Abstract
Aims
We investigated potential mechanisms by which a seed microbiome recruited from vermicomposted dairy manure alters Pythium aphanidermatum zoospore mediated pathogenesis in cucumber.
Methods
Bioassays were conducted to measure arrival of zoospores at the seed surface via qPCR and subsequent seedling disease incidence. Seed exudates were collected at relevant time points for use in zoospore microscopy assays. Metabolomic analysis was used to characterize seed exudates.
Results
Microbes recruited by the germinating seed from a disease suppressive substrate within 8 hours of sowing prevented zoospore arrival at the seed surface, modified seed exudates and reduced disease incidence. In vitro exposure to microbially modified seed exudates altered zoospore homing responses and reduced both encystment and germination compared to control exudates. Combining modified and control exudates failed to restore zoospore attraction to levels observed with control exudates. Observed zoosporolytic activity of the modified exudates was unique to the ethyl acetate fraction and metabolomic analysis revealed several putative zoosporolytic compounds present at higher relative abundance when compared to control exudates.
Conclusions
The observed disease suppression was likely due to the production of a specific zoosporolytic compound or set of compounds in the spermosphere by one or more members of the seed-recruited vermicompost microbiome.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Studies of plant microbiomes continue to reinforce the importance of host-associated microbes in plant health, especially as they relate to plant immunity and pathogen protection (Berg et al. 2014; Rout 2014; Vandenkoornhuyse et al. 2015). Although many host-associated microbes may be transmitted either vertically or horizontally from adult plants to seeds, epiphytic seed- and root-associated microbes are commonly recruited from soil at the moment of seed-soil contact (Barret et al. 2015; Nelson 2004; Ofek et al. 2011). This is followed by successions of microbes over the course of seedling development (Barret et al. 2015; Liu et al. 2012; Ofek et al. 2011) that include microbes that protect plants from soil-borne pathogens (Mendes et al. 2011).
There has been much interest in elucidating mechanisms by which soil microbes prevent pathogen infection, particularly in agricultural soils that become suppressive through continuous cropping (Mazzola 2002; Thomashow and Bakker 2015; Weller et al. 2002) or application of organic amendments (Benitez et al. 2007; Kowalchuk et al. 2003; Litterick et al. 2004; van Os and van Ginkel 2001). Despite clear microbial involvement in many suppressive soils, the mechanisms of disease suppression remain elusive (Janvier et al. 2007). This is due, in part, because pathogenesis is a spatially and temporally-dynamic process that involves the plant, pathogen, and associated microbes.
Given that soil microbes associate with plants in a highly specific manner (Hartmann et al. 2009), an appropriate approach for understanding disease suppression is to focus on host-associated as opposed to soil-associated microbes (Mendes et al. 2011) that are present and active at the precise time of plant infection. Such a targeted approach can increase the likelihood of discovering microbes more directly associated with changes in pathogen behavior that reduce plant infection. Such a strategy has proven valuable in understanding how spermosphere bacteria suppress pathogen infections of seeds (Heungens and Parke 2000; Windstam and Nelson 2008a, 2008b) as well as in providing insight into mechanisms of compost-induced disease suppression (Chen et al. 2012; Chen and Nelson 2012, 2008; McKellar and Nelson 2003).
In our current work, we adopt this approach to understand how cucumber seedling infections by Pythium aphanidermatum are suppressed in vermicomposted dairy manure (VDM). A number of important aspects of this system facilitate the use of this approach. First, many vermicomposts are suppressive to diseases caused by many major soil-borne pathogens (Jack 2011). Vermicomposts produced in a highly engineered flow-through system are chemically and physically uniform and consistently disease suppressive (Table 1, Fig. 1). Second, P. aphanidermatum is one of the most important seed- and root-infecting plant pathogens with a host range of over 650 species (Farr and Rossman 2015) and inherently sensitive to microbial interference (Martin and Loper 1999), making it a valuable model to assess disease suppression. Third, P. aphanidermatum is believed to infect seeds and roots via the formation of motile zoospores, which display a complex but well-characterized homing response (Deacon and Donaldson 1993; Nelson 2006; Walker and van West 2007).
Representative 7 d-old cucumber (Cucumis sativus cv. “Marketmore 76”) seedlings from disease suppression bioassays described in SOM. Surface disinfested cucumber seeds were sown in a sterile quartz sand amended with vermicomposted dairy manure (40% v:v), b sterile quartz sand, and c sterile quartz sand amended with autoclaved vermicomposted dairy manure (40% v:v). Each group of 10 inoculated seedlings received 6 × 105 Pythium aphanidermatum zoospores. Matric potential was held constant at −3.5 kPa. Scale bar = 5 cm
During pathogenesis, zoospores respond to chemical cues from the host in the form of seed or root exudates to detect and swim towards the infection court (Deacon 1996). There is evidence to suggest that P. aphanidermatum and other oomycete zoospores may also respond to electrical gradients around roots (Morris and Gow 1993; van West et al. 2002). However, the presence of these gradients around seeds is unknown. Upon arrival at the host surface, zoospores attach, encyst on the seed, radical, or root surface, then subsequently germinate and penetrate the host. Although homing cues are largely unidentified, they appear to be species- and developmental stage specific (Donaldson and Deacon 1993a, 1993b). Interference with these cues can be an effective means of suppressing infection by zoosporic pathogens (Heungens and Parke 2000; Islam 2010; Lioussanne et al. 2008; Shang et al. 1999). However, because homing interference may result either from the degradation of a zoospore attractant or the production of a zoospore repellant/toxin (Zhou and Paulitz 1993), or a combination of both (Heungens and Parke 2000), the specific mechanisms of homing interference may be obscured. This makes it necessary to examine each homing response stage along with the chemical cues that elicit these responses to better understand pathogen suppression.
