ and reverse primer 5′-AGA TCA TGA TGA TGA TTT GTT TAT AAA TTA A-3′, and PMtSEO3 was amplified using forward primer 5′-AGA GAA TTC CAA TCC ATT GCT TGT TAT CGC AAT C-3′ and reverse primer 5′-AGA CCA TGG TAG GCC AAG AAA GTT ATT ATA TCT CTC TTT GG-3′ (restriction sites underlined). For transformation, both promoter-GUS constructs were inserted into the EcoRI/HindIII sites of binary vector pBIN19 (Bevan 1984) to obtain pBPMtSEO2-GUS and pBPMtSEO3-GUS, respectively.
Promoter-GFPER constructs were generated using the same strategy, with forward primer 5′-AGA GGT ACC CAA ACA ATG CAA ATG AAC AAA-3′ and reverse primer 5′-AGA CTC GAG TGA TGA TTT GTT TAT AAA TTA A-3′ for PMtSEO2, and forward primer 5′-AGA GGT ACC CAA TCC ATT GCT TGT TAT CGC AAT C-3′ and reverse primer 5′-AGA CTC GAG TAG GCC AAG AAA GTT ATT ATA TCT CTC TTT GG-3′ for PMtSEO3. Both products were digested with KpnI and XhoI (restriction sites underlined) and inserted into the corresponding sites of pBSGFPER (Noll et al. 2007) to obtain PMtSEO2-GFPER and PMtSEO3-GFPER, respectively. For plant transformation, both promoter-GFPER constructs were inserted into the KpnI/HindIII sites of binary vector pBIN19 (Bevan 1984) to obtain pBPMtSEO2-GFPER and pBPMtSEO3-GFPER, respectively.
Plant and Root Transformation and Expression Analysis
The binary vectors described above were transferred into Agrobacterium rhizogenes strain NCPPB2659 and Agrobacterium tumefaciens strain LBA4404. Transgenic M. truncatula roots were generated according to established protocols (Collier et al. 2005) with A. rhizogenes. Genetic transformation of N. tabacum cv. SR1 plants was carried out by A. tumefaciens-mediated leaf-disk transformation as described (Horsch et al. 1986). GUS activity was detected using 5-bromo-4-chloro-3-indolyl-β-d-glucuronide without an oxidative catalyst according to established protocols (Jefferson 1987). GFPER expression in immature sieve elements was monitored with a confocal laser scanning microscope (CLSM; excitation 488 nm, emission 509 nm; Leica DMIRE2, Wetzlar, Germany). Prior to analysis, the abaxial epidermis of the petiole midrib was removed. The nuclei of immature sieve elements were visualized by vacuum infiltration of Bisbenzimide H 33342 (5 μg/ml; Fluka, Munich, Germany) solubilised in sodium phosphate buffer (pH 7) supplemented with 0.1% Tween-20 (Fang and Spector 2007). Nuclei were visualized by CLSM (excitation 355 nm, emission 470 nm). Sieve plates were counterstained with a 0.01% aniline-blue solution according to established protocols (Thompson and Wolniak 2008) and visualized by CLSM (excitation 364 nm, emission 470 nm).
