Abstract
Background and aims
Soil microbial communities influence nutrient cycling, chemistry and structure of soil, and plant productivity. In turn, agronomic practices such as fertilization and crop rotation alter soil physical and chemical properties and consequently soil microbiomes. Understanding the long-term effects of agronomic practices on soil microbiomes is essential for improving agronomic practices to optimize these microbial communities for agricultural sustainability. We examine the composition and substrate-utilization profiles of microbial communities at the Morrow Plots in Illinois.
Methods
Microbial community composition is assessed with 16S rRNA gene sequencing and subsequent bioinformatic analyses. Community- level substrate utilization is characterized with the BIOLOG EcoPlate.
Results
Fertilizer and rotation treatments significantly affected microbial community structure, while substrate utilization was affected by fertilizer, but not crop-rotation treatments. Differences in relative abundance and occurrence of bacterial taxa found in fertilizer treatments can explain the observed differences in community level substrate utilization.
Conclusion
Long-term fertilization and crop-rotation treatments affect soil microbial community composition and physiology, specifically through chronic nutrient limitation, long-term influx of microbes and organic matter via manure application, as well as through changes in soil chemistry. Relatively greater abundance of Koribacteraceae and Solibacterales taxa in soils might prove useful as indicators of soil degradation.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Long-term agricultural productivity and sustainability relies on positive interactions between three key components with which crops interact – soils, microbial communities in the soils, and agricultural management practices. The soil environment plays a role in selecting the resident microbial community. In turn, microbial communities in soils drive biogeochemical cycles of critical nutrients such as nitrogen and carbon and can have positive or negative interactions with crops. Agricultural management practices affect the chemical and physical characteristics of soils over the long term and can also directly influence the composition of the microbial communities in these soils. Agricultural soils are a critical but constrained resource that is under increasing pressure due to climate change and population growth, as well as susceptible to deterioration through suboptimal agronomic management practices (Doran and Smith 1987; Greenland 1981; Tilman et al. 2002). Agronomic practices, including fertilizer treatments and crop rotations, can positively or negatively affect properties of soils that influence agricultural productivity (Angers et al. 1993a; Aref and Wander 1997; Dick 1992; Havlin et al. 1990; Khan et al. 2007; Lupwayi et al. 1998; Mitchell et al. 1991; Mulvaney et al. 2009; Six et al. 2006; van Diepeningen et al. 2006). In contrast to processes like soil erosion and compaction that can occur rapidly, changes in the chemical properties of soils (e.g. organic matter content, capacity to retain moisture, aggregate structure, etc.), often manifest themselves far more slowly – on the order of decades – even under sharply contrasting management practices (Aref and Wander 1997; Darmody and Norton 1993; Khan et al. 2007; Mitchell et al. 1991; Mulvaney et al. 2009; Nafziger and Dunker 2011). While changes in the amount and chemical forms of key plant nutrients as well as pH can be altered by fertilization in the short term without causing substantive changes in the soils’ chemical and physical properties, these properties can change with agronomic management over decades (Rivard et al. 2015; Velde and Peck 2002). Therefore, it is likely that the feedback effects between the soil microbiome and soil chemistry also manifest on similarly long time scales. Well-documented and consistently maintained long-term agricultural research plots can provide important information about the long-term effects of agronomic practices on agricultural soils (Enwall et al. 2007; Hallin et al. 2009; Neumann et al. 2013; Powlson et al. 2014; Widmer et al. 2006). The Morrow Plots at University of Illinois were established in 1876 with continuous-corn treatment and unfertilized treatments that date back to establishment. The current suite of modern fertilizer and crop-rotation treatment combinations have been in place since 1968. Therefore, the Morrow Plots are a unique and valuable resource for investigating the long-term effects of agronomic management on soil microbiomes and soil quality.
Microbial communities exhibit immense diversity in terms of structure (i.e. community composition) as well as function (i.e. physiology of all species combined) that vary from place to place as well as over time, but the interactions between microbial communities in the soil and their environment remain poorly understood (Giller et al. 1997; Horner-Devine et al. 2004; McGuire and Treseder 2010). This is in part due to the fact that the vast majority of the highly diverse microbes in soils cannot be cultured and characterized individually (Kirk et al. 2004). As complex microbial communities in soils are a critical component of the agricultural ecosystem, we must improve our understanding of how they interact with agronomic practices, soil, crops, and the environment. This knowledge is critical because microbial communities drive the biogeochemical processes that affect nutrient and organic matter composition of the soil (Balser and Firestone 2005; Jackson et al. 2003; Nannipieri et al. 2002) as well as the soils’ environmental functions. Improved knowledge of microbial communities could help reduce nitrate runoff and emission of greenhouse gases (Cavigelli and Robertson 2000; Kramer et al. 2006). Characterizing the structure and functions of microbial communities from long-term agronomic research plots can provide insights into the mechanisms underlying the effects of agronomic practices on soil properties (Chu et al. 2007b; Jackson et al. 2003; Neumann et al. 2013; Wessén et al. 2010b) and may also lead to development of indicators of soil health, environmental function, and agricultural productivity (Franchini et al. 2007; Mummey et al. 2002; Otto et al. 2005; Pascual et al. 2000; Powlson et al. 1987).
Fertilizer treatments affect microbial communities through direct influence on soil nutrient content and chemistry (Peacock et al. 2001; Zhong et al. 2010). Crop rotations can affect soil microbial communities through differences in plant-microbe interactions (Larkin 2008; Navarro-Noya et al. 2013). Both fertilizer treatments and crop-rotations also influence soil microbial communities through differences in the quantity and composition of root exudates (Baudoin et al. 2003; el Zahar Haichar et al. 2008), rhizodeposits and above-ground crop residues (Bending et al. 2002; Benizri et al. 2002; Butler et al. 2003; Miller et al. 2008; Navarro-Noya et al. 2013; Nelson and Mele 2006; Paterson et al. 2007). While modern techniques have been employed in numerous studies to understand the effects of medium to long-term (10 years or more) fertilizer treatments on soil microbiota (Börjesson et al. 2012; Chu et al. 2007a; Clark and Hirsch 2008; Hirsch et al. 2010; Kamaa et al. 2011; Neumann et al. 2013; Ogilvie et al. 2008; Wessén et al. 2010a; Wessén et al. 2010b; Widmer et al. 2006; Wu et al. 2011), the effect of medium-term or long-term combined fertilizer and crop-rotations treatments on microbial communities appears to have been studied with modern methods in very few instances (Navarro-Noya et al. 2013; Souza et al. 2015; Xuan et al. 2012; Yin et al. 2010) primarily because long-term experiments with crop-rotation and fertilizer treatment combinations are extremely rare. To improve agronomic practices for enhancing long-term sustainability, it is therefore essential to be able to predict the potential long-term effects of existing and improved practices on agricultural soils and their microbial ecosystems. Extant, well-maintained long-term field-experiments with a range of crop-rotation and fertilizer treatments are therefore a valuable resource for building our understanding of long-term effects of agronomic management.
In this study, we characterize the composition, diversity, and richness as well as community-level substrate utilization of soil microbiomes under contrasting fertilizer and crop-rotation treatments. While chemical and structural characteristics of arable soils under long-term management treatments have been previously characterized (Aref and Wander 1997; Darmody and Norton 1993; Khan et al. 2007; Mitchell et al. 1991; Mulvaney et al. 2009; Nafziger and Dunker 2011), understanding the long-term effects of agricultural management on the structure and diversity in microbial community function is a critical knowledge gap. Characterizing soil microbial communities under contrasting management practices will enable us to discover the drivers of differences between the microbial communities as well as the potential long-term effects of divergent microbial communities on the productivity and sustainability of agricultural soils.
