Abstract
Aim
Few studies have analyzed life-history strategies of arbuscular mycorrhizal fungi (AMF), in terms of the different propagule types they produce, and their ability to colonize new seedlings. The aim was to assess whether life-history strategies influence AMF successional dynamics and assemblages.
Methods
Rosemary (Rosmarinus officinalis L.) seedlings, grown in a mesocosm system, were colonized by either the AMF hyphae coming from a living rosemary plant, or from spores germinating in soil. The AMF community established in the plantlets was monitored every 3 months during 2 years, using terminal restriction fragment length polymorphism of genes coding for rDNA.
Results
The two different sources of AMF propagules resulted in a different initial community colonizing rosemary roots. AMF propagating from hyphae attached to living mycorrhizal-roots seemed to colonize faster and were season-dependent. AMF taxa originating from soil-borne propagules were most frequent over time and exhibit the dominant colonization strategy in this system. The evolution of the AMF community also revealed different strategies in succession.
Conclusions
AMF associated with rosemary evidenced contrasting life-history strategies in terms of source of inoculum for new colonization and hence survival. The observed successional dynamics of AMF have implications for understanding the ecological processes in Mediterranean environments and seasonality of colonization processes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Glomeromycota are a widespread group of obligate mutualistic fungi which symbiotically colonize the roots of approximately two-thirds of plant species to form arbuscular mycorrhizas (Barea and Azcón-Aguilar 2013). Arbuscular mycorrhizal (AM) fungi (AMF) increase nutrient and water uptake by the plant through exploration of soil resources by the fungal hyphae, and the host plant supplies the fungus with carbon compounds from photosynthesis (Koide and Mosse 2004). At an ecosystem level, these symbioses benefit plant diversity, productivity and soil structure (Barea et al. 2011; Rillig 2004; van der Heijden et al. 1998, 2006; Vogelsang et al. 2006). Traditionally, AMF have been considered as generalists, with little or no partner specificity, but recent more sophisticated studies have shown a considerable level of partner selectivity for particular plant species or ecological plant groups (Davison et al. 2011).
Functional characteristics of AMF are important determinants of the formation and significance of AM symbioses in natural ecosystems (Helgason and Fitter 2009; van der Heijden and Scheublin 2007). Large contrasts have been shown in colonization strategies (Hart and Reader 2002, 2005), life cycles (Boddington and Dodd 1999) or benefits conferred to the host plants (Avio et al. 2006; Munkvold et al. 2004). Recent research suggests a differentiation in the life-history of AMF in terms of their propagative strategies (Denison and Kiers 2011). Fresh root colonization occurs either from hyphae of existing mycelial networks attached to living plants or from hyphae from germinating soil-borne spores or growing from old root fragments. The presence of glomeromycotan DNA sequences in roots but not in spores from the same habitat suggests that both sources of inocula are important for AMF having different life-history strategies (Dumbrell et al. 2011; Hempel et al. 2007; Husband et al. 2002; Merryweather and Fitter 1998; Sánchez-Castro et al. 2012a).
To investigate such ecological aspects, a representative and well-conserved Mediterranean shrub community, located at the Sierra de Baza (Granada, Southeast Spain), was selected. The diversity of habitats in Mediterranean environments, together with the climatic regime of long, dry, hot summers and low winter temperatures, with irregular rain events (López-Bermúdez et al. 1990; Vallejo et al. 1999), has resulted in a large number of plant strategies (Médail and Quézel 1997; Myers et al. 2000). As AMF diversity was shown to promote plant diversity (van der Heijden et al. 1998), it seems feasible that a high plant diversity can facilitate a high taxonomical and functional AMF diversity (Pringle and Bever 2002). Thus, Mediterranean environments are good candidates for the dissection of contrasting life-history strategies in AMF and their significance in AMF community successional dynamics. Rosemary (Rosmarinus officinalis L.), a widespread evergreen sclerophyllus shrub, was chosen as host plant. Rosemary plants form naturally extended vegetation patches (rosemary groves), associated with herbaceous, annual, plant species (Sánchez-Castro et al. 2012a). This plant community provides an AM inoculum pool responsible for colonizing new plantlets joining the community. We hypothesized that the AMF taxa involved might have contrasting life-history strategies with regards to their ability to colonize new plants from either mycelia emanating from plants that were already colonized, or from soil-borne propagules stimulated by the presence of new roots in their vicinity.
Our objectives were thus (i) to test the hypothesis that there were AMF taxa with contrasting life-history strategies and (ii) to ascertain experimentally how these contrasting life-history strategies influence the successional dynamics of the AMF community.
Materials and methods
Experimental design
The study comprised a mesocosm experiment using rosemary (Rosmarinus officinalis L.) as host plant. The mesocosm units were established in 15 l circular containers (40 cm diameter, 15 cm deep). Nine rosemary plants were sown in each unit: one in the centre (donor plant) and the other eight (receiver plants) at equal intervals around a 12 cm radius circle surrounding the central one. Two different sources of mycorrhizal inoculum were tested to distinguish between the two AMF propagative strategies: natural soil from the target area having only soil-borne AMF propagules (either spores or mycorrhizal root fragments), or field-grown rosemary plants, colonized by the AMF taxa naturally associated with rosemary. Three different treatments were established: 1) soil containing mycorrhizal propagules with a non-mycorrhizal plant as donor plant (‘soil-borne propagules’); 2) sterilized soil with a field rosemary plant as mycorrhizal donor plant (‘plant-derived propagules’) and 3) the combination of the previous treatments (‘soil+plant propagules’).