The progression of MtSEO2 and MtSEO3 gene expression in M. truncatula was monitored by reverse transcriptase (RT)-PCR on total RNA from leaves at different developmental stages (young, medium sized, fully developed). After first-strand cDNA synthesis by the reverse transcription of 1 µg total RNA in the presence of an oligo(dT) primer and 1 µl SuperScriptTM II reverse transcriptase (Invitrogen, Karlsruhe, Germany), the remaining RNA was digested using two units of RNase H (Roche, Penzberg, Germany) for 20 min at 37°C. A 1,329-bp MtSEO2-specific fragment was then amplified with forward primer 5′-AAC CTC GTC GGT ATC TCG-3′ and reverse primer 5′-AAT GCA GCA ACT ATC TGG ATC ATC-3′, and a 1,419-bp MtSEO3-specific fragment was amplified with forward primer 5′-TAT TGG GCT ATC ATT ACT GC-3′ and reverse primer 5′-TCA AGA CCT TTT CTC AAT CTG-3′. The housekeeping gene GAPDH was amplified as a control using forward primer 5′-AGC TCA AGG TTA AGG ATG AC-3′ and reverse primer 5′-TGG CCA AGG GAG CAA GGC AA-3′. For the analysis of MtSEO expression in roots, cDNA was obtained from total root RNA as described above. A 1,314-bp MtSEO1-specific fragment was amplified with the forward primer 5′-GTA TCG GAC TAT ACA CTA TC-3′ and reverse primer 5′-TCA TAT CTT GCC ATT CTG-3′. MtSEO2 and MtSEO3 mRNA was monitored in roots by RT-PCR with the same gene-specific primers described above. RT-PCR products were fractionated by non-denaturing gel electrophoresis and visualized by ethidium bromide staining.
Computer-assisted Analysis of Promoter Sequences
The MtSEO promoter sequences were aligned using the Lasergene software package v7.1 (DNAstar, Madison, WI, USA). Putative TATA box sequences were identified by analyzing the corresponding genomic clone using the FGENESH program (http://www.softberry.com). Core binding sites for DNA-binding-with-one-finger (DOF) transcription factors were identified using the PLACE Web Signal Scan software (http://www.dna.affrc.go.jp/PLACE/index.html; Higo et al. 1999), whereas core homeodomain-leucine-zipper (HD-ZIP) binding sites were identified by direct comparison with the elements described for the AtSUC2 promoter sequence (Schneidereit et al. 2008).
Results and Discussion
Cloning and Sequence Analysis of the MtSEO2 and MtSEO3 Promoter Regions
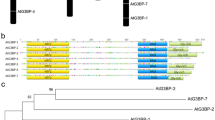
A 1-kb promoter region of the MtSEO1 gene was isolated recently and shown to drive gene expression specifically in immature sieve elements (Noll, 2005; Noll et al. 2007). We cloned similar sized fragments of the MtSEO2 and MtSEO3 promoters (designated PMtSEO2 and PMtSEO3) and compared the three sequences (Fig. 1). Computer-assisted comparison of the PMtSEO1, PMtSEO2, and PMtSEO3 sequences (Fig. 1a) revealed an average of 46% nucleotide sequence identity. Identical nucleotides were not concentrated in any particular region of the promoters but were distributed evenly throughout the sequence, with the exception of the putative TATA box. In addition (Fig. 1b), the MtSEO promoters also shared sequence similarities with other vascular tissue-specific promoters, such as the AtSUC2 (Truernit and Sauer 1995), CmGAS1 (Ayre et al. 2003), rolC (Schmülling et al. 1989), and CFDV (Hehn and Rohde 1998) promoters.
a Computer-assisted comparison of the nucleotide sequences of PMtSEO1, PMtSEO2, and PMtSEO3. For each individual promoter, the adenosine residue of the putative translational start codon ATG (indicated in bold letters) is designated as +1. Gray boxes mark the putative TATA box. Potential binding sites are shown for DOF (underlined and bold italic letters) and HD-ZIP (white letters in black boxes) transcription factors. Asterisks indicate identical nucleotides in the promoter sequences. b Percentage sequence identities between MtSEO promoters and other vascular-specific promoters
Recent in silico analysis of the companion cell-specific AtSUC2 promoter identified potential binding sites for HD-ZIP and DOF transcription factors, and site-directed mutagenesis has shown that these binding sites are essential for promoter activity (Schneidereit et al. 2008). Similar core binding domains for DOF and HD-ZIP transcription factors are also present in the MtSEO promoters (Fig. 1a). The core binding site of HD-ZIP transcription factors is cAAT(A/T)ATTg (Sessa et al. 1998; Schneidereit et al. 2008), and similar motifs are found in PMtSEO2 (AATAATT, position −305) and PMtSEO3 (AATAATT, position −984; AATTATT, position −291) but not in PMtSEO1. In contrast, the typical core motifs for DOF binding sites, AAAG and CTTT (Yanagisawa 1995; Yanagisawa and Schmidt 1999) are present many times in all three MtSEO promoters. HD-ZIP and DOF transcription factors are unique to the plant kingdom, and corresponding genes have been identified in EST libraries prepared from vascular tissue mRNA (Zhao et al. 2004; Pommerrenig et al. 2006). However, further detailed mutation analysis will be necessary to determine whether the HD-ZIP and DOF motifs in the MtSEO promoters play an essential role in limiting promoter activity to immature sieve elements.