Materials and methods
Research site
The Morrow Plots, located on the campus of the University of Illinois at Urbana-Champaign, are the oldest continuously maintained agricultural research plots in the United States. The plots were established in 1876 with the intention of conducting experiments useful to farmers. While the plots have generated valuable crop-yields and soil chemistry data for over 130 years, microbial communities in the plots have not yet been systematically examined. The fertilizer and crop rotations have changed gradually since establishment of the plots. The current basic rotations and fertility treatments have been in place since 1968; small modifications have been made to tillage (with conversion from moldboard plowing to chisel plowing), planting density, crop variety, and herbicide choice, during that ~50 year period. Briefly, the current experiment (Fig. S1) consists of three blocks of crop rotation treatments – one block each of continuous corn (Zea mays) (C), a two-year corn and soybean rotation (CS), and a three-year corn-oats-alfalfa rotation (COA), with an 8 ft margin between blocks. It is important to note that the continuous corn and unfertilized treatments date back to 1876, making these the oldest continuously managed agricultural research plots in the western hemisphere. Even though the rotation blocks are not replicated, they are one of the best resource available for understanding the effects of very long-term agronomic treatments on soils and the microbial communities in them. Each crop rotation block is split into eight plots comprising replicated fertilizer treatments: unfertilized (UF); inorganic fertilizers (IN) with nitrogen (as urea), phosphorus (as P2O5), potassium (as K2O), and limestone; and organic fertilization (OR) with cattle manure, limestone, and phosphorus. Corn yields from fertilized plots and from blocks with crop rotations have been consistently higher (Fig. S2). For a comprehensive description of the treatment history, site conditions, and yields, please refer to the description by Aref and Wander (Aref and Wander 1997) and Odell et al. (Odell et al. 1982).
Sample collection and processing
Soil samples were collected on a single day late in the growing season in 2009 – a year when all the plots were planted in corn. Sampling during the all-corn year controlled for transient effects of plant-microbe interactions from the different crops, enabling the investigation of long-term effects of crop-rotation on the microbial communities. Each sample consisted of five cores (1.9 cm dia × 12 cm deep). All cores were collected at least 3 m from the edge of the plot in order to minimize edge effects. The samples represent bulk soil, not rhizosphere, as the cores were collected in the planting rows, but away from the root zone. The cores from each plot were placed in sealed plastic bags on ice while in the field and transported back to the lab and processed within two hours of collection. The cores from each plot were composited and homogenized with a 2 mm sieve. Subsamples from each the composited, homogenized soil sample were processed as appropriate for chemical analyses, community level physiological profiling with BIOLOG, and DNA extraction for bacterial community bioinformatic analyses as described below.
Microbial community physiological profiling
Substrate utilization-based fingerprints of the soil microbial communities were generated with the BIOLOG assay using EcoPlates™ (Biolog®, Hayward, CA, USA) – a 96 well plate with three replicates of 31 individual substrates and one control (no substrate). A suspension of the soil microbial community was created by vortexing 0.5 g fresh soil with glass beads in 2 mL sterile PBS for 2 min and then centrifuging the mixture at 4 °C and 750 x g for 6 min to precipitate large soil particles and debris. The supernatant was diluted 1:25 with sterile PBS and 150 μL of the diluted supernatant was added to each well of the EcoPlate™. The plates were incubated in the dark at room temperature. Metabolic response of the community to available substrate in each well was measured by recording the absorbance at 590 nm daily for 6 days.
DNA extraction
DNA from the microbial community of the Morrow Plots soil samples was extracted using FastDNA for Soil kit from MP Biomedicals (Solon, OH, USA). The extracted DNA was purified using CTAB (cetyl trimethyl ammonium bromide) purification process to remove humic contaminants (Sambrook et al. 1989). Extracted DNA was stored at −20 °C until used in DNA-based analyses described below.
Bacterial community 16S rRNA gene sequencing
Bacterial community composition was assessed by sequencing the V4 – V5 region of 16S rRNA gene using the PCR primers 515F and 926R(Baker et al. 2003; Dojka et al. 1998; Haas et al. 2011). Primers were modified with adapter sequences required for Illumina sequencing (Caporaso et al. 2011) and a unique dual-index barcode was assigned to each sample (Hamady et al. 2008). To prepare the templates for sequencing, 50 μl PCR reactions with following composition were conducted: 25uL 2X KAPA HiFi HotStart ReadyMix (KAPA Biosystems, Woburn, MA, USA), 500 μM of each primer and 50 ng DNA. Thermal cycling was carried out with an initial denaturation at 98 °C for 45 s. followed by 25 cycles of 98 °C for 15 s., 65 °C for 30 s., 72 °C for 30 s., and ended with a final extension step at 72 °C for 2 min. Amplicons from each sample were cleaned with 0.8 X volume of AMPure® XP beads (Agencourt Bioscience, Beverly, MA, USA) and then quantified on the Qubit® Fluorometer (Invitrogen-Molecular Probes, Eugene, OR, USA) using the Quant-iT™ dsDNA HS Assay kit (Invitrogen, Carlsbad, CA, USA). Equal concentrations of amplicons from each sample were pooled for sequencing. The amplicon pool was sequenced at W. M. Keck Center for Comparative and Functional Genomics at the University of Illinois at Urbana-Champaign (Urbana, IL, USA) on Illumina MiSeq platform with a 2 × 250 bp reads configuration using the MiSeq V2 Nano flowcell (Illumina, San Diego, CA, USA).
Each sequence was assigned to its original sample according to the barcode. After de-multiplexing, the forward and reverse read of each paired-end sequence were merged using software FLASH (Fast Length Adjustment of SHort reads) (Magoč and Salzberg 2011). Sequences with more than 10 % bases with quality score lower than 30 and sequences containing ambiguous bases “N″ were removed from downstream processing. Filtered sequences were clustered into operational taxonomic units (OTUs) using USEARCH (Edgar 2010). USEARCH was used to (1) de-replicate sequences and remove singletons to reduce the data size and calculating time; (2) remove chimeras contained in the sequences using GOLD (Reddy et al. 2014) as a reference database (http://drive5.com/uchime/uchime_download.html); (3) form clusters of 97 % identity sequences and represent each OTU by consensus sequences (representative sequences). The cluster file was converted into an OTU table using a customized script derived from QIIME (Kuczynski et al. 2012). These consensus sequences were used as representative sequences in each OTU and the taxonomic attribution of filtered sequences was assigned in QIIME with uclust using 97 % similarity score and 51 % consensus against the August 2013 Greengenes database. Fewer than 1 % of sequences across all samples received no taxonomic assignments and were removed from further analyses. The OTU table was subsampled to compensate for differences in total copies sequenced between samples. The OTU table was rarefied to equal abundance (6800 reads/sample) for further analyses. The sample from one of the OR plots from the COA block exhibited very low reads and was excluded from further analyses.
Chemical analysis
Soil chemical parameters, including pH, total nitrogen, NO3-nitrogen, NH4-nitrogen, total carbon, and total organic matter were quantified at the Iowa State Soil and Plant Analysis Laboratory (Ames, IA). Soil moisture was quantified using gravimetric soil moisture analysis.
Data analysis
Data were analyzed using multivariate correlational and ordination methods in the R statistical environment (R Core Team 2016), using R packages vegan (version 2.0–10) (Oksanen et al. 2015) and labdsv (version 1.6–1) (Roberts 2012).
The effect size and statistical significance of rotation and fertility treatments on each of the experimental analyses were determined with Permutational Multivariate Analysis of Variance (PerMANOVA, function adonis) (Anderson 2005). In addition, since fertilization and crop-rotation can affect microbial communities independently, treatment effects on microbial communities were visualized with partial ordination analyses that controlled for the effect of the other treatment category - i.e. the effect of rotation treatments was visualized while controlling for the effect of fertility treatments and vice versa. Differences in community structure (16S rRNA gene sequencing) were also visualized with partial correspondence analysis (pCA) (Legendre and Legendre 2012). Where necessary, the ranges of the ordination axes were manually set to exclude an extreme outlier. Correlation of soil chemical parameters with each of the ordinations was quantified with the function envfit and plotted on the pCA plots.