Mycorrhizal propagule source
Both the soil (a calcaric cambisol) and the donor plants used as source of AMF propagules in the experiment were obtained from the Sierra de Baza Natural Park, where rosemary plants grow naturally. For this purpose, an area of about 100 m2 comprising a representative, well-conserved, Mediterranean shrub community was selected. The vegetation was dominated by patchily-distributed Lamiaceae shrubs: Rosmarinus officinalis L., Lavandula latifolia Medik., Thymus zigys Loefl. ex L. and T. mastichina (L.) L. associated with herbaceous, annual, plant species. On March 2008, six points were randomly selected to collect 300 l of soil. The soil was collected at a depth of ca. 5–30 cm and sieved through a 5 mm sieve.
Before setting up the experiment, the soil was dried to eliminate the AMF propagules with poor survival capacity. Soil was placed as a 3 cm layer in a glasshouse without temperature control from March to November and mixed weekly to ensure uniform desiccation. The mycorrhizal colonization potential of the soil was measured weekly by quantifying the number of AMF entry points in a trap plant, Sorghum vulgare Pers. (Franson and Bethlenfalvay 1989). Prior to drying, one-third of the collected soil was steam-sterilized (1 h for 3 consecutive days) to be used as soil without mycorrhizal propagules. During drying, the mycorrhizal inoculum potential of the soil did not vary significantly, maintaining an average value of 0.68 (±0.02 SE) colonization units per cm of root during the 8 months of drying (Supporting Information Fig. S1). The soil moisture, after an initial decreasing, was maintained around 1.73 % (±0.02) (Fig. S1). The loss of the most sensitive mycorrhizal propagules to soil alteration occurred probably during the soil sieving.
Two to 3-year-old rosemary plants were randomly chosen from the target area in Sierra de Baza on November 2008. Twelve of the plants were used as mycorrhizal donor plants and another six to analyze the natural AMF assemblages colonizing their roots. The root system of the field plants was carefully washed with tap water to remove extraradical hyphae and spores before sowing.
Set up of the experiment
The experiment was set up on November 2008. A field-grown rosemary plant was established in the center of each mesocosm unit as the mycorrhizal donor plant. In soil-borne mycorrhizal propagule treatments, the donor plant was replaced by a non-mycorrhizal plant, obtained from sterile rooted rosemary cuttings. All mesocosm units containing steam-sterilized soil were supplied with a filtrate of the non-steamed soil to restore natural microbial communities, free from AMF propagules. Donor plants were allowed to grow for 1 month before planting the receiver plants to ensure the spreading out of the AMF hyphae present in the roots into the soil.
In December 2008, eight non-mycorrhizal rooted rosemary cuttings (receiver plants) were established in each mesocosm unit. Six replicate mesocosm units were established per treatment. The mesocosm units were maintained for 2 years in a glasshouse without supplementary light. Day:night temperatures were adjusted to 25 °C:18 °C in summer using a cooler system. One receiver plant was harvested every 3 months, beginning in March 2009, to determine the mycorrhizal colonization level and to characterize the associated AMF community.
Mycorrhizal colonization
The entire rosemary root system of receiver plants was cut in 1 cm pieces and thoroughly mixed. Two g of roots were stained with trypan blue (Phillips and Hayman 1970) and examined under a compound light microscope to determine the percentage mycorrhizal root colonization and the frequency of intraradical fungal structures, arbuscules and vesicles (McGonigle et al. 1990). The remaining roots, in aliquots of 200 mg, were immediately frozen in liquid nitrogen and stored at −80 °C until molecular analysis to determine AMF diversity.
AMF gene library
To apply the TRFLP database approach (see Dickie and Fitzjohn 2007), a previous gene library containing a major part of the sequences of glomeromycotan fungi included in the whole study was constructed. For that purpose, DNA was extracted from 200 mg of mycorrhizal roots of each sample (individual plant) using a DNeasy plant mini-kit (Qiagen Inc., Mississauga, ON, Canada) and eluted in 75 μl ddH2O. Partial ribosomal 18S DNA fragments were amplified using the primer set AML1-AML2 (Lee et al. 2008), a specific pair of primers for AMF DNA amplification. The Polymerase Chain Reactions (PCR) were carried out using Illustra Pure-Taq Ready-To-Go PCR beads (GE Healthcare UK Limited, Buckinghamshire, UK) and 1 μl (5 μM) of each primer. PCR conditions were: 1 min at 94 °C, 30 cycles at 94 °C for 1 min, 62 °C for 1 min, and 72 °C for 1 min, followed by a final extension period at 72 °C for 5 min. As a template, 1 μl of extracted DNA previously diluted 1 : 10 was used in all reactions.