We designed our current work to understand how seed-colonizing microbes recruited from a disease suppressive substrate alter zoospore responses of Pythium aphanidermatum to germinating seeds and subsequently suppress disease. We attempt to explain disease suppression by relating reductions in seed colonization by P. aphanidermatum with changes in zoospore behavior that result from direct chemical alterations of seed exudates by seed-recruited microbes. This approach allows us to answer the following questions; 1) can the seed-recruited microbiome from a suppressive substrate explain the observed disease suppression?, 2) which stages of the homing response are altered by the recruited seed microbiota?, 3) are altered zoospore responses due to the modification of seed exudates by the seed microbiota?, and if so, 4) does this modification of seed exudates involve (a) the degradation of a chemotactic cue, (b) the production of a zoospore repellant/lytic agent or both? Answers to these questions will further establish the role of the host-associated microbiome in the disruption of chemical signaling between hosts and pathogens, thereby successfully preventing the development of disease.
Materials and methods
Experimental materials, detailed methods and results of the basic disease suppression bioassay along with the methods and results of experiments designed to confirm that the point source inoculation method in the bioassay apparatus can only cause seedling disease via actively swimming zoospores are included in the Supporting Online Materials [SOM]. In addition, the SOM contain qPCR methods used to assess P. aphanidermatum colonization of germinating cucumber seeds. The experiments described below are based on two types of samples collected from the same design of point source transplant bioassays; 1) germinating seeds or 2) seed exudates. Sampling timepoints are summarized in Fig. 2 which lays out the broader experimental design where, for example, seed exudate samples collected at 12 h post transplant (hpt) and used in in vitro zoospore assays are roughly equivalent to the spermosphere environment experienced by inoculated germinating seeds collected 12 h post inoculation (hpi) and used to assess pathogen colonization via qPCR (Fig. 2).
Schematic linking in vivo and in vitro experiments. Time points for the collection of seed exudates are indicated in the top portion of the figure and time points for zoospore inoculation and harvest of germinating seeds for DNA extraction are indicated in the bottom portion of the figure, hpt: hours post-transplant, hpi: hours post inoculation
In situ zoospore swimming bioassay
Point source transplant bioassays similar to those described previously (Chen and Nelson 2008; Heungens and Parke 2000) were used to assess zoospore attraction to germinating seeds and determine whether seed-recruited VDM microbes can protect seeds from zoospore infection. See the supplementary online materials [SOM] for a description of experiments to eliminate mass flow as a potential mechanism for zoospore movement in the point source transplant bioassays. All bioassays were conducted in a Büchner funnel apparatus to facilitate the strict control of substrate matric potentials. Seeds were embedded into nylon mesh in a 4 cm diam circle before sowing to ensure their position would not be disturbed during flooding. After sowing, substrates were flooded from below through the fritted glass in the Büchner funnels (Fig. S1). After ~5 min, matric potentials were adjusted to −3.5 kPa and allowed to equilibrate. Seeds were sown in sand and in VDM-amended sand (40% v:v) as described above and allowed to germinate 8 h before transplanting to sterile sand and point source inoculating with a zoospore suspension (5 mL, 8 × 104 zoospores mL−1). The funnels were then covered with ventilated Parafilm™ to create a moist chamber. Two thirds of the funnels were destructively harvested at 12, 18 and 24 hpi for assessments of Pa58 biomass on seeds via quantitative PCR and collection of seed exudates for in vitro zoospore assays (Fig. 2). One third of the seeds were assessed for seedling survival and disease symptoms at 9 d to assure the viability of zoospore inoculum. For 9-d-old seedlings, disease incidence (presence or absence of symptoms) was analyzed in SAS using binary logistic regression with Bonferroni’s correction for multiple comparisons. Differences in Pa58 DNA on seed surfaces were analyzed using an ANOVA in the general linear model of SAS with sliced interactions for treatment*hpi to generate a means separation.
Quantitative assessment of Pa58 biomass associated with seeds
Cucumber seeds were removed from their respective substrates at 12, 18 and 24 hpi and gently tapped to remove adhering sand and VDM particles. Ten seeds were placed in initial DNA extraction buffers (UltraClean® Soil DNA Isolation Kit, MoBio, USA) and frozen overnight at -20 °C before sample processing. Manufacturer’s protocol for samples with high humic acids was used for DNA extraction. Pa58-specific primer sets were designed using a consensus sequence generated from an alignment of 42 ITS sequences from the NCBI database and our laboratory strain Pa58 (Lasergene® Megalign, DNASTAR, USA). One primer pair was selected for use in quantitative PCR analysis (PaITS-F 5′ AATGTACGTTCGCTCTTTCTTG 3′, PaITS-R 5′ GGTTGCTTCCTTTAATGTCCTA 3′). Quantitative PCR (qPCR) was carried out using an iQ™5 thermocycler (Bio-Rad, USA), using protocols outlined in the SOM.
In vitro zoospore responses to seed exudates
Zoospore encystment assay
A zoospore suspension was prepared as described in the SOM and 100 mL (1.2 × 104 zoospores mL−1) was added to a 15 cm diam glass petri dish. Rubber gaskets (Grace BioLabs, Bend OR) were adhered to microscope slides, filled with 305 μL 0.01% agarose which was allowed to set for 25 min. Ten uL of 35 X reconstituted seed exudates from each treatment was added to the agarose discs and allowed to dry for 3 min. Treatments are defined as follows: Control exudate (CE) or sand treatment consists of exudates collected from seeds pre-germinated in sand for 8 h, then transplanted to sand for 24 h, Microbially modified exudate (MME) or vermicompost treatment consists of exudates collected from seeds pre-germinated in 2 vermicompost amended sand for 8 h then transplanted to sand for 24 h. See SOM for seed exudate collection details. Slides were then immersed in the zoospore suspension and incubated in the dark at room temperature for 30 min. Slides were removed and 4 images were acquired at 10X magnification for each treatment and used for zoospore enumeration (DP25 digital camera with DP2-BSW software, Olympus, USA). A mixture of CE and MME exudate samples was prepared for an additional assay to determine whether observed differences in zoospore encystment were due to the absence of an attractant or the addition of a repellant/lytic agent in the VDM MME samples. For the mixture treatment, freeze-dried seed exudates were re-suspended at 70 X and then mixed at a 1:1 ratio so that their individual concentration in the “mixture” treatment is equivalent to their concentration when tested individually (35 X). Assay and imaging were carried out as described above with an additional imaging step 1 h after initial imaging to monitor the fate of zoospore cysts at higher magnification (304 X); lysis, encytment or encystment and germination). Data were analyzed using an ANOVA with a Tukey’s test for means separation (Minitab 16, USA).