MtSEO2 and MtSEO3 Expression in M. truncatula
Forisomes are known to form during the very early stages of phloem development when sieve elements are clearly immature and the nucleus and vacuoles are still present (Wergin and Newcomb 1970; Palevitz and Newcomb 1971). The expression of MtSEO1 is similarly restricted, reflecting the highly regulated spatiotemporal activity of its promoter (Noll et al. 2007). We carried out complementary experiments for MtSEO2 and MtSEO3 by analyzing endogenous gene expression in M. truncatula leaves at various ages, by semi-quantitative RT-PCR. As shown in Fig. 2, MtSEO2 (column b) and MtSEO3 (column c) mRNA were detected in young and medium-aged leaves but not in old leaves. The level of GAPDH mRNA remained constant in all leaves, excluding a general decline in mRNA levels as the basis for our observations (column d). In addition, the expression of MtSEO genes in M. truncatula roots was confirmed by RT-PCR (Fig. 3a).
Analysis of MtSEO expression in wild-type and transgenic M. truncatula roots. a Detection of MtSEO1, MtSEO2, and MtSEO3 mRNA in wild-type roots by semi-quantitative RT-PCR. b PMtSEO1-driven GFPER expression in several “pipeline-like” assemblies of fairly narrow, elongated cells located within the phloem of the vascular bundle of a pBPMtSEO1-GFPER-transgenic M. truncatula root. c A row of cells in a pBPMtSEO1-GFPER-transgenic M. truncatula root displaying the characteristic end-to-end connection of sieve elements. Fluorescent sieve elements from pBPMtSEO1-GFPER-transgenic (d), pBPMtSEO2-GFPER-transgenic (e), and pBPMtSEO3-GFPER-transgenic (f) roots with aniline-blue stained sieve plates (indicated by red arrows). g Immature sieve element with vacuoles (indicated by a blue arrow) from a pBPMtSEO2-GFPER-transgenic root. Scale bars = 10 µm
Next, MtSEO2 and MtSEO3 promoter activity in M. truncatula roots was analyzed by expressing an ER-targeted version of the GFP reporter gene in order to ensure that the spatial distribution of the endogenous protein was not modified by diffusion or transportation. Roots were co-cultivated with A. rhizogenes harboring either pBPMtSEO2-GFPER or pBPMtSEO3-GFPER in addition to the positive control pBPMtSEO1-GFPER. After 3 weeks, microscopic analysis of transgenic root sections revealed the presence of GFP exclusively in the root phloem (Fig. 3b) in cells showing the typical end-to-end connection of sieve elements (Fig. 3c). As expected, fluorescent sieve elements of pBPMtSEO1-GFPER (Fig. 3d), pBPMtSEO2-GFPER (Fig. 3e), and pBPMtSEO3-GFPER (Fig. 3f) transgenic roots possess typical sieve plates. Moreover, large vacuoles could be identified within these cells (Fig. 3g, showing pBPMtSEO2-GFPER as an example), indicating their immature developmental state (see also Fig. 4). However, a more detailed analysis of the regulation of the MtSEO2 and MtSEO3 promoters in M. truncatula was not possible since it proved impossible to regenerate transgenic plants from the transformed root sections.