An equimolecular mix of all PCR products was cleaned up (Illustra GFX PCR DNA and Gel Band Purification Kit, GE Healthcare Life Sciences, Freiburg, Germany), cloned into the pCR®2.1 vector following the protocol recommended by the manufacturer of the TA Cloning® Kit (Invitrogen Life Technologies, Karlsruhe, Germany) and transformed into One Shot® TOP10F’ Chemically Competent Escherichia coli cells (Invitrogen Life Technologies). Each clone was reamplified using NS31 (Simon et al. 1992), a universal eukaryotic primer, and Glo1, a primer designed for glomeromycotan fungi (Cornejo et al. 2004). The resulting PCR product of around 230 bp has an ideal size to be discriminated by Single Strand Conformational Polymorphism (SSCP). This internal region of AML1-AML2 is a highly variable area allowing the discrimination of a majority of sequences of AMF (Cornejo et al. 2004). Clones for SSCP were prepared and analyzed following the protocol described by Kjøller and Rosendahl (2000). To ensure a good coverage of the AMF diversity in the rosemary plants, the AML1-AML2 fragment of around 40 % of the clones exhibiting the same SSCP profile was sequenced by the sequencing services of the Estación Experimental del Zaidín (Granada, Spain) using the set of vector binding primers M13F-M13R. The abundance of DNA sequences indicated by their profiles in the SSCP fingerprinting was used to carry out the rarefaction analysis.
Phylogenetic analysis
The phylogenetic analysis was carried out using the sequences obtained in the study and a representative group of sequences of all major glomeromycotan groups from GenBank (references KC665640-KC665707). Sequences were aligned with MAFFT version 6 and similarities excluding the primer sequence were determined using BioEdit software. The outgroup used was Mortierella polycephala X89436. The phylogenetic analysis was computed in MEGA4 (Tamura et al. 2007) using the neighbor-joining algorithm with Kimura-2 parameters as the model of substitution and 1,000 replications to obtain bootstrap values. Phylotypes were determined using a minimum of 97 % sequence similarity and a high bootstrap value as indicators (Öpik et al. 2010). A blast search in the MaarjAM database (Öpik et al. 2010) was used to name phylotypes giving them the number code of the closest virtual taxon (sharing always a minimum of 97 % similarity). The prefix of the phylotype name used corresponds with the glomeromycotan family (Krüger et al. 2012): Cla-Claroideoglomeraceae, Div-Diversisporaceae, Glo-Glomeraceae, Pac-Pacisporaceae, Par-Paraglomeraceae and Sac-Sacculosporaceae (Oehl et al. 2011).
TRFLP analyses
REPK online software (Collins and Rocap 2007) was used to select a combination of four enzymes able to discriminate between the phylotypes found in the gene library. These enzymes were: HinfI, MboI, AvaII and BstXI (New England Biolabs). In order to carry out a Multiplex TRFLP analysis, two PCR reactions were performed per sample as described in the “AMF gene library” section, one with the fluorescent label 6-FAM and the other with HEX, both attached to the forward primer AML1. Forty ng of each PCR product were digested with the corresponding restriction enzyme for 2 h at 37 °C (HEX PCR products with HinfI and AvaII separately and 6-FAM PCR products with MboI and BstXI). Digestions with MboI and HinfI were mixed as done for BstXI and AvaII when they were cleaned (Illustra GFX PCR DNA and Gel Band Purification Kit, GE Healthcare Life Sciences, Freiburg, Germany) before fragment length determination analysis. TRF determination was done by the Unidad de Genómica y Síntesis de DNA, Instituto de Biomedicina y Parasitología López Neyra (Granada, Spain). Data were processed using GeneMapper software version 3.7 (Applied Biosystems 2004). In order to get the empirical TRF profiles of the detected sequences in the database, the clones selected for the gene library construction were re-amplified, digested and analyzed in the same way as explained for the samples. The R (R Development Core Team 2010) package, TRAMPR (FitzJohn and Dickie 2007) was used for matching TRFLP profiles of the samples with the sequence database profiles, which allowed us to allocate the phylotypes present in the roots of each plant.
Statistical analyses
Repeated measure ANOVA with Greenhouse-Geisser adjusted probability was used to analyze the mycorrhizal colonization data in order to test propagule source and time effects. A split-plot design with time as the within-subject factor and mycorrhizal propagule source as the between-subject factor was used; mesocosm units were considered as subjects. Due to the lack of normality of phylotype richness data, non parametric Kruskal-Wallis tests were carried out to check for effects of time and propagule source.
The diversity of sequences resulting from the cloning was analyzed by rarefaction analysis using the Analytic Rarefaction freeware software (Holland 2008). A Non-metric Multidimensional Scaling (NMDS) ordination, using Jaccard distance as community dissimilarity measurement, was carried out to allow for a representation of AMF community variation. Effects of the three categorical variables (both types of propagule sources -soil or plant- and season) and the continuous variable (time) were investigated by permutational multivariate analysis of variance (permanova, McArdle and Anderson 2001) embedded in the function adonis from vegan package in R (Oksanen et al. 2011), using Jaccard distance as a measure of community dissimilarity. These analyses were stratified according to the date of sampling (time). Consequently the effect of time was extracted from the effect of the rest of variables.
Since data obtained from the TRFLP database consisted of a presence: absence matrix, only richness (S) as diversity index was calculated. In order to assess the differences in phylotype composition found in the rosemary roots colonized from different propagule sources during the experiment, the frequency of detection was calculated as the percentage of replicates containing each phylotype.