Zoospore germination assay
Zoospore cyst germination percentages were calculated for pre-encystment and post-encystment exposure of zoospores to seed exudates and fractionated seed exudates (Please see section D of the SOM for details on seed exudate fractionation). For pre-encystment exposure, 10 μL of the test substance (exudate or exudate fraction) was mixed with 6 mL swimming zoospore suspension for 15 min after which suspensions were mechanically encysted via vigorous agitation, and poured into a tissue culture well (Nunc 8 well square tissue culture plates, Thermo Scientific, USA) containing a thin layer of molecular grade low melt agarose and incubated for 1 h prior to imaging. For post-encystment exposure, 10 μL of the test substance was mixed with 6 mL mechanically encysted zoospore suspension, immediately added to the tissue culture well and incubated for 1 h prior to imaging. The proportion of germinated cysts (either via germ tubes or secondary zoospores) and germ tube lengths were calculated through image analysis (Olympus DP2-BSW software) for a total of 4 fields of view (~4 mm2) with a water immersion objective (20 X 0.5 W Ph2, Zeiss). Germination percentages were analyzed using binary logistic regression and Bonferonni’s adjustment for multiple comparisons (SAS v.9.3). Germ tube lengths were analyzed using an ANOVA with Tukey’s test for multiple comparisons (Minitab 16).
Metabolomic analysis of seed exudates
Six replicate samples of MME and CE were prepared from seeds germinated for 24 h in sand or VDM and fractionated as described in the SOM. Samples were resuspended in a small volume of methanol and shipped to Metabolon (http://metabolon.com) for metabolomic analysis as described previously using GC/MS (Lawton et al. 2008) and LC/MS (Evans et al. 2009), except that samples were analyzed directly rather than being subjected to the primary sample extraction procedure. Integrated peak ion count data for each identified compound was used to represent the relative amount of compound in each sample. Missing data were imputed using the minimum observed value for each compound; test groups were compared by statistical analysis (R; Welch’s two-sample t-test) using log-transformed imputed data. False Discovery Rates (FDR), expressed as q-values, were calculated (Storey and Tibshirani 2003). Ratios of the group means (from imputed data) were used to construct a fold-change heat map, with fold-change values >1 as yellow, and fold-change values <1 as blue.
Results
Zoospore homing response in the presence and absence of VDM
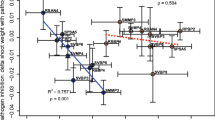
To determine when a suppressive microbiota develops on seed surfaces, seeds were sown in vermicompost for 8 h, then transplanted to sterile sand, inoculated and monitored for pathogen colonization at 12, 18 and 24 hpi and disease development at 9 dpi (Fig. 1). In the absence of VDM, motile zoospores colonized seeds rapidly (within 12 h), causing high seedling mortality (Fig. 3a). At 18 and 24 hpi, significantly fewer zoospores (i.e., less P. aphanidermatum DNA detected) reached seeds exposed to VDM rather than sand (Fig. 3a). Correspondingly, seedling mortality was lower in seeds exposed for 8 h to VDM rather than sand prior to transplant and inoculation. The standard curve equations for DNA extracted from pure lyophilized Pa58 mycelia (Ct = 28.9 + 3.15 log ng DNA, R2 = 99.7%) and for DNA extracted from seeds sown in VDM combined with pure lyophilized Pa58 mycelia (Ct = 28.2 + 3.13 log ng DNA, R2 = 98.9%) had near identical slopes, indicating that any residual VDM on the seed surface did not appreciably affect our ability to detect Pa58 DNA on seeds sown in VDM.