Analysis of MtSEO2 and MtSEO3 promoter activity in transgenic tobacco plants. ER-tagged GFP-expression in immature sieve elements controlled either by PMtSEO2 (a) or PMtSEO3 (b). The developing sieve plates are indicated with red arrows, nuclei with yellow arrows. SE immature sieve element, CC companion cell. Note that a and b represent an overlay of two individual pictures in order to show GFP expression and the stained nuclei. Histochemical analysis of GUS activity (indicated by black arrows) in pBPMtSEO2 (c) and pBPMtSEO3 (e) T1-transgenic seedlings. GUS staining of the maturation region of a pBPMtSEO2-transgenic primary root (d) and pBPMtSEO3-transgenic secondary root (f) is marked by yellow arrows. Scale bars = 10 µm
MtSEO2 and MtSEO3 Promoter Analysis in N. tabacum
Next, tobacco plants transgenic for pBPMtSEO2-GFPER and pBPMtSEO3-GFPER were produced in order to characterize the regulation of MtSEO2 and MtSEO3 in the phloem during plant development. A detailed analysis of the pBPMtSEO1-GFPER-transgenic tobacco plants has already been published (Noll et al. 2007). Ten individual transgenic lines were regenerated for each construct and verified by PCR to establish transgene integration and RT-PCR to confirm transgene expression (data not shown). GFP fluorescence in transgenic tobacco plants expressing an ER-tagged version of GFP under the control of either PMtSEO2 (PMtSEO2-GFPER; Fig. 4a) or PMtSEO3 (PMtSEO3-GFPER; Fig. 4b) was exclusively restricted to cells located within the vascular bundle of young leaves, specifically in cells with morphological characteristics typical of immature sieve elements. Subsequently, and as expected, PMtSEO2 and PMtSEO3 activities were also temporally regulated. In the seedlings of pBPMtSEO2-GUS (Fig. 4c) and pBPMtSEO3-GUS (Fig. 4e) T1-transgenic tobacco plants, GUS activity was apparent only in the veins of young leaves and disappeared soon after veins reached functional maturity in older leaves. In addition, only the maturation region of primary and secondary roots showed strong GUS activity (Fig. 4d, f).
Conclusion
The MtSEO1, MtSEO2, and MtSEO3 promoters are similar in sequence, and the corresponding genes have very similar, if not identical, spatiotemporal expression profiles as determined by RT-PCR and the analysis of reporter-transgenic M. truncatula and N. tabacum plants. This suggests that the promoters may have one or more transcription factors in common, which could belong to the DOF or HD-ZIP families based on the presence of appropriate binding sites in the promoter sequences. The ability of the MtSEO promoters to drive transgene expression correctly in transgenic tobacco plants that lack forisomes indicates that the regulation of phloem development is highly conserved in diverse plants.