The taxonomic information obtained from 16S rRNA gene sequencing was used to quantify Shannon-Weaver diversity index (Hill et al. 2003) for each community (function diversity) and to estimate community richness (function estimateR) with Abundance-based coverage estimate (ACE) and the chao-1 estimate (Chao et al. 2005).
Indicator Value Analysis (function indval in R package labdsv) (Dufrêne and Legendre 1997) was employed to identify microbial taxa that were indicators of a specific group of samples, and hence, the treatments that these samples represented. Indicator value analysis assesses differences in relative abundance and frequency of presence of microbial OTUs between sets of samples. We focused this analysis on the high-abundance, or top 19.24 % (586 out of 3045) of the operational taxonomic units with assigned taxonomy, which constituted 81.53 % of all fragment reads. Each of these OTUs comprised a total of at least 50 sequence reads across the rarified (i.e. equal-abundance) sample pool. Indicator value analysis was iterated 10,000 times and OTUs that cleared the significance threshold of 0.05 were accepted as indicators of a treatment. OTUs were assessed for indicator status of one of the fertilizer treatments, controlling for the effects of crop-rotation treatments and vice versa. Indicator value analysis was primarily conducted for higher abundance OTUs for two reasons. First, because high-abundance OTUs are likely to be responsible for the bulk of the functional attributes of microbial communities, examining abundance and frequency differences among higher abundance OTUs is more relevant than those of rarer taxa for investigating functional differences between microbial communities. Second, indicator value analysis can overestimate the indicator status of lower abundance OTUs since their low total reads across all samples heightens the probability of large differences in abundance between sample classes. However, in order to examine the effects of agronomic treatments on rare taxa we also present indicator value analysis of all OTUs in the supplementary materials (Fig. S3, Tables S6 and S7).
Differences between substrate consumption profiles (BIOLOG assay) of the microbial communities were visualized using partial Principal Components Analysis (PCA) (Härdle and Simar 2012). Where necessary, the ranges of the ordination axes were manually set to exclude an extreme outlier. Average well-color development (AWCD) from the BIOLOG assay was calculated for all substrates as well as for the subsets of substrates containing nitrogen and/or phosphorus following conventional procedures (Garland and Mills 1991). This allowed us to test the availability of carbon, nitrogen, and phosphorus as drivers of community-level microbial physiology, i.e. consumption of all substrates, or substrates containing nitrogen, or substrates containing nitrogen and phosphorus.
Similarities between community diversity, community richness, and BIOLOG AWCD data from pairs of treatments were examined using Tukey’s Honest Significant Difference test to correct for multiple comparisons. A significance threshold of p ≤ 0.05 was used to determine significance for all analyses.
Results
Bacterial community composition
3045 bacterial OTUs with two or more 16S rRNA gene reads were assigned taxonomic classification, amounting to a total of over 156,000 reads. The broad metrics of total-community diversity and richness, with adjustments for multiple comparisons, of the communities were generally not significantly different (p ≤ 0.05), with only the communities under OR treatment being richer (ACE) than those under IN among fertilizer treatments (Fig. 1). Shannon diversity and chao1 richness estimates were not significantly different between communities under either the fertilizer or rotation treatments. (Table S1).
Bacterial communities from OR plots exhibit significantly greater diversity (left panel) compared to communities from IN (p = 0.028) or UF (p = 0.052) plots as well as richness (right panel) compared to communities from IN (p = 0.016) or UF (p = 0.048) plots. Differences in community diversity or richness between the crop-rotation treatments were not statistically significant. Box-plots are plotted with the usual convention - with boxes demarcating the middle two quartiles and filled circles denoting the median datum. Whiskers extend to the farthest data-points still within 1.5 times inter-quartile range (IQR). Outliers beyond 1.5 IQR are shown as empty circles. (IN: inorganic fertilizer treatment, OR: manure fertilizer treatment, UF: unfertilized treatment)
However, microbial community structures inferred from 16S rRNA gene sequences were significantly dissimilar between the rotation treatments as well as fertility treatments, with PerMANOVA R2 = 0.27, p < 0.0001 for comparisons among fertility treatments, and R2 = 0.16, p = 0.0007 for comparisons among rotation treatments. The interaction of rotation and fertilizer treatments also significantly affected community composition (R2 = 0.17, p = 0.04). Partial correspondence analysis clearly demonstrates that communities obtained from the same treatment are similar to each other and distinct from communities under other treatments (Fig. 2) reinforcing the finding based on community diversity metrics that long-term fertilizer and crop-rotation treatments are associated with distinct microbial communities.
Partial correspondence analysis ordinations of the microbial community diversity as revealed by 16S rRNA gene sequencing. Community structure was affected by both fertilizer (left panel, PerMANOVA R2 = 0.27, p < 0.0001) controlling for the effect of rotation and by rotation (right panel, R2 = 0.16, p = 0.0007) controlling for the effect of fertilizer treatments. Soil chemical parameters significantly correlated with each treatment are plotted on each ordination including pH, percent carbon (% C), total percent nitrogen (% N), and parts per million nitrates (μg/g NO3 N). Greater values of soil chemical parameters are significantly correlated with longer rotation and organic amendment. Ellipses circumscribe the 99 % confidence limit around the centroid of each treatment group. In the left panel, CA 2 range is set manually to exclude an extreme outlier UF sample, while in the right panel, CA 1 range is set manually to exclude an extreme outlier C sample. (IN: inorganic fertilizer treatment, OR: manure fertilizer treatment, UF: unfertilized treatment, COA: corn-oats-alfalfa rotation, CS: corn-soybean rotation, C: continuous corn rotation)
Indicator value of OTUs by treatments
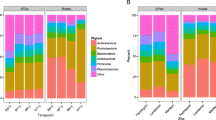
Among high abundance microbial OTUs, differences in relative abundance and frequency of microbial taxa were observed to a greater extent among the fertilizer treatments (a total of 179 OTUs, with 74 in UF, 63 in OR, and 42 in IN) than among crop-rotation treatments (a total of 75 OTUs, with 32 in COA, 30 in C, and 13 in CS) (Fig. 3). Between fertilizer treatments, more OTUs belonging to Actinobacteria and Planctomycetes were indicators of UF treatment than OR or IN treatments, while fewer Acidobacteria taxa were indicators of IN treatment compared to UF and OR treatments. Among crop-rotation treatments, more taxa belonging to Actinobacteria and Proteobacteria were indicators of communities under COA rotation, while a greater number of taxa belonging to Acidobacteria and Bacteroidetes were indicators of communities in the continuous corn (C) treatment.
Abundant (top 80 % of sequenced 16S rRNA gene fragment reads) bacterial OTUs that are indicators of each treatment, binned together by phylum. The greatest number of OTUs were significant indicators of UF communities, followed by OR communities and IN communities. (Figure S3 displays indicator taxa across all OTUs.) (IN: inorganic fertilizer treatment, OR: manure fertilizer treatment, UF: unfertilized treatment, COA: corn-oats-alfalfa rotation, CS: corn-soybean rotation, C: continuous corn rotation)
Groups of taxonomically related and functionally notable OTUs that were found to be indicators of the treatments are described below and listed in Tables S4-S7. The OTUs known through sequencing and culture-based studies to be adapted to low-nutrient conditions (Eichorst et al. 2011; Kanokratana et al. 2010; Ward et al. 2009), including eleven OTUs belonging to the family Koribacteraceae, and three OTUs of the genus Candidatus Koribacter, were indicators of UF communities and were also the most highly abundant OTUs in these communities. In addition, five OTUs of the order Solibacterales, including two OTUs belonging to the genus Candidatus Solibacter were indicators of UF communities. All of these bacteria belonged to the Acidobacteria phylum.
The OTUs of the order Syntrophobacterales, which are obligate anaerobic chemolithotrophs (Kuever 2014), were indicators of OR communities (3 OTUs), UF communities (2 OTUs) and COA communities (1 OTU).