a In vivo and b-c in vitro responses of Pythium aphanidermatum (Pa58) zoospores to microbially modified cucumber seed exudates; overview of the experimental design (Fig. 1). a) Least squared means of Pa58 DNA on seeds removed from a disease suppression bioassay at 12, 18 and 24 hpi for seeds pre-germinated in sand or 40% v:v vermicomposted dairy manure (VDM) for 8 h prior to transplanting to sand and zoospore inoculation. Each point is an average of 2 Büchner funnels within 3 full repetitions of the qPCR assay (n = 6), asterix indicates significant difference between treatments at selected time point (treatment p < 0.0001, total treatment*hpi p = 0.101, significant individual treatment*hpi interactions at 18 and 24 hpi all p < 0.001). Seedling disease incidence (DI %) at 9 d indicated on secondary y axis. b) Zoospore arrival and encystment response to seed exudates modified by the 8 h seed colonizing microbial community derived from VDM and harvested 12, 18 or 24 h after transplanting colonized seeds to sterile sand and zoospore inoculation. Each value is an average of 4 fields of view from 3 replications respectively for 3 different MME batches (n = 9, p < 0.0001). Asterix indicates significant difference between treatments at selected time point. c) Percent germination of Pa58 zoospores exposed pre- or post-mechanical encystment. For pre-encystment exposure only exudates collected at one timepoint (24 hpt) ware tested, while for post-encystment exposure exudates collected at three timepoints (12, 18 and 24 hpt) were tested. Binary logistic regression for both pre-encystment exposure (n = 3, p < 0.0001, sand significantly higher than VDM and the water control) and post-encystment exposure (n = 3, p < 0.0001), asterix indicates significant difference between sand and VDM treatments at selected time point. An average of 400 total zoospores from 4 fields of view and 3 replications were used to calculate germination percentages for each treatment – time point combination with a total of over 9000 individual zoospores scored
Zoospore homing responses to seed exudates
Zoospore chemotaxis & encystment
Exudates from seeds sown in sand for 8 h before transplant to sand (control exudates = CE) attracted high numbers of zoospores that subsequently encysted and germinated. Significantly greater numbers of zoospores encysted in the presence of CE from later rather than earlier time points (24 h post transplant (hpt) > 12 and 18 hpt, Fig. 3b). Numbers from zoospores arriving and encysting in response to exudates from seeds sown in vermicompost for 8 h before transplant to sand (microbially modified exudates = MME) did not differ from those exposed to water for any of the timepoints tested. Combining CE with MME failed to restore zoospore chemotaxis and encystment (Table 2). Instead, a higher proportion of zoospores lysed following exposure to MME or to a mixture of MME and CE compared to those exposed only to CE or water (Fig. 4, Table 2).
Representative P. aphanidermatum zoospore germlings exposed to microbially modified seed exudate (MME) for 30 min in the zoospore encystment assay. Horizontal panels represent different microscopy images from the same treatment. Treatments include; a 24 h MME from seeds pre-germinated in sand for 8 h, then transplanted to sand for 24 h, b 24 h MME from seeds pre-germinated in VDM for 8 h, then transplanted to sand for 24 h and, c a 1:1 mixture of A & B. Scale bar = 25 μm
Ethyl acetate (EtOAc) fractionation of CE and MME significantly impacted zoospore responses to the corresponding fractions. For CE, higher numbers of zoospores swam to and encysted on the EtOAc fraction (EFrac) compared to the aqueous fraction (AFrac) (Table 3). Numbers of encysted zoospores did not differ between the MME AFrac and EFrac compared to the respective water and EtOAc controls. Highest percentages of zoospore cyst germination were observed in response to the CE AFrac; lower levels were observed in water (no seeds) and the MME AFrac. Germination rates in the MME EFrac were significantly lower than those in the EtOAc and water controls. A significantly higher proportion of zoospores lysed in the MME EFrac compared to the AFrac (Table 3). Zoospores exposed to the MME EFrac lacked germ tubes and showed signs of membrane disruption (Fig. 5), both of which were not observed in the MME AFrac, either fraction of control exudates, or in the water or EtOAc controls (Fig. 5).
Representative Pythium aphanidermatum zoospore germlings exposed to fractionated seed exudates in the zoospore encystment assay. Horizontal panels represent different microscopy images from the same treatment. a aqueous (AFrac) and b EtOAc fractions (EFrac) of exudates from seeds pre-germinated in sand for 8 h and transplanted to sand for 24 h; c AFrac and d EFrac respectively of MME from seeds pre-germinated in VDM for 8 h and transplanted to sand for 24 h. Scale bar = 25 μm
Zoospore germination
Pre-encystment incubation of zoospores with control exudates (CE) collected at 24 hpt resulted in a higher germination rate than incubation with microbially modified exudates (MME) collected at 24 hpt or water (Fig. 3c). For post-encystment exposure, cyst germination rates declined over time for all exudates tested (12 hpt > 18 hpt > 24 hpt). However, cyst germination rates after incubation in MME were significantly lower than those observed after incubation in control exudates (Fig. 3c). No significant differences in germ tube lengths were observed between exudates collected 24 hpt from either treatment (p = 0.299, data not shown).
Partial characterization of CE and MME
A comparative analysis of the EFrac of CE and MME (Fig. S2) was performed using LC/GC-MS. A total of 286 individual compounds were identified among all seed exudate samples, of which 146 were of an unknown structure and 41 had not previously appeared in any previous analysis at Metabolon. Identified compounds fell into the following classes: amino acids, carbohydrates, cofactors, lipids, nucleotides, products of secondary metabolism, and xenobiotics. A total of 106 compounds differed significantly in relative concentration between CE and MME (p < 0.05). Of those compounds, only 20 were at a higher concentration in MME (Fig. 6) whereas 86 had a higher concentration in CE (Fig. S2). Of the compounds present at higher prevalence in MME, unknown compound X-19501 was detected at 9X higher relative abundance in MME than in CE, whereas other compounds were only 1X - 2X more prevalent. Additionally, higher concentrations of free fatty acids and xenobiotics were found in MME relative to CE. The xenobiotics hydrochlorothiazide (a high blood pressure medication registered for use in dairy cattle) and sucralose (an artificial sweetener), are both assumed to be carry overs from the dairy manure and not likely to be involved in the observed changes in zoospore responses. Compounds at higher concentrations in CE were mainly amino acids, sugars and lipids, all of which are typical seed exudate components (Nelson 2004). The baseline false discovery rates (FDR) for a dataset of this size (n = 286) was calculated to be 14.3%, indicating that q-values under 10% offer high confidence in the result.