Abbreviations
- CC:
-
companion cell
- GFP:
-
green fluorescent protein
- GUS:
-
β-glucuronidase
- Mt:
-
Medicago truncatula
- SE:
-
sieve element
- SEO:
-
sieve element occlusion
References
Arsanto JP (1982) Observations on P-protein in dicotyledons. Substructural and developmental features. Am J Bot 69:1200–1212. doi:10.2307/2443095
Ayre BG, Blair JE, Turgeon R (2003) Functional and phylogenetic analyses of a conserved regulatory program in the phloem of minor veins. Plant Physiol 133:1229–1239. doi:10.1104/pp. 103.027714
Behnke HD (1991) Nondispersive protein bodies in sieve elements: a survey and review of their origin, distribution and taxonomic significance. IAWA Bull 12:143–175
Bevan M (1984) Binary Agrobacterium vectors for plant transformation. Nucleic Acids Res 12:8711–8721. doi:10.1093/nar/12.22.8711
Collier R, Fuchs B, Walter N, Lutke WK, Taylor CG (2005) Ex vitro composite plants: an inexpensive, rapid method for root biology. Plant J 43:449–457. doi:10.1111/j.1365-313X.2005.02454.x
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Fang Y, Spector DL (2007) Identification of nuclear dicing bodies containing proteins for microRNA biogenesis in living Arabidopsis plants. Curr Biol 17:818–823. doi:10.1016/j.cub.2007.04.005
Hehn A, Rohde W (1998) Characterization of cis-acting elements affecting strength and phloem specificity of the coconut foliar decay virus promoter. J Gen Virol 79:1495–1499
Higo K, Ugawa Y, Iwamoto M, Korenaga T (1999) Plant cis-acting regulatory DNA elements (PLACE) database:1999. Nucleic Acids Res 27:297–300. doi:10.1093/nar/27.1.297
Horsch RB, Klee HJ, Stachel S, Winans SC, Nester EW, Rogers SG, Fraley RT (1986) Analysis of Agrobacterium tumefaciens virulence mutants in leaf discs. Proc Natl Acad Sci USA 83:2571–2575. doi:10.1073/pnas.83.8.2571
Jaeger M, Uhlig K, Clausen-Schaumann H, Duschl C (2008) The structure and functionality of contractible forisome protein aggregates. Biomaterials 29:247–256. doi:10.1016/j.biomaterials.2007.09.020
Jefferson RA (1987) Assaying chimeric genes in plants: the GUS gene fusion system. Plant Mol Biol Rep 5:387–405. doi:10.1007/BF02667740
Knoblauch M, Peters WS, Ehlers K, van Bel AJE (2001) Reversible calcium-regulated stopcocks in legume sieve tubes. Plant Cell 13:1221–1230
Knoblauch M, Noll GA, Müller T, Prüfer D, Schneider-Hüther I, Scharner D, van Bel AJE, Peters WS (2003) ATP-independent contractile proteins from plants. Nat Mater 2:600–603. doi:10.1038/nmat960 Erratum in Nature Mater 2005;4:353
Lawton DM (1978a) Ultrastructural comparison of the tailed and tailless P-protein crystals respectively of runner bean (Phaseolus multiflorus) and garden pea (Pisum sativum) with tilting stage electron microscopy. Protoplasma 97:1–11. doi:10.1007/BF01276386
Lawton DM (1978b) P-protein crystals do not disperse in uninjured sieve elements in roots of runner bean (Phaseolus multiflorus) fixed in glutaraldehyde. Ann Bot (Lond) 42:353–361
Noll GA (2005) Molekularbiologische Charakterisierung der Forisome. Doctoral thesis, Justus-Liebig Universität, Gießen
Noll GA, Fontanellaz ME, Rüping B, Ashoub A, van Bel AJE, Fischer R, Knoblauch M, Prüfer D (2007) Spatial and temporal regulation of the forisome gene for1 in the phloem during plant development. Plant Mol Biol 65:285–294. doi:10.1007/s11103-007-9217-0
Palevitz BA, Newcomb EH (1971) The ultrastructure and development of tubular and crystalline P-protein in the sieve elements of certain papilionaceous legumes. Protoplasma 72:399–426. doi:10.1007/BF01289511
Pélissier HC, Peters WS, Collier R, van Bel AJ, Knoblauch M (2008) GFP tagging of sieve element occlusion (SEO) proteins results in green fluorescent forisomes. Plant Cell Physiol 49:1699–1710. doi:10.1093/pcp/pcn141