Five OTUs of the order Xanthomonadales, including one of the genus Lysobacter, were indicators of OR communities, IN communities (2 OTUs) and UF and CS communities (1 OTU each).
When low-abundance taxa – i.e. the bottom 80 % of OTUs (2459 out of 3045) that constituted only 20 % of sequencing reads – are included in the analysis, the general trends described above for the abundant OTUs persist - with the exception of far more OTUs being indicators of OR communities (310 OTUs for OR communities, compared to 162 of UF communities) (Fig. S3). Conducting indicator value analysis for all taxa highlights the unique rare taxa in the communities of OR plots. In all, 310 OTUs were found to be indicators of OR treatment, 162 of UF treatment, and 69 of IN treatment. Relatively fewer were found to be indicators of each of the crop-rotation treatments, with 98 of COA, 76 of C, and 35 of CS.
Community level physiological profiles
The ability of the microbial communities to utilize substrates, as assessed by BIOLOG substrate utilization assays, was affected by fertility treatments but not by crop rotation treatments. PerMANOVA analysis showed a stronger and statistically significant effect from fertility treatments on substrate utilization (R2 = 0.20, p = 0.0002) while the effect of rotation treatments was not statistically significant (R2 = 0.10, p = 0.13). The interaction of rotation and fertility treatments was also statistically non-significant (R2 = 0.12, p = 0.76). The dissimilarity in physiological profiles between unfertilized treatments compared to the organic and inorganic fertility treatments is notable and apparent from the PCA ordination (Fig. 4).
Partial principle components analysis ordinations of BIOLOG profiles. Substrate consumption activity of communities was affected by fertilizer treatments (controlling for effect of rotation) (left panel, PerMANOVA R2 = 0.20, p = 0.0002) with communities from UF plots substantially dissimilar to communities from OR and IN plots. Rotation treatments (controlling for the effect of fertilizer treatments) did not significantly affect substrate consumption profiles (right panel, R2 = 0.10, p = 0.13). Soil chemical parameters significantly correlated with each treatment are plotted on each ordination including pH and percent soil moisture. Ellipses circumscribe the 99 % confidence limit around the centroid of each treatment group. (IN: inorganic fertilizer treatment, OR: manure fertilizer treatment, UF: unfertilized treatment, COA: corn-oats-alfalfa rotation, CS: corn-soybean rotation, C: continuous corn rotation)
Utilization of nitrogen-containing substrates varied among fertilizer treatments to a greater extent than utilization of the complete substrate panel (Fig. S4). Communities from unfertilized treatments and inorganic fertility treatments differed the most, though the effect was less than statistically significant (AWCD 1.02 vs. 0.94; p = 0.16). Similarly, utilization of nitrogen-containing substrates differed somewhat among communities from unfertilized and organic fertilized plots (AWCD 1.02 vs. 0.94; p = 0.23). Notably, communities from inorganic and organic fertilizer treatment plots showed statistically similar (p = 0.99) consumption of nitrogen-containing substrates (Table S2).
Soil chemical parameters
The majority of soil fertility parameters were highest in manure-amended plots and/or under the longest rotation (Corn-Oats-Alfalfa). Total carbon, total nitrogen, organic matter content, and soil moisture were affected by both rotation and fertilizer treatments, with highest values under OR and COA treatments (Table S3). Soil pH and ammonia-N content were significantly affected by fertilizer treatments but not by rotation treatments. Soil pH was highest in plots under organic fertilizer treatment and lowest in unfertilized plots. Soil ammonium was highest in plots receiving inorganic fertilizer and lowest in plots receiving organic fertilizer. Soil nitrate content was significantly affected by rotation treatments, being highest in plots under the corn-oats-alfalfa rotation, and lowest in plots under continuous-corn rotation. This likely reflects differences in soil moisture content, which follow the same order and are a by-product structural enhancement associated with soil organic carbon. Soil nitrate content, which was extremely low (~3.7 μg/g), did not differ among fertilizer treatments, which is not unexpected given that sampling was carried out late in the growing season and the soil was quite dry (gravimetric moisture content ~22 %) so nitrification was suppressed (NH4 values were ~4.3 μg/g).
Discussion
The results presented above demonstrate that the structure and function of microbial communities can be differentially affected by fertilizer and rotation treatments. Long-term fertilizer treatments are correlated with differences in both composition and community-level physiological profile of soil microbial communities. On the other hand, while long-term crop rotation treatments are correlated with soil microbial communities that were structurally distinct from each other, the community-level dissimilarities arising from rotation treatments did not translate to significant differences in their substrate-utilization profiles.
While the effects of long-term fertilizer treatments on bacterial community substrate-induced respiration and structure have been elucidated previously (Enwall et al. 2007; Widmer et al. 2006), the effect of crop-rotation treatments on community structure, but not on substrate utilization, is one of the notable results of the present study. This difference between the effect of crop-rotations on community structure and substrate utilization profiles suggests that functional redundancy plays a role in how management practices affect community structure and function in agricultural soil microbiomes. Microbial functions related to substrate utilization may be redundant across the taxa whose relative abundance is affected by crop-rotations. Conversely, between fertilizer treatments, microbial functional diversity related to substrate-utilization may play a greater role in shaping microbial community structure. Indicator value analysis of microbial OTUs, which assesses the differential relative abundance and frequency of presence of microbial taxa between communities, offers a plausible mechanism for this disconnect between microbial community structure and physiology. At the Morrow Plots, the long-term chronic nutrient deficit in the UF plots appears to have resulted in an enrichment of taxa adapted for low-nutrient conditions in microbial communities of these plots. These taxa are capable of acquiring and utilizing available substrates more effectively (Eichorst et al. 2011; Kanokratana et al. 2010; Ward et al. 2009), and are less abundant in communities from plots that received inorganic or manure fertilizer. The oligotrophic taxa may be particularly efficient utilizers of nitrogen-containing substrates, as reflected in the somewhat greater AWCD of N-containing substrates for UF communities compared to for IN and OR communities (Fig. S3, Table S2).
These differences in the abundance as well as physiology of key taxa between communities may have manifested in the observed differences in community-level physiological profiles of the microbial communities under the fertilizer treatments. Conversely, taxa adapted to nutrient-depletion exhibit smaller differences between crop-rotation treatments. The smaller contribution of oligotrophic taxa to the differences in community structure under crop-rotation treatments compared to communities under fertilizer treatments is congruent with the distinct substrate utilization profile of UF communities compared to IN and OR communities (Fig. 4 , left panel) and the not-statistically distinct substrate utilization profiles of the C, CS, and COA communities. It is also worth noting that communities associated with the continuous-Corn (C) rotation have a substantial number of oligotrophic indicator taxa (Tables S5 and S7) and their substrate utilization profiles appears somewhat distinct from those of CS and COA communities (Fig. 4 , right panel).
OTUs adapted for nutrient depleted conditions
One of the strongest patterns of indicator species observed was that OTUs adapted to nutrient-depleted conditions were indicators of communities from low-nutrient plots. Strains of Candidatus Koribacter and Candidatus Solibacter have been characterized with genomic sequencing, culture-based studies, and community sequencing, and are known to be adapted for low nutrient conditions (Eichorst et al. 2011; Kanokratana et al. 2010; Ward et al. 2009). The low-nutrient adaptations of these taxa is congruent with their greater relative abundance in UF communities which are subjected to low organic-matter, carbon, and nitrogen content of the UF soils. The greater relative abundance of these OTUs in UF communities may also have contributed to the distinct BIOLOG activity of the UF communities relative to IN and OR communities. These taxa happen to belong to the phylum Acidobacteria.
Koribacteraceae and Solibacterales OTUs were also indicators of communities under the continuous-Corn treatment suggesting that regardless of the fertilizer treatment, the high nutrient requirement of corn depletes soil nutrients, as is also evident from the soil chemistry measurements presented. The higher relative abundance of taxa adapted to nutrient-depleted environment in unfertilized and continuous-Corn treatments reinforces previous observations that these practices degrade soil quality (Havlin et al. 1990; Karlen et al. 2006; Ketcheson 1980).