Individual compounds detected at a significantly higher relative abundance in VDM microbially modified seed exudate (MME) compared to control seed exudate. Six biological replicates per treatment were considered, as indicated by column headings. Relative abundances derived from gas and liquid chromatography combined with mass spectrometry (GC/LC-MS). Scaled imputed data derived from raw integrated peak ion counts for individual compounds were natural log transformed and subjected to a Welch’s two way t-test to calculate p-values for treatment differences. Heat map colors as follows; blue = lower relative abundance, black = no change, yellow = higher relative abundance
Discussion
Our analysis of the interactions among germinating seeds, seed-recruited microbiota, and the pathogenic responses of P. aphanidermatum has offered several insights into the ecology of pathogenesis. Our results point to the important role that host-recruited microbes play in disease suppression. This is consistent with growing evidence that microbes recruited to and rapidly established on plant surfaces early in seed germination may directly modulate the activities of soil pathogens (Chen et al. 2012; Chen and Nelson 2012, 2008; McKellar and Nelson 2003). In the absence of these microbes, zoospores were able to swim towards and colonize seed surfaces within 12 h. In contrast, the presence of a rapidly-developing seed microbiome recruited from a VDM substrate greatly reduced pathogen colonization and infection of germinating seeds, providing a level of disease protection equivalent to sowing directly in the solid VDM substrate. Our data further indicate that disease suppression in VDM correlates with reductions in the seed-associated biomass of P. aphanidermatum within hours after sowing. Although other microbes or compounds not associated with seeds may be present in VDM that may impact other stages of the P. aphanidermatum life cycle (Mondal et al. 1996), microbial interactions that alter zoospore homing responses are likely to have a more direct and significant impact on disease development as has been demonstrated in other pathosystems (Chen et al. 2012; Chen and Nelson 2008, 2012).
Monitoring the pathogen’s homing response to MME to assess the suppressive activity of the seed microbiota led to several important observations. We had hypothesized that because chemotaxis and encystment of P. aphanidermatum zoospores occur in direct response to chemical compounds in host seed exudates (Donaldson and Deacon 1993a, 1993b), any interference with these homing responses by the seed microbiota would result from the microbial modification of the seed exudate chemical profile rather than the direct microbial attack of incoming zoospores. This was supported by the observation that zoospores responded differently to cell-free exudates collected from seeds colonized by a suppressive microbiota than from seeds sown in sand and presumably colonized only by their vertically transmitted seed endophytes. A possible explanation for this pattern is that both chemotaxis and encystment were inhibited by microbial modifications to exudates. Alternatively, even if zoospores swam chemotactically to MME in in vitro assays, they may not have encysted or attached to the agarose and would not have been recorded in our assays. Although we have no direct evidence for the inhibition of zoospore chemotaxis, it is clear that encystment, germination and lysis were significantly impacted.
Previously this “pathogen as biosensor” approach has led to important biological insights about the interactions between zoospores, plant hosts and individual host-associated microbes. For example, exudates collected from roots colonized with Pseudomonas spp. attracted fewer P. aphanidermatum zoospores than exudates from non-colonized roots (Zhou and Paulitz 1993). Similarly, exudates from roots colonized with Glomus intraradices attracted fewer Phytophthora nicotianae zoospores than water (Lioussanne et al. 2008), presumably because of the zoospore repellants isocitric acid and proline. Others have demonstrated that microbially modified root exudates can have dual modes of action on homing responses of zoospores. For example, reduced numbers of Pythium torulosum zoospores actively swam towards and encysted on tobacco root exudates treated with an antibiotic-producing as opposed to an antibiotic-deficient strain of Bacillus cereus (Shang et al. 1999). However, only the antibiotic-producing strain reduced zoospore cyst germination, again suggesting multiple mechanisms of homing response interference (Shang et al. 1999). Similarly, attraction of P. aphanidermatum zoospores to pea seed exudates was eliminated when seeds were treated either with an antibiotic-producing or an antibiotic-deficient strain of Burkholderia cepacia (Heungens and Parke 2000). As before, only the antibiotic-producing strain caused zoospore lysis, prevented cyst germination, and reduced germ tube growth (Heungens and Parke 2000), indicating that B. cepacia not only reduced chemoattractants but also produced a zoosporocidal toxin.
Exposure of P. aphanidermatum zoospores to mixtures of CE and MME point to zoospore repellant(s)/toxin(s) as the predominant factor preventing P. aphanidermatum zoospores from reaching seeds. Given that zoospore chemotaxis and encystment in a mixture of CE and MME was no different than responses to MME alone points to the overriding role of a potential zoosporocidal toxin or repellant in homing interference. Others have demonstrated that bacterial compounds may repel, but not damage fungal zoospores (Lam et al. 2011). However, in our system significant zoospore damage was observed in the MME EFrac, pointing to compound X-19501 as a likely candidate for zoospore toxicity. Although X-19501 has not been identified, it shares similarities with hydroxylated dioic acids and may be a structural isomer of 3-hydroxytetradecanediote (Alexander, D. Metabolon Report), a class of lipids similar to the anti-fungal compound 3-hydroxydecanoic acid identified in MME (Sjogren et al. 2003). This compound may also be present as a fatty acid moiety of cyclic lipopeptides, also known to be zoosporolytic (Raaijmakers et al. 2006).
Additional evidence for the presence of a zoosporocidal toxin came from observations of germinating zoospore cysts. A high proportion of zoospores exposed to MME from VDM either lysed during encystment or, if they ultimately encysted, did not subsequently germinate. This response is similar to other zoosporolytic molecules affecting other oomycete pathogens, including zwittermycin on Phytophthora cactorum (Gilbert et al. 1990), xanthobaccin A on Aphanomyces cochlioides (Islam et al. 2005), cyclic lipopeptide Mass A on Phytophthora infestans (de Bruijn et al. 2007), rhamnolipid B on Phytophthora capsici (Kim et al. 2000) and oat root saponins on Pythium spp. (Deacon and Mitchell 1985). Collectively, these observations indicate that a wide range of compounds could be potentially responsible for lysis in our pathosystem. Zoospores, vesicles formed during zoosporogenesis, and zoospore cysts in early stages of development would be most susceptible to lysis because they all lack a cell wall. P. aphanidermatum vesicles formed during zoosporogenesis were also observed to lyse in the present of liquid extracts from the same source of VDM used in this study (Carr and Nelson 2014). It should be kept in mind that not all compounds known to interfere with zoospore pathogenesis are necessarily lytic (Folman et al. 2004). Our observation of reduced germination when mechanically encysted zoospores with fully formed cell walls were exposed to MME, suggests that multiple compounds altering zoospore behavior may also be produced by VDM microbes colonizing seeds.