Pommerrenig B, Barth I, Niedermeier M, Kopp S, Schmid J, Dwyer RA, McNair RJ, Klebl F, Sauer N (2006) Common plantain. A collection of expressed sequence tags from vascular tissue and a simple and efficient transformation method. Plant Physiol 142:1427–1441. doi:10.1104/pp. 106.089169
Sanger F, Nickler S, Coulson AR (1977) DNA sequencing with chain termination inhibitors. Proc Natl Acad Sci USA 74:5463–5467. doi:10.1073/pnas.74.12.5463
Schmülling T, Schell J, Spena A (1989) Promoters of the rolA, B and C genes of Agrobacterium rhizogenes are differentially regulated in transgenic plants. Plant Cell 1:665–670
Schneidereit A, Imlau A, Sauer N (2008) Conserved cis-regulatory elements for DNA-binding-with-one-finger and homeo-domain-leucine-zipper transcription factors regulate companion cell-specific expression of the Arabidopsis thaliana SUCROSE TRANSPORTER 2 gene. Planta 228:651–662. doi:10.1007/s00425-008-0767-4
Schwan S, Fritzsche M, Cismak A, Heilmann A, Spohn U (2007a) In vitro investigation of the geometric contraction behavior of chemo-mechanical P-protein aggregates (forisomes). Biophys Chem 125:444–452. doi:10.1016/j.bpc.2006.10.008
Schwan S, Fritzsche M, Cismak A, Noll GA, Prüfer D, Spohn U, Heilmann A (2007b) Micromechanical measurements on chemo-mechanical protein aggregates. MRS Proc 0975:DD03–DD10
Sessa G, Steindler C, Morelli G, Ruberti I (1998) The Arabidopsis Athb-8, -9 and -14 genes are members of a small gene family coding for highly related HD-ZIP proteins. Plant Mol Biol 38:609–622. doi:10.1023/A:1006016319613
Thompson MV, Wolniak SM (2008) A plasma membrane-anchored fluorescent protein fusion illuminates sieve element plasma membranes in Arabidopsis and tobacco. Plant Physiol 146:1599–1610. doi:10.1104/pp. 107.113274
Truernit E, Sauer N (1995) The promoter of the Arabidopsis thaliana SUC2 sucrose-H + symporter gene directs expression of beta-glucuronidase to the phloem: evidence for phloem loading and unloading by SUC2. Planta 196:564–570. doi:10.1007/BF00203657
Wergin WP, Newcomb EH (1970) Formation and dispersal of crystalline P-protein in sieve elements of soybean (Glycine max L.). Protoplasma 71:365–388. doi:10.1007/BF01279682
Will T, Tjallingii WF, Thönnessen A, van Bel AJE (2007) Molecular sabotage of plant defense by aphids. Proc Natl Acad Sci USA 104:10536–10541. doi:10.1073/pnas.0703535104
Yanagisawa S (1995) A novel DNA-binding domain that may form a single zinc finger motif. Nucleic Acids Res 23:3403–3410. doi:10.1093/nar/23.17.3403
Yanagisawa S, Schmidt RJ (1999) Diversity and similarity among recognition sequences of Dof transcription factors. Plant J 17:209–214. doi:10.1046/j.1365-313X.1999.00363.x
Zhao Y, Liu Q, Davis RE (2004) Transgene expression in strawberries driven by a heterologous phloem-specific promoter. Plant Cell Rep 23:224–230. doi:10.1007/s00299-004-0812-0
Acknowledgements
The technical assistance of Ann-Christin Müller is gratefully acknowledged. This work was partially supported by the BMBF grant 0312014, the Fraunhofer MAVO program and by the VolkswagenStiftung, contract no. I/82 075.
Author information
Authors and Affiliations
Corresponding author
Additional information
Gundula A. Noll and Boris Rüping contributed equally to this work.
Rights and permissions
About this article
Cite this article
Noll, G.A., Rüping, B., Ernst, A.M. et al. The Promoters of Forisome Genes MtSEO2 and MtSEO3 Direct Gene Expression to Immature Sieve Elements in Medicago truncatula and Nicotiana tabacum . Plant Mol Biol Rep 27, 526–533 (2009). https://doi.org/10.1007/s11105-009-0120-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-009-0120-5