Finally, Solibacterales OTUs were found to be indicators of IN communities, suggesting that while inorganic fertilizers provide sufficient plant nutrients to enable high agricultural yields, they may promote conditions that result in deterioration of the soil over the long-term. This is supported by the observed soil chemistry of IN soils – which had lower organic-matter contents and pH than OR plots but values were similar to those of UF plots. The indicator status of such taxa in the synthetically fertilized IN plots suggests that conventional fertilizer practices should be carefully managed to reduce deleterious effects on arable soils as previously suggested by others (Khan et al. 2007; Mulvaney et al. 2009).
In contrast, no Koribacteraceae or Solibacterales OTUs or other OTUs with known nutrient depletion adaptations were indicators of OR, CS, and COA communities, in agreement with the generally higher values of soil chemical parameters measured from these plots. The lack of competitive advantage to oligotrophic taxa in these plots is also expected from studies that have shown improved soil quality from manure fertilization and crop rotation treatments (Edmeades 2003; Karlen et al. 2006; Manna et al. 2005). The greater relative abundance of Koribacteraceae or Solibacterales OTUs in UF and IN soils compared to OF soils containing higher OM levels suggests that the relative proportion and abundance of these and other functionally similar taxa might be used as markers for soil deterioration and soil quality.
OTUs that are obligate anaerobes
Syntrophobacterales taxa, which are obligate anaerobic chemolithotrophs (Kuever 2014), were indicators of OR and COA communities - suggesting a greater prevalence of anaerobic microsites in the treatments. Anaerobic sites are known to occur more frequently in larger soil aggregates (Sexstone et al. 1985) which have been previously characterized at the Morrow Plots in the CS and COA plots (Darmody and Norton 1993). Larger soil aggregates with greater water-stability and erosion-resistance are also known to be associated with greater soil organic matter content (Tisdall and Oades 1982) and conservation agriculture practices such as no-till, manure application, and crop-rotation (Angers et al., 1993b; Beare et al. 1994; Mikha and Rice 2004; Wortmann and Shapiro 2008). Obligate anaerobes may therefore be useful biomarkers of beneficial soil structures in agricultural soils.
Disease suppressive OTUs
Xanthomonadales OTUs, including one of the genus Lysobacter, which are known to be are abundant in disease suppressive soils (Islam et al. 2005; Mendes et al. 2011; Postma et al. 2010; Rosenzweig et al. 2012) were found to be indicators of OR communities. While many Xanthomonadales taxa are known to be plant pathogens, phenology and yield data at the Morrow Plots provide no indication that the prevalent Xanthomonadales taxa in the Morrow Plots were interacting pathogenically with the plants. This suggests that manure application may increase the abundance of potential plant-beneficial bacteria in agricultural soils.
Indicator taxa including rare OTUs
While the discussion above has focused on relative enrichment among higher abundance OTUs, it is worth noting that the OR communities contain a highly distinctive rare microbiome. This heightened diversity in the community more than four months after manure application indicates that the presence of these taxa is not a short-term effect of inputs, but reflects a long-term shift in the community structure. This shift may be driven by the greater diversity of niches provided by recalcitrant substrates introduced via the manure compared to synthetic fertilizers, as well as the influx of microbial taxa through the manure (Doan et al. 2014; Nair and Ngouajio 2012; Zhen et al. 2014). The distinctive and higher-diversity rare-microbiome in OR communities as well as their overall greater richness and diversity may confer benefits on the microbial communities in soils receiving manure-based amendment, including greater resistance to disturbances as well as functional stability and resilience (Allison and Martiny 2008; Girvan et al. 2005; Griffiths and Philippot 2013; Lehman et al. 2015; Shade et al. 2014).
The differential abundance of OTUs with known functional or habitat characteristics among contrasting soil environments and treatments thus appear to have the potential to serve as microbial biomarkers of soil health, nutrient cycling, and agricultural productivity in conjunction with differences in soil-chemistry as well as management history.
Effects of crop rotation on community structure
The differences in enriched OTUs between the higher-diversity COA rotation and the continuous corn rotation indicates that soil microbial community structure retains certain legacy effects from the long-term crop rotation treatment. The relatively fewer indicator OTUs between crop-rotations compared to fertilizer treatments may be in part due to the all-corn sampling year, which presented a uniform rhizodeposit and exudate environment for the microbial community. The lack of statistically significant differences in the diversity of the microbial community subjected to the different crop-rotation treatments at the Morrow Plots is in contrast to the finding by Lupwayi et al. (1998) of greater diversity in communities under diversified crop-rotations compared to continuous cropping of wheat. While the substantial differences between these two experiments complicate direct comparison of the two data sets, it may also be true that the effects of crop-rotation treatments on soil microbial communities reported here may not be easily generalizable, perhaps in part due to the legacy design of the Morrow Plots experiment. Additionally, comparing the crop-rotations at the Morrow Plots necessitates the integration of nutrient-exhausted and fully-fertilized treatments within each rotation. These fertilization treatments each have significantly different plant residue returns to the soil, which are as notable as differences in carbon flows resulting from the different cropping regimes. By contrast, while the experimental plots studied by Lupwayi et al. are better replicated, they were sampled just three to four years post-establishment and the fertility treatments did not include a no-fertilization treatment.
Conclusions
Long-term fertilizer and crop-rotation treatments have led to differences in structure and function of the soil microbial communities at the Morrow Plots, and are also reflected in differential abundance of taxa adapted to conditions generated by the specific treatments. The greater community level substrate utilization in unfertilized communities compared to those subjected to either inorganic or manure fertilizers suggests that chronic nutrient limitation is a more significant force than type of fertilizer in determining the substrate consumption activity of microbial communities. However, higher-diversity rare microbiome, and greater relative abundance of potential plant-beneficial bacteria in manure-fertilized soils suggests that the long-term influx of organic matter and microbial biodiversity through manure application may serve to enhance beneficial microbial diversity in agricultural soils in addition to improved soil fertility parameters. A more complete understanding of the long-term interactions and feedback between agronomic practices, agricultural soil, and structure and function of microbial communities derived from and building on these results is essential for enabling the design of optimal agronomic practices that improve agricultural sustainability. Finally, it is important to note that the differences in community structure were not broadly reflected in metrics such as Shannon diversity and community richness estimation, suggesting that these metrics are not always sufficient for understanding the differences between microbial communities. Conversely, the use of indicator taxa as biomarkers may have the potential to enable assessment of wide variety of soil characteristics, agricultural productivity potential, and environmental impacts of agricultural soils with a single and increasing affordable modern analytical tool.