Dufrêne-Legendre indicator species analysis (Dufrêne and Legendre 1997) was carried out using the function indval from the labdsv package in R statistics (Roberts 2010) to identify AMF tied to specific levels of the tested variables that could serve as indicator species. As a reference value, 0.20 was used as the threshold to consider a species indicator of a certain treatment. The different levels of each variable were tested separately. In the case of the time variable, different periods were tested.
Results
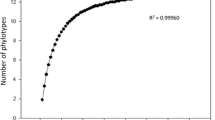
Sequencing of 70 clones of the gene library resulted in the phylogenetic tree shown in Fig. S2. Phylotypes of 15 members of six glomeromycotan families were detected (Glomeraceae -9-, Pacisporaceae -1-, Claroideoglomeraceae -2-, Diversisporaceae -1-, Sacculosporaceae -1- and Paraglomeraceae -1-). The rarefaction analysis, based on the SSCP profile of 162 clones, showed a maximum of 17 phylotypes (Fig. S3) if 1,300 clones were analyzed (r 2 = 0.996). The most abundant glomeromycotan DNA sequences in this study corresponded to the Glomeraceae. Only 2.7 % of non-specific amplifications were detected, including rosemary sequences.
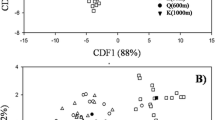
Mycorrhizal propagule source significantly affected the AMF community composition in the rosemary roots (F = 2.43, p = 0.009 for soil-borne propagules and F = 6.66, p = 0.001 for plant-derived propagules). In addition, the effect of time revealed a successional pattern (F = 9.26, p = 0.001) influenced by season (F = 3.44, p = 0.002). When observing the evolution of the AMF community composition (Fig. 1), the soil+plant propagules treatment showed a tendency to be close, and in consequence similar, to the treatment including only soil-borne propagules from the second sampling time. However in the three last sampling times the AMF community appeared very similar in all treatments. Phylotype richness was influenced by propagule source (χ 2 = 9.2556, df = 2, P < 0.01) as well as time (χ 2 = 19.3504, df = 7, P < 0.01). In general, plants colonized by soil-borne propagules had a higher richness than those colonized from plant-derived propagules (Fig. 2). Plants colonized by soil+plant propagules had values closer to those colonized from soil-borne propagules during the first year of the study; however, at the end of the experiment they became more similar to those colonized from plant-derived propagules.
Time-course evolution of the composition of the AMF community. NMDS ordination (k = 2, stress = 0.121) was represented by date of harvesting. Standard error bars are shown for each treatment and axis. Black circles represent soil-borne propagule treatment, white circle plant-derived propagule treatment and black triangle soil+plant propagule treatment
Dufrêne-Legendre species indicator analysis (Dufrêne and Legendre 1997) showed that two phylotypes were indicators of the presence of soil-borne propagules and the absence of plant-derived propagules (Cla056 and Glo064, Table 1). Other two phylotypes, Div062 and Par336, were indicators of the absence of propagules coming from the soil or the plant, respectively (Table 1). In general, members of the Diversisporales (Diversisporaceae, Sacculosporaceae and Pacisporaceae) appear as early colonizers with a high frequency of detection during the first sampling times (Fig. 3, Table 1). Although Pac284 did not show a statistically significant trend, probably due to its low level of detection, it was similar to the other Diversisporalean phylotypes. Members of the Claroideoglomeraceae appeared principally in the central time periods, while during the second year of the study, the dominating phylotypes were members of Glomeraceae. With respect to season, the Diversisporales appear as indicators of March, while Glo166, the most abundant phylotype in this study, appeared as an indicator of June and September. It is noteworthy that phylogenetic conservatism of fungal traits among glomeromycotan families (Powell et al. 2009) appears well represented by the positions of phylotypes in the NMDS ordination shown in Fig. 4.
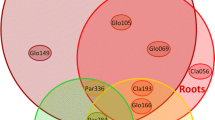
NMDS ordination of AMF communities (k = 2, stress = 0.121). The relative positions of the different phylotypes is represented: White squares represent AMF phylotypes belonging to the Diversisporales, black squares to the Glomeraceae (Glomerales), white triangles to the Claroideoglomeraceae (Glomerales) and the black triangle corresponds to the member from the Paraglomerales. Numbers accompanying phylotypes indicate the code of the closest virtual taxa (Öpik et al. 2010, see also Fig. S2). Each line represents equal sampling dates (time variable)
With regard to the AMF community associated with plants growing in natural conditions (‘donor plants’), it is noteworthy that the community was dominated by members of the Glomeraceae, being Glo105 present in every analyzed plant. Glo109 was the only Glomeraceae phylotype found in the receiver plants not detected in the donor plants. The only member of the Paraglomeraceae detected in this study (Par336) was found in a 50 % of the donor plants and the rest of glomeromycotan families were not detected. Apparently there was no correspondence among the AMF phylotypes colonizing donor plants and those colonizing preferentially from plant-derived propagules. In fact, Div062, the AMF phylotype colonizing preferably from the plant was not detected in any of the analyzed donor plants, while Par336 and Glo064, which colonize almost exclusively from the soil, were detected in 50 % of the plants growing in field conditions (data not shown). A higher level of concordance was found for other phylotypes: for instance Cla056, colonizing almost exclusively from the soil was not detected in the donor plants, and Glo166 and Glo105, the most abundant phylotypes in the ‘receiver plants’ were detected in 50 and 100 % of the donor plants, respectively.