While zoosporolytic compounds may be an important factor inhibiting P. aphanidermatum homing responses, a dual mechanism of both direct interference via the presence of a toxin as well as a minor, but significant, homing response interference via the degradation of attractants is suggested from our MME fractionations. Only low levels of zoospore attraction and encystment were observed in the MME AFrac compared to CE AFrac, indicating that chemoattractants may have been degraded. Because the total dry mass of CE and MME was roughly equivalent across multiple samplings, it is unlikely that the changes in zoospore response result from a general reduction in seed exudation rates, but rather from the degradation of specific compounds that serve as chemotaxis cues. Whereas none of the individual 86 compounds present at lower concentrations in MME than in CE are known zoospore attractants or cyst germination stimulants, only a very narrow range of compounds have ever been tested with P. aphanidermatum zoospores (Donaldson and Deacon 1993b), making it possible that some of these compounds are indeed chemoattractants. Considering that over half of the compounds present at lower concentration in MME are unknown, it is clear that seed exudate and spermosphere chemistry is a frontier for future exploration. In terms of general chemical classes, amino acids are important zoospore chemoattractants (Donaldson and Deacon 1993b), free fatty acids and their variants can trigger sporangial germination (Windstam and Nelson 2008a, 2008b) in Pythium spp., and both these chemical classes were well represented among the compounds found to be in lower relative abundance in MME.
While we only indirectly measured the functional attributes of the recruited seed microbiome via its impact on seed exudates, a wide range of compost-derived bacteria could be responsible for the observed impacts on seeds. For example, Gammaproteobacteria (largely Pseudomonas spp.) on the surface of cucumber seeds were associated with the suppression of Pythium ultimum (Chen et al. 2012) and Firmicutes (especially the Paenobacillaceae) were associated with the suppression of Pythium aphanidermatum (Ofek et al. 2011). Furthermore, fatty acid-metabolizing bacteria and Actinobacteria on cotton seeds were associated with the suppression of Pythium ultimum (McKellar and Nelson 2003).
Seed microbiomes, at least during the earliest stages of germination, are of relatively low complexity compared to those of the rhizosphere (Barret et al. 2015). For example, while roots of plants grown in soil can harbor a diversity of organisms estimated between ~18,700 (Lundberg et al. 2012) and ~30,000 OTUs (Mendes et al. 2011), cucumber seeds germinating for 8 h in Pythium suppressive composts have ~350 (Chen et al. 2012) and by 24 h ~ 550 bacterial OTUs (Ofek et al. 2011). These bacteria colonize primarily the intercellular crevices in emerging radicles, but are also present on the seed coat (Ofek et al. 2011). We found that the 8 h seed microbiome, which is present long before the development of a root system, can contribute to the observed suppression of disease. These results further identify the spermosphere as a crucial microbial habitat supporting important microbial interactions influencing host survival and coexistence with soil pathogens. Future work focusing on how bacterial taxa in soils respond to specific chemical components present in seed exudates and how these taxa prevent zoospore-mediated infections will further elucidate how host-associated microbiota lead to the suppression of disease.
References
Barret M et al (2015) Emergence shapes the structure of the seed microbiota. Appl Environ Microbiol 81:1257–1266. doi:10.1128/Aem.03722-14
Benitez MS et al (2007) Multiple statistical approaches of community fingerprint data reveal bacterial populations associated with general disease suppression arising from the application of different organic field management strategies. Soil Biol Biochem 39:2289–2301
Berg G, Grube M, Schloter M, Smalla K (2014) The plant microbiome and its importance for plant and human health. Front Microbiol 5:491. doi:10.3389/Fmicb.2014.00491
Carr EA, Nelson EB (2014) Disease-suppressive vermicompost induces a shift in germination mode of Pythium aphanidermatum zoosporangia. Plant Dis 98:361–367. doi:10.1094/PDIS-05-13-0466-RE
Chen M-H, Jack ALH, McGuire IC, Nelson EB (2012) Seed-colonizing bacterial communities associated with the suppression of Pythium seedling disease in a municipal biosolids compost. Phytopathology 102:478–489. doi:10.1094/phyto-08-11-0240-r
Chen M-H, Nelson EB (2008) Seed-colonizing microbes from municipal biosolids compost suppress Pythium ultimum damping-off on different plant species. Phytopathology 98:1012–1018
Chen M-H, Nelson EB (2012) Microbial-induced carbon competition in the spermosphere leads to pathogen and disease suppression in a municipal biosolids compost. Phytopathology 102:588–596
de Bruijn I, de Kock MJ, Yang M, de Waard P, van Beek TA, Raaijmakers JM (2007) Genome-based discovery, structure prediction and functional analysis of cyclic lipopeptide antibiotics in Pseudomonas species. Mol Microbiol 63:417–428
Deacon JW (1996) Ecological implications of recognition events in the pre-infection stages of root pathogens. New Phytol 133:135–145