References
Allison SD, Martiny JBH (2008) Resistance, resilience, and redundancy in microbial communities. Proc Natl Acad Sci 105:11512–11519. doi:10.1073/pnas.0801925105
Anderson MJ (2005) Permutational multivariate analysis of variance. University of Auckland, Auckland, Department of Statistics
Angers D, Bissonnette N, Legere A, Samson N (1993a) Microbial and biochemical changes induced by rotation and tillage in a soil under barley production. Can J Soil Sci 73:39–50
Aref S, Wander MM (1997) Long-term trends of corn yield and soil organic matter in different crop sequences and soil fertility treatments on the morrow plots. Adv Agron 62:153–197
Baker GC, Smith JJ, Cowan DA (2003) Review and re-analysis of domain-specific 16S primers. J Microbiol Methods 55:541–555
Balser TC, Firestone MK (2005) Linking microbial community composition and soil processes in a California annual grassland and mixed-conifer forest. Biogeochemistry 73:395–415
Baudoin E, Benizri E, Guckert A (2003) Impact of artificial root exudates on the bacterial community structure in bulk soil and maize rhizosphere. Soil Biol Biochem 35:1183–1192
Beare MH, Hendrix PF, Coleman DC (1994) Water-stable aggregates and organic matter fractions in conventional- and no-tillage soils. Soil Sci Soc Am J 58:777. doi:10.2136/sssaj1994.03615995005800030020x
Bending GD, Turner MK, Jones JE (2002) Interactions between crop residue and soil organic matter quality and the functional diversity of soil microbial communities. Soil Biol Biochem 34:1073–1082
Benizri E, Dedourge O, Dibattista-Leboeuf C, Piutti S, Nguyen C, Guckert A (2002) Effect of maize rhizodeposits on soil microbial community structure. Appl Soil Ecol 21:261–265
Börjesson G, Menichetti L, Kirchmann H, Kätterer T (2012) Soil microbial community structure affected by 53 years of nitrogen fertilisation and different organic amendments. Biol Fertil Soils 48:245–257. doi:10.1007/s00374-011-0623-8
Butler JL, Williams MA, Bottomley PJ, Myrold DD (2003) Microbial community dynamics associated with rhizosphere carbon flow. Appl Environ Microbiol 69:6793–6800
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Lozupone CA, Turnbaugh PJ, Fierer N, Knight R (2011) Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci 108:4516–4522
Cavigelli MA, Robertson GP (2000) The functional significance of denitrifier community composition in a terrestrial ecosystem. Ecology 81:1402–1414
Chao A, Chazdon RL, Colwell RK, Shen T-J (2005) A new statistical approach for assessing similarity of species composition with incidence and abundance data. Ecol Lett 8:148–159
Chu H, Fujii T, Morimoto S, Lin X, Yagi K, Hu J, Zhang J (2007a) Community structure of ammonia-oxidizing bacteria under long-term application of mineral fertilizer and organic manure in a Sandy loam soil. Appl Environ Microbiol 73:485–491. doi:10.1128/AEM.01536-06
Chu H, Lin X, Fujii T, Morimoto S, Yagi K, Hu J, Zhang J (2007b) Soil microbial biomass, dehydrogenase activity, bacterial community structure in response to long-term fertilizer management. Soil Biol Biochem 39:2971–2976. doi:10.1016/j.soilbio.2007.05.031
Clark IM, Hirsch PR (2008) Survival of bacterial DNA and culturable bacteria in archived soils from the Rothamsted Broadbalk experiment. Soil Biol Biochem 40:1090–1102. doi:10.1016/j.soilbio.2007.11.021
Darmody R.G., Norton L.D. (1993) Structural degradation of a prairie soil from long-term management. Developments in Soil Science 22:641–649 %@ 0166–2481.
Dick R.P. (1992) A review: long-term effects of agricultural systems on soil biochemical and microbial parameters. Agriculture, Ecosystems \& Environment 40:25–36.
Doan TT, Bouvier C, Bettarel Y, Bouvier T, Henry-des-Tureaux T, Janeau JL, Lamballe P, Van Nguyen B, Jouquet P (2014) Influence of buffalo manure, compost, vermicompost and biochar amendments on bacterial and viral communities in soil and adjacent aquatic systems. Appl Soil Ecol 73:78–86
Dojka MA, Hugenholtz P, Haack SK, Pace NR (1998) Microbial diversity in a hydrocarbon- and chlorinated-solvent-contaminated aquifer undergoing intrinsic bioremediation. Appl Environ Microbiol 64:3869–3877
Doran JW, Smith MS (1987) Organic matter management and utilization of soil and fertilizer nutrients. Soil fertility and organic matter as critical components of production systems. Soil Science Society of America and American Society of Agronomy, pp 53–72
Dufrêne M, Legendre P (1997) Species assemblages and indicator species: the need for a flexible asymmetrical approach. Ecol Monogr 67:345–366
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461
Edmeades DC (2003) The long-term effects of manures and fertilisers on soil productivity and quality: a review. Nutr Cycl Agroecosyst 66:165–180. doi:10.1023/A:1023999816690
Eichorst SA, Kuske CR, Schmidt TM (2011) Influence of plant polymers on the distribution and cultivation of bacteria in the phylum Acidobacteria. Appl Environ Microbiol 77:586–596. doi:10.1128/AEM.01080-10
el Zahar Haichar F, Marol C, Berge O, Rangel-Castro JI, Prosser JI, Balesdent J, Heulin T, Achouak W (2008) Plant host habitat and root exudates shape soil bacterial community structure. The ISME journal 2:1221–1230
Enwall K, Nyberg K, Bertilsson S, Cederlund H, Stenström J, Hallin S (2007) Long-term impact of fertilization on activity and composition of bacterial communities and metabolic guilds in agricultural soil. Soil Biol Biochem 39:106–115. doi:10.1016/j.soilbio.2006.06.015
Franchini J, Crispino C, Souza R, Torres E, Hungria M (2007) Microbiological parameters as indicators of soil quality under various soil management and crop rotation systems in southern Brazil. Soil Tillage Res 92:18–29
Garland JL, Mills AL (1991) Classification and characterization of heterotrophic microbial communities on the basis of patterns of community-level sole-carbon-source utilization. Appl Environ Microbiol 57:2351–2359
Giller K, Beare M, Lavelle P, Izac A, Swift M (1997) Agricultural intensification, soil biodiversity and agroecosystem function. Appl Soil Ecol 6:3–16
Girvan MS, Campbell CD, Killham K, Prosser JI, Glover LA (2005) Bacterial diversity promotes community stability and functional resilience after perturbation. Environ Microbiol 7:301–313. doi:10.1111/j.1462-2920.2005.00695.x
Greenland DJ (1981) Soil management and soil degradation. J Soil Sci 32:301–322
Griffiths BS, Philippot L (2013) Insights into the resistance and resilience of the soil microbial community. FEMS Microbiol Rev 37:112–129. doi:10.1111/j.1574-6976.2012.00343.x
Haas BJ, Gevers D, Earl AM, Feldgarden M, Ward DV, Giannoukos G, Ciulla D, Tabbaa D, Highlander SK, Sodergren E, Methé B, DeSantis TZ, Consortium THM, Petrosino JF, Knight R, Birren BW (2011) Chimeric 16S rRNA sequence formation and detection in sanger and 454-pyrosequenced PCR amplicons. Genome Res 21:494–504. doi:10.1101/gr.112730.110
Hallin S, Jones CM, Schloter M, Philippot L (2009) Relationship between N-cycling communities and ecosystem functioning in a 50-year-old fertilization experiment. The ISME Journal 3:597–605. doi:10.1038/ismej.2008.128
Hamady M, Walker JJ, Harris JK, Gold NJ, Knight R (2008) Error-correcting barcoded primers for pyrosequencing hundreds of samples in multiplex. Nat Methods 5:235–237
Härdle W.K., Simar L. (2012) Principal components analysis, applied multivariate statistical analysis, Springer pp 269-305.