Propagule source and time also influenced significantly mycorrhizal colonization. The effect of the propagule source on total mycorrhizal colonization (F = 8.51, df = 2, P < 0.01, ε = 0.45) was mainly due to differences in the first sampling time since plants colonized only from plant-derived propagules had a low amount of mycorrhizal colonization (6.40 %, Fig. 5). At the second sampling time mycorrhizal colonization levels were similar among treatments. The extent of arbuscular colonization in the roots was influenced by time (F = 38.48, df = 7, P < 0.001, ε = 0.51) while the source of propagules had no significant effects. The frequency of arbuscules in the root system over time showed a cyclic pattern which can be easily attributed to seasonality (Fig. 5). Finally, the frequency of vesicle detection was clearly influenced by both time (F = 11.73, df = 7, P < 0.001, ε = 0.36) and propagule source (F = 5.10, df = 2, P < 0.05, ε = 0.36). Plants colonized only from plant-derived propagules had higher levels of vesicle formation than those colonized only from soil-borne propagules (Fig. 5).
Discussion
This study used an experimental approach to reveal different life-history strategies of AMF in relation to their colonization dynamics, the way in which they persist during unfavourable seasons and, in consequence, the temporal succession of their communities. Basically, AMF could follow two contrasting life-history strategies: (i) those colonising from soil-borne sources and (ii) those which colonise from mycelium emanating from neighbouring root colonisation. The former strategy is based on resistant AMF propagules which can survive adverse conditions between seasons thereby ensuring the survival of AMF populations in the absence of host plants until the next favourable season (Tommerup and Abbott 1981). For ectomycorrhizal fungi, a high rate of hyphal death during the dry season in Mediterranean environments has been shown (Allen and Kitajima 2013). In our study, soil drying treatment was strong enough to select only spores and small fragments of colonized roots. Hyphal networks are much more sensitive to soil disturbance and desiccation (Jasper et al. 1989).
The type of propagule clearly influenced the resultant AMF community that colonized new plants during the first year of the study. Only one phylotype (Div062) was preferentially detected when mycorrhizal plants were the only source of propagules, suggesting that this phylotype disperses through mycelium rather than spores. As described for ectomycorrhizal fungi (Peay et al. 2011), the formation and maintenance of hyphal networks is a way of dispersion not dependent on spore formation (Denison and Kiers 2011). Mycelial growth is favoured by the permanent growth of the roots of perennial plants (Rosendahl and Stukenbrock 2004), such as rosemary. When attached to roots, hyphal networks are generally more efficient than spores in colonizing new roots because they have greater availability of carbon that can be used to explore the soil and encounter uncolonized roots (Peay et al. 2011). It is noteworthy that plants colonized from plant-derived propagules showed a significant increase in the abundance of AMF vesicles in the roots. This increase was especially prominent in December, during early winter, which may be an indication of resource hoarding by these AMF for the upturn in growth of the following spring.
The rest of AMF phylotypes associated with the rosemary rhizosphere in field-grown plants seemed to colonize roots more effectively from soil-borne propagules. This could be explained by the dominance in these environments of AMF that preferentially colonize through spores (‘spore colonizers’) as it has been suggested for some degraded habitats (Jansa et al. 2003; Oehl et al. 2011). In general, the ecology of the rosemary shrublands supports the patterns found here. Although rosemary is a perennial woody species, the ecological characteristics of its communities could be unfavourable to the development of AMF spreading by mycelial networks as there is scarce plant cover and poor soil development (Rosendahl and Stukenbrock 2004; Schnoor et al. 2011).
Par336, Glo064 and Cla056 were mainly detected in the receiver plants when they grew in the soil containing only soil-borne propagules. This suggests that these AMF phylotypes could have difficulties in spreading from living roots, probably due to a poorly developed external mycelium as suggested for some AMF, particularly the Glomeraceae (Hart and Reader 2005). However it is possible that the mycorrhizal donor plants were not colonized with these fungal phylotypes, but this is unlikely, at least for Par336 and Glo064 which were detected in 50 % of the plants growing naturally in the field.
A clear pattern of AMF taxonomic succession was found. Three different colonization strategies appear to drive the succession of AMF phylotypes in roots:
Diversisporalean fungi seem to be spring colonizers. It maybe that their strategy consists of being earlier colonizers in each growing season before later competitors can displace them. These phylotypes are subsequently replaced by members of the Glomerales. This seasonal trend was described for other Diversispora sp. in a previous study in Sierra de Baza Natural Park (Sánchez-Castro et al. 2012b).
Members of the Claroideoglomeraceae appear somewhat later than Diversisporales in the succession. Their dynamics showed a plateau in the central periods of the study. They were also later displaced by members of the Glomeraceae.