Deacon JW, Donaldson SP (1993) Molecular recognition in the homing responses of zoosporic fungi, with special reference to Pythium and Phytophthora. Mycol Res 97:1153–1171
Deacon JW, Mitchell RT (1985) Toxicity of oat roots, oat root extracts, and saponins to zoospores of Pythium spp. and other fungi. Trans Br Mycol Soc 84:479–487. doi:10.1016/S0007-1536(85)80010-3
Donaldson SP, Deacon JW (1993a) Differential encystment of zoospores of Pythium species by saccharides in relation to establishment on roots. Physiol Mol Plant Pathol 42:177–184
Donaldson SP, Deacon JW (1993b) Effects of amino acids and sugars on zoospore taxis, encystment and cyst germination in Pythium aphanidermatum (Edson) Fitzp, P. catenulatum Matthews and P. dissotocum Drechs. New Phytol 123:289–295
Evans AM, DeHaven CD, Barrett T, Mitchell M, Milgram E (2009) Integrated, nontargeted ultrahigh performance liquid chromatography/electrospray ionization tandem mass spectrometry platform for the identification and relative quantification of the small-molecule complement of biological systems. Anal Chem 81:6656–6667
Farr DF, Rossman AY (2015) Fungal databases, systematic mycology and microbiology laboratory. ARS, USDA. Retrieved from http://nt.ars-grin.gov/fungaldatabases/ http://nt.ars-grin.gov/fungaldatabases/
Folman LB, De Klein MJEM, Postma J, van Veen JA (2004) Production of antifungal compounds by Lysobacter enzymogenes isolate 3.1T8 under different conditions in relation to its efficacy as a biocontrol agent of Pythium aphanidermatum in cucumber. Biol Control 31:145–154. doi:10.1016/j.biocontrol.2004.03.008
Gilbert GS, Handelsman J, Parke JL (1990) Role of ammonia and calcium in lysis of zoospores of Phytophthora cactorum by Bacillus cereus strain UW85. Exp Mycol 14:1–8
Hartmann A, Schmid M, van Tuinen D, Berg G (2009) Plant-driven selection of microbes. Plant Soil 321:235–257
Heungens K, Parke JL (2000) Zoospore homing and infection events: effects of the biocontrol bacterium Burkholderia cepacia AMMDR1 on two oomycete pathogens of pea (Pisum sativum L.) Appl Environ Microbiol 66:5192–5200
Islam MT (2010) Morphology and behavior of the successive generations of secondary zoospores of the damping-off pathogen Aphanomyces cochlioides. J Plant Pathol 92:471–478
Islam MT, Hashidoko Y, Deora A, Ito T, Tahara S (2005) Suppression of damping-off disease in host plants by the rhizoplane bacterium Lysobacter sp. strain SB-K88 is linked to plant colonization and antibiosis against soilborne Peronosporomycetes. Appl Environ Microbiol 71:3786–3796
Jack ALH (2011) The suppression of plant pathogens by vermicomposts. In: Sherman RL, Arancon NQ, Edwards CA (eds) Vermiculture technology: earthworms, organic wastes, and environmental management. CRC Press, Boca Raton, FL, pp 165–181
Janvier C, Villeneuve F, Alabouvette C, Edel-Hermann V, Mateille T, Steinberg C (2007) Soil health through soil disease suppression: which strategy from descriptors to indicators? Soil Biol Biochem 39:1–23
Kim BS, Lee JY, Hwang BK (2000) In vivo control and in vitro antifungal activity of rhamnolipid B, a glycolipid antibiotic, against Phytophthora capsici and Colletotrichum orbiculare. Pest Manag Sci 56:1029–1035. doi:10.1002/1526-4998(200012)56:12<1029::AID-PS238>3.0.CO;2-Q
Kowalchuk GA, van Os GJ, van Aartrijk J, van Veen JA (2003) Microbial community responses to disease management soil treatments used in flower bulb cultivation. Biol Fertil Soils 37:55–63. doi:10.1007/s00374-002-0561-6
Lam BA, Walton DB, Harris RN (2011) Motile zoospores of Batrachochytrium dendrobatidis move away from antifungal metabolites produced by amphibian skin bacteria. EcoHealth 8:36–45
Lawton KA et al (2008) Analysis of the adult human plasma metabolome. Pharmacogenomics 9:383–397
Lioussanne L, Jolicoeur M, St-Arnaud M (2008) Mycorrhizal colonization with Glomus intraradices and development stage of transformed tomato roots significantly modify the chemotactic response of zoospores of the pathogen Phytophthora nicotianae. Soil Biol Biochem 40:2217–2224. doi:10.1016/j.soilbio.2008.04.013
Litterick AM, Harrier L, Wallace P, Watson CA, Wood M (2004) The role of uncomposted materials, composts, manures, and compost extracts in reducing pest and disease incidence and severity in sustainable temperate agricultural and horticultural crop production - a review. CRC Crit Rev Plant Sci 23:453–479. doi:10.1080/07352680490886815
Liu Y, Zuo S, Zou Y, Wang J, Song W (2012) Investigation on diversity and population succession dynamics of indigenous bacteria of the maize spermosphere. World J Microbiol Biotechnol 28:391–396. doi:10.1007/s11274-011-0822-3
Lundberg DS et al (2012) Defining the core Arabidopsis thaliana root microbiome. Nature 488:86–93. doi:10.1038/Nature11237
Martin FN, Loper JE (1999) Soilborne plant diseases caused by Pythium spp: ecology, epidemiology, and prospects for biological control. CRC Crit Rev Plant Sci 18:111–181. doi:10.1016/s0735-2689(99)00389-5
Mazzola M (2002) Mechanisms of natural soil suppressiveness to soilborne diseases. Antonie Van Leeuwenhoek 81:557–564
McKellar ME, Nelson EB (2003) Compost-induced suppression of Pythium damping-off is mediated by fatty-acid-metabolizing seed-colonizing microbial communities. Appl Environ Microbiol 69:452–460
Mendes R et al (2011) Deciphering the rhizosphere microbiome for disease-suppressive bacteria. Science 332:1097–1100. doi:10.1126/science.1203980