Havlin JL, Kissel DE, Maddux LD, Claassen MM, Long JH (1990) Crop rotation and tillage effects on soil organic carbon and nitrogen. Soil Sci Soc Am J 54:448–452
Hill TCJ, Walsh KA, Harris JA, Moffett BF (2003) Using ecological diversity measures with bacterial communities. FEMS Microbiol Ecol 43:1–11
Hirsch PR, Mauchline TH, Clark IM (2010) Culture-independent molecular techniques for soil microbial ecology. Soil Biol Biochem 42:878–887. doi:10.1016/j.soilbio.2010.02.019
Horner-Devine MC, Carney KM, Bohannan BJM (2004) An ecological perspective on bacterial biodiversity. Proceedings of the Royal Society of London. Series B: Biological Sciences 271:113–122
Islam MT, Hashidoko Y, Deora A, Ito T, Tahara S (2005) Suppression of damping-off disease in host plants by the rhizoplane bacterium Lysobacter sp. strain SB-K88 is linked to plant colonization and antibiosis against soilborne peronosporomycetes. Appl Environ Microbiol 71:3786–3796
Jackson LE, Calderon FJ, Steenwerth KL, Scow KM, Rolston DE (2003) Responses of soil microbial processes and community structure to tillage events and implications for soil quality. Geoderma 114:305–317
Kamaa M, Mburu H, Blanchart E, Chibole L, Chotte J-L, Kibunja C, Lesueur D (2011) Effects of organic and inorganic fertilization on soil bacterial and fungal microbial diversity in the Kabete long-term trial, Kenya. Biol Fertil Soils 47:315–321. doi:10.1007/s00374-011-0539-3
Kanokratana P, Uengwetwanit T, Rattanachomsri U, Bunterngsook B, Nimchua T, Tangphatsornruang S, Plengvidhya V, Champreda V, Eurwilaichitr L (2010) Insights into the phylogeny and metabolic potential of a primary tropical peat swamp Forest microbial community by metagenomic analysis. Microb Ecol 61:518–528
Karlen DL, Hurley EG, Andrews SS, Cambardella CA, Meek DW, Duffy MD, Mallarino AP (2006) Crop rotation effects on soil quality at three northern corn/soybean belt locations. Agron J 98:484. doi:10.2134/agronj2005.0098
Ketcheson JW (1980) Long-range effects of intensive cultivation and monoculture on the quality of southern Ontario soils. Can J Soil Sci 60:403–410
Khan SA, Mulvaney RL, Ellsworth TR, Boast CW (2007) The myth of nitrogen fertilization for soil carbon sequestration. J Environ Qual 36:1821–1832
Kirk JL, Beaudette LA, Hart M, Moutoglis P, Klironomos JN, Lee H, Trevors JT (2004) Methods of studying soil microbial diversity. J Microbiol Methods 58:169–188
Kramer SB, Reganold JP, Glover JD, Bohannan BJ, Mooney HA (2006) Reduced nitrate leaching and enhanced denitrifier activity and efficiency in organically fertilized soils. Proc Natl Acad Sci U S A 103:4522–4527
Kuczynski J., Stombaugh J., Walters W.A., González A., Caporaso J.G., Knight R. (2012) Using QIIME to analyze 16S rRNA gene sequences from microbial communities. Current protocols in microbiology:1E-5 doi:10.1002/9780471729259.mc01e05s27
Kuever J. (2014) The family Syntrophorhabdaceae, the prokaryotes, Springer pp 301--303.
Larkin RP (2008) Relative effects of biological amendments and crop rotations on soil microbial communities and soilborne diseases of potato. Soil Biol Biochem 40:1341–1351
Legendre P, Legendre LFJ (2012) Numerical ecology, vol 24. Elsevier.
Lehman RM, Acosta-Martinez V, Buyer JS, Cambardella CA, Collins HP, Ducey TF, Halvorson JJ, Jin VL, Johnson JMF, Kremer RJ, Lundgren JG, Manter DK, Maul JE, Smith JL, Stott DE (2015) Soil biology for resilient, healthy soil. J Soil Water Conserv 70:12A–18A. doi:10.2489/jswc.70.1.12A
Lupwayi N, Rice W, Clayton G (1998) Soil microbial diversity and community structure under wheat as influenced by tillage and crop rotation. Soil Biol Biochem 30:1733–1741
Magoč T, Salzberg SL (2011) FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27:2957–2963
Manna MC, Swarup A, Wanjari RH, Ravankar HN, Mishra B, Saha MN, Singh YV, Sahi DK, Sarap PA (2005) Long-term effect of fertilizer and manure application on soil organic carbon storage, soil quality and yield sustainability under sub-humid and semi-arid tropical India. Field Crop Res 93:264–280. doi:10.1016/j.fcr.2004.10.006
McGuire KL, Treseder KK (2010) Microbial communities and their relevance for ecosystem models: decomposition as a case study. Soil Biol Biochem 42:529–535
Mendes R, Kruijt M, Bruijn Id, Dekkers E, Voort Mvd, Schneider JHM, Piceno YM, DeSantis TZ, Andersen GL, Bakker PAHM, Raaijmakers JM (2011) Deciphering the rhizosphere microbiome for disease-suppressive bacteria. Science 332:1097–1100
Mikha MM, Rice CW (2004) Tillage and manure effects on soil and aggregate-associated carbon and nitrogen. Soil Sci Soc Am J 68:809. doi:10.2136/sssaj2004.8090
Miller MN, Zebarth B, Dandie CE, Burton DL, Goyer C, Trevors JT (2008) Crop residue influence on denitrification, N 2 O emissions and denitrifier community abundance in soil. Soil Biol Biochem 40:2553–2562
Mitchell C, Westerman R, Brown J, Peck T (1991) Overview of long-term agronomic research. Agron J 83:24–29
Mulvaney RL, Khan SA, Ellsworth TR (2009) Synthetic nitrogen fertilizers deplete soil nitrogen: a global dilemma for sustainable cereal production. J Environ Qual 38:2295–2314
Mummey DL, Stahl PD, Buyer JS (2002) Microbial biomarkers as an indicator of ecosystem recovery following surface mine reclamation. Appl Soil Ecol 21:251–259
Nafziger ED, Dunker RE (2011) Soil organic carbon trends over 100 years in the morrow plots. Agron J 103:261–267
Nair A, Ngouajio M (2012) Soil microbial biomass, functional microbial diversity, and nematode community structure as affected by cover crops and compost in an organic vegetable production system. Appl Soil Ecol 58:45–55
Nannipieri P, Kandeler E, Ruggiero P (2002) Enzyme activities and microbiological and biochemical processes in soil. Enzymes in the environment. Marcel Dekker, New York, pp. 1–33
Navarro-Noya YE, Gómez-Acata S, Montoya-Ciriaco N, Rojas-Valdez A, Suárez-Arriaga MC, Valenzuela-Encinas C, Jiménez-Bueno N, Verhulst N, Govaerts B, Dendooven L (2013) Relative impacts of tillage, residue management and crop-rotation on soil bacterial communities in a semi-arid agroecosystem. Soil Biol Biochem 65:86–95
Nelson DR, Mele PM (2006) The impact of crop residue amendments and lime on microbial community structure and nitrogen-fixing bacteria in the wheat rhizosphere. Soil Research 44:319–329
Neumann D, Heuer A, Hemkemeyer M, Martens R, Tebbe CC (2013) Response of microbial communities to long-term fertilization depends on their microhabitat. FEMS Microbiol Ecol 86:71–84. doi:10.1111/1574-6941.12092
Odell RT, Walker WM, Boone LV, Oldham MG (1982) The Morrow Plots: a century of learning. Bulletin-Agricultural Experiment Station, College of Agriculture, University of Illinois at Urbana-Champaign. Agricultural Experiment Station, College of Agriculture, University of Illinois at Urbana-Champaign, Urbana, IL, p 775
Ogilvie LA, Hirsch PR, Johnston AWB (2008) Bacterial diversity of the Broadbalk `Classical' winter wheat experiment in relation to long-term fertilizer inputs. Microb Ecol 56:525–537. doi:10.1007/s00248-008-9372-0
Oksanen J, Blanchet FG, Kindt R, Legendre P, Minchin PR, O'Hara RB, Simpson GL, Solymos P, Stevens MHH, Wagner H, others. (2015) Community Ecology Package. R package version 2.0-10. 2013
Otto A, Shunthirasingham C, Simpson MJ (2005) A comparison of plant and microbial biomarkers in grassland soils from the prairie Ecozone of Canada. Org Geochem 36:425–448
Pascual JA, Garcia C, Hernandez T, Moreno JL, Ros M (2000) Soil microbial activity as a biomarker of degradation and remediation processes. Soil Biol Biochem 32:1877–1883
Paterson E, Gebbing T, Abel C, Sim A, Telfer G (2007) Rhizodeposition shapes rhizosphere microbial community structure in organic soil. New Phytol 173:600–610
Peacock Ag, Mullen MD, Ringelberg DB, Tyler DD, Hedrick DB, Gale PM, White DC (2001) Soil microbial community responses to dairy manure or ammonium nitrate applications. Soil Biol Biochem 33:1011–1019
Postma J, Nijhuis EH, Yassin AF (2010) Genotypic and phenotypic variation among Lysobacter capsici strains isolated from Rhizoctonia suppressive soils. Syst Appl Microbiol 33:232–235
Powlson DS, Prookes PC, Christensen BT (1987) Measurement of soil microbial biomass provides an early indication of changes in total soil organic matter due to straw incorporation. Soil Biol Biochem 19:159–164. doi:10.1016/0038-0717(87)90076-9
Powlson DS, MacDonald AJ, Poulton PR (2014) The continuing value of long-term field experiments: insights for achieving food security and environmental integrity, in: D. Dent (Ed.). Soil as World Heritage, Springer Netherlands, pp. 131–157
R Core Team (2016) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria
Reddy T.B.K., Thomas A.D., Stamatis D., Bertsch J., Isbandi M., Jansson J., Mallajosyula J., Pagani I., Lobos E.A., Kyrpides N.C. (2014) The Genomes OnLine Database (GOLD) v. 5: a metadata management system based on a four level (meta) genome project classification. Nucleic acids research doi:10.1093/nar/gku950.