The phylotypes showing highest persistence in this study were Glomeraceae. Despite frequently being considered r-strategists (IJdo et al. 2010), they were not the first colonizers in this study. However, once they had colonized roots, they persisted longest. This might be explained by the fact that the Glomeraceae have been repeatedly described as extensive root colonizers (Hart and Reader 2002), and may have introduced a bias and hindered the detection of the less abundant fungi colonizing the roots. In addition, it is also possible that the experimental conditions during this study have restricted root growth, especially during the second year, and a more constrained niche space can exclude species with poor rates of root colonization (Maherali and Klironomos 2012; van der Heijden and Scheublin 2007), as may have been the case for the Diversisporales in this study.
The AMF community colonizing the donor plants fits quite well with that hypothetically associated with 2-year-old rosemary plants sampled in November. According to our results, it should be associated with a community dominated by Glomeraceae and with a minor presence of Diversisporales (due to their seasonal behaviour) and Claroideoglomeraceae, which were undetectable at the end of the second year.
In general, it seems clear that the initial AMF community was gradually replaced such that all treatments resulted in similar communities over time. This convergence of AMF assemblages could be driven by two mechanisms: either selection by the host plant of the most compatible/beneficial AMF through the preferential allocation of resources to the preferred fungal partners (Bever et al. 2009; Grman 2012), or a natural succession of AMF communities based on their life-history strategies (Hart et al. 2001; Chagnon et al. 2013). The results reported here favour the latter explanation, although they could have been influenced by the restriction imposed by the experimental conditions of the study, i.e. the size of the mesocosm units or the lack of other host plants usually present in the target ecosystem (Sýkorová et al. 2007).
Further studies are needed in order to fully understand the diversity of life-history strategies among AMF. Additional experiments designed to equalize the inoculum potential of AMF taxa with contrasting life-history strategies will help to get further insights into AMF competition and the ecological processes directing succession in AMF communities.
References
Allen MF, Kitajima K (2013) In situ high-frequency observations of mycorrhizas. New Phytol 200:222–228
Avio L, Pellegrino E, Bonari E, Giovannetti M (2006) Functional diversity of arbuscular mycorrhizal fungal isolates in relation to extraradical mycelial networks. New Phytol 172:347–357
Barea JM, Azcón-Aguilar C (2013) Evolution, biology and ecological effects of arbuscular mycorrhizas. In: Comisão AF, Pedroso CC (eds) Symbiosis: evolution, biology and ecological effects. Nova, New York, pp 1–34
Barea JM, Palenzuela J, Cornejo P, Sánchez-Castro I, Navarro-Fernández C, Lopéz-García A, Estrada B, Azcón R, Ferrol N, Azcón-Aguilar C (2011) Ecological and functional roles of mycorrhizas in semi-arid ecosystems of Southeast Spain. J Arid Environ 75:1292–1301
Bever JD, Richardson SC, Brandy ML, Holmes J, Watson M (2009) Preferential allocation to beneficial symbiont with spatial structure maintains mycorrhizal mutualism. Ecol Lett 12:13–21
Boddington CL, Dodd JC (1999) Evidence that differences in phosphate metabolism in mycorrhizas formed by species of Glomus and Gigaspora might be related to their life-cycle strategies. New Phytol 142:531–538
Chagnon PL, Bradley RL, Maherali H, Klironomos JN (2013) A trait-based framework to understand life history of mycorrhizal fungi. Trends Plant Sci 18:484–491
Collins RE, Rocap G (2007) REPK: an analytical web server to select restriction endonucleases for terminal restriction fragment length polymorphism analysis. Nucleic Acids Res 35:W58–W62
Cornejo P, Azcón-Aguilar C, Barea JM, Ferrol N (2004) Temporal temperature gradient gel electrophoresis (TTGE) as a tool for the characterization of arbuscular mycorrhizal fungi. FEMS Microbiol Lett 241:265–270
Davison J, Öpik M, Daniell TJ, Moora M, Zobel M (2011) Arbuscular mycorrhizal fungal communities in plant roots are not random assemblages. FEMS Microbiol Ecol 178:103–115
Denison RF, Kiers ET (2011) Life histories of symbiotic rhizobia and mycorrhizal fungi. Curr Biol 21:775–785
Dickie IA, FitzJohn RG (2007) Using terminal restriction fragment length polymorphism (T-RFLP) to identify mycorrhizal fungi: a methods review. Mycorrhiza 17:259–270
Dufrêne M, Legendre P (1997) Species assemblages and indicator species: the need for a flexible asymmetrical approach. Ecol Monogr 67:345–366
Dumbrell AJ, Ashton PD, Aziz N, Feng G, Nelson M, Dytham C, Fitter AH, Helgason T (2011) Distinct seasonal assemblages of arbuscular mycorrhizal fungi revealed by massively parallel pyrosequencing. New Phytol 190:794–804