Mondal SN, Kageyama K, Hyakumachi M (1996) Decreased germinability and virulence of oospores of Pythium aphanidermatum in relation to loss of endogenous carbon during incubation in soil. Soil Biol Biochem 28:545–553
Morris BM, Gow NAR (1993) Mechanism of electrotaxis of zoospores of phytopathogenic fungi. Phytopathology 83:877–882. doi:10.1094/Phyto-83-877
Nelson EB (2004) Microbial dynamics and interactions in the spermosphere. Annu Rev Phytopathol 42:271–309
Nelson EB (2006) Rhizosphere regulation of preinfection behavior of oomycete plant pathogens. In: Mukerji KG, Manoharachary C, Singh J (eds) Microbial activity in the Rhizosphere. Springer-Verlag, Berlin, pp 311–341
Ofek M, Hadar Y, Minz D (2011) Colonization of cucumber seeds by bacteria during germination. Environ Microbiol 13:2794–2807. doi:10.1111/j.1462-2920.2011.02551.x
Raaijmakers JM, de Bruijn I, de Kock MJD (2006) Cyclic lipopeptide production by plant-associated Pseudomonas spp.: diversity, activity, biosynthesis, and regulation. Mol Plant-Microbe Interact 19:699–710. doi:10.1094/mpmi-19-0699
Rout ME (2014) The plant microbiome. Adv Bot Res Volume 69:279–309. doi:10.1016/B978-0-12-417163-3.00011-1
Shang HZ, Chen JJ, Handelsman J, Goodman RM (1999) Behavior of Pythium torulosum zoospores during their interaction with tobacco roots and Bacillus cereus. Curr Microbiol 38:199–204. doi:10.1007/pl00006787
Sjogren J, Magnusson J, Broberg A, Schnurer J, Kenne L (2003) Antifungal 3-hydroxy fatty acids from Lactobacillus plantarum MiLAB 14. Appl Environ Microbiol 69:7554–7557. doi:10.1128/aem.69.12.7554-7557.2003
Storey JD, Tibshirani R (2003) Statistical significance for genomewide studies. Proc Natl Acad Sci U S A 100:9440–9445
Thomashow L, Bakker PAHM (2015) Microbial control of root-pathogenic fungi and oomycetes. In: Lugtenberg B (ed) principles of plant-microbe interactions. Springer international publishing, pp 165-173. doi:10.1007/978-3-319-08575-3_18
van Os GJ, van Ginkel JH (2001) Suppression of Pythium root rot in bulbous Iris in relation to biomass and activity of the soil microflora. Soil Biol Biochem 33:1447–1454. doi:10.1016/S0038-0717(01)00053-0
van West P et al (2002) Oomycete plant pathogens use electric fields to target roots. Mol Plant-Microbe Interact 15:790–798
Vandenkoornhuyse P, Quaiser A, Duhamel M, Le Van A, Dufresne A (2015) The importance of the microbiome of the plant holobiont. The New Phytologist
Walker CA, van West P (2007) Zoospore development in the oomycetes. Fungal Biol Rev 21:10–18
Weller DM, Raaijmakers JM, Gardener BB, Thomashow LS (2002) Microbial populations responsible for specific soil suppressiveness to plant pathogens. Annu Rev Phytopathol 40:309–348
Windstam S, Nelson EB (2008a) Differential interference with Pythium ultimum sporangial activation and germination by Enterobacter cloacae in the corn and cucumber spermospheres. Appl Environ Microbiol 74:4285–4291. doi:10.1128/aem.00263-08
Windstam S, Nelson EB (2008b) Temporal release of fatty acids and sugars in the spermosphere: impacts on Enterobacter cloacae-induced biological control. Appl Environ Microbiol 74:4292–4299. doi:10.1128/aem.00264-08
Zhou T, Paulitz TC (1993) In-vitro and in-vivo effects of Pseudomonas spp. on Pythium aphanidermatum zoospore behavior in exudates and on the rhizoplane of bacteria-treated cucumber roots. Phytopathology 83:872–876. doi:10.1094/Phyto-83-872
Acknowledgements
The authors wish to thank Mary Ann Karp, Eric Carr, Monica Minson, Hilary Davis and Lauren Nelson for general technical support. Chemical fractionation of seed exudate samples: Donna Gibson, Bioassay apparatus and seedling photo credits: Kent Loeffler and Claire Smith, statistical consulting: Francoise Vermeylen, qPCR technical support: Eric Markel, manuscript feedback: Emilie Chapelle, Irene de Bruijn, Xu Cheng, Ellen Crocker and the anonymous reviewers for Plant and Soil.
This work was supported by grants from the USDA SBIR [2008-33610-19027 and 2009-33610-20277] as a subcontract to the USDA SBIR principal investigator Thomas Herlihy with matching funds to E. Nelson as principal investigator from NYSTAR CAT (http://www.biotech.cornell.edu/cat); the NY Farm Viability Institute (www.nyfvi.org), the Organic Farming Research Foundation (www.ofrf.org), and USDA NIFA Hatch Funds [NYC-153543]. Additional support was provided to ALH Jack as a scholarship from the Organic Crop Improvement Association (www.ocia.org) and an Andrew W. Mellon fellowship through the Cornell University College of Agriculture and Life Sciences (http://cals.cornell.edu/academics/student-research/graduate-grants-proposal/). Commercial metabolomic analysis was funded by RT Solutions, LLC (www.wormpower.net).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
A. Jack had a consulting contract with RT Solutions, LLC from January 1, 2013 to January 31, 2013. No other conflicts to report.
Additional information
Responsible Editor: Matthieu Barret.
Rights and permissions
About this article
Cite this article
Jack, A.L.H., Nelson, E.B. A seed-recruited microbiome protects developing seedlings from disease by altering homing responses of Pythium aphanidermatum zoospores. Plant Soil 422, 209–222 (2018). https://doi.org/10.1007/s11104-017-3257-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-017-3257-2