Rivard C., Lanson B., Cotte M. (2015) Phosphorus speciation and micro-scale spatial distribution in North-American temperate agricultural soils from micro X-ray fluorescence and X-ray absorption near-edge spectroscopy. Plant and Soil: 1–16 %@ 0032-079X.
Roberts DW (2012) Package ‘labdsv’.
Rosenzweig N, Tiedje JM, Quensen Iii JF, Meng Q, Hao JJ (2012) Microbial communities associated with potato common scab-suppressive soil determined by pyrosequencing analyses. Plant Dis 96:718–725
Sambrook J., Fritsch E.F., Maniatis T. (1989) Molecular cloning Cold spring harbor laboratory press New York.
Sexstone AJ, Revsbech NP, Parkin TB, Tiedje JM (1985) Direct measurement of oxygen profiles and denitrification rates in soil Aggregates1. Soil Sci Soc Am J 49:645. doi:10.2136/sssaj1985.03615995004900030024x
Shade A., Jones S.E., Caporaso J.G., Handelsman J., Knight R., Fierer N., Gilbert J.A. (2014) Conditionally Rare Taxa Disproportionately Contribute to Temporal Changes in Microbial Diversity. mBio 5:e01371–14. doi:10.1128/mBio.01371–14.
Six J, Frey S, Thiet R, Batten K (2006) Bacterial and fungal contributions to carbon sequestration in agroecosystems. Soil Sci Soc Am J 70:555–569
Souza RC, Hungria M, Cantão ME, Vasconcelos ATR, Nogueira MA, Vicente VA (2015) Metagenomic analysis reveals microbial functional redundancies and specificities in a soil under different tillage and crop-management regimes. Appl Soil Ecol 86:106–112. doi:10.1016/j.apsoil.2014.10.010
Tilman D, Cassman KG, Matson PA, Naylor R, Polasky S (2002) Agricultural sustainability and intensive production practices. Nature 418:671–677
Tisdall JM, Oades JM (1982) Organic matter and water-stable aggregates in soils. J Soil Sci 33:141–163. doi:10.1111/j.1365-2389.1982.tb01755.x
van Diepeningen AD, de Vos OJ, Korthals GW, van Bruggen AH (2006) Effects of organic versus conventional management on chemical and biological parameters in agricultural soils. Appl Soil Ecol 31:120–135
Velde B., Peck T. (2002) Clay mineral changes in the Morrow experimental plots, University of Illinois. Clays and Clay Minerals 50:364–370 %@ 0009–8604.
Ward NL, Challacombe JF, Janssen PH, Henrissat B, Coutinho PM, Wu M, Xie G, Haft DH, Sait M, Badger J, Barabote RD, Bradley B, Brettin TS, Brinkac LM, Bruce D, Creasy T, Daugherty SC, Davidsen TM, DeBoy RT, Detter JC, Dodson RJ, Durkin AS, Ganapathy A, Gwinn-Giglio M, Han CS, Khouri H, Kiss H, Kothari SP, Madupu R, Nelson KE, Nelson WC, Paulsen I, Penn K, Ren Q, Rosovitz MJ, Selengut JD, Shrivastava S, Sullivan SA, Tapia R, Thompson LS, Watkins KL, Yang Q, Yu C, Zafar N, Zhou L, Kuske CR (2009) Three genomes from the phylum Acidobacteria provide insight into the lifestyles of these microorganisms in soils. Appl Environ Microbiol 75:2046–2056. doi:10.1128/AEM.02294-08
Wessén E, Hallin S, Philippot L (2010a) Differential responses of bacterial and archaeal groups at high taxonomical ranks to soil management. Soil Biol Biochem 42:1759–1765. doi:10.1016/j.soilbio.2010.06.013
Wessén E, Nyberg K, Jansson JK, Hallin S (2010b) Responses of bacterial and archaeal ammonia oxidizers to soil organic and fertilizer amendments under long-term management. Appl Soil Ecol 45:193–200. doi:10.1016/j.apsoil.2010.04.003
Widmer F, Rasche F, Hartmann M, Fliessbach A (2006) Community structures and substrate utilization of bacteria in soils from organic and conventional farming systems of the DOK long-term field experiment. Appl Soil Ecol 33:294–307. doi:10.1016/j.apsoil.2005.09.007
Wortmann CS, Shapiro CA (2008) The effects of manure application on soil aggregation. Nutr Cycl Agroecosyst 80:173–180
Wu M, Qin H, Chen Z, Wu J, Wei W (2011) Effect of long-term fertilization on bacterial composition in rice paddy soil. Biol Fertil Soils 47:397–405. doi:10.1007/s00374-010-0535-z
Xuan DT, Guong VT, Rosling A, Alström S, Chai B, Högberg N (2012) Different crop rotation systems as drivers of change in soil bacterial community structure and yield of rice, Oryza sativa. Biol Fertil Soils 48:217–225. doi:10.1007/s00374-011-0618-5
Yin C, Jones KL, Peterson DE, Garrett KA, Hulbert SH, Paulitz TC (2010) Members of soil bacterial communities sensitive to tillage and crop rotation. Soil Biol Biochem 42:2111–2118. doi:10.1016/j.soilbio.2010.08.006
Zhen Z, Liu H, Wang N, Guo L, Meng J, Ding N, Wu G, Jiang G (2014) Effects of manure compost application on soil microbial community diversity and soil microenvironments in a temperate cropland in China. PLoS One 9:e108555. doi:10.1371/journal.pone.0108555
Zhong W, Gu T, Wang W, Zhang B, Lin X, Huang Q, Shen W (2010) The effects of mineral fertilizer and organic manure on soil microbial community and diversity. Plant Soil 326:511–522
Acknowledgments
We would like to acknowledge Robert Dunker and previous managers and the University of Illinois Agricultural Experiment Station for long-term support and maintenance of the Morrow Plots. This work was also supported by the Cooperative State Research, Education, and Extension Service, US Department of Agriculture, under project number ILLU 875-374. The authors are grateful to students enrolled in the Fall 2009 Microbial Ecology Methods course taught by A. Kent for assistance with sample collection and processing. Chinmay Soman is supported by the National Science Foundation through the Science, Engineering, and Education for Sustainability Post-Doctoral Fellowship, Award # 1314064.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Jeff R. Powell.
Rights and permissions
About this article
Cite this article
Soman, C., Li, D., Wander, M.M. et al. Long-term fertilizer and crop-rotation treatments differentially affect soil bacterial community structure. Plant Soil 413, 145–159 (2017). https://doi.org/10.1007/s11104-016-3083-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-016-3083-y