FitzJohn RG, Dickie IA (2007) TRAMPR: an R package for analysis and matching of terminal-restriction fragment length polymorphism (TRFLP) profiles. Mol Ecol Notes 7:583–587
Franson RL, Bethlenfalvay GJ (1989) Infection unit method of vesicular-arbuscular mycorrhizal propagule determination. Soil Sci Soc Am J 53:754–756
Grman E (2012) Plant species differ in their ability to reduce allocation to non-beneficial arbuscular mycorrhizal fungi. Ecology 93:711–718
Hart MM, Reader JR (2002) Taxonomic basis for variation in the colonization strategy of arbuscular mycorrhizal fungi. New Phytol 153:335–344
Hart MM, Reader RJ (2005) The role of the external mycelium in early colonization for three arbuscular mycorrhizal fungal species with different colonization strategies. Pedobiologia 49:269–279
Hart MM, Reader JR, Klironomos JN (2001) Life-history strategies of arbuscular mycorrhizal fungi in relation to their successional dynamics. Mycologia 93:1186–1194
Helgason T, Fitter AH (2009) Natural selection and the evolutionary ecology of the arbuscular mycorrhizal fungi (Phylum Glomeromycota). J Exp Bot 60:2465–2480
Hempel S, Renker C, Buscot F (2007) Differences in the species composition of arbuscular mycorrhizal fungi in spore, root and soil communities in a grassland ecosystem. Environ Microbiol 9:1930–1938
Holland SM (2008) Analytic rarefaction 1.3. UGA Stratigraphy Lab. http://strata.uga.edu/software/anRareReadme.html. Accessed 1 March 2012
Husband R, Herre EA, Turner SL, Gallery R, Young JPW (2002) Molecular diversity of arbuscular mycorrhizal fungi and patterns of host association over time and space in a tropical forest. Mol Ecol 11:2669–2678
IJdo M, Schtickzelle N, Cranenbrouck S, Declerck S (2010) Do arbuscular mycorrhizal fungi with contrasting life-history strategies differ in their responses to repeated defoliation? FEMS Microbiol Ecol 72:114–122
Jansa J, Mozafar A, Kuhn G, Anken T, Ruh R, Sanders IR, Frossard E (2003) Soil tillage affects the community structure of mycorrhizal fungi in maize roots. Ecol Appl 13:1164–1176
Jasper DA, Abbott LK, Robson AD (1989) Soil disturbance reduces the infectivity of external hyphae of vesicular arbuscular mycorrhizal fungi. New Phytol 112:93–99
Kjøller R, Rosendahl S (2000) Detection of arbuscular mycorrhizal fungi (Glomales) in roots by nested PCR and SSCP (Single Strand Conformation Polymorphism). Plant Soil 226:189–196
Koide RT, Mosse B (2004) A history of research on arbuscular mycorrhiza. Mycorrhiza 14:145–163
Krüger M, Krüger C, Walker C, Stockinger H, Schüssler A (2012) Phylogenetic reference data for systematics and phylotaxonomy of arbuscular mycorrhizal fungi from phylum to species level. New Phytol 193:970–984
Lee J, Lee S, Young JPW (2008) Improved PCR primers for the detection and identification of arbuscular mycorrhizal fungi. FEMS Microbiol Ecol 65:339–349
López-Bermúdez F, Albadalejo J, Stocking MA, Díaz E (1990) Factores ambientales de la degradación del suelo en el área mediterránea. In: Albadalejo J, Stocking MA, Díaz E (eds) Degradation and rehabilitation of soil in Mediterranean environmental conditions. CSIC, Murcia, pp 15–45
Maherali H, Klironomos JN (2012) Phylogenetic and trait-based assembly of arbuscular mycorrhizal fungal communities. PLoS ONE 7:e36695
McArdle BH, Anderson MJ (2001) Fitting multivariate models to community data: a comment on distance-based redundancy analysis. Ecology 82:290–297
McGonigle T, Miller M, Evans D, Fairchild G, Swan J (1990) A new method which gives an objective-measure of colonization of roots. New Phytol 115:495–501
Médail F, Quézel P (1997) Hot-spots analysis for conservation of plant biodiversity in the Mediterranean basin. Ann Mo Bot Gard 84:112–127
Merryweather J, Fitter AH (1998) The arbuscular mycorrhizal fungi of Hyacinthoides non-scripta–II. Seasonal and spatial patterns of fungal populations. New Phytol 138:131–142
Munkvold L, Kjøller R, Vestberg M, Rosendahl S, Jakobsen I (2004) High functional diversity within species of arbuscular mycorrhizal fungi. New Phytol 164:357–364
Myers N, Mittermeier RA, Mittermeier CG, da Fonseca GAB, Kent J (2000) Biodiversity hotspots for conservation priorities. Nature 403:853–858
Oehl F, Sieverding E, Palenzuela J, Ineichen K, da Silva GA (2011) Advances in Glomeromycota taxonomy and classification. IMA Fungus 2:191–199
Oksanen J, Blanchet FG, Kindt R, Legendre P, Minchin PR, O’Hara RB, Simpson GL, Solymos P, Henry M, Stevens H et al. (2011) Vegan: community ecology package. R package version 2.0-1. R Project. http://CRAN.R-project.org/package=vegan. Accessed 1 March 2012
Öpik M, Vanatoa E, Moora M, Davison J, Kalwij JM, Reier Ü, Zobel M (2010) The online database MaarjAM reveals global and ecosystemic distribution patterns in arbuscular mycorrhizal fungi (Glomeromycota). New Phytol 188:223–241
Peay KG, Kennedy PG, Bruns TD (2011) Rethinking ectomycorrhizal succession: are root density and hyphal exploration types drivers of spatial and temporal zonation? Fungal Ecol 4:233–240
Phillips JM, Hayman DS (1970) Improved procedures for clearing roots and staining parasitic and vesicular-arbuscular mycorrhizal fungi for rapid assessment of infection. Trans Br Mycol Soc 55:158–161
Powell JR, Parrent JL, Hart MM, Klironomos JN, Rillig MC, Maherali H (2009) Phylogenetic trait conservatism and the evolution of functional trade-offs in arbuscular mycorrhizal fungi. Proc R Soc B 276:4237–4245
Pringle A, Bever JD (2002) Divergent phenologies may facilitate the coexistence of arbuscular mycorrhizal fungi in a North Carolina grassland. Am J Bot 89:1439–1446
R Development Core Team (2010) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Rillig MC (2004) Arbuscular mycorrhizae and terrestrial ecosystem processes. Ecol Lett 7:740–754
Roberts DW (2010) labdsv: ordination and multivariate analysis for ecology. R package version 1.4-1. R Project. http://CRAN.R-project.org/package=labdsv. Accessed 1 March 2012
Rosendahl S, Stukenbrock EH (2004) Community structure of arbuscular mycorrhizal fungi in undisturbed vegetation revealed by analyses of LSU rDNA sequences. Mol Ecol 13:3179–3186
Sánchez-Castro I, Ferrol N, Barea JM (2012a) Analyzing the community composition of arbuscular mycorrhizal fungi colonizing the roots of representative shrubland species in a Mediterranean ecosystem. J Arid Environ 80:1–9
Sánchez-Castro I, Ferrol N, Cornejo P, Barea JM (2012b) Temporal dynamics of arbuscular mycorrhizal fungi colonizing roots of representative shrub species in a semi-arid Mediterranean ecosystem. Mycorrhiza 22:449–460
Schnoor TK, Lekberg Y, Rosendahl S, Olsson PA (2011) Mechanical soil disturbance as a determinant of arbuscular mycorrhizal fungal communities in semi-natural grassland. Mycorrhiza 21:211–220
Simon LM, Lalonde TD, Bruns TD (1992) Specific amplification of 18S fungal ribosomal genes from vesicular arbuscular endomycorrhizal fungi colonising roots. Appl Environ Microbiol 58:291–295
Sýkorová Z, Ineichen K, Wiemken A, Redecker D (2007) The cultivation bias: different communities of arbuscular mycorrhizal fungi detected in roots from the field, from bait plants transplanted to the field, and from a greenhouse trap experiment. Mycorrhiza 18:1–14
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Tommerup IC, Abbott LK (1981) Prolonged survival and viability of VA mycorrhizal hyphae after root death. Soil Biol Biochem 13:431–433
Vallejo VR, Bautista S, Cortina J (1999) Restoration for soil protection after disturbances. In: Trabaud L (ed) Life and environment in the Mediterranean. Advances in ecological sciences. WIT, Wessex, pp 301–343
van der Heijden MGA, Scheublin TR (2007) Functional traits in mycorrhizal ecology: their use for predicting the impact of arbuscular mycorrhizal fungal communities on plant growth and ecosystem functioning. New Phytol 174:244–250
van der Heijden MGA, Klironomos JN, Ursic M, Moutoglis P, Streitwolf-Engel R, Boller T, Wiemken A, Sanders IR (1998) Mycorrhizal fungal diversity determines plant biodiversity, ecosystem variability and productivity. Nature 396:69–72
van der Heijden MGA, Streitwolf-Engel R, Riedl R, Siegrist S, Neudecker A, Ineichen K, Boller T, Wiemken A, Sanders IR (2006) The mycorrhizal contribution to plant productivity, plant nutrition and soil structure in experimental grassland. New Phytol 172:739–752
Vogelsang KM, Reynolds HL, Bever JD (2006) Mycorrhizal fungal identity and richness determine the diversity and productivity of a tallgrass prairie system. New Phytol 172:554–562
Acknowledgements
A. López-García thanks the Formación de Personal Investigador Programme (Ministerio de Ciencia e Innovación, Spain) for financial support. This research was supported by the Spanish Government under the Plan Nacional de I+D+I (project CGL-2009-08825). We sincerely thank Professor Peter Jeffries (Univ. of Kent) for editing comments and grammatical corrections to the manuscript, Dr. Nuria Ferrol for helpful discussions, Dr. Søren Rosendahl and Dr. Alicia Barroso for advices on optimizing the SSCP and TRFLP protocols. Additionally, we would like to thank the two anonymous reviewers and the Section Editor for their valuable comments and suggestions to improve the manuscript. We also thank the Consejería de Medio Ambiente, Junta de Andalucía (Spain) for permission to work in Sierra de Baza Natural Park.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Thom W. Kuyper.
Rights and permissions
About this article
Cite this article
López-García, Á., Palenzuela, J., Barea, J.M. et al. Life-history strategies of arbuscular mycorrhizal fungi determine succession into roots of Rosmarinus officinalis L., a characteristic woody perennial plant species from Mediterranean ecosystems. Plant Soil 379, 247–260 (2014). https://doi.org/10.1007/s11104-014-2060-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-014-2